Abstract
The novel SH2-containing protein 3 (NSP3) is an oncogenic molecule that has been concomitantly associated with T cell trafficking. However, its oncological role in lung cancer and whether it plays a role in modulating the tumor immune microenvironment is not properly understood. In the present in silico study, we demonstrated that NSP3 (SH2D3C) is associated with advanced stage and poor prognoses of lung cancer cohorts. Genetic alterations of NSP3 (SH2D3C) co-occurred inversely with Epidermal Growth Factor Receptor (EGFR) alterations and elicited its pathological role via modulation of various components of the immune and inflammatory pathways in lung cancer. Our correlation analysis suggested that NSP3 (SH2D3C) promotes tumor immune evasion via dysfunctional T-cell phenotypes and T-cell exclusion mechanisms in lung cancer patients. NSP3 (SH2D3C) demonstrated a high predictive value and association with therapy resistance in lung cancer, hence serving as an attractive target for therapy exploration. We evaluated the in silico drug-likeness and NSP3 (SH2D3C) target efficacy of six organosulfur small molecules from Allium sativum using a molecular docking study. We found that the six organosulfur compounds demonstrated selective cytotoxic potential against cancer cell lines and good predictions for ADMET properties, drug-likeness, and safety profile. E-ajoene, alliin, diallyl sulfide, 2-vinyl-4H-1,3-dithiin, allicin, and S-allyl-cysteine docked well into the NSP3 (SH2D3C)-binding cavity with binding affinities ranging from −3.5~−6.70 Ă and random forest (RF) scores ranging from 4.31~5.26 pKd. In conclusion, our study revealed that NSP3 is an important onco-immunological biomarker encompassing the tumor microenvironment, disease staging and prognosis in lung cancer and could serve as an attractive target for cancer therapy. The organosulfur compounds from A. sativum have molecular properties to efficiently interact with the binding site of NSP3 and are currently under vigorous preclinical study in our laboratory.
1. Introduction
Lung cancer is the second most commonly diagnosed cancer, with an estimated 2.2 million new cases (11.4%) worldwide, and remains the leading cause of cancer deaths, with an estimated 1.8 million deaths (18%) in 2020 [1]. With higher incidences and mortality rates in men than in women, the global burden of lung cancer is ranked first in men, whereas, in women, it ranks second behind breast cancer [1,2]. This pattern is largely attributed to air pollution and increased tobacco consumption [3,4]. Histologically, the majority (80%) of lung cancers are non-small-cell lung cancer (NSCLC) while the remaining 20% are small-cell lung cancer (SCLC) [5,6]. NSCLC is further divided into three pathological subtypes of lung adenocarcinoma (LUAD), squamous cell carcinoma, and large-cell carcinoma, with LUAD being the most common subtype [7,8]. Despite advances in treatment strategies such as chemotherapy, immunotherapy, radiotherapy, and surgery, the prognosis of lung cancer is still disappointing with fewer than 20% of patients reaching an overall 5-year survival period [9,10].
A tumor-driven immune imbalance plays an important role in tumor initiation and progression. The tumor microenvironment (TME) is a complex system that encompasses various stroma cell types, immune cell types, and other extracellular components [11]. Interactions among components of the TME play pivotal roles in tumor occurrence and progression [12]. In addition, the efficacy of immune checkpoint blockades is regulated by the interplay between components of the TME [13,14]. It was initially thought that tumor immune infiltration was strictly associated with tumor-induced immune suppression. However, increasing evidence suggested that both genetic and epigenetic alterations within the TME mediate tumor immune evasion and tumor progression via distinct mechanisms involving T-cell anergy and dysfunctional T-cell phenotypes in various cancer types. In addition, tumor infiltrations of immunosuppressive cells, such as cancer-associated fibroblasts (CAFs), M2-macrophages, and regulatory T (Treg) cells, are known to mediate tumor evasion of immune cells by inhibiting the activities of cytotoxic T cells via a T-cell exclusion mechanism [15,16]. Previous studies reported that CAFs induce the formation of a pre-metastatic niche and increase metastasis in lung cancer patients [17]. CAFs decrease drug sensitivity by blocking the delivery of drugs and thus contribute to the poor prognosis of cancer patients [18,19].
The novel SH2-containing protein 3 (NSP3) is an oncogenic molecule that regulates T cell receptor signaling [20]. It acts as an adapter protein that mediates cell signaling pathways involved in cellular functions such as cell adhesion and migration, tissue organization, and the regulation of the immune response [20,21,22]. It thus could be an attractive target for the development of therapeutic strategies for treating cancer.
In drug discovery pipelines, natural products, particularly medicinal plants, serve as an important source of natural therapeutic agents for treating several diseases [23,24,25,26]. Garlic (Allium sativum L. of the Amaryllidaceae family) is an aromatic herbaceous annual spice with various biological activities [27,28,29,30,31]. It contains hundreds of phytochemicals, among which sulfur-containing compounds are of therapeutic interest in the field of oncology [30,32,33,34,35,36,37,38,39]. Previous studies appraised the uses of in silico approaches to assessing biomarkers and the prognostic relevance of genes and gene signatures in a diseased condition based on the available clinical information of cohorts, and also aid in identifying drug-like molecules for targeted therapy [40,41,42]. In this study, we used a bioinformatics approach to evaluate the effect of NSP3 expression as well as genetic and epigenetic alterations on the TME with respect to tumor immune evasion mechanisms and prognoses in lung cancer cohorts. In addition, we reported the drug-likeness, pharmacokinetics (PKs), toxicity, and potential therapeutic properties of some sulfur-containing compounds from A. sativum. Lastly, we also demonstrated the potential of the compounds to regulate the activity of NSP3 using molecular docking studies of receptor−ligand interactions.
2. Materials and Methods
2.1. Differential Expression Analysis of NSP3 (SH2D3C) in Lung Cancer Cohorts
We used the Tumor IMmune Estimation Resource (TIMER2.0) algorithm (http://timer.cistrome.org/ accessed on 23 July 2021) [43], and the Gene Expression Profiling Interactive Analysis (GEPIA) database (http://gepia.cancer-pku.cn/ accessed on 23 July 2021) [44] for the differential expression analysis of NSP3 (SH2D3C) in tumor vs. normal tissues from various cancer types of The Cancer Genome Atlas (TCGA) database. In addition, we used the TNMplot server [45] to conduct a differential gene expression analysis of NSP3 (SH2D3C) between the pathological free (normal), primary cancer, and metastatic lung cancer tissues.
2.2. Prognostic Analysis of NSP3 (SH2D3C) in Lung Cancer Cohorts
To analyze the prognostic value of NSP3, the expression level and survival information of lung cancer cohorts from GEO, EGA, and TCGA were integrated via PostgreSQL server. We split the patient samples into high and low expression levels based on the quantile expressions levels of the gene. The survival differences between the two groups was visualized using the Kaplan−Meier survival analysis (https://kmplot.com/analysis/ accessed on 21 August 2021) [46]. The hazard ratio with 95% confidence intervals and logrank p value are calculated.
2.3. Analysis of NSP3 (SH2D3C) Genetic Alterations Frequency and Co-occurrence in Lung Cancer Cohorts
We explore the cancer genomic data set through the cBioPortal tool (http://www.cbioportal.org/ accessed on 20 August 2021) [47,48] to analyze the types and frequencies of NSP3 genetic alterations in lung cancer cohorts. The survival differences between cohorts with and without alterations in NSP3 (SH2D3C) genome was also analyzed through the server and visualized using the Kaplan−Meier plots. In addition, we analyzed the gene mutation co-occurrence patterns between NSP3 (SH2D3C) and other genes in TCGA lung cancer cohorts. Alteration co-occurrence was considered significant only at p < 0.05 and q < 0.05.
2.4. Analysis of the SH2D3C Association with Immune and Immunosuppressive Cell Infiltration in Lung Cancer
We explore the immune module of the TIMER algorithm (http://timer.comp-genomics.org/ accessed on 5 August 2021) to comprehensively analyze correlations between NSP3 (SH2D3C) expressions and tumor infiltration levels of six immune cell types (B cells, cluster of differentiation-positive (CD8+) T cells, CD4+ T cells, macrophages, neutrophils, and dendritic cells (DCs)) and immunosuppressive cells (cancer-associated fibroblasts (CAFs), T reg cells, and tumor-associated macrophages (TAMs)) in TCGA cohorts of lung cancer. We used the Tumor Immune Dysfunction and Exclusion (TIDE) (http://tide.dfci.harvard.edu accessed on 3 August 2021) tools [49] to analyze correlations of the expression, methylation, and copy number alterations (CNAs) of NSP3 (SH2D3C) with cytotoxic T lymphocyte (CTL) infiltration, dysfunctional T-cell phenotypes, and T-cell exclusion phenotypes in lung cancer cohorts.
2.5. SH2D3C Interaction Network, Functional Enrichment, and Disease-Specific Associations Analysis
The protein−protein (PPI) interaction network of NSP3 was constructed and analyzed via the search tool for retrieval of interacting genes/proteins (STRING) server (http://string-db.org/, v10.5 accessed on 21 July 2021) [50]. The official gene symbols of NSP3 were loaded to the single protein modules of the server and analyzed for both predictive and known interactions in Homo sapiens under the confidence search of 0.70 and at p < 0.05. In addition, the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways and gene ontology (GO) of the biological process enrichment of the network was downloaded via the server and was further confirmed using the Enrichr platform, an online gene set enrichment analysis (GSEA) server [51,52]. The enrichment plots were generated using the ImageGP (http://www.ehbio.com/ImageGP/ accessed on 27 August 2021) platform. Analysis and visualization of the gene−gene interaction (GGI) network of the NSP3 was performed through the GeneMANIA platform, a real-time multiple association network integration algorithm for predicting gene functions [53]. The gene disease-specific associations of NSP3 was analyzed at the confident levels of 0.4 by using the OPENTARGET platform (https://www.targetvalidation.org/ accessed on 27 August 2021), a bio-web algorithm that integrates genetic, omic, and chemical data to identify the involvement of genes in diseases and aid systematic drug target identification and prioritization [54].
2.6. SH2D3 Knockdown (shSH2D3C) Efficacy and Gene Expression Correlation Analysis in NSCLC
We evaluated the efficacy of SH2D3 knockdown (shSH2D3C) in association with the expression of a number gene. A total of 58 samples are classified into two groups by the median value of shSH2D3C and target genes. Results were presented in the form of predictivity and descriptivity. The predictivity is defined as the fold change (FC) of shSH2D3C efficacy between samples of high and low expression of the gene. Descriptivity is defined as the FC of gene expression between samples of high and low shSH2D3C efficacy.
2.7. In Silico Analysis of the Druglikeness, Pharmacokinetics, and Pharmacology of Some Organosulfur Small Molecules from Allium Sativum
We analyzed the drug-likeness, pharmacokinetics, and medicinal chemistry of six organosulfur small molecules from A. sativum (E-ajoene, alliin, diallyl sulfide, 2-vinyl-4H-1,3-dithiin, allicin and S-allyl-cysteine) using the SwissADME algorithm [55]. We used the blood−brain barrier (BBB) Prediction Server (https://www.cbligand.org/BBB/ accessed on 21 July 2021), which operates based on the support vector machine (SVM) and LiCABEDS algorithms to analyze the BBB permeation ability of the compounds [56]. We evaluated the in silico cytotoxicity of the compounds against the cancer cell lines and normal cell lines by using the Cell Line Cytotoxicity Predictor (CLC-Pred) modules of the computer-aided Prediction of Biological Activity Spectra (PASS) web resources [57] created based on the training set of data on cytotoxicity retrieved from ChEMBLdb (version 23).
2.8. In Silico Acute Toxicity Analysis of the Organosulfur Compounds
We predicted the 50% lethal dose (LD50) of the compounds for the different routes of administrations (intraperitoneal, intravenous, oral, and subcutaneous) in rats using the GUSAR software for quantitative structure-activity relationship (QSAR)/quantitative structure-property relationship (QSPR) modeling [58]. The GUSAR software was developed based on training sets of data from the SYMYX MDL Toxicity Database, consisting of approximately 104 chemical structures with data on acute rat toxicity represented by the LD50 values (log10 (mmol/kg)). In addition, we also evaluated the ecotoxicity properties of the compounds using the bioaccumulation factor Log10(BCF), Daphnia magna 50% lethal concentration (LC50) -Log10(mol/L), Fathead Minnow LC50 Log10(mmol/L), and Tetrahymena pyriformis 50% inhibitory growth concentration (IGC50) -log10(mol/L) as indicators.
2.9. Molecular Docking of Receptor−Ligand Interaction between SH2D3C and the Organosulfur Compound
The crystal structures of NSP3 (PDB: 6W6Y) were downloaded from the Protein Data Bank (PDB) (https://www.rcsb.org/ accessed on 15 August 2021) in PDB file format and subsequently converted into the Auto Dock PDBQT format using AutoDock Vina (ver. 0.8, Scripps Research Institute, La Jolla, CA, USA) [59]. Three-dimensional (3D) molecular ball-and-stick models of some organosulfur compounds from A. sativum (alliin, allicin, E-ajoene, Z-ajoene, 2-vinyl-4H-1,3-dithiin, diallyl sulfide, and allyl methyl sulfide) were drawn using the Avogadro molecular builder and visualization tool ver. 1.XX and subsequently converted to PDB format using the PyMOL tool and then to PDBQT format using AutoDock Vina. The receptor was prepared by pre-docking removal of water (H20) molecules and the addition of hydrogen atoms (polar only) and Kolman charges [60,61]. Molecular docking was performed using AutoDock Vina as described previously [40,60,62,63]. Docking outcomes were visualized using the Discovery Studio Visualizer ver. 19.1.0.18287 (BIOVIA, San Diego, CA, USA) [64].
2.10. Data Analysis
Spearman’s rank correlations were used to assess correlations between NSP3 expression and infiltration of immunosuppressive cells. A KM curve was employed to visualize survival differences between lung cancer cohorts with high and those with low expression levels of NSP3 (SH2D3C). The statistical significance of the differential expression of NSP3 between lung cancer tumor and adjacent normal tissue was evaluated using the Wilcoxon test. Statistical significance was indicated by * p < 0.05; ** p < 0.01; *** p < 0.001.
3. Results
3.1. NSP3 (SH2D3C) Is Associated with Advance Stage and Poor Prognoses of Lung Cancer Cohorts
Our differential expression profile of NSP3 (SH2D3C) between tumor and adjacent normal tissue revealed that NSP3 (SH2D3C) has deregulatory expression in tumor samples of TCGA cancer types (Figure 1A). However, our focus is on lung cancer, and we found that NSP3 (SH2D3C) is expressed at a lower level in lung cancer tumors than the adjacent normal tissue (Figure 1B). On a contrary note, we found that cohorts with metastatic lung cancer exhibited higher NSP3 (SH2D3C) expression levels than the primary tumor (Figure 1C). Our survival analysis also revealed that patients with higher gene expression levels exhibited a shorter overall and progressive free survival duration than cohorts with lower expression levels (Figure 1D). Furthermore, we queried the differential protein expression levels of NSP3 (SH2D3C) between lung cancer tissue and disease-free tissue. Interestingly, we found that lung cancer tumors exhibited higher intensity of NSP3 (SH2D3C) than the pathological free tissue (Figure 1E). Moreover, cohorts with high mRNA expression levels of NSP3 (SH2D3C) exhibited shorter survival duration than cohorts with low mRNA expression levels (Figure 1F). Collectively, our results suggest that NSP3 (SH2D3C) is associated with the advanced stage and poor prognosis of lung cancer and thus could serve as a prognostic biomarker of lung cancer progression and follow-up.
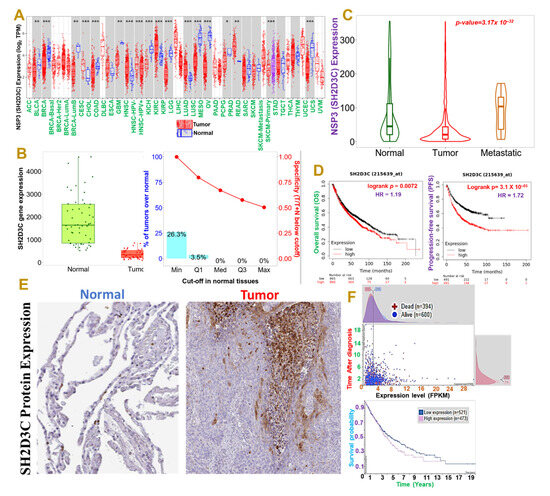
Figure 1.
Non-structural protein 3 (NSP3; SH2D3C) is associated with advanced stages and poor prognoses of lung cancer cohorts. (A) Box plots showing differential expression profile of NSP3 (SH2D3C) between tumor and adjacent normal tissue across all TCGA cancer types and (B) between tumor and adjacent normal tissues in lung cancer cohort. (C) Violine plots showing differential expression profile of NSP3 (SH2D3C) between normal tissue, tumor, and metastatic lung cancer patients. (D) Kaplan–Meier plot of the overall and progressive-free survival between lung adenocarcinoma patients with low and high expression levels of NSP3 (SH2D3C). (E) Representative immunohistochemistry staining of SH2D3C between lung cancer tumor and pathological free tissue in the Human Protein Atlas (HPA) database. (F) Interactive survival scatter plots and Kaplan–Meier plot of the survival probability between high and low mRNA expression levels in lung cancer cohorts. * p < 0.05; ** p < 0.01; *** p < 0.001.
3.2. Genetic Alterations of NSP3 (SH2D3C) Are Associated with Poorer Prognosis and Inversely Associated with EGFR Alterations in Lung Cancer Patients
We used the cBioPortal database to query the genetic alteration profile of NSP3 (SH2D3C) in 566 lung adenocarcinoma cases (TCGA, PanCancer Atlas) and found that 2.1% of the cohorts harbored genetically altered NSP3 (SH2D3C) (Figure 2A). However, mutation and multiple alterations were the only genetic alterations of NSP3 (SH2D3C) in lung adenocarcinoma cohorts (Figure 2B). Specific mutation profiling indicated that out of the total NSP3 (SH2D3C) mutation in the database, 83.33% were missense (Figure 2B, C) and only a few truncating mutations occur, while no in-frame or fusion mutations were recorded (Figure 2C). Furthermore, we found that the genetic alterations of NSP3 (SH2D3C) are associated with shorter overall survival (Figure 2D), disease-free survival (Figure 2E), and progression-free survival (Figure 2F) of the cohorts. We queried the co-occurrence of NSP3 (SH2D3C) alterations with other genetic alterations and found that NSP3 (SH2D3C) alteration is inversely associated with Epidermal Growth Factor Receptor (EGFR) alterations in lung cancer, i.e., lung cancer patients with NSP3 (SH2D3C) alteration are devoid of EGFR alterations (Figure 2G).
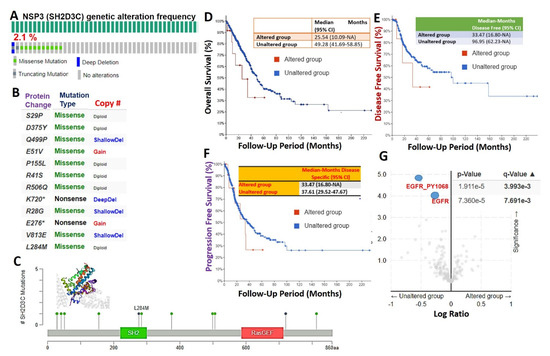
Figure 2.
Genetic alterations of non-structural protein 3 (NSP3; SH2D3C) are associated with poorer prognosis and inversely associated with Epidermal Growth Factor Receptor (EGFR) alterations in lung cancer patients. Genetic alteration profile of NSP3 (SH2D3C) in lung adenocarcinoma. (A) Prevalence and distribution of NSP3 (SH2D3C) genetic alterations in lung adenocarcinoma cohort. (B) Specific mutation types of NSP3 (SH2D3C) in lung adenocarcinoma cohorts. (C) Lollipop plot of NSP3 (SH2D3C) mutation location in lung adenocarcinoma cohort across the cBioPortal for Cancer Genomics dataset. Mutations are color-coded as missense, truncating, and in-frame mutations. (D) Kaplan–Meier plot of the (D) shorter overall survival (E), disease-free survival, and (F) progression-free survival between lung cancer cohorts with genetically altered and cohorts without altered NSP3 (SH2D3C). (G) Heat map of NSP3 (SH2D3C) mutation co-occurrence in lung cancer patients.
3.3. NSP3 Elicits Its Pathological Role via Modulation of Various Components of the Immune and Inflammatory Pathways in Lung Cancer
The GGI network revealed that the NSP3 gene interacts with a number of oncogenic, immune, and inflammatory genes including AR, BCAR1, BCAR3, CAV1, CSRNP1, EFS, EGFR, EPHB2, GIMAP6, ICAM2, IL7R, KIT, MET, MYO1G, NAALADL1, NEDD9, PTK2B, RGL4, SEPTIN1, and TNNT (Figure 3A). Similarly, the PPI network analysis revealed 21 nodes, 48 edges, an average local clustering coefficient of 0.747 and a PPI enrichment p-value of 0.000159 (Figure 3B). The nodes with the highest degree in the network included SH2D3C (11), G3BP1 (8), EIF4G1 (7), PABPC1 (7), BCAR1 (5), CAPRIN1 (5), EIF4A1 (5), and EIF4A2 (5) (Figure 3B). To analyze the functional enrichment of the gene network, we examined the GO biological processes and KEGG pathways, and our results revealed the biological enrichment of the vascular endothelial growth factor receptor (VEGFR) signaling pathway, immune response-activating cell surface receptor signaling, Fc receptor signaling pathway, cytoplasmic translational initiation, viral process, transmembrane receptor protein tyrosine kinase signaling pathway, post-transcriptional regulation of gene expression, regulation of substrate adhesion-dependent cell spreading, I-kappaB kinase/NF-kappaB signaling, and Fc-gamma receptor signaling pathway involved in phagocytosis (Figure 3C), while the KEGG pathway analysis revealed enrichment of Yersinia infection, shigellosis, chemokine signaling pathway, human cytomegalovirus infection, bacterial invasion of epithelial cells, chronic myeloid leukemia, human immunodeficiency virus 1 infection, and small cell lung cancer (Figure 3D). Collectively, our results suggested that NSP3 is involved in the pathology of bacterial and viral infections, immune and inflammatory diseases, and cell proliferation and lung cancer development.
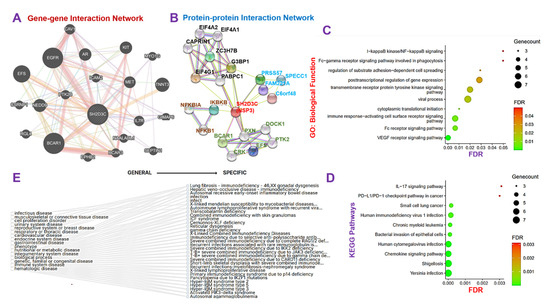
Figure 3.
Non-structural protein 3 (NSP3; SH2D3C) elicits its pathological role via modulation of various components of the immune and inflammatory pathways in lung cancer. (A) Gene−gene interaction (GGI) and (B) protein−protein interaction (PPI) network of NSP3. (C) The Kyoto Encyclopedia of Genes and Genomes and (D) G.O. biological function enrichments of the NSP3 network proteins. (E) Gene-diseases associated tree plot of NSP3.
3.4. NSP3 (SH2D3C) Promotes Tumor-Immune Evasion via Dysfunctional T-Cell Phenotypes and T-Cell Exclusion Mechanism in Lung Cancer Patients
We analyzed the relationship of NSP3 (SH2D3C) expression with tumor infiltration by immune cells and found that NSP3 (SH2D3C) expression showed strong correlations with the infiltration of six immune cell types, including the B cells (r = 0.292–0.3220 all p < 0.1 × 10−10), CD8+ T cells (r = 0.18–0.33 all p < 0.05), CD4+ T cells (r = 0.54–0.64 all p < 0.1 × 10−30), M1-macrophages (r = 0.295–0.44 all p < 0.1 × 10−10), neutrophils (r = 0.34–0.40 all p < 0.1 × 10−10), and DCs (r = 0.35–0.59 all p < 0.1 × 10−10) (Figure 4A). Interestingly, we found that NSP3 (SH2D3C) expression was inversely associated with CTL levels in lung cancer patients (Figure 4B). In addition, we found that a higher expression profile of NSP3 (SH2D3C) in cohorts with higher CTL levels exhibited a shorter survival period, while a lower expression profile of NSP3 (SH2D3C) in cohorts with higher CTL levels exhibited a longer survival period, suggesting a correlation with dysfunctional T-cell phenotypes (Figure 4C). Furthermore, we queried associations of NSP3 (SH2D3C) expressions with infiltration levels of immunosuppressive cells and found that NSP3 (SH2D3C) expression was positively correlated with tumor infiltration of CAFs (r = 0.205–0.395 all p < 0.1 × 10−5), T reg cells (r = 0.231–0.143 all p < 0.05), and M2 macrophages (r = 0.535–0.414 all p < 0.1 × 10−20) in LUAD and LUSC, respectively (Figure 4D). Altogether, our findings strongly suggest that NSP3 (SH2D3C) promotes tumor immune evasion via dysfunctional T-cell phenotypes and T-cell exclusion mechanisms in lung cancer patients.
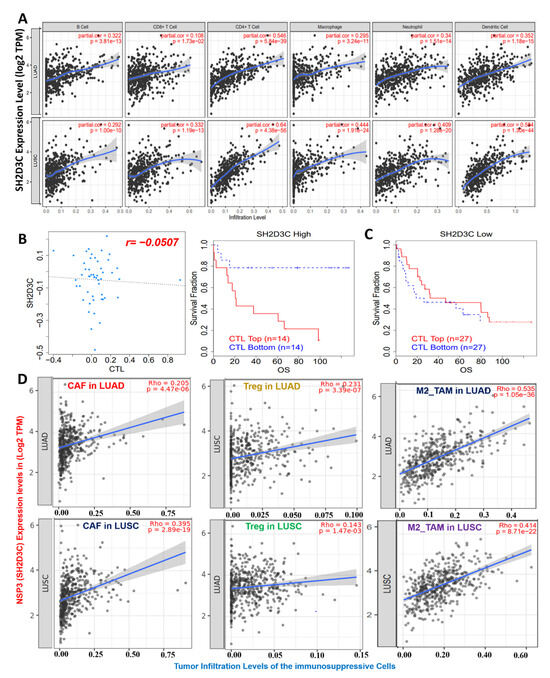
Figure 4.
Non-structural protein 3 (NSP3; SH2D3C) promotes dysfunctional T-cell phenotypes in lung cancer patients. (A) Scatterplot showing correlations between NSP3 (SH2D3C) expression and infiltration of various immune cells in lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC). (B) Correlation analysis between NSP3 (SH2D3C) expression and cytotoxic T lymphocyte (CTL) levels in lung cancer patients. (C) Kaplan–Meier plot of the overall survival of lung cancer patients with different NSP3 (SH2D3C) and CTL levels. (D) Boxplot showing NSP3 (SH2D3C) expression correlations with infiltration levels of immunosuppressive cells, including cancer-associated fibroblasts (CAFs), M2 macrophages, and regulatory T cells in LUAD patients.
3.5. NSP3 (SH2D3C) Methylation and Copy Number Alterations (CNA) Are Associated with Infiltration of Immune Cells and Poorer Prognosis of Lung Cancer Cohorts
We evaluated the effect of CNA and methylation of NSP3 (SH2D3C) in lung cancer (Figure 5A–F), and the results revealed that NSP3 (SH2D3C) is hypermethylated in tumors of lung cancer patients compared to the normal tissue (Figure 5D). In addition, lung cancer patients with high methylation levels of NSP3 (SH2D3C) exhibited shorter survival duration than cohorts with low methylation levels (Figure 5E). Analysis of CTL level and methylation status of NSP3 (SH2D3C) revealed that cohorts with high levels of NSP3 (SH2D3C) exhibited dysfunctional T-cell phenotypes and poor prognosis (Figure 5F). Nevertheless, we found that the CNA of NSP3 (SH2D3C) are a risk factor for LUAD and LUSC cohorts (Figure 5A). Analysis of the patients’ survival revealed that both LUAD and LUSC patients with CNA of NSP3 (SH2D3C) exhibited shorter overall survival duration than the cohort with low CNA (Figure 5B). In addition, we found that SCNA of NSP3 (SH2D3C) including deep deletion, arm-level deletion, arm-level gain, and high amplification also mediated the infiltrations of the tumor-immune cells in both LUAD and LUSC (Figure 5C). Collectively, these findings prove that hyper-methylation and copy number alterations of NSP3 (SH2D3C) are associated with infiltration of immune cells and poorer prognosis of lung cancer cohorts.
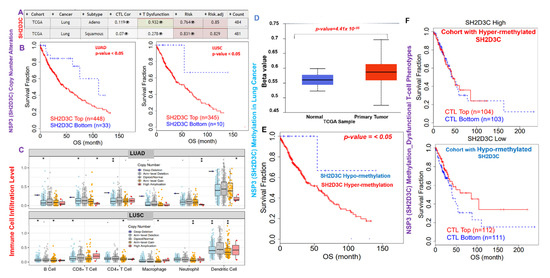
Figure 5.
Non-structural protein 3 (NSP3; SH2D3C) methylation and copy number alterations (CNA) are associated with infiltration of immune cells and poorer prognosis of lung cancer cohorts. (A) Heatmap plot of the NSP3 (SH2D3C) expression and T-cell profile in LUAD and LUSC cohorts. (B) Kaplan–Meier plot of the overall survival differences between high and low CNA of NSP3 (SH2D3C) in LUAD and LUSC cohorts. (C) Boxplot showing the association between different somatic copy number alterations and infiltration of various immune cells in LUAD and LUSC. (D) Boxplot showing differential NSP3 (SH2D3C) methylation between lung cancer tumors and adjacent normal tissue. (E) Kaplan–Meier plot of the overall survival differences between lung cancer cohorts with high and low methylation levels of NSP3 (SH2D3C). (F) Kaplan–Meier plot of the overall survival of lung cancer patients with different NSP3 (SH2D3C) methylation status and CTL levels. The “eye” sign represent the hidden view of the CTL, dysfunctional and risk plots of the cohorts.
3.6. SH2D3C Is Associated with Therapy Resistance in NSCLC
We evaluated the cross-correlation between the expression levels of SH2D3C and the sensitivity of NSCLC cell lines to various clinical drugs. Our results revealed that high expression level of SH2D3C is associated with the resistance of NSCLC cell lines to doxorubicin, mitoxantrone, GSK1070916, obatoclax mesylate, alisertib, NSC319726, mitomycin-C, bleomycin, etoposide, pelitinib, and BX-912 (Figure 6A). Furthermore, we evaluated the efficacy of SH2D3 knockdown (shSH2D3C) in association with the expression of a number of target genes. Interestingly, we found that higher expression levels of the target genes are associated with the low efficacy of shSH2D3C in NSCLC cell lines (Figure 6B). Analysis of the biomarker relevance revealed a clinical predictive value (AUC > 0.5) of SH2D3C in 12 datasets of immunotherapy cohorts. Interestingly, SH2D3C achieved a higher clinical predictive score in a larger number of datasets when compared with TMB, T-cell clonality, and B-cell clonality with a clinical predictive score in 7, 8, and 8 datasets of immunotherapy cohorts (Figure 6C). Summing up the results obtained so far, our study suggested that SH2D3C is an important player in immune and inflammatory events in the carcinogenesis of NSCLC. It is associated with response to therapy in NSCLC and thus serves as an attractive target for exploration.
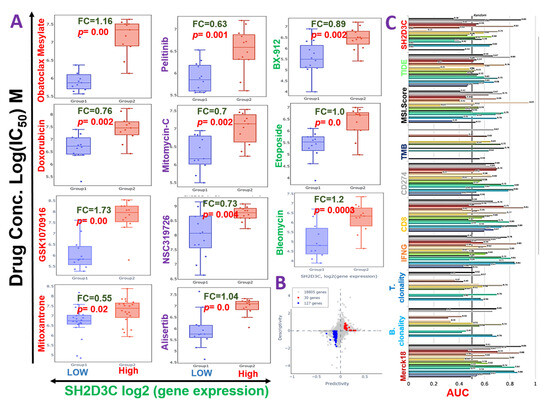
Figure 6.
Non-structural protein 3 (NSP3; SH2D3C) is associated with chemotherapy resistance in NSCLC. (A) Bar plots showing the sensitivity of various clinical drugs between samples with low and high expression levels of SH2D3C. (B) Scatter plot of the efficacy of SH2D3 knockdown (shSH2D3C) in association with the expression of a number gene. The predictivity is defined as the fold change (FC) of shSH2D3C efficacy between samples of high and low expression of the gene. Descriptivity is defined as the FC of gene expression between samples of high and low shSH2D3C efficacy. (C) Bar plot of the comparative biomarker relevance between the SH2D3C and standardized biomarkers.
3.7. The Organosulfur Compounds in Allium sativum Exhibited Desirable Physicochemical and Pharmacokinetics Properties of a Drug-Like Candidate
Our in silico analysis of the drug-likeness, physicochemical, and pharmacokinetic properties of the organosulfur small molecules from A. sativum (Table 1, Table S1, Figure S1) revealed good predictions of the six compounds for ADMET properties, drug-likeness, adherence to Lipinskís rules, and no alerts for PAINS (Table 1). Our analysis of the human gastrointestinal absorption and blood−brain barrier penetration ability of the compounds indicated that all the six compounds evaluated demonstrated BBB penetration ability (Figure 7A), with E-ajoene allicin and diallyl disulfide having higher brain-penetrant scores. QSAR modeling of acute toxicity in rats revealed estimated LD50 concentrations ranged between 74.190~937.700 mg/kg, 50.130~772.600 mg/kg BW, 429.00~3155.00 mg/kg, and 25.850~798.300 mg/kg for intraperitoneal, intravenous, oral, and subcutaneous route of administrations of the six compounds, respectively (Table 1), suggesting a high safety profile of the compounds especially when administered via the oral route.

Table 1.
Drug likeness, medicinal chemistry, and physicochemical and ADMET properties of the organosulfur small molecules from Allium sativum.
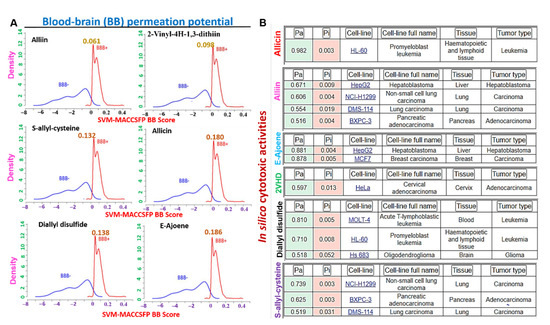
Figure 7.
Blood−brain barrier permeation and in silico cytotoxic properties of some organosulfur compounds in garlic. (A) Blood−brain barrier penetration (BBB) permeation curve of some organosulfur compounds in A. sativum. The BBB permeation ability was measured by the support vector machine (SVM) and LiCABEDS algorithms of the BBB prediction server. (B) In silico cytotoxic properties of some organosulfur compounds in garlic against cancer cell lines. Pa = activity, pi = inactivity.
3.8. The Organosulfur Compounds from Allium sativum Exhibited Selective in Silico Cytotoxic Activities against Cancer Cell Lines and Are not Cytotoxic to Normal Human Cell Lines
We used the in silico approach to evaluate the possible cytotoxic activities of E-ajoene, alliin, diallyl sulfide, 2-vinyl-4H-1,3-dithiin, allicin and S-allyl-cysteine on cancer cell lines as well as normal cell lines. Interestingly, we found that none of the compounds exhibit cytotoxic activities on the normal human cell lines; however, our results revealed a very strong cytotoxic activity (pa = 0.982) of allicin against leukemia cell line (HL-60), and strong activity of E-ajoene on the liver (pa = 0.881) and breast (pa = 0.878) cancer cell lines. Alliin was identified to be active against lung, liver, and pancreatic cancer cells (pa = 0.51~0.671). 2-vinyl-4H-1,3-dithiin demonstrated in silico cytotoxic activity against leukemia and brain cancer cell lines (pa = 0.518~0.810), while S-allyl-cysteine demonstrated activity against the lung and pancreatic cancer cell lines (pa = 0.519~0.739). Collectively, this study revealed that the six organosulfur compounds in garlic evaluated in this study exhibited selective cytotoxic activity on cancer cell lines and are not cytotoxic to normal human cell lines (Figure 7B).
3.9. Molecular Docking Profile of NSP3 (SH2D3C) with Some Organosulfur Compounds in Allium Sativum
Molecular docking studies revealed that E-ajoene, alliin, diallyl sulfide, 2-vinyl-4H-1,3-dithiin, and allicin docked well into the NSP3 (SH2D3C) binding cavity with binding affinities ranging −3.5~−6.70 Ă and random forest (RF) ranging 4.31~5.26 pKd (Table 2). The complexes are bounded by a number of conventional H-bonding and alkyl interactions. Furthermore, the complexes are stabilized by several hydrophobic contacts and van der Waals forces with several amino acids, surrounding the compound backbones in the receptor-binding pocket (Figure 8 and Figure 9).

Table 2.
Docking profile of NSP3 (SH2D3C) with the organosulfur small molecule from Allium sativum.
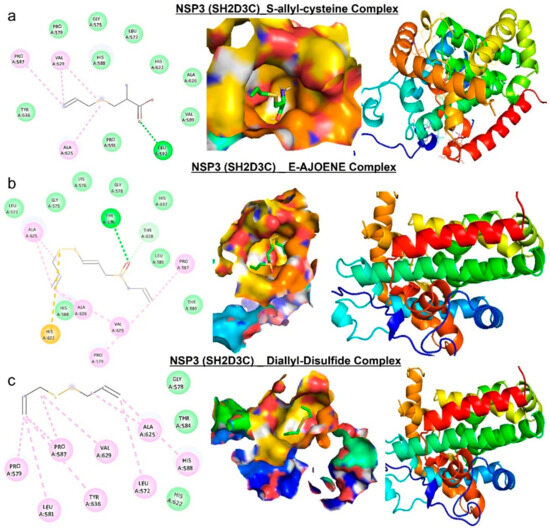
Figure 8.
Molecular docking profile on NSP3 with the organosulfur small molecule from Allium sativum. Two-dimensional (2D) structure and binding surface flip of the ligand−receptor interactions between NSP3 (SH2D3C) and (a) S-allyl-cysteine, (b) E-ajoene, and (c) diallyl sulfide.
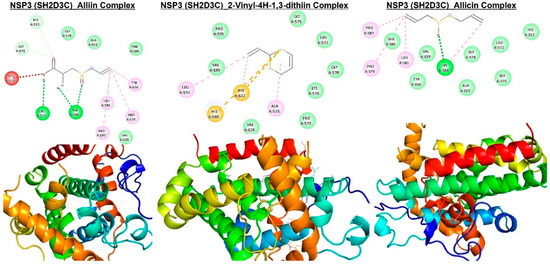
Figure 9.
Molecular docking profile on novel SH2-containing protein 3 (NSP3) with the organosulfur small molecule from Allium sativum. Three- (3D) and two-dimensional (2D) structure of the ligand−receptor interactions between NSP3 (SH2D3C) and alliin, allicin, and 2-vinyl-4H-1,3-dithiin.
4. Discussion
The impacts of cancer incidences and mortality are increasing at a global level. An estimated 19.3 million new cancer cases and almost 10.0 million cancer deaths occurred in 2020 [1]. Messenger (m)RNA expressions play important roles in cancer cell progression and survival, and comparative gene expression profiles between tumors and adjacent tissues have been very useful in identifying potential cancer biomarkers [65,66]. Our results suggest that NSP3 (SH2D3C) is differentially expressed in different cancer types and thus could serve different prognostic purposes in different cancer types. As such, different therapeutic approaches must be considered when targeting NSP3 (SH2D3C) in different cancer types. Controversial roles of proteins in different cancer types have been reported in previous studies; for instance, EPHB4 was described as an oncogene in some cancers and as a tumor suppressor in other cancers [67,68,69]. These contradictory findings suggest a complex role of NSP3 signaling in cancer, characterized by multi-domain and multifunctional properties. Specifically, our differential expression and survival analyses strongly suggested that NSP3 (SH2D3C) is a potential biomarker of advanced stages of lung cancer.
Genes with similar expression patterns share similar functions. Therefore, gene functional enrichment analysis is important in an in silico approach and has been widely used in the analysis of gene-related pathological processes [70,71]. Our interaction network revealed a number of onco-functional genes including G3BP1, EIF4G1, PABPC1, BCAR1, CAPRIN1, EIF4A1, and EIF4A2, and HIRA as the functional partners associated with the immune-oncogenic roles of SH2D3C. Consequently, GO and KEGG pathway enrichment analyses were conducted to identify functional processes and pathways that may be mediated by these genes. Interestingly, our results suggested that NSP3-associated networks were involved in several pathways and biological processes associated with bacterial and viral infections, immune and inflammatory diseases, and cell proliferation and lung cancer development.
Tumor immune cell infiltrations are indicators of host immune responses to cancer cells [72,73]. Several studies revealed that some tumors have a high level of infiltration by cytotoxic T cells in a dysfunctional state and could not elicit tumor growth, while in other tumors, immunosuppressive factors may prevent T cells from infiltrating the tumors [74,75,76]. These immunosuppressive cells, including T reg cells, M2-TAMs, and CAFs, are known to promote tumorigenic and metastatic properties by inhibiting T-cell expansion, secreting cytokines, and remodeling the extracellular matrix (ECM) [77,78]. Interestingly, our analysis of gene expression correlations with tumor infiltrations of immune cells and immunosuppressive cells revealed that NSP3 could regulate lung cancer tumor evasion of immune cells via both T-cell exclusion and induction of dysfunctional T-cell phenotypes. In addition, both genetic alterations and gene methylation of NSP3 were also found to regulate dysfunctional T-cell phenotypes and are associated with the early death of lung cancer patients. Altogether, our findings strongly suggest that NSP3 is an important onco-immunological biomarker encompassing the TME, disease staging, and prognosis in lung cancer, and can serve as an attractive target for cancer therapies. However, subsequent development of NSP3 as a reliable therapeutic target in lung cancer requires preclinical and clinical validation.
Drug-likeness is a complex balance of molecular properties and structural features which determine whether a molecule is like known drugs [79,80]. Interestingly, our in silico analysis revealed that the six organosulfur compounds from A. sativum are good drug-like candidates and exhibited selective in silico cytotoxic activities against lung, brain, liver, and pancreatic cancers with no cytotoxic effects on normal cell lines. The compounds are BBB-penetrant and therefore will be very useful in treating glioblastomas and other diseases associated with the central nervous system (CNS). Evaluation of toxicity of natural products and active compounds is an important factor in the development of drugs for therapeutic applications [81,82,83]. Our in silico acute toxicity and ecotoxicity studies revealed that different routes of administration of the sulfur-containing compounds produced different LD50 levels in rats. The overall analysis, however, revealed that the oral route was the safest route for administering these compounds; this route should be well tolerated when used for acute administration and is currently being employed for in vivo evaluations of these compounds against lung cancer and glioblastomas in our laboratory.
Molecular docking has become an increasingly valuable tool for depicting the possible interaction between a small-molecule drug candidate and a protein target during the early stage of drug development [84,85,86]. Our molecular docking analysis indicated that the organosulfur compounds of Allium sativum have the molecular properties to interact efficiently with the binding site of NSP3 and are currently under rigorous experimental validation in our laboratory. E-ajoene, alliin, diallyl sulfide, 2-vinyl-4H-1,3-dithiin, and allicin docked well into the binding cavity of NSP3 (SH2D3C). Hydrogen bonds and other noncovalent interactions, such as hydrophobic and ionic interactions and van der Waals forces, play important roles in stabilizing ligand−protein complexes [87]. We found that the interactions of the compounds with NSP3 predominantly involved hydrogen bonds, van der Waals forces, π-alkyl, and various hydrophobic interactions. The van der Waals forces created around the compound’s backbone with the several amino acids create a strong cohesive environment, which further stabilized the complexes [88]. However, S-allyl-cysteine interaction with NSP3 (SH2D3C) is unfavorable and hence less susceptible to NSP3 ligandability. Altogether, our molecular docking experiments suggested that the organosulfur compounds from A. sativum have the molecular properties to interact efficiently with the binding site of NSP3. This interaction is under rigorous experimental validation in in vitro as well as in vivo models of GBM and lung cancer.
5. Conclusions
In conclusion, our in silico study suggested that NSP3 is an important onco-immunological biomarker encompassing the tumor microenvironment, disease staging, and prognoses in lung cancer and could serve as an attractive target for cancer therapies. Our molecular docking experiments suggested that the organosulfur compounds from Allium sativum, including alliin, allicin, E-ajoene, Z-ajoene, 2-vinyl-4H-1,3-dithiin, diallyl sulfide, and allyl methyl sulfide, have molecular properties allowing them to efficiently interact with the binding site of NSP3 and are currently under rigorous experimental validation in our laboratory.
Supplementary Materials
The following are available online at https://www.mdpi.com/2227-9059/9/11/1582/s1, Figure S1: The bioavailability radar of organosulfur compounds from Allium sativum; Table S1: The chemical structures of the main organosulfur compounds in garlic.
Author Contributions
Conceptualization, B.L., Y.-C.Y., T.-H.H., M.H. and C.-Y.F.H.; methodology, B.L., Y.-C.Y., T.-H.H., M.H. and C.-Y.F.H.; software, B.L., Y.-C.Y., T.-H.H., M.H. and C.-Y.F.H.; validation, Y.-C.Y., T.-H.H., M.H. and C.-Y.F.H.; investigation, B.L., Y.-C.Y., T.-H.H., M.H. and C.-Y.F.H.; resources, Y.-C.Y., T.-H.H. and C.-Y.F.H.; data curation, B.L., Y.-C.Y., T.-H.H. and C.-Y.F.H.; writing—original draft preparation, B.L. and Y.-C.Y.; writing—review and editing, T.-H.H. and C.-Y.F.H.; visualization, B.L., Y.-C.Y.; supervision, M.H., T.-H.H. and C.-Y.F.H.; funding acquisition, Y.-C.Y., T.-H.H. and C.-Y.F.H. All authors have read and agreed to the published version of the manuscript.
Funding
Tse Hung Huang is supported by the following grants: CMRPG2K0031, CMRPG2L0111, CMRPG2J0043, CMRPG2H0362, CORPG2J0042, MOST108-2320-B-182A-010-MY3, and MOST109-2320-B-182A-013-MY2. Chi-Ying F Huang is supported by the ministry of Science and Technology (MOST 109-2327-B-010-005 and MOST 110-2320-B-A49A-541. Yuan-Chieh Yeh is supported by the ministry of Science and Technology (MOST 107-2320-B-182A-019-MY3).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA A Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef]
- Mu, L.; Liu, L.; Niu, R.; Zhao, B.; Shi, J.; Li, Y.; Swanson, M.; Scheider, W.; Su, J.; Chang, S.C.; et al. Indoor air pollution and risk of lung cancer among Chinese female non-smokers. Cancer Causes Control 2013, 24, 439–450. [Google Scholar] [CrossRef]
- Thun, M.; Peto, R.; Boreham, J.; Lopez, A.D. Stages of the cigarette epidemic on entering its second century. Tob. Control 2012, 21, 96–101. [Google Scholar] [CrossRef] [PubMed]
- Leung, A.W.Y.; de Silva, T.; Bally, M.B.; Lockwood, W.W. Synthetic lethality in lung cancer and translation to clinical therapies. Mol. Cancer 2016, 15, 61. [Google Scholar] [CrossRef] [PubMed]
- Blann, A.; Nation, B. British Journal of Biomedical Science in 2015: What have we learned? Taylor Fr. 2016, 4–9. [Google Scholar] [CrossRef]
- Pao, W.; Girard, N. New driver mutations in non-small-cell lung cancer. Lancet Oncol. 2011, 12, 175–180. [Google Scholar] [CrossRef]
- Sullivan, J.P.; Minna, J.D.; Shay, J.W. Evidence for self-renewing lung cancer stem cells and their implications in tumor initiation, progression, and targeted therapy. Cancer Metastasis Rev. 2010, 29, 61–72. [Google Scholar] [CrossRef]
- Chen, Z.; Fillmore, C.M.; Hammerman, P.S.; Kim, C.F.; Wong, K.-K. Non-small-cell lung cancers: A heterogeneous set of diseases. Nat. Rev. Cancer 2014, 14, 535–546. [Google Scholar] [CrossRef] [PubMed]
- David, S.E.; Wallace, A.; Hossein, B.; Andrew, C.C.; Richard, T.C.; Lucian, R.C.; Thomas, A.D.A.; Todd, L.D.; Ramaswamy, G.; Frederic, W.G.; et al. Non–Small Cell Lung Cancer, Version 2.2013. J. Natl. Compr. Cancer Netw. 2013, 11, 645–653. [Google Scholar] [CrossRef]
- Yue, C.; Ma, H.; Zhou, Y. Identification of prognostic gene signature associated with microenvironment of lung adenocarcinoma. PeerJ 2019, 7, e8128. [Google Scholar] [CrossRef] [PubMed]
- Fan, T.; Zhu, M.; Wang, L.; Liu, Y.; Tian, H.; Zheng, Y.; Tan, F.; Sun, N.; Li, C.; He, J. Immune profile of the tumor microenvironment and the identification of a four-gene signature for lung adenocarcinoma. Aging 2021, 13, 2397. [Google Scholar] [CrossRef] [PubMed]
- Bao, X.; Shi, R.; Zhang, K.; Xin, S.; Li, X.; Zhao, Y.; Wang, Y. Immune Landscape of Invasive Ductal Carcinoma Tumor Microenvironment Identifies a Prognostic and Immunotherapeutically Relevant Gene Signature. Front. Oncol. 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Lawal, B.; Lin, L.-C.; Lee, J.-C.; Chen, J.-H.; Bekaii-Saab, T.S.; Wu, A.T.H.; Ho, C.-L. Multi-Omics Data Analysis of Gene Expressions and Alterations, Cancer-Associated Fibroblast and Immune Infiltrations, Reveals the Onco-Immune Prognostic Relevance of STAT3/CDK2/4/6 in Human Malignancies. Cancers 2021, 13, 954. [Google Scholar] [CrossRef]
- Song, Y.; Sun, Y.; Sun, T.; Tang, R. Comprehensive bioinformatics analysis identifies tumor microenvironment and immune-related genes in small cell lung cancer. Comb. Chem. High Throughput Screen. 2020, 23, 381–391. [Google Scholar] [CrossRef]
- Jerby-Arnon, L.; Shah, P.; Cuoco, M.S.; Rodman, C.; Su, M.-J.; Melms, J.C.; Leeson, R.; Kanodia, A.; Mei, S.; Lin, J.-R. A cancer cell program promotes T cell exclusion and resistance to checkpoint blockade. Cell 2018, 175, 984–997.e24. [Google Scholar] [CrossRef]
- Kong, J.; Tian, H.; Zhang, F.; Zhang, Z.; Li, J.; Liu, X.; Li, X.; Liu, J.; Li, X.; Jin, D.; et al. Extracellular vesicles of carcinoma-associated fibroblasts creates a pre-metastatic niche in the lung through activating fibroblasts. Mol. Cancer 2019, 18, 175. [Google Scholar] [CrossRef]
- Li, K.; Kang, H.; Wang, Y.; Hai, T.; Rong, G.; Sun, H. Letrozole-induced functional changes in carcinoma-associated fibroblasts and their influence on breast cancer cell biology. Med. Oncol. 2016, 33, 64. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Zhou, X.; Liu, X.; Jia, H.H.; Zhao, X.H.; Wang, Q.X.; Han, L.; Song, X.; Zhu, Z.Y.; Sun, T.; et al. Reprogramming carcinoma associated fibroblasts by AC1MMYR2 impedes tumor metastasis and improves chemotherapy efficacy. Cancer Lett. 2016, 374, 96–106. [Google Scholar] [CrossRef]
- Dodelet, V.C.; Pazzagli, C.; Zisch, A.H.; Hauser, C.A.; Pasquale, E.B. A novel signaling intermediate, SHEP1, directly couples Eph receptors to R-Ras and Rap1A. J. Biol. Chem. 1999, 274, 31941–31946. [Google Scholar] [CrossRef]
- Sakakibara, A.; Ohba, Y.; Kurokawa, K.; Matsuda, M.; Hattori, S. Novel function of Chat in controlling cell adhesion via Cas-Crk-C3G-pathway-mediated Rap1 activation. J. Cell Sci. 2002, 115, 4915–4924. [Google Scholar] [CrossRef]
- Wang, L.; Vervoort, V.; Wallez, Y.; Coré, N.; Cremer, H.; Pasquale, E.B. The SRC homology 2 domain protein Shep1 plays an important role in the penetration of olfactory sensory axons into the forebrain. J. Neurosci. 2010, 30, 13201–13210. [Google Scholar] [CrossRef]
- Adesina, D.A.; Adefolalu, S.F.; Jigam, A.A.; Lawal, B. Antiplasmodial effect and sub-acute toxicity of alkaloid, flavonoid and phenolic extracts of Sida acuta leaf on Plasmodium berghei-infected animals. J. Taibah Univ. Sci. 2020, 14, 943–953. [Google Scholar] [CrossRef]
- Ibrahim, J.; Kabiru, A.Y.; Abdulrasheed-Adeleke, T.; Lawal, B.; Adewuyi, A.H. Antioxidant and hepatoprotective potentials of curcuminoid isolates from turmeric (Curcuma longa) rhizome on CCl4-induced hepatic damage in Wistar rats. J. Taibah Univ. Sci. 2020, 14, 908–915. [Google Scholar] [CrossRef]
- Lawal, B.; Shittu, O.K.; Oibiokpa, F.I.; Berinyuy, E.B.; Mohammed, H. African natural products with potential antioxidants and hepatoprotectives properties: A review. Clin. Phytoscience 2016, 2, 23. [Google Scholar] [CrossRef]
- Lawal, B.; Shittu, O.K.; Kabiru, A.Y.; Jigam, A.A.; Umar, M.B.; Berinyuy, E.B.; Alozieuwa, B.U. Potential antimalarials from African natural products: A reviw. J. Intercult. Ethnopharmacol. 2015, 4, 318–343. [Google Scholar] [CrossRef]
- Martins, N.; Petropoulos, S.; Ferreira, I.C.F.R. Chemical composition and bioactive compounds of garlic (Allium sativum L.) as affected by pre- and post-harvest conditions: A review. Food Chem. 2016, 211, 41–50. [Google Scholar] [CrossRef]
- Barnes, J.; Anderson, L.A.; Phillipson, J.D. Herbal Medicines: A Guide for Healthcare Professionals; Pharmaceutical Press: London, UK, 2003. [Google Scholar]
- Rahman, K. Historical perspective on garlic and cardiovascular disease. J. Nutr. 2001, 131, 977S–979S. [Google Scholar] [CrossRef]
- El-Saber Batiha, G.; Magdy Beshbishy, A.; Wasef, L.G.; Elewa, Y.H.A.; Al-Sagan, A.A.; Abd El-Hack, M.E.; Taha, A.E.; Abd El-Hack, Y.M.; Devkota, H.P. Chemical Constituents and Pharmacological Activities of Garlic (Allium sativum L.): A Review. Nutrients 2020, 12, 872. [Google Scholar] [CrossRef]
- Tesfaye, A.; Mengesha, W. Traditional uses, phytochemistry and pharmacological properties of garlic (Allium Sativum) and its biological active compounds. Int. J. Sci. Res. Eng. Technol 2015, 1, 142–148. [Google Scholar]
- Al-Snafi, A.E. Pharmacological effects of Allium species grown in Iraq. An overview. Int. J. Pharm. Health Care Res. 2013, 1, 132–147. [Google Scholar]
- Zeng, Y.; Li, Y.; Yang, J.; Pu, X.; Du, J.; Yang, X.; Yang, T.; Yang, S. Therapeutic role of functional components in alliums for preventive chronic disease in human being. Evid.-Based Complementary Altern. Med. 2017, 2017, 9402849. [Google Scholar] [CrossRef]
- Souza, G.A.; Ebaid, G.X.; Seiva, F.R.; Rocha, K.H.; Galhardi, C.M.; Mani, F.; Novelli, E.L. N-acetylcysteine an allium plant compound improves high-sucrose diet-induced obesity and related effects. Evid.-Based Complementary Altern. Med. 2011, 2011, 643269. [Google Scholar] [CrossRef]
- Liu, Y.; Yan, J.; Han, X.; Hu, W. Garlic-derived compound S-allylmercaptocysteine (SAMC) is active against anaplastic thyroid cancer cell line 8305C (HPACC). Technol. Health Care 2015, 23, S89–S93. [Google Scholar] [CrossRef]
- Cao, X.; Cao, L.; Ding, L.; Bian, J.-s. A new hope for a devastating disease: Hydrogen sulfide in Parkinson’s disease. Mol. Neurobiol. 2018, 55, 3789–3799. [Google Scholar] [CrossRef]
- Miron, T.; Rabinkov, A.; Mirelman, D.; Wilchek, M.; Weiner, L. The mode of action of allicin: Its ready permeability through phospholipid membranes may contribute to its biological activity. Biochim. Et Biophys. Acta (BBA)-Biomembr. 2000, 1463, 20–30. [Google Scholar] [CrossRef]
- Borlinghaus, J.; Albrecht, F.; Gruhlke, M.C.; Nwachukwu, I.D.; Slusarenko, A.J. Allicin: Chemistry and biological properties. Molecules 2014, 19, 12591–12618. [Google Scholar] [CrossRef]
- Shimon, L.J.; Rabinkov, A.; Shin, I.; Miron, T.; Mirelman, D.; Wilchek, M.; Frolow, F. Two structures of alliinase from Alliium sativum L.: Apo form and ternary complex with aminoacrylate reaction intermediate covalently bound to the PLP cofactor. J. Mol. Biol. 2007, 366, 611–625. [Google Scholar] [CrossRef]
- Lawal, B.; Liu, Y.-L.; Mokgautsi, N.; Khedkar, H.; Sumitra, M.R.; Wu, A.T.H.; Huang, H.-S. Pharmacoinformatics and Preclinical Studies of NSC765690 and NSC765599, Potential STAT3/CDK2/4/6 Inhibitors with Antitumor Activities against NCI60 Human Tumor Cell Lines. Biomedicines 2021, 9, 92. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-C.; Wu, A.T.H.; Chen, J.-H.; Huang, W.-Y.; Lawal, B.; Mokgautsi, N.; Huang, H.-S.; Ho, C.-L. HNC0014, a Multi-Targeted Small-Molecule, Inhibits Head and Neck Squamous Cell Carcinoma by Suppressing c-Met/STAT3/CD44/PD-L1 Oncoimmune Signature and Eliciting Antitumor Immune Responses. Cancers 2020, 12, 3759. [Google Scholar] [CrossRef]
- Oshevire, D.B.; Mustapha, A.; Alozieuwa, B.U.; Badeggi, H.H.; Ismail, A.; Hassan, O.N.; Ugwunnaji, P.I.; Ibrahim, J.; Lawal, B.; Berinyu, E.B. In-silico investigation of curcumin drug-likeness, gene-targets and prognostic relevance of the targets in panels of human cancer cohorts. GSC Biol. Pharm. Sci. 2021, 14. [Google Scholar] [CrossRef]
- Li, T.; Fu, J.; Zeng, Z.; Cohen, D.; Li, J.; Chen, Q.; Li, B.; Liu, X.S. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 2020, 48, W509–W514. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Li, C.; Kang, B.; Gao, G.; Li, C.; Zhang, Z. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017, 45, W98–W102. [Google Scholar] [CrossRef] [PubMed]
- Bartha, Á.; Győrffy, B. TNMplot.com: A Web Tool for the Comparison of Gene Expression in Normal, Tumor and Metastatic Tissues. Int. J. Mol. Sci. 2021, 22, 2622. [Google Scholar] [CrossRef]
- Győrffy, B.; Surowiak, P.; Budczies, J.; Lánczky, A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS ONE 2013, 8, e82241. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013, 6, 2347–2348. [Google Scholar] [CrossRef]
- Jiang, P.; Gu, S.; Pan, D.; Fu, J.; Sahu, A.; Hu, X.; Li, Z.; Traugh, N.; Bu, X.; Li, B.; et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 2018, 24, 1550–1558. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Chen, E.Y.; Tan, C.M.; Kou, Y.; Duan, Q.; Wang, Z.; Meirelles, G.V.; Clark, N.R.; Ma’ayan, A. Enrichr: Interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 2013, 14, 128. [Google Scholar] [CrossRef]
- Kuleshov, M.V.; Jones, M.R.; Rouillard, A.D.; Fernandez, N.F.; Duan, Q.; Wang, Z.; Koplev, S.; Jenkins, S.L.; Jagodnik, K.M.; Lachmann, A.; et al. Enrichr: A comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016, 44, W90–W97. [Google Scholar] [CrossRef] [PubMed]
- Mostafavi, S.; Ray, D.; Warde-Farley, D.; Grouios, C.; Morris, Q. GeneMANIA: A real-time multiple association network integration algorithm for predicting gene function. Genome Biol. 2008, 9, S4. [Google Scholar] [CrossRef] [PubMed]
- Carvalho-Silva, D.; Pierleoni, A.; Pignatelli, M.; Ong, C.; Fumis, L.; Karamanis, N.; Carmona, M.; Faulconbridge, A.; Hercules, A.; McAuley, E.; et al. Open Targets Platform: New developments and updates two years on. Nucleic Acids Res. 2019, 47, D1056–D1065. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, L.; Lv, M.; Pei, R.; Li, P.; Pei, Z.; Wang, Y.; Su, W.; Xie, X.-Q. AlzPlatform: An Alzheimer’s Disease Domain-Specific Chemogenomics Knowledgebase for Polypharmacology and Target Identification Research. J. Chem. Inf. Modeling 2014, 54, 1050–1060. [Google Scholar] [CrossRef] [PubMed]
- Poroikov, V.V.; Filimonov, D.A.; Gloriozova, T.A.; Lagunin, A.A.; Druzhilovskiy, D.S.; Rudik, A.V.; Stolbov, L.A.; Dmitriev, A.V.; Tarasova, O.A.; Ivanov, S.M.; et al. Computer-aided prediction of biological activity spectra for organic compounds: The possibilities and limitations. Russ. Chem. Bull. 2019, 68, 2143–2154. [Google Scholar] [CrossRef]
- Lagunin, A.; Zakharov, A.; Filimonov, D.; Poroikov, V. QSAR Modelling of Rat Acute Toxicity on the Basis of PASS Prediction. Mol. Inform. 2011, 30, 241–250. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Lawal, B.; Lee, C.-Y.; Mokgautsi, N.; Sumitra, M.R.; Khedkar, H.; Wu, A.T.H.; Huang, H.-S. mTOR/EGFR/iNOS/MAP2K1/FGFR/TGFB1 Are Druggable Candidates for N-(2,4-Difluorophenyl)-2′,4′-Difluoro-4-Hydroxybiphenyl-3-Carboxamide (NSC765598), With Consequent Anticancer Implications. Front. Oncol. 2021, 11. [Google Scholar] [CrossRef]
- Wu, A.T.H.; Lawal, B.; Wei, L.; Wen, Y.-T.; Tzeng, D.T.W.; Lo, W.-C. Multiomics Identification of Potential Targets for Alzheimer Disease and Antrocin as a Therapeutic Candidate. Pharmaceutics 2021, 13, 1555. [Google Scholar] [CrossRef]
- Lawal, B.; Kuo, Y.-C.; Tang, S.-L.; Liu, F.-C.; Wu, A.T.H.; Lin, H.-Y.; Huang, H.-S. Transcriptomic-Based Identification of the Immuno-Oncogenic Signature of Cholangiocarcinoma for HLC-018 Multi-Target Therapy Exploration. Cells 2021, 10, 2873. [Google Scholar] [CrossRef]
- Lawal, B.; Kuo, Y.-C.; Sumitra, M.R.; Wu, A.T.; Huang, H.-S. In vivo Pharmacokinetic and Anticancer Studies of HH-N25, a Selective Inhibitor of Topoisomerase I, and Hormonal Signaling for Treating Breast Cancer. J. Inflamm. Res. 2021, 14, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Visualizer, I.T.S. BIOVIA, Dassault Systèmes, BIOVIA Workbook, Release 2020; BIOVIA Pipeline Pilot, Release 2020, San Diego: Dassault Systèmes. 2020. Available online: https://biovianotebook.com/products/collaborative-science/biovia-discovery-studio/ (accessed on 23 August 2021).
- Sartor, I.T.S.; Recamonde-Mendoza, M.; Ashton-Prolla, P. TULP3: A potential biomarker in colorectal cancer? PLoS ONE 2019, 14, e0210762. [Google Scholar] [CrossRef]
- Cristescu, R.; Lee, J.; Nebozhyn, M.; Kim, K.M.; Ting, J.C.; Wong, S.S.; Liu, J.; Yue, Y.G.; Wang, J.; Yu, K.; et al. Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat. Med. 2015, 21, 449–456. [Google Scholar] [CrossRef]
- Davalos, V.; Dopeso, H.; Castaño, J.; Wilson, A.J.; Vilardell, F.; Romero-Gimenez, J.; Espín, E.; Armengol, M.; Capella, G.; Mariadason, J.M.; et al. EPHB4 and survival of colorectal cancer patients. Cancer Res. 2006, 66, 8943–8948. [Google Scholar] [CrossRef]
- Noren, N.K.; Foos, G.; Hauser, C.A.; Pasquale, E.B. The EphB4 receptor suppresses breast cancer cell tumorigenicity through an Abl-Crk pathway. Nat. Cell. Biol. 2006, 8, 815–825. [Google Scholar] [CrossRef]
- Dopeso, H.; Mateo-Lozano, S.; Mazzolini, R.; Rodrigues, P.; Lagares-Tena, L.; Ceron, J.; Romero, J.; Esteves, M.; Landolfi, S.; Hernández-Losa, J.; et al. The receptor tyrosine kinase EPHB4 has tumor suppressor activities in intestinal tumorigenesis. Cancer Res. 2009, 69, 7430–7438. [Google Scholar] [CrossRef]
- Chen, J.-H.; Wu, A.T.H.; Lawal, B.; Tzeng, D.T.W.; Lee, J.-C.; Ho, C.-L.; Chao, T.-Y. Identification of Cancer Hub Gene Signatures Associated with Immune-Suppressive Tumor Microenvironment and Ovatodiolide as a Potential Cancer Immunotherapeutic Agent. Cancers 2021, 13, 3847. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.-Y.; Lin, K.-C.; Lawal, B.; Wu, A.T.H.; Wu, C.-Z. MXD3 as an onco-immunological biomarker encompassing the tumor microenvironment, disease staging, prognoses, and therapeutic responses in multiple cancer types. Comput. Struct. Biotechnol. J. 2021, 19, 4970–4983. [Google Scholar] [CrossRef] [PubMed]
- Ino, Y.; Yamazaki-Itoh, R.; Shimada, K.; Iwasaki, M.; Kosuge, T.; Kanai, Y.; Hiraoka, N. Immune cell infiltration as an indicator of the immune microenvironment of pancreatic cancer. Br. J. Cancer 2013, 108, 914–923. [Google Scholar] [CrossRef]
- Hiraoka, N.; Yamazaki–Itoh, R.; Ino, Y.; Mizuguchi, Y.; Yamada, T.; Hirohashi, S.; Kanai, Y. CXCL17 and ICAM2 Are Associated with a Potential Anti-Tumor Immune Response in Early Intraepithelial Stages of Human Pancreatic Carcinogenesis. Gastroenterology 2011, 140, 310–321.e4. [Google Scholar] [CrossRef] [PubMed]
- Spranger, S.; Gajewski, T.F. Tumor-intrinsic oncogene pathways mediating immune avoidance. Oncoimmunology 2016, 5, e1086862. [Google Scholar] [CrossRef] [PubMed]
- Gajewski, T.F.; Schreiber, H.; Fu, Y.-X. Innate and adaptive immune cells in the tumor microenvironment. Nat. Immunol. 2013, 14, 1014–1022. [Google Scholar] [CrossRef]
- Kaderbhaï, C.; Tharin, Z.; Ghiringhelli, F. The Role of Molecular Profiling to Predict the Response to Immune Checkpoint Inhibitors in Lung Cancer. Cancers 2019, 11, 201. [Google Scholar] [CrossRef] [PubMed]
- Giannoni, E.; Bianchini, F.; Masieri, L.; Serni, S.; Torre, E.; Calorini, L.; Chiarugi, P. Reciprocal Activation of Prostate Cancer Cells and Cancer-Associated Fibroblasts Stimulates Epithelial-Mesenchymal Transition and Cancer Stemness. Cancer Res. 2010, 70, 6945–6956. [Google Scholar] [CrossRef]
- Erez, N.; Truitt, M.; Olson, P.; Arron, S.T.; Hanahan, D. Cancer-Associated Fibroblasts Are Activated in Incipient Neoplasia to Orchestrate Tumor-Promoting Inflammation in an NF-kappaB-Dependent Manner. Cancer Cell 2010, 17, 135–147. [Google Scholar] [CrossRef] [PubMed]
- Lipinski, C.A. Drug-like properties and the causes of poor solubility and poor permeability. J. Pharmacol. Toxicol. Methods 2000, 44, 235–249. [Google Scholar] [CrossRef]
- Matlock, M.K.; Hughes, T.B.; Dahlin, J.L.; Swamidass, S.J. Modeling small-molecule reactivity identifies promiscuous bioactive compounds. J. Chem. Inf. Modeling 2018, 58, 1483–1500. [Google Scholar] [CrossRef] [PubMed]
- Lawal, B.; Shittu, O.K.; Oibiokpa, F.I.; Mohammed, H.; Umar, S.I.; Haruna, G.M. Antimicrobial evaluation, acute and sub-acute toxicity studies of Allium sativum. J. Acute Dis. 2016, 5, 296–301. [Google Scholar] [CrossRef]
- Shittu, O.K.; Lawal, B.; Alozieuwa, B.U.; Haruna, G.M.; Abubakar, A.N.; Berinyuy, E.B. Alteration in biochemical indices following chronic administration of methanolic extract of Nigeria bee propolis in Wistar rats. Asian Pac. J. Trop. Dis. 2015, 5, 654–657. [Google Scholar] [CrossRef]
- Yusuf, A.A.; Lawal, B.; Yusuf, M.A.; Adejoke, A.O.; Raji, F.H.; Wenawo, D.L. Free radical scavenging, antimicrobial activities and effect of sub-acute exposure to Nigerian Xylopia Aethiopica seed extract on liver and kidney functional indices of albino rat. Iran. J. Toxicol. 2018, 12, 51–58. [Google Scholar] [CrossRef]
- Kitchen, D.B.; Decornez, H.; Furr, J.R.; Bajorath, J. Docking and scoring in virtual screening for drug discovery: Methods and applications. Nat. Rev. Drug Discov. 2004, 3, 935–949. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.-Y.; Zhang, H.-X.; Mezei, M.; Cui, M. Molecular docking: A powerful approach for structure-based drug discovery. Curr. Comput. Aided Drug Des. 2011, 7, 146–157. [Google Scholar] [CrossRef]
- Jorgensen, W.L. The many roles of computation in drug discovery. Science 2004, 303, 1813–1818. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Huang, D. Hydrogen bonding penalty upon ligand binding. PLoS ONE 2011, 6, e19923. [Google Scholar] [CrossRef] [PubMed]
- Arthur, D.E.; Uzairu, A. Molecular docking studies on the interaction of NCI anticancer analogues with human Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit. J. King Saud Univ.—Sci. 2019, 31, 1151–1166. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).