Structural and Computational Study of the GroEL–Prion Protein Complex
Abstract
1. Introduction
2. Materials and Methods
2.1. Purification of Chaperonin GroEL
2.2. Expression and Purification of the Ovine Prion Protein (PrP)
2.3. Preparation of Cryo-EM Samples
2.4. Single-Particle Data Collection and Processing
2.5. Molecular Dynamics
3. Results
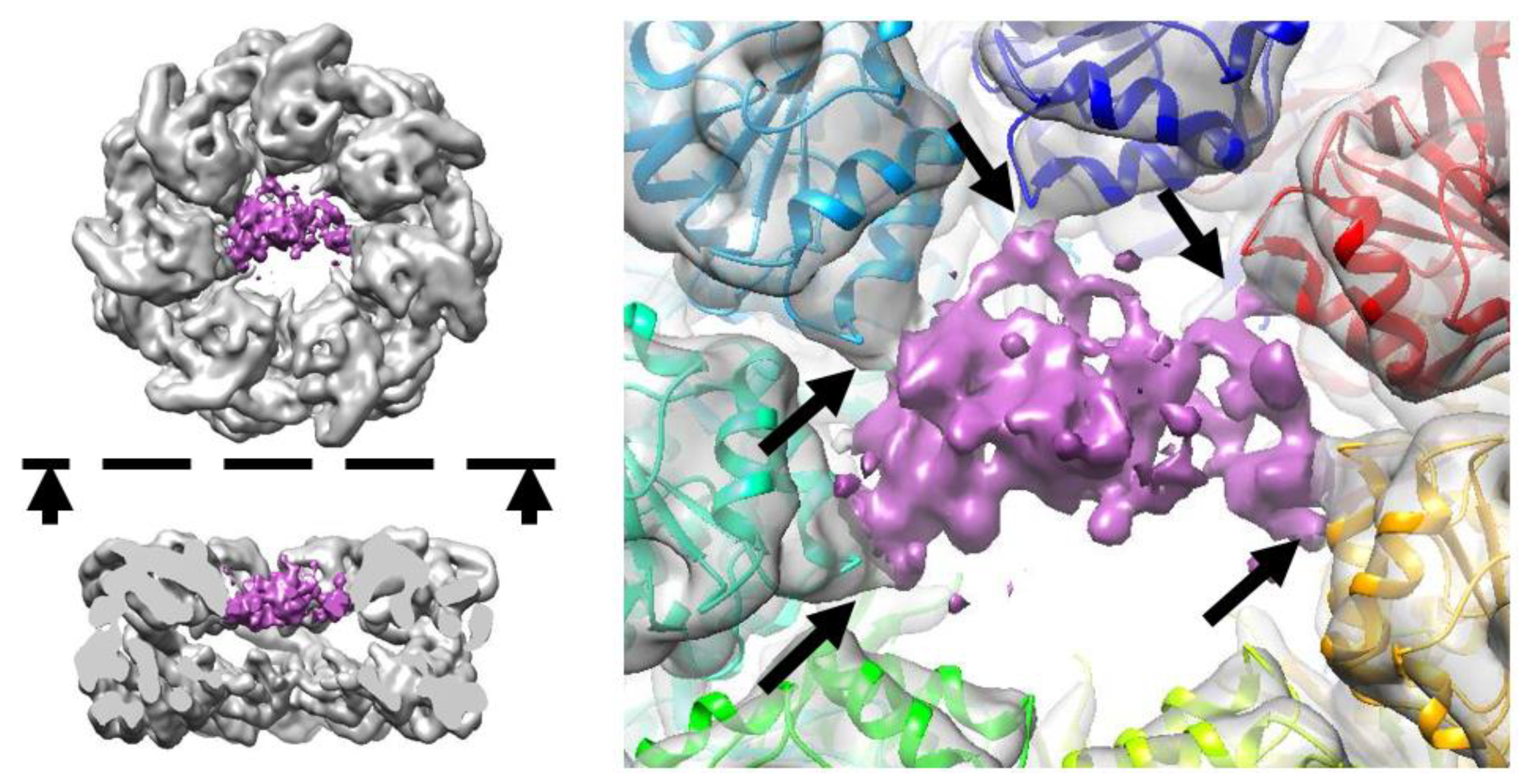
3.1. Cryo-EM Structure of the GroEL–PrP Complex
3.2. Correlation between Cryo-EM and MD Simulations
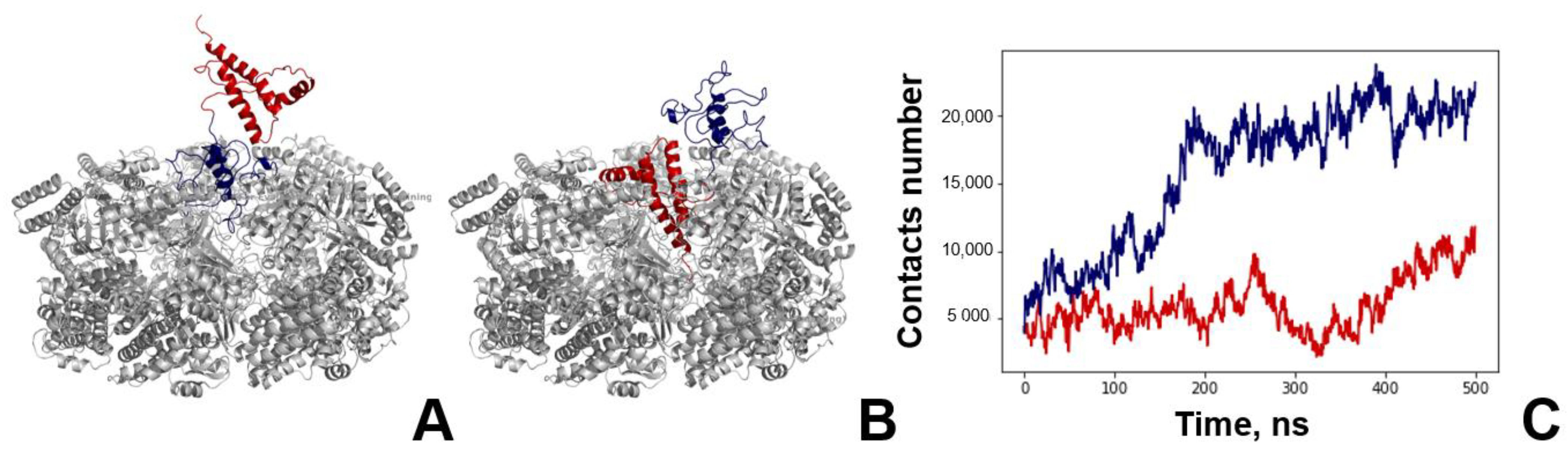
3.3. Analysis of MD Simulation of the GroEL–PrP Complex
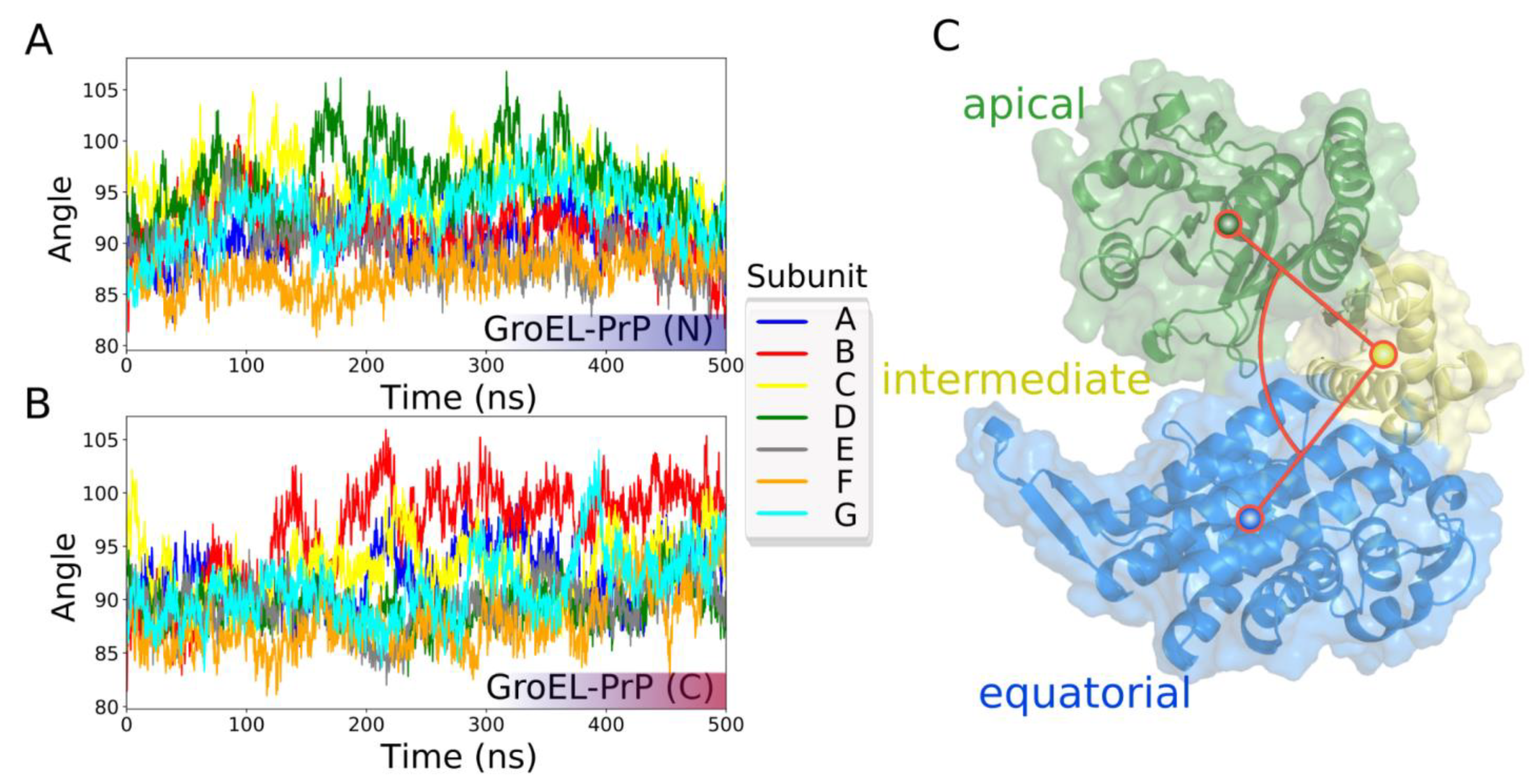
3.4. GroEL Conformation Dynamics
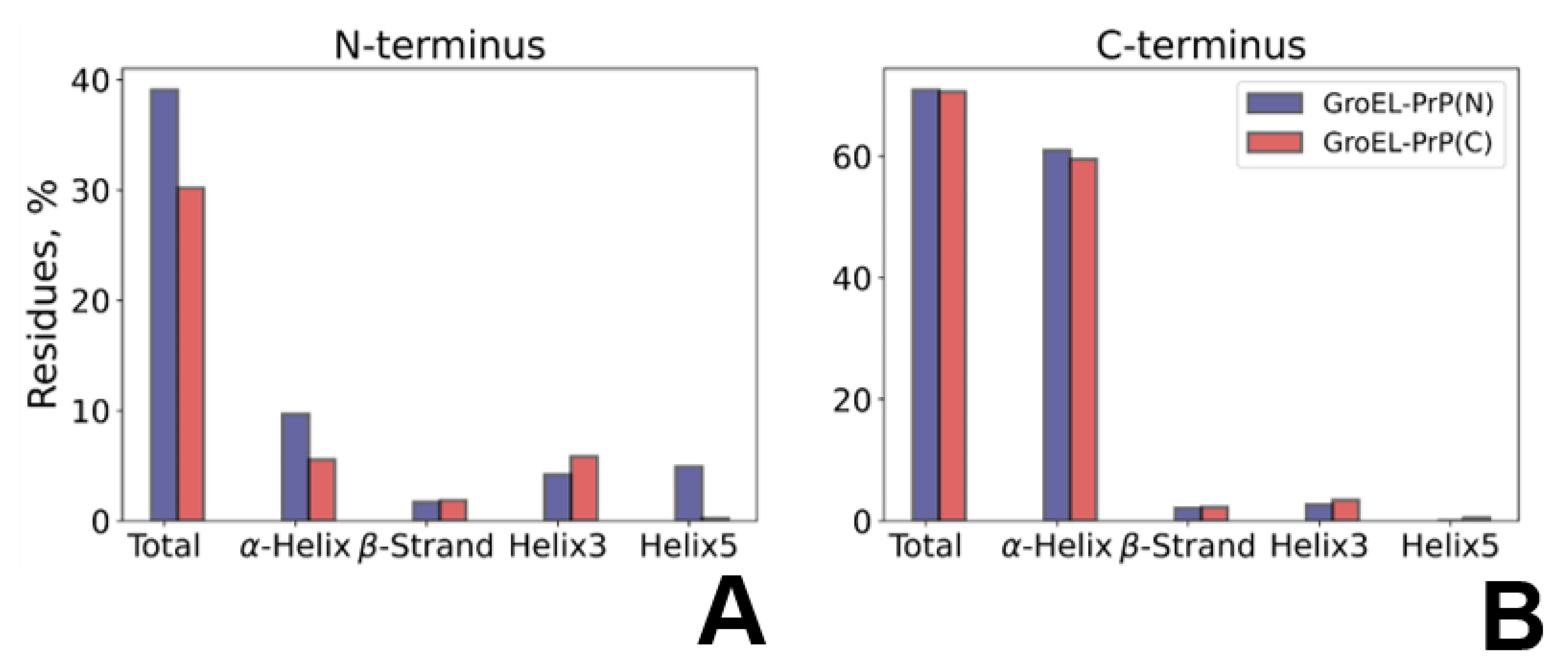
3.5. PrP Conformation Dynamics
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sparrer, H.; Lilie, H.; Buchner, J. Dynamics of the GroEL—Protein Complex: Effects of Nucleotides and Folding Mutants. J. Mol. Biol. 1996, 258, 74–87. [Google Scholar] [CrossRef] [PubMed]
- Grallert, H.; Buchner, J. Review: A Structural View of the GroE Chaperone Cycle. J. Struct. Biol. 2001, 135, 95–103. [Google Scholar] [CrossRef]
- Horwich, A.L.; Farr, G.W.; Fenton, W.A. GroEL-GroES-mediated protein folding. Chem. Rev. 2006, 106, 1917–1930. [Google Scholar] [CrossRef]
- Kudryavtseva, S.S.; Pichkur, E.B.; Yaroshevich, I.A.; Mamchur, A.A.; Panina, I.S.; Moiseenko, A.V.; Sokolova, O.S.; Muronetz, V.I.; Stanishneva-Konovalova, T.B. Novel cryo-EM structure of an ADP-bound GroEL–GroES complex. Sci. Rep. 2021, 11, 18241. [Google Scholar] [CrossRef] [PubMed]
- Fenton, W.A.; Kashi, Y.; Furtak, K.; Norwich, A.L. Residues in chaperonin GroEL required for polypeptide binding and release. Nature 1994, 371, 614–619. [Google Scholar] [CrossRef]
- Buckle, A.M.; Zahn, R.; Fersht, A.R. A structural model for GroEL-polypeptide recognition. Proc. Natl. Acad. Sci. USA 1997, 94, 3571–3575. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Sigler, P.B. The crystal structure of a GroEL/peptide complex: Plasticity as a basis for substrate diversity. Cell 1999, 99, 757–768. [Google Scholar] [CrossRef]
- Chen, D.H.; Madan, D.; Weaver, J.; Lin, Z.; Schröder, G.F.; Chiu, W.; Rye, H.S. Visualizing GroEL/ES in the act of encapsulating a folding protein. Cell 2013, 153, 1354. [Google Scholar] [CrossRef]
- Piana, S.; Shaw, D.E. Atomic-Level Description of Protein Folding inside the GroEL Cavity. J. Phys. Chem. 2018, 122, 11440–11449. [Google Scholar] [CrossRef]
- Machida, K.; Kono-Okada, A.; Hongo, K.; Mizobata, T.; Kawata, Y. Hydrophilic residues 526KNDAAD531 in the flexible C-terminal region of the chaperonin GroEL are critical for substrate protein folding within the central cavity. J. Biol. Chem. 2008, 283, 6886–6896. [Google Scholar] [CrossRef]
- Weaver, J.; Rye, H.S. The C-terminal tails of the bacterial chaperonin GroEL stimulate protein folding by directly altering the conformation of a substrate protein. J. Biol. Chem. 2014, 289, 23219–23232. [Google Scholar] [CrossRef]
- Ishino, S.; Kawata, Y.; Taguchi, H.; Kajimura, N.; Matsuzaki, K.; Hoshino, M. Effects of C-terminal truncation of chaperonin GroEL on the yield of in-cage folding of the green fluorescent protein. J. Biol. Chem. 2015, 290, 15042–15051. [Google Scholar] [CrossRef]
- Oesch, B.; Westaway, D.; Wälchli, M.; McKinley, M.P.; Kent, S.B.; Aebersold, R.; Barry, R.A.; Tempst, P.; Teplow, D.B.; Hood, L.E.; et al. A cellular gene encodes scrapie PrP 27-30 protein. Cell 1985, 40, 735–746. [Google Scholar] [CrossRef]
- Wulf, M.A.; Senatore, A.; Aguzzi, A. The biological function of the cellular prion protein: An update. BMC Biol. 2017, 15. [Google Scholar] [CrossRef] [PubMed]
- Basler, K.; Oesch, B.; Scott, M.; Westaway, D.; Wälchli, M.; Groth, D.F.; McKinley, M.P.; Prusiner, S.B.; Weissmann, C. Scrapie and cellular PrP isoforms are encoded by the same chromosomal gene. Cell 1986, 46, 417–428. [Google Scholar] [CrossRef]
- Pan, K.M.; Baldwin, M.; Nguyen, J.; Gasset, M.; Serban, A.; Groth, D.; Mehlhorn, I.; Huang, Z.; Fletterick, R.J.; Cohen, F.E. Conversion of α-helices into β-sheets features in the formation of the scrapie prion proteins. Proc. Natl. Acad. Sci. USA 1993, 90, 10962–10966. [Google Scholar] [CrossRef] [PubMed]
- Samson, A.O.; Levitt, M. Normal modes of prion proteins: From native to infectious particle. Biochemistry 2011, 50, 2243–2248. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kiselev, G.G.; Naletova, I.N.; Sheval, E.V.; Stroylova, Y.Y.; Schmalhausen, E.V.; Haertlé, T.; Muronetz, V.I. Chaperonins induce an amyloid-like transformation of ovine prion protein: The fundamental difference in action between eukaryotic TRiC and bacterial GroEL. Biochim. Biophys. Acta Proteins Proteom. 2011, 1814, 1730–1738. [Google Scholar] [CrossRef]
- Edenhofer, F.; Rieger, R.; Famulok, M.; Wendler, W.; Weiss, S.; Winnacker, E.L. Prion protein PrPc interacts with molecular chaperones of the Hsp60 family. Virol. J. 1996, 70, 4724–4728. [Google Scholar] [CrossRef] [PubMed]
- DebBurman, S.K.; Raymond, G.J.; Caughey, B.; Lindquist, S. Chaperone-supervised conversion of prion protein to its protease-resistant form. Proc. Natl. Acad. Sci. USA 1997, 94, 13938–13943. [Google Scholar] [CrossRef]
- Stöckel, J.; Hartl, F.U. Chaperonin-mediated de novo generation of prion protein aggregates. J. Mol. Biol. 2001, 313, 861–872. [Google Scholar] [CrossRef]
- Wälti, M.A.; Schmidt, T.; Murray, D.T.; Wang, H.; Hinshaw, J.E.; Clore, G.M. Chaperonin GroEL accelerates protofibril formation and decorates fibrils of the Het-s prion protein. Proc. Natl. Acad. Sci. USA 2017, 114, 9104–9109. [Google Scholar] [CrossRef]
- Kudryavtseva, S.S.; Stroylova, Y.Y.; Zanyatkin, I.A.; Haertle, T.; Muronetz, V.I. Inhibition of chaperonin GroEL by a monomer of ovine prion protein and its oligomeric forms. Biochemistry 2016, 81, 1213–1220. [Google Scholar] [CrossRef] [PubMed]
- Corrales, F.J.; Fersht, A.R. Kinetic significance of GroEL14·(GroES7)2 complexes in molecular chaperone activity. Fold. Des. 1996, 1, 265–273. [Google Scholar] [CrossRef][Green Version]
- Rezaei, H.; Marc, D.; Choiset, Y.; Takahashi, M.; Hui Bon Hoa, G.; Haertlé, T.; Grosclaude, J.; Debey, P. High yield purification and physico-chemical properties of full-length recombinant allelic variants of sheep prion protein linked to scrapie susceptibility. Eur. J. Biochem. 2000, 267, 2833–2839. [Google Scholar] [CrossRef] [PubMed]
- Wagner, T.; Merino, F.; Stabrin, M.; Moriya, T.; Antoni, C.; Apelbaum, A.; Hagel, P.; Sitsel, O.; Raisch, T.; Prumbaum, D.; et al. SPHIRE-crYOLO is a fast and accurate fully automated particle picker for cryo-EM. Commun. Biol. 2019, 2, 218. [Google Scholar] [CrossRef]
- Scheres, S.H.W. RELION: Implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 2012, 180, 519–530. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Fang, J.; Chittuluru, J.; Asturias, F.J.; Penczek, P.A. Iterative stable alignment and clustering of 2D transmission electron microscope images. Structure 2012, 20, 237–247. [Google Scholar] [CrossRef]
- Mamchur, A.A.; Panina, I.S.; Yaroshevich, I.A.; Kudryavtseva, S.S.; Stanishneva-Konovalova, T.B. Molecular Dynamics Simulations of the Full-Length Prion Protein. Lobachevskii J. Math. 2020, 41, 1502–1508. [Google Scholar] [CrossRef]
- Panina, I.S.; Mamchur, A.A.; Yaroshevich, I.A.; Zlenko, D.V.; Pichkur, E.B.; Kudryavtseva, S.S.; Muronetz, V.I.; Sokolova, O.S.; Stanishneva-Konovalova, T.B. Study of GroEL Conformational Mobility by Cryo-Electron Microscopy and Molecular Dynamics. Crystallogr. Rep. 2021, 66, 846–853. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. Gromacs: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Robustelli, P.; Piana, S.; Shaw, D.E. Developing a molecular dynamics force field for both folded and disordered protein states. Proc. Natl. Acad. Sci. USA 2018, 115, E4758–E4766. [Google Scholar] [CrossRef] [PubMed]
- Bussi, G.; Donadio, D.; Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 2007, 126, 014101. [Google Scholar] [CrossRef]
- Parrinello, M.; Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 1981, 52, 7182–7190. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Piana, S.; Donchev, A.G.; Robustelli, P.; Shaw, D.E. Water dispersion interactions strongly influence simulated structural properties of disordered protein states. J. Phys. Chem. 2015, 119, 5113–5123. [Google Scholar] [CrossRef]
- Gowersk, R.J.; Linke, M.; Barnoud, J.; Reddy, T.J.E.; Melo, M.N.; Seyler, S.L.; Domanski, J.; Dotson, D.L.; Buchoux, S.; Kenney, I.M.; et al. MDAnalysis: A Python Package for the Rapid Analysis of Molecular Dynamics Simulations. In Proceedings of the 15th Python in Science Conference, Los Alamos, NM, USA, 11 September 2016; pp. 98–105. [Google Scholar]
- Michaud-Agrawal, N.; Denning, E.J.; Woolf, T.B.; Beckstein, O. MDAnalysis: A toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 2011, 32, 2319–2327. [Google Scholar] [CrossRef] [PubMed]

| GroEL–PrP(N) | ||||||||
| Subunit\Time, ns | A | B | C | D | E | F | G | Sum |
| 0 | 1452 | 408 | 444 | 0 | 20 | 298 | 1333 | 3955 |
| 50 | 2442 | 142 | 1535 | 555 | 64 | 0 | 2642 | 7380 |
| 100 | 3270 | 15 | 2222 | 1011 | 1563 | 459 | 1527 | 10,067 |
| 150 | 2427 | 962 | 2667 | 1673 | 705 | 200 | 3082 | 11,716 |
| 200 | 3454 | 80 | 5675 | 2344 | 381 | 3155 | 2743 | 17,832 |
| 250 | 1974 | 9 | 5977 | 3479 | 744 | 2826 | 3179 | 18,188 |
| 300 | 2738 | 129 | 4480 | 2620 | 798 | 4092 | 1867 | 16,724 |
| 350 | 2465 | 864 | 5790 | 3179 | 680 | 3646 | 3087 | 19,711 |
| 400 | 1970 | 425 | 7387 | 2319 | 610 | 3832 | 4370 | 20,913 |
| 450 | 2410 | 277 | 6781 | 4137 | 524 | 2989 | 2835 | 19,953 |
| 500 | 1880 | 444 | 8282 | 4240 | 487 | 2966 | 4125 | 22,424 |
| average | 2407 | 341 | 4658 | 2323 | 598 | 2224 | 2799 | |
| GroEL–PrP(C) | ||||||||
| Subunit\Time, ns | A | B | C | D | E | F | G | Sum |
| 0 | 1955 | 319 | 883 | 0 | 13 | 780 | 456 | 4406 |
| 50 | 1427 | 43 | 631 | 0 | 2 | 1166 | 448 | 3717 |
| 100 | 1896 | 0 | 32 | 0 | 0 | 90 | 1117 | 3135 |
| 150 | 4133 | 123 | 31 | 0 | 1 | 52 | 92 | 4432 |
| 200 | 4140 | 33 | 8 | 0 | 19 | 359 | 94 | 4653 |
| 250 | 3649 | 593 | 2000 | 114 | 637 | 683 | 166 | 7842 |
| 300 | 2852 | 373 | 9 | 0 | 55 | 722 | 270 | 4281 |
| 350 | 1713 | 78 | 3 | 0 | 122 | 1917 | 469 | 4302 |
| 400 | 1480 | 62 | 3 | 0 | 220 | 2262 | 2474 | 6501 |
| 450 | 3634 | 0 | 125 | 0 | 678 | 2417 | 1759 | 8613 |
| 500 | 4345 | 0 | 893 | 0 | 577 | 4788 | 1172 | 11,775 |
| average | 2839 | 148 | 420 | 10 | 211 | 1385 | 774 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mamchur, A.A.; Moiseenko, A.V.; Panina, I.S.; Yaroshevich, I.A.; Kudryavtseva, S.S.; Pichkur, E.B.; Sokolova, O.S.; Muronetz, V.I.; Stanishneva-Konovalova, T.B. Structural and Computational Study of the GroEL–Prion Protein Complex. Biomedicines 2021, 9, 1649. https://doi.org/10.3390/biomedicines9111649
Mamchur AA, Moiseenko AV, Panina IS, Yaroshevich IA, Kudryavtseva SS, Pichkur EB, Sokolova OS, Muronetz VI, Stanishneva-Konovalova TB. Structural and Computational Study of the GroEL–Prion Protein Complex. Biomedicines. 2021; 9(11):1649. https://doi.org/10.3390/biomedicines9111649
Chicago/Turabian StyleMamchur, Aleksandra A., Andrei V. Moiseenko, Irina S. Panina, Igor A. Yaroshevich, Sofia S. Kudryavtseva, Evgeny B. Pichkur, Olga S. Sokolova, Vladimir I. Muronetz, and Tatiana B. Stanishneva-Konovalova. 2021. "Structural and Computational Study of the GroEL–Prion Protein Complex" Biomedicines 9, no. 11: 1649. https://doi.org/10.3390/biomedicines9111649
APA StyleMamchur, A. A., Moiseenko, A. V., Panina, I. S., Yaroshevich, I. A., Kudryavtseva, S. S., Pichkur, E. B., Sokolova, O. S., Muronetz, V. I., & Stanishneva-Konovalova, T. B. (2021). Structural and Computational Study of the GroEL–Prion Protein Complex. Biomedicines, 9(11), 1649. https://doi.org/10.3390/biomedicines9111649






