Curcumin Nanoparticles and Their Cytotoxicity in Docetaxel-Resistant Castration-Resistant Prostate Cancer Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals and Reagents
2.2. HPLC Analysis for Curcumin
2.3. Screening of Oils and Surfactants
2.4. Preparation of Curcumin Nanoparticles
2.5. Solubility of Curcumin in Water and the Cell Medium
2.6. Particle Size and Zeta Potential Measurement
2.7. Morphology of CUR MT NPs
2.8. Determination of Drug Loading and Entrapment Efficiency
2.9. Particle Size Stability of Nanoparticles at 4 °C and 37 °C
2.10. Differential Scanning Calorimetry
2.11. In Vitro Release Study
2.12. Lyophilization of CUR MT NPs
2.13. Cytotoxicity Studies
2.14. Statistical Analysis
3. Results
3.1. Selection of Oils and Surfactants and Preparation of CUR MT NPs
3.2. Solubility
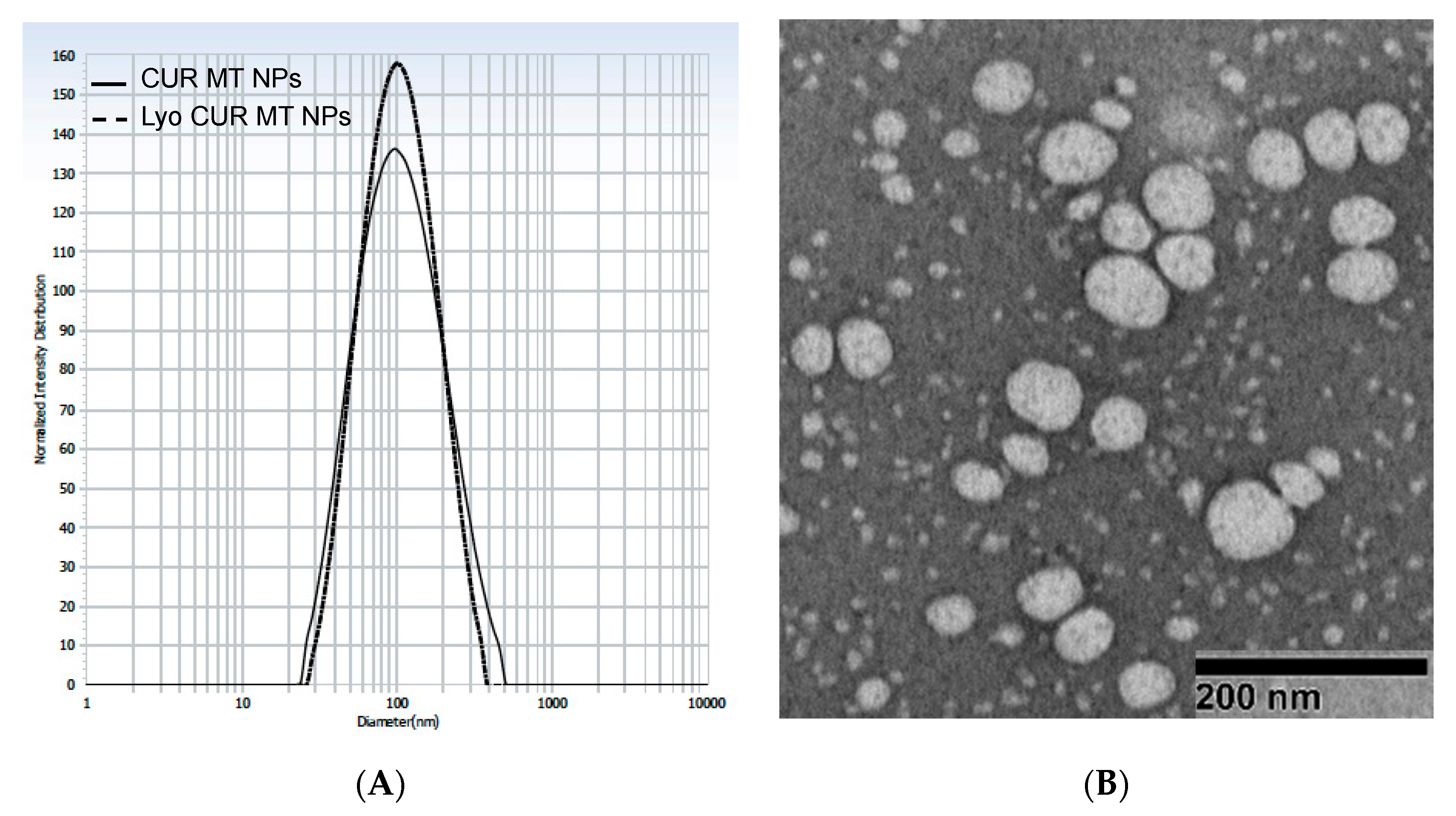
3.3. Particle Size and Entrapment Efficiency
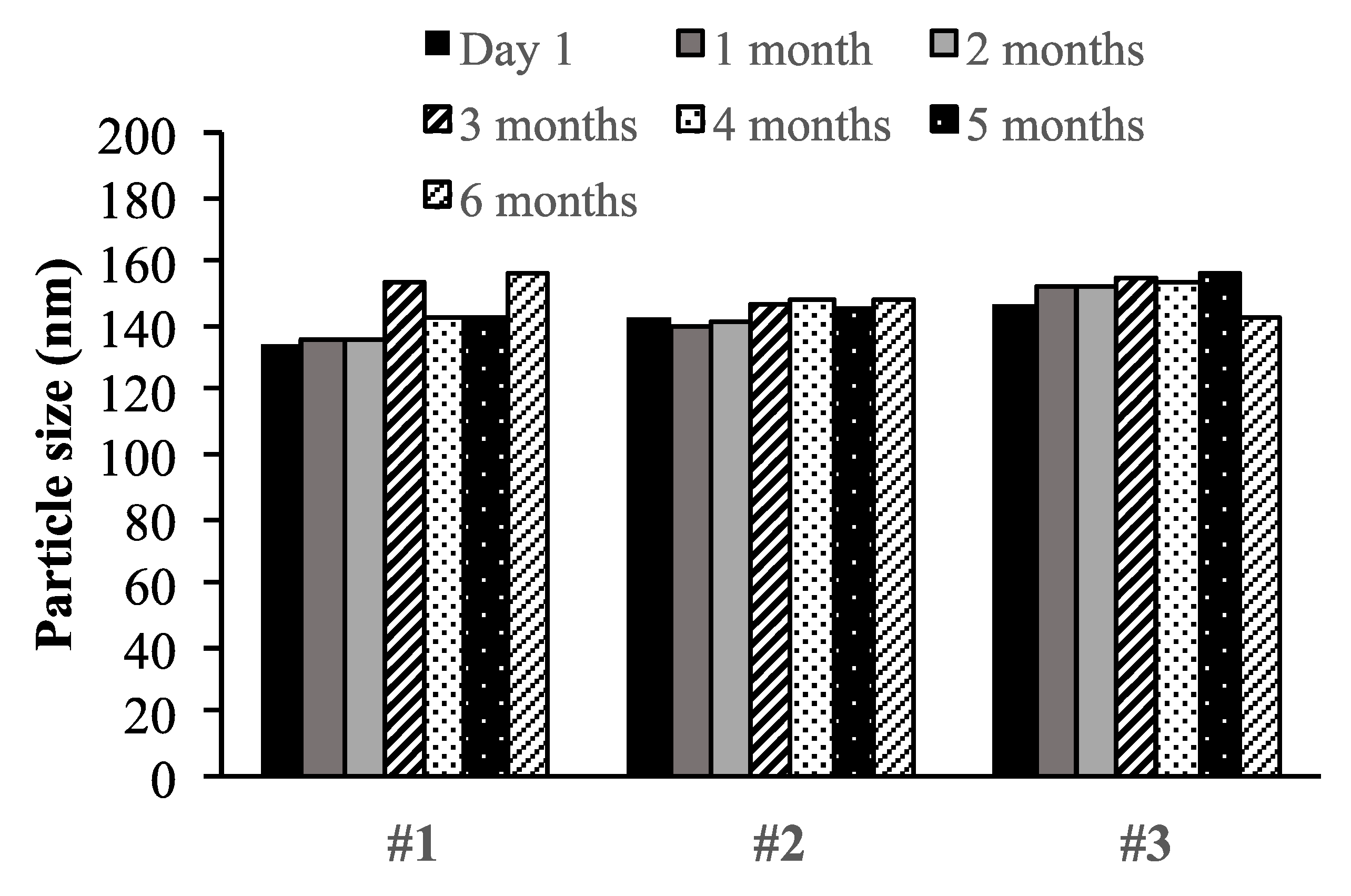
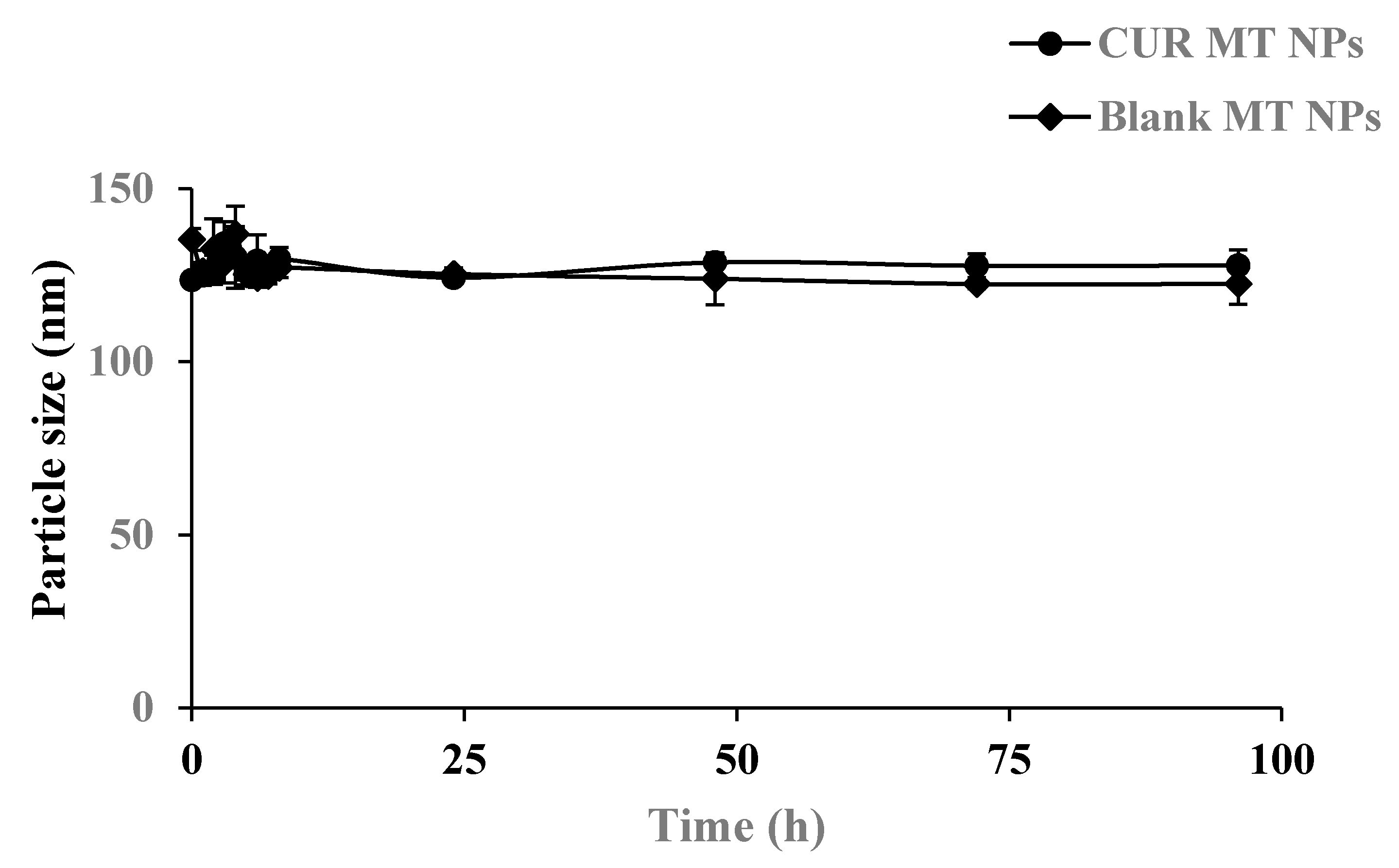
3.4. Physical Stability
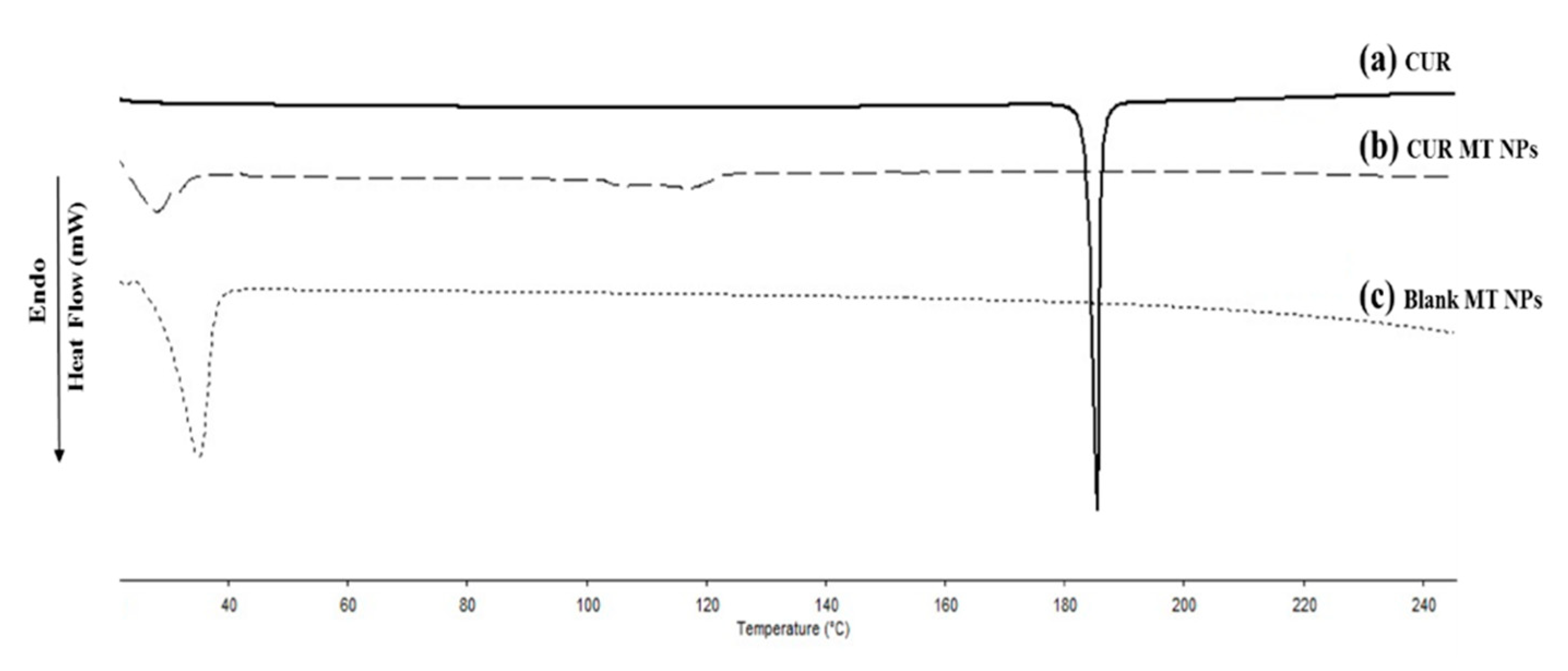
3.5. Differential Scanning Calorimetry Analysis
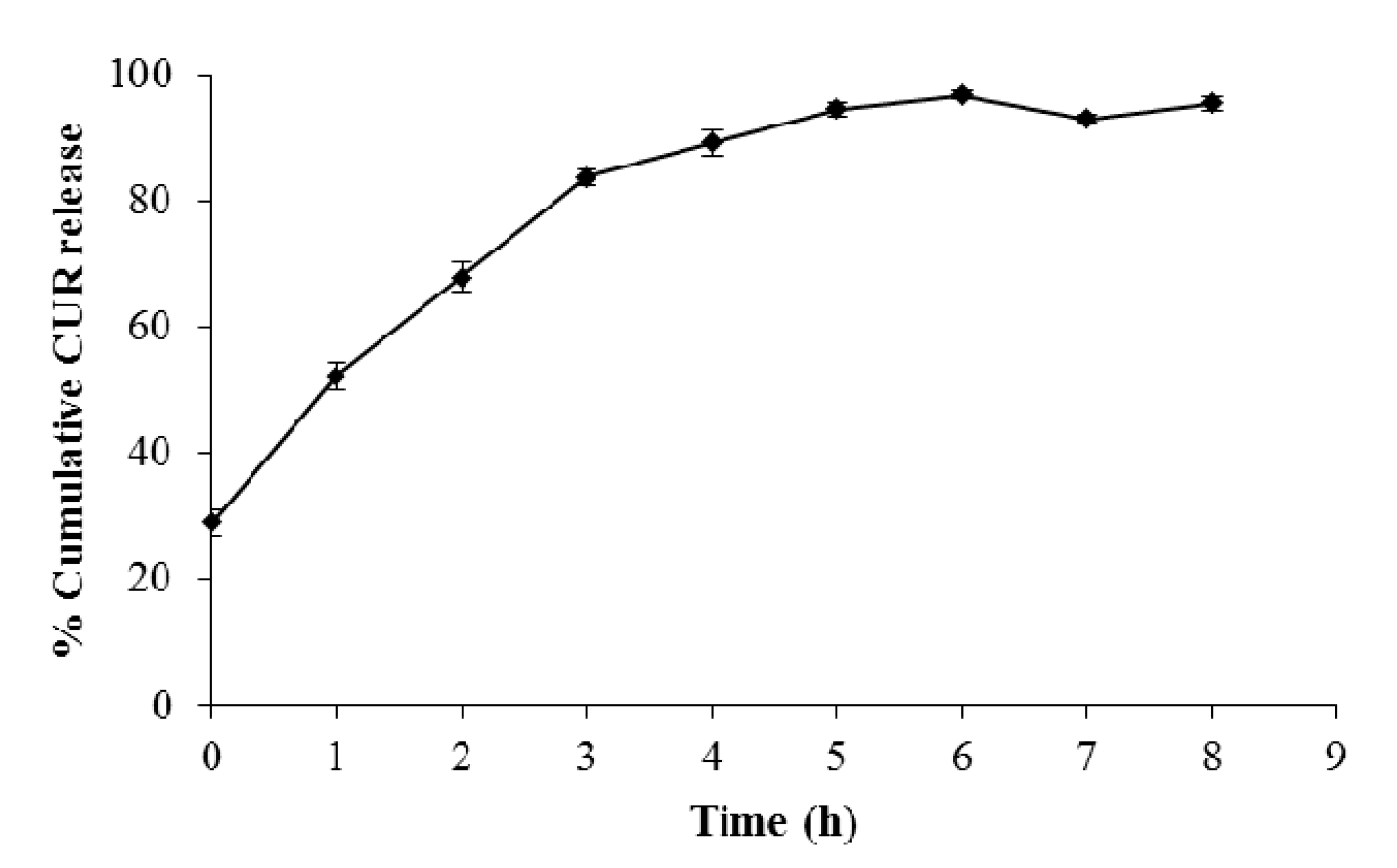
3.6. In Vitro Release of Curcumin from the Nanoparticles
3.7. Lyophilization
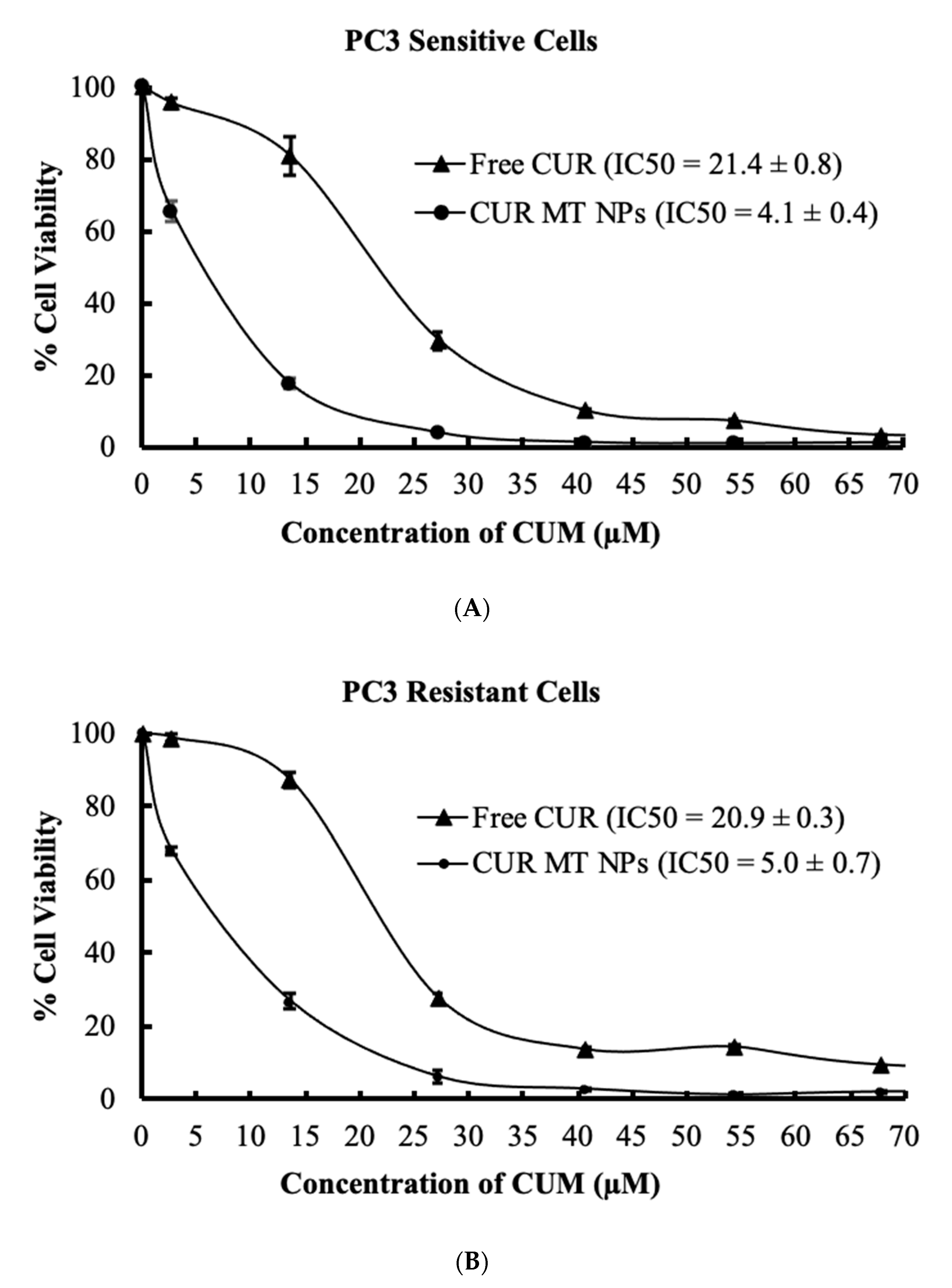
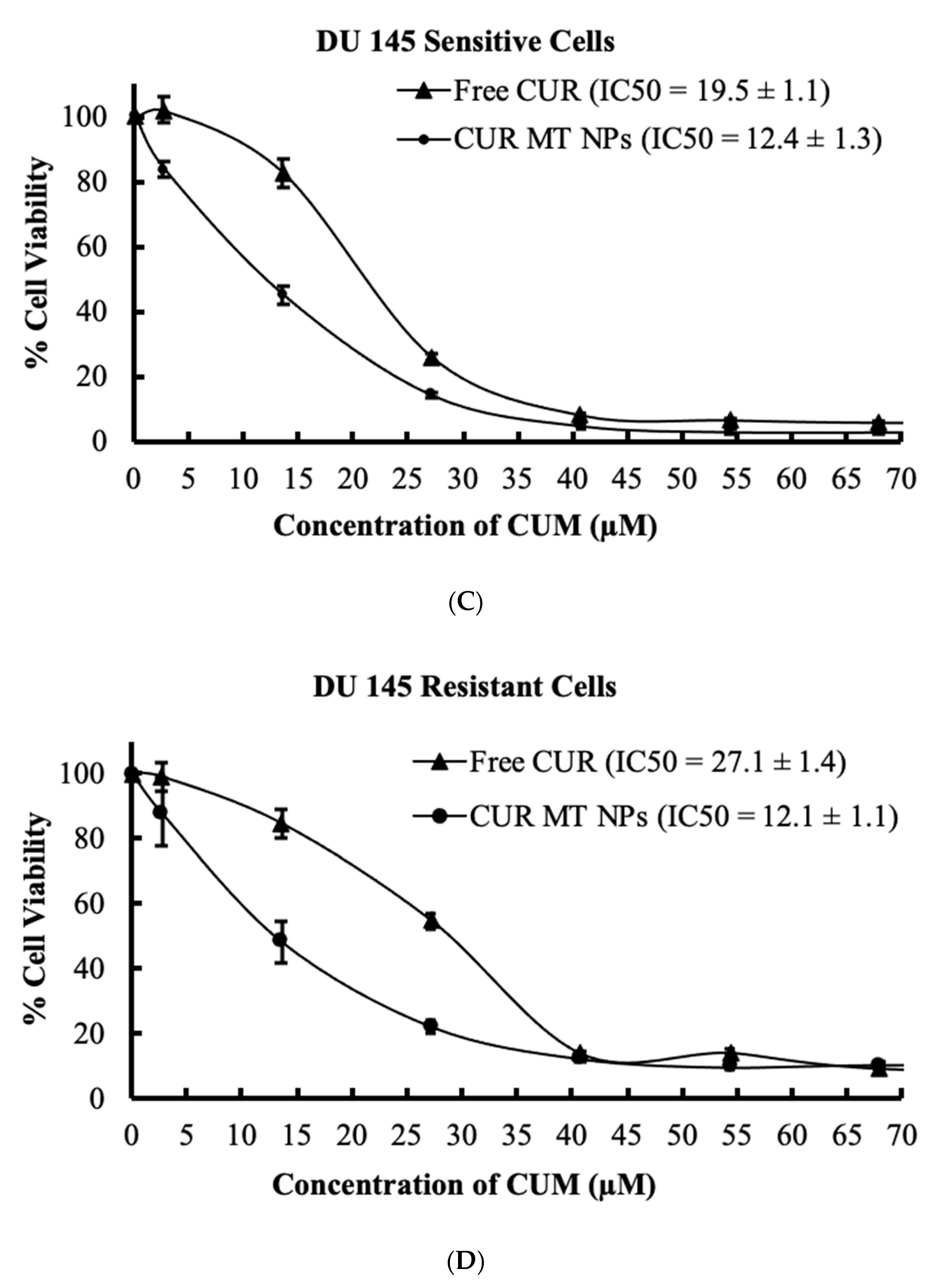
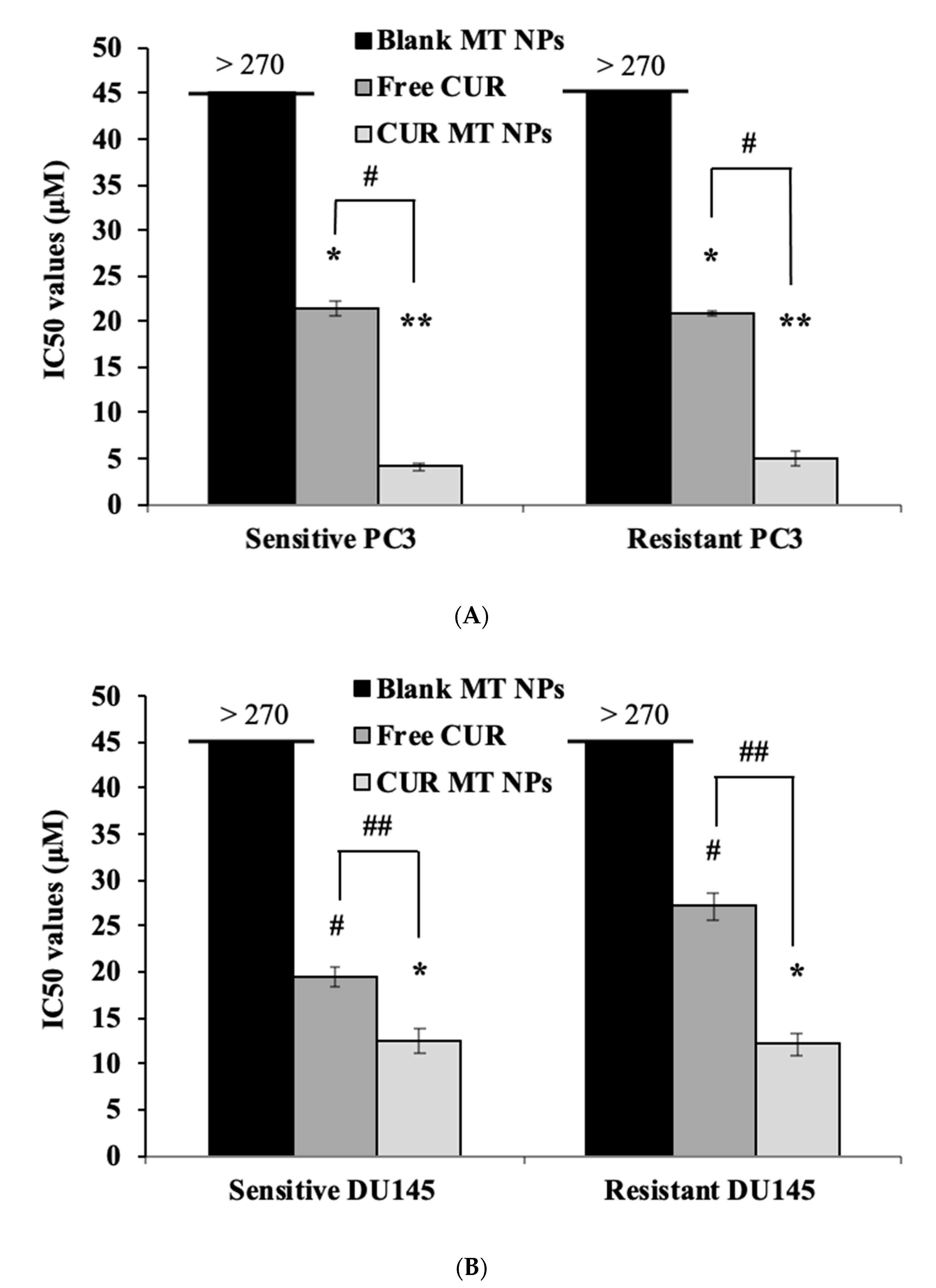
3.8. In Vitro Cytotoxicity Studies
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomatraram, I.; Siegel, R.L.; Torr, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Karantanos, T.; Corn, P.G.; Thompson, T.C. Prostate cancer progression after androgen deprivation therapy; mechanisms of castrate resistance and novel therapeutic approaches. Oncogene 2013, 32, 5501–5511. [Google Scholar] [CrossRef] [PubMed]
- Tannock, I.F.; De Wit, R.; Berry, W.R.; Horti, J.; Pluzanska, A.; Chi, K.N.; Oudard, S.; Theodore, C.; James, N.D.; Turesson, I.; et al. Docetaxel plus prednisone or mitoxantrone plus prednisone for advanced prostate cancer. N. Engl. J. Med. 2004, 351, 1502–1512. [Google Scholar] [CrossRef] [PubMed]
- Brooks, T.; Minderman, H.; O’Loughlin, K.L.; Pera, P.; Ojima, I.; Baer, M.R.; Bernacki, R.J. Taxane-based reversal agents modulate drug resistance mediated by P-glycoprotein, multidrug resistance protein, and breast cancer resistance protein. Mol. Cancer Ther. 2003, 2, 1195–1205. [Google Scholar]
- Goldman, B. Multidrug resistance: Can new drugs help chemotherapy score against cancer? J. Natl. Cancer Inst. 2003, 95, 255–257. [Google Scholar] [CrossRef]
- Thomas, H.; Coley, H.M. Overcoming multidrug resistance in cancer: An update on the clinical strategy of inhibiting p-glycoprotein. Cancer Control. 2003, 10, 159–165. [Google Scholar] [CrossRef]
- Takeda, M.; Mizokami, A.; Mamiya, K.; Li, Y.Q.; Zhang, J.; Keller, E.T.; Namiki, M. The establishment of two paclitaxel-resistant prostate cancer cell lines and the mechanisms of paclitaxel resistance with two cell lines. Prostate 2007, 67, 955–967. [Google Scholar] [CrossRef]
- Tiwari, A.K.; Sodani, K.; Dai, C.L.; Ashby, C.R., Jr.; Chen, Z.S. Revisiting the ABCs of multidrug resistance in cancer chemotherapy. Curr. Pharm. Biotechnol. 2011, 12, 570–594. [Google Scholar] [CrossRef]
- Aggarwal, B.B.; Shishodia, S. Molecular targets of dietary agents for prevention and therapy of cancer. Biochem. Pharmacol. 2006, 71, 1397–1421. [Google Scholar] [CrossRef]
- Shishodia, S.; Chaturvedi, M.M.; Aggarwal, B.B. Role of curcumin in cancer therapy. Curr. Prob. Cancer 2007, 31, 243–305. [Google Scholar] [CrossRef]
- Wilken, R.; Wang, V.M.; Srivatsan, E. Curcumin: A review of anti-cancer properties and therapeutic activity in head and neck squamous cell carcinoma. Mol. Cancer 2011, 10, 12. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Yu, X.; Qi, R.; Shang, F.; Su, Z. Inhibitory Effects of curcumin in combination with paclitaxel on prostate cancer xenografted model. Progress Mod. Biomed. 2010, 10, 823–827. [Google Scholar]
- Wang, Y.J.; Pan, M.H.; Cheng, A.L.; Lin, L.; Ho, Y.S.; Lin, J.K. Stability of curcumin in buffer solutions and characterization of its degradation products. J. Pharm. Biomed. Anal. 1997, 15, 1867–1876. [Google Scholar] [CrossRef]
- Bar-Sela, G.; Epelbaum, R.; Schaffer, M. Curcumin as an anti-cancer agent: Review of the gap between basic and clinical applications. Curr. Med. Chem. 2010, 17, 190–197. [Google Scholar] [CrossRef]
- Shoba, G.; Joy, D.; Joseph, T.; Majeed, M.; Rajendran, R.; Srinivas, P.S. Influence of piperine on the pharmacokinetics of curcumin in animals and human volunteers. Planta Med. 1998, 64, 353–356. [Google Scholar] [CrossRef]
- De Jong, W.H.; Borm, P.J. Drug delivery and nanoparticles: Applications and hazards. Int. J. Nanomed. 2013, 3, 133–149. [Google Scholar] [CrossRef]
- Markman, J.L.; Rekechenetskiy, A.; Holler, E.; Ljubimova, J.Y. Nanomedicine therapeutic approaches to overcome cancer drug resistance. Adv. Drug Deliv. Rev. 2013, 65, 1866–1879. [Google Scholar] [CrossRef]
- Grama, C.; Ankola, D.; Kumar, M. Poly (lactide-co-glycolide) nanoparticles for peroral delivery of bioactives. Curr. Opin. Colloid Interface Sci. 2011, 16, 238–245. [Google Scholar] [CrossRef]
- Collnot, E.M.; Baldes, C.; Schaefer, U.F.; Edgar, K.J.; Wempe, M.F.; Lehr, C.M. Vitamin E-TPGS p-glycoprotein inhibition mechanism: Influence on conformational flexibility, intracellular ATP levels, and role of time and site of access. Mol. Pharmaceut. 2010, 7, 642–651. [Google Scholar] [CrossRef]
- Tanaudommongkon, A.; Tanaudommongkon, I.; Prathipati, P.; Nguyen, J.T.; Dong, X. Development and Characterization of Docetaxel-Loaded Nanoparticles for Docetaxel-Resistant Castration-Resistant Prostate Cancer. J. Nanosci. Nanotech. 2017, 17, 3920–3926. [Google Scholar] [CrossRef]
- Dong, X.; Mattingly, C.A.; Tseng, M.; Cho, M.; Adams, V.; Mumper, R.J. Development of new lipid-based paclitaxel nanoparticles using sequential simplex optimization. Eur. J. Pharm. Biopharm. 2008, 72, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Neves, A.R.; Lucio, M.; Martins, S.; Lima, J.L.; Reis, S. Novel resveratrol nanodelivery systems based on lipid nanoparticles to enhance its oral bioavailability. Int. J. Nanomed. 2013, 8, 177–187. [Google Scholar] [CrossRef]
- Huang, L.; Liu, Y. In vivo delivery of RNAi with lipid-based nanoparticles. Annu. Rev. Biomed. Eng. 2011, 13, 507–530. [Google Scholar] [CrossRef] [PubMed]
- Maheshwari, R.K.; Singh, A.K.; Gaddipati, J.; Srimal, R.C. Multiple biological activities of curcumin: A short review. Life Sci. 2006, 78, 2081–2087. [Google Scholar] [CrossRef] [PubMed]
- Jurenka, J.S. Anti-inflammatory properties of curcumin, a major constituent of curcuma longa: A review of preclinical and clinical research. Altern. Med. Rev. 2009, 14, 141–153. [Google Scholar] [PubMed]
- Banerjee, S.; Singh, S.K.; Chowdhury, I.; Lillard, J.W.; Singh, R. Combinatorial effect of curcumin with docetaxel modulates apoptotic and cell survival molecules in prostate cancer. Front. Biosci. 2017, 9, 235–245. [Google Scholar] [CrossRef]
- Dong, X.; Mattingly, C.; Tseng, M.T.; Cho, M.J.; Liu, Y.; Adams, V.R.; Mumper, R.J. Doxorubicin and paclitaxel-loaded lipid-based nanoparticles overcome multidrug resistance by inhibiting p-glycoprotein and depleting ATP. Cancer Res. 2009, 69, 3918–3926. [Google Scholar] [CrossRef]
- Xue, X.; Liang, X.-J. Overcoming drug efflux-based multidrug resistance in cancer with nanotechnology. Chi. J. Cancer 2012, 31, 100–109. [Google Scholar] [CrossRef]
- Niazi, M.; Zakeri-Milani, P.; Najafi Hajivar, S.; Soleymani Goloujeh, M.; Ghobakhlou, N.; Shahbazi Mojarrad, J.; Valizadeh, H. Nano-based strategies to overcome p-glycoprotein-mediated drug resistance. Expert Opin. Drug Metab. Toxicol. 2016, 12, 1021–1033. [Google Scholar] [CrossRef]
| Oil | Solubility (w/w) | Surfactant | Solubility (w/w) |
|---|---|---|---|
| Miglyol 812 | 0.0153 | Tween 20 | 0.166 |
| Imwitor 491 | 0.0261 | Tween 80 | 0.141 |
| Kollisolv MCT 70 | 0.0179 | Labrasol | 0.173 |
| Labrafac | 0.0120 | Poloxamer 188 | 0.153 |
| Imwitor 900K | 0.0512 | TPGS | 0.164 |
| Imwitor 960 | 0.0488 | Gelucire 44/14 | 0.153 |
| Compritol 888 | 0.0160 |
| Formulations | CUR (µg/mL) | Particle size a (nm) * | P.I. * | Zeta Potential (mV) * | EE% * | DL% * |
|---|---|---|---|---|---|---|
| CUR MT NPs | 150 | 138.7 ± 5.4 | 0.268 ± 0.033 | −24.4 ± 3.3 | 96.3 ± 6.0 | 3.6 |
| Blank MT NPs | n/a | 119.9 ± 3.5 | 0.246 ± 0.016 | −20.6 ± 1.9 | n/a | n/a |
| Lyo CUR MT NPs | 150 | 101.3 ± 11.1 | 0.268 ± 0.033 | −16.6 ± 3.7 | 93.1 ± 5.8 | 3.5 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tanaudommongkon, I.; Tanaudommongkon, A.; Prathipati, P.; Nguyen, J.T.; Keller, E.T.; Dong, X. Curcumin Nanoparticles and Their Cytotoxicity in Docetaxel-Resistant Castration-Resistant Prostate Cancer Cells. Biomedicines 2020, 8, 253. https://doi.org/10.3390/biomedicines8080253
Tanaudommongkon I, Tanaudommongkon A, Prathipati P, Nguyen JT, Keller ET, Dong X. Curcumin Nanoparticles and Their Cytotoxicity in Docetaxel-Resistant Castration-Resistant Prostate Cancer Cells. Biomedicines. 2020; 8(8):253. https://doi.org/10.3390/biomedicines8080253
Chicago/Turabian StyleTanaudommongkon, Irin, Asama Tanaudommongkon, Priyanka Prathipati, Joey Trieu Nguyen, Evan T. Keller, and Xiaowei Dong. 2020. "Curcumin Nanoparticles and Their Cytotoxicity in Docetaxel-Resistant Castration-Resistant Prostate Cancer Cells" Biomedicines 8, no. 8: 253. https://doi.org/10.3390/biomedicines8080253
APA StyleTanaudommongkon, I., Tanaudommongkon, A., Prathipati, P., Nguyen, J. T., Keller, E. T., & Dong, X. (2020). Curcumin Nanoparticles and Their Cytotoxicity in Docetaxel-Resistant Castration-Resistant Prostate Cancer Cells. Biomedicines, 8(8), 253. https://doi.org/10.3390/biomedicines8080253






