Effect of Iron Oxide Nanoparticles and Amoxicillin on Bacterial Growth in the Presence of Dissolved Organic Carbon
Abstract
1. Introduction
2. Experimental Section
2.1. Materials
2.2. Iron Oxide Nanoparticle (IONP) Synthesis
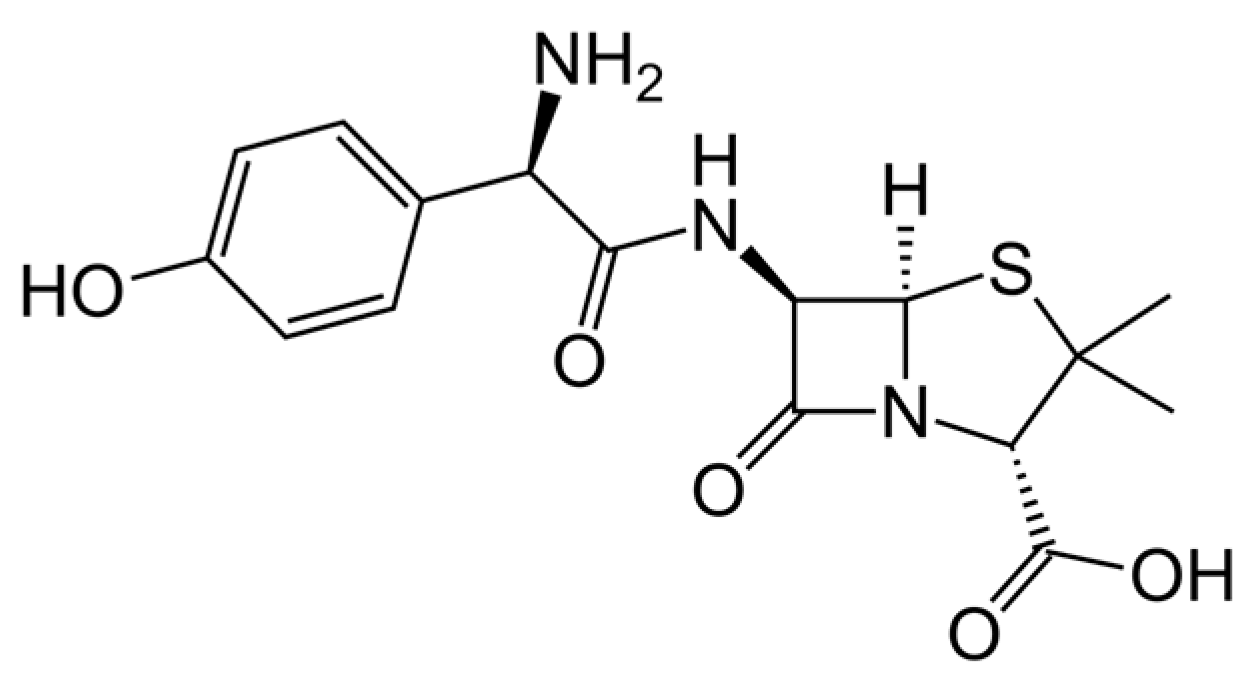
2.3. Amoxicillin Functionalized Iron Oxide Nanoparticle (IONP-Amox) Synthesis
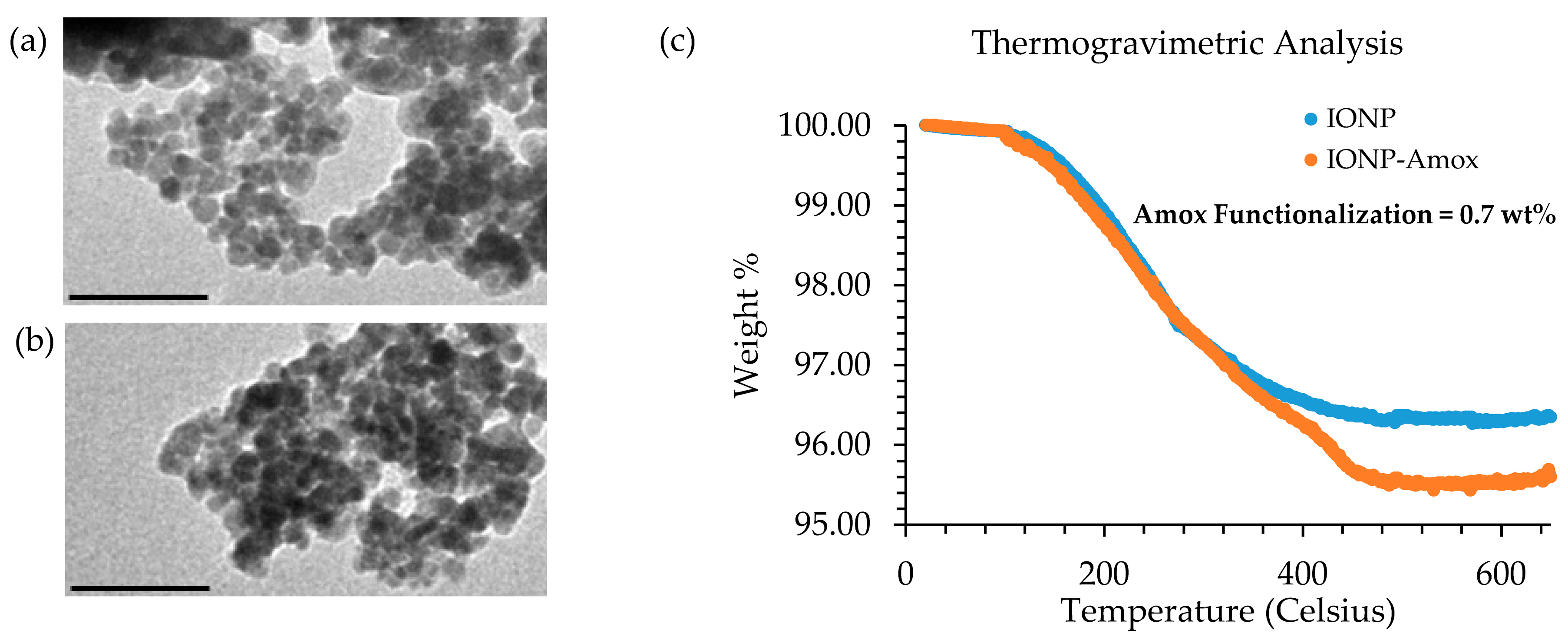
2.4. Nanoparticle Characterization: Transmission Electron Microscopy (TEM) & Thermogravimetric (TG) Analysis
2.5. Bacterial Culture Conditions
2.6. OD600 Time Series Study Outline
2.7. OD600 to CFU/mL Calibration Plot
2.8. Statistical Analysis
3. Results and Discussion
3.1. TG Analysis and TEM Comparison of IONP and IONP-Amox
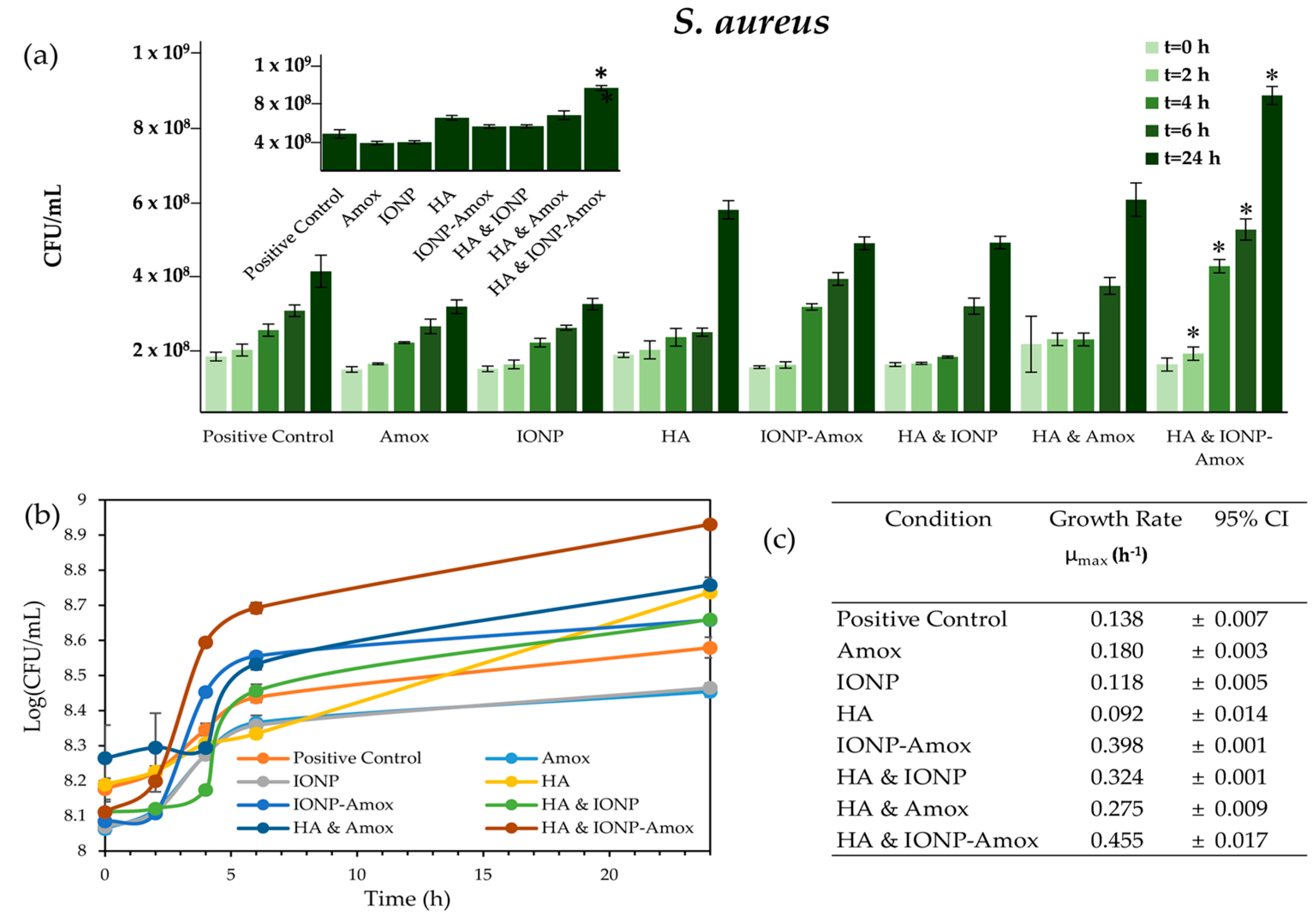
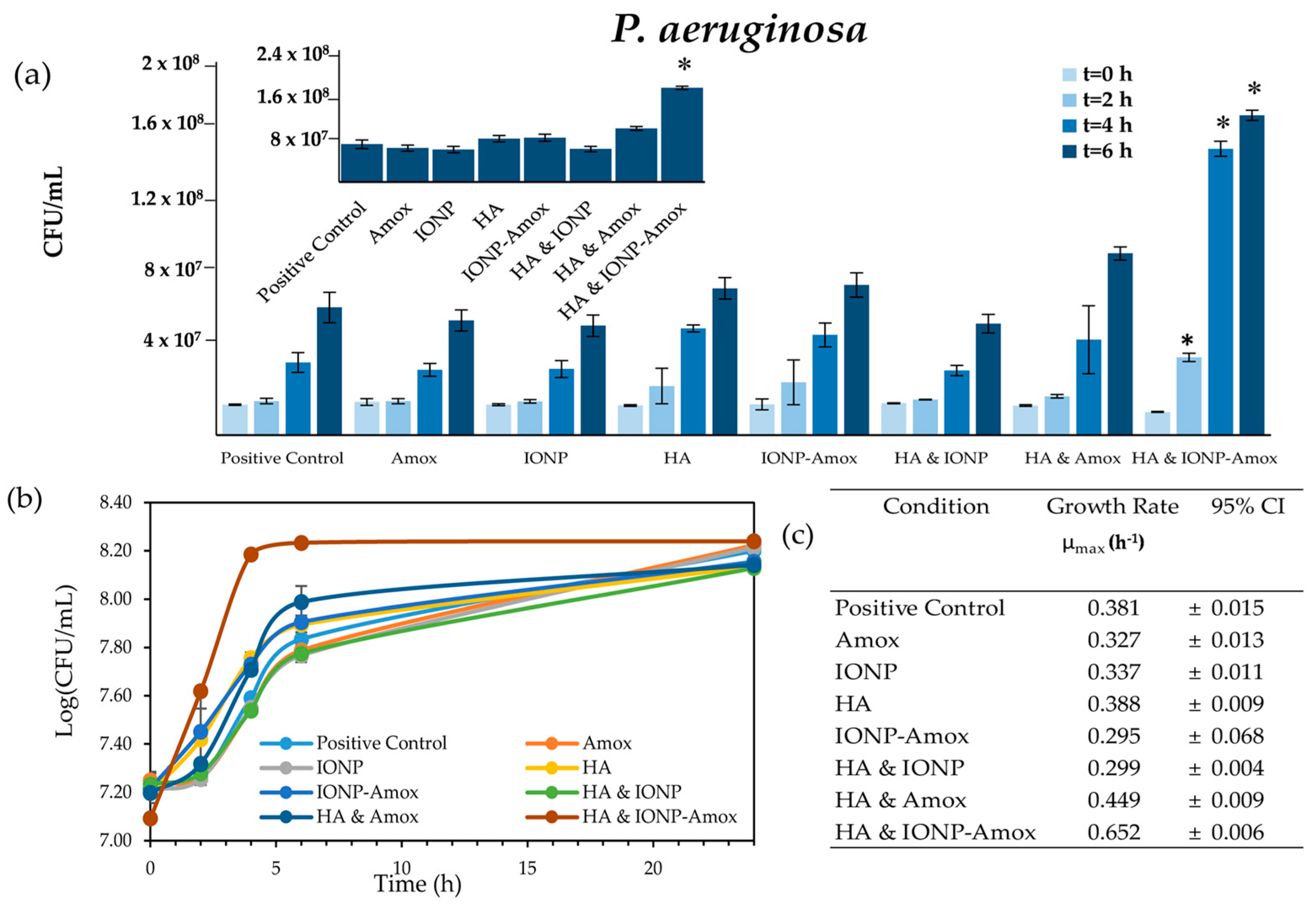
3.2. Impact of Amox Containing Paired Stressors
3.2.1. Bacterial Growth Impacts: IONP-Amox
3.2.2. Bacterial Growth Impacts: HA & Amox
3.3. Impact of all Combined Stressors: IONP-Amox & HA
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lee, C.; Kim, J.Y.; Lee, W.I.; Nelson, K.L.; Yoon, J.; Sedlak, D.L. Bactericidal effect of zero-valent iron nanoparticles on Escherichia coli. Environ. Sci. Technol. 2008, 42, 4927–4933. [Google Scholar] [CrossRef] [PubMed]
- Morones, J.R.; Elechiguerra, J.L.; Camacho, A.; Holt, K.; Kouri, J.B.; Ramírez, J.T.; Yacaman, M.J. The bactericidal effect of silver nanoparticles. Nanotechnology 2005, 16, 2346. [Google Scholar] [CrossRef] [PubMed]
- Sondi, I.; Salopek-Sondi, B. Silver nanoparticles as antimicrobial agent: A case study on E. coli as a model for Gram-negative bacteria. J. Colloid Interf. Sci. 2004, 275, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Ireland, J.C.; Klostermann, P.; Rice, E.W.; Clark, R.M. Inactivation of Escherichia coli by titanium dioxide photocatalytic oxidation. Appl. Environ. Microbiol. 1993, 59, 1668–1670. [Google Scholar] [PubMed]
- Knoll, A.H.; Canfield, D.E.; Konhauser, K.O. Fundamentals of Geobiology; John Wiley & Sons: Hoboken, NJ, USA, 2012; pp. 1–84. [Google Scholar]
- Wellington, E.M.; Boxall, A.B.; Cross, P.; Feil, E.J.; Gaze, W.H.; Hawkey, P.M.; Johnson-Rollings, A.S.; Jones, D.L.; Lee, N.M.; Otten, W. The role of the natural environment in the emergence of antibiotic resistance in Gram-negative bacteria. Lancet Infec. Dis. 2013, 13, 155–165. [Google Scholar] [CrossRef]
- Linares, J.F.; Gustafsson, I.; Baquero, F.; Martinez, J.L. Antibiotics as intermicrobial signaling agents instead of weapons. Proc. Natl. Acad. Sci. USA 2006, 103, 19484–19489. [Google Scholar] [CrossRef] [PubMed]
- Goh, E.B.; Yim, G.; Tsui, W.; McClure, J.; Surette, M.G.; Davies, J. Transcriptional modulation of bacterial gene expression by subinhibitory concentrations of antibiotics. Proc. Natl. Acad. Sci. USA 2002, 99, 17025–17030. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Z.; Yu, Y.; Chen, Z.; Jin, M.; Yang, D.; Zhao, Z.; Wang, J.; Shen, Z.; Wang, X.; Qian, D.; et al. Nanoalumina promotes the horizontal transfer of multiresistance genes mediated by plasmids across genera. Proc. Natl. Acad. Sci. USA 2012, 109, 4944–4949. [Google Scholar] [CrossRef] [PubMed]
- Fux, C.; Costerton, J.; Stewart, P.; Stoodley, P. Survival strategies of infectious biofilms. Trends Microbiol. 2005, 13, 34–40. [Google Scholar] [CrossRef] [PubMed]
- The Center for Disease Dynamics Economics and Policy: Global Antibiotic Resistance Partnership, Final Report. 2015. Available online: Http://cddep.org/publications/state_worlds_antibiotics_2015 (accessed on 11 October 2016).
- Peng, P.C.; Wang, Y.; Liu, L.Y.; Zou, Y.D.; Liao, X.D.; Liang, J.B.; Wu, Y.B. The excretion and environmental effects of amoxicillin, ciprofloxacin, and doxycycline residues in layer chicken manure. Poult. Sci. 2016, 95, 1033–1041. [Google Scholar] [CrossRef] [PubMed]
- Lienert, J.; Bürki, T.; Escher, B.I. Reducing micropollutants with source control: Substance flow analysis of 212 pharmaceuticals in faeces and urine. Water Sci. Technol. 2007, 56, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Executive Agency for Health Consumers: Study on the Environmental Risks of Medicinal Products, Final Report. 2013. Available online: Https://ec.europa.eu/health//sites/health/files/files/environment/study_environment.pdf (accessed on 10 October 2016).
- Landers, T.F.; Cohen, B.; Wittum, T.E.; Larson, E.L. A review of antibiotic use in food animals: Perspective, policy, and potential. Public Health Rep. 2012, 127, 4–22. [Google Scholar] [CrossRef] [PubMed]
- Ray, P.C.; Yu, H.; Fu, P.P. Toxicity and environmental risks of nanomaterials: Challenges and future needs. J. Environ. Sci. Health C 2009, 27, 1–35. [Google Scholar] [CrossRef] [PubMed]
- Dissanayake, N.M.; Current, K.M.; Obare, S.O. Mutagenic effects of Iron oxide nanoparticles on biological cells. Int. J. Mol. Sci. 2015, 16, 23482–23516. [Google Scholar] [CrossRef] [PubMed]
- Nick, S.T.; Bolandi, A.; Samuels, T.A.; Obare, S.O. Advances in understanding the transformation of engineered nanoparticles in the environment. Pure Appl. Chem. 2014, 86, 1129–1140. [Google Scholar] [CrossRef]
- Kapse-Mistry, S.; Govender, T.; Srivastava, R.; Yergeri, M. Nanodrug delivery in reversing multidrug resistance in cancer cells. Front. Pharmacol. 2014, 5, 2–22. [Google Scholar]
- Laurent, S.; Forge, D.; Port, M.; Roch, A.; Robic, C.; Vander Elst, L.; Muller, R.N. Magnetic iron oxide nanoparticles: Synthesis, stabilization, vectorization, physicochemical characterizations, and biological applications. Chem. Rev. 2008, 108, 2064–2110. [Google Scholar] [CrossRef] [PubMed]
- Tepluchin, M.; Kureti, S.; Casapu, M.; Ogel, E.; Mangold, S.; Grunwaldt, J. Study on the hydrothermal and SO2 stability of Al2O3-supported manganese and iron oxide catalysts for lean CO oxidation. Catal. Today 2015, 258, 498–506. [Google Scholar] [CrossRef]
- Ghasemi, E.; Ziyadi, H.; Afshar, A.M.; Sillanpää, M. Iron oxide nanofibers: A new magnetic catalyst for azo dyes degradation in aqueous solution. Chem. Eng. J. 2015, 264, 146–151. [Google Scholar] [CrossRef]
- Pereira, M.; Oliveira, L.; Murad, E. Iron oxide catalysts: Fenton and Fenton-like reactions—A review. Clay Miner. 2012, 47, 285–302. [Google Scholar] [CrossRef]
- He, F.; Li, F. Perovskite promoted iron oxide for hybrid water-splitting and syngas generation with exceptional conversion. Energy Environ. Sci. 2015, 8, 535–539. [Google Scholar] [CrossRef]
- Pastrana-Martínez, L.M.; Pereira, N.; Lima, R.; Faria, J.L.; Gomes, H.T.; Silva, A.M. Degradation of diphenhydramine by photo-Fenton using magnetically recoverable iron oxide nanoparticles as catalyst. Chem. Eng. J. 2015, 261, 45–52. [Google Scholar] [CrossRef]
- Hong, Y.; Honda, R.J.; Myung, N.V.; Walker, S.L. Transport of iron-based nanoparticles: Role of magnetic properties. Environ. Sci. Technol. 2009, 43, 8834–8839. [Google Scholar] [CrossRef] [PubMed]
- Philippe, A.; Schaumann, G.E. Interactions of dissolved organic matter with natural and engineered inorganic colloids: A review. Environ. Sci. Technol. 2014, 48, 8946–8962. [Google Scholar] [CrossRef] [PubMed]
- Aiken, G.R.; Hsu-Kim, H.; Ryan, J.N. Influence of dissolved organic matter on the environmental fate of metals, nanoparticles, and colloids. Environ. Sci. Technol. 2011, 45, 3196–3201. [Google Scholar] [CrossRef] [PubMed]
- Bian, S.; Mudunkotuwa, I.A.; Rupasinghe, T.; Grassian, V.H. Aggregation and dissolution of 4 nm ZnO nanoparticles in aqueous environments: Influence of pH, ionic strength, size, and adsorption of humic acid. Langmuir 2011, 27, 6059–6068. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.; Zhang, Y.; Yu, W.; Shen, H.; Zhang, H.; Gu, N. Preparation and characterization of magnetite nanoparticles coated by amino silane. Colloids Surf. Physicochem. Eng. Asp. 2003, 212, 219–226. [Google Scholar] [CrossRef]
- Rehana, D.; Haleel, A.K.; Rahiman, A.K. Hydroxy, carboxylic and amino acid functionalized superparamagnetic iron oxide nanoparticles: Synthesis, characterization and in vitro anti-cancer studies. J. Chem. Sci. 2015, 127, 1155–1166. [Google Scholar] [CrossRef]
- Grumezescu, A.M.; Gestal, M.C.; Holban, A.M.; Grumezescu, V.; Vasile, B.Ș.; Mogoantă, L.; Iordache, F.; Bleotu, C.; Mogoșanu, G.D. Biocompatible Fe3O4 increases the efficacy of amoxicillin delivery against Gram-positive and Gram-negative bacteria. Molecules 2014, 19, 5013–5027. [Google Scholar] [CrossRef] [PubMed]
- Hauser, A.K.; Mathias, R.K.; Anderson, K.W.; Hilt, J.Z. The effects of synthesis method on the physical and chemical properties of dextran coated iron oxide nanoparticles. Mater. Chem. Phys. 2015, 160, 177–186. [Google Scholar] [CrossRef] [PubMed]
- Maurer-Jones, M.A.; Gunsolus, I.L.; Murphy, C.J.; Haynes, C.L. Toxicity of engineered nanoparticles in the environment. Anal. Chem. 2013, 85, 3036–3049. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.N.; Smith, K.; Samuels, T.A.; Lu, J.; Obare, S.O.; Scott, M.E. Nanoparticles functionalized with ampicillin destroy multiple-antibiotic-resistant isolates of Pseudomonas aeruginosa and Enterobacter aerogenes and methicillin-resistant Staphylococcus aureus. Appl. Environ. Microbiol. 2012, 78, 2768–2774. [Google Scholar] [CrossRef] [PubMed]
- Pirt, S.J. Principles of Microbe and Cell Cultivation; Blackwell Scientific Publications: Oxford, UK, 1975. [Google Scholar]
- Collins, C.H.; Lyne, P.M.; Grange, J.M.; Falkinham, J.O., III. Microbiological Methods, 8th ed.; Arnold: London, UK, 2004. [Google Scholar]
- Nebbioso, A.; Piccolo, A. Molecular characterization of dissolved organic matter (DOM): A critical review. Anal. Bioanal. Chem. 2013, 405, 109–124. [Google Scholar] [CrossRef] [PubMed]
- Pettit, R.E. Organic Matter, Humus, Humate, Humic Acid, Fulvic Acid and Humin: Their Importance in Soil Fertility and Plant Health. CTI Research, 2004; pp. 1–17. Available online: https://static1.squarespace.com/static/55c8cff5e4b0af53827c3795/t/56084ddbe4b0da24d92b6e73/1443384795781/Texas+A%26M+Study.pdf (accessed on 17 May 2017).
- Fabrega, J.; Fawcett, S.R.; Renshaw, J.C.; Lead, J.R. Silver nanoparticle impact on bacterial growth: Effect of pH, concentration, and organic matter. Environ. Sci. Technol. 2009, 43, 7285–7290. [Google Scholar] [CrossRef] [PubMed]
- Asik, B.B.; Turan, M.A.; Celik, H.; Katkat, A.V. Effects of humic substances on plant growth and mineral nutrients uptake of wheat (Triticum durum cv. Salihli) under conditions of salinity. Asian J. Crop Sci. 2009, 1, 87–95. [Google Scholar] [CrossRef]
- Kulikova, N.; Stepanova, E.; Koroleva, O. Mitigating activity of humic substances: Direct influence on biota. In Use of Humic Substances to Remediate Polluted Environments: From Theory to Practice; Springer: Berlin/Heidelberg, Germany, 2005; pp. 285–309. [Google Scholar]
- Nardi, S.; Pizzeghello, D.; Muscolo, A.; Vianello, A. Physiological effects of humic substances on higher plants. Soil Biol. Biochem. 2002, 34, 1527–1536. [Google Scholar] [CrossRef]
- Dalgaard, P.; Koutsoumanis, K. Comparison of maximum specific growth rates and lag times estimated from absorbance and viable count data by different mathematical models. J. Microbiol. Methods 2001, 43, 183–196. [Google Scholar] [CrossRef]
- Calabrese, E.J.; Baldwin, L.A. Hormesis: The dose-response revolution. Annu. Rev. Pharmacol. Toxicol. 2003, 43, 175–197. [Google Scholar] [CrossRef] [PubMed]
- Calabrese, E.J. Paradigm lost, paradigm found: The re-emergence of hormesis as a fundamental dose response model in the toxicological sciences. Environ. Pollut. 2005, 138, 378–411. [Google Scholar] [CrossRef] [PubMed]
- Migliore, L.; Godeas, F.; De Filippis, S.P.; Mantovi, P.; Barchi, D.; Testa, C.; Rubattu, N.; Brambilla, G. Hormetic effect(s) of tetracyclines as environmental contaminant on Zea mays. Environ. Pollut. 2010, 158, 129–134. [Google Scholar] [CrossRef] [PubMed]
- Mathieu, A.; Fleurier, S.; Frénoy, A.; Dairou, J.; Bredeche, M.; Sanchez-Vizuete, P.; Song, X.; Matic, I. Discovery and Function of a General Core Hormetic Stress Response in E. coli Induced by Sublethal Concentrations of Antibiotics. Cell Rep. 2016, 17, 46–57. [Google Scholar] [CrossRef] [PubMed]
- Robbins, A.H.; Stout, C.D. Structure of activated aconitase: Formation of the [4Fe-4S] cluster in the crystal. Proc. Natl. Acad. Sci. USA 1989, 86, 3639–3643. [Google Scholar] [CrossRef] [PubMed]
- Oglesby, A.G.; Farrow, J.M., III; Lee, J.H.; Tomaras, A.P.; Greenberg, E.P.; Pesci, E.C.; Vasil, M.L. The influence of iron on Pseudomonas aeruginosa physiology: A regulatory link between iron and quorum sensing. J. Biol. Chem. 2008, 283, 15558–15567. [Google Scholar] [CrossRef] [PubMed]
- Somerville, G.; Mikoryak, C.A.; Reitzer, L. Physiological characterization of Pseudomonas aeruginosa during exotoxin A synthesis: Glutamate, iron limitation, and aconitase activity. J. Bacteriol. 1999, 181, 1072–1078. [Google Scholar] [PubMed]
- Borcherding, J.; Baltrusaitis, J.; Chen, H.; Stebounova, L.; Wu, C.; Rubasinghege, G.; Mudunkotuwa, I.A.; Caraballo, J.C.; Zabner, J.; Grassian, V.H. Iron oxide nanoparticles induce Pseudomonas aeruginosa growth, induce biofilm formation, and inhibit antimicrobial peptide function. Environ. Sci. Nano 2014, 1, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Haley, K.P.; Skaar, E.P. A battle for iron: Host sequestration and Staphylococcus aureus acquisition. Microbes Infect. 2012, 14, 217–227. [Google Scholar] [CrossRef] [PubMed]
- Hassett, D.J.; Sokol, P.A.; Howell, M.L.; Ma, J.F.; Schweizer, H.T.; Ochsner, U.; Vasil, M.L. Ferric uptake regulator (Fur) mutants of Pseudomonas aeruginosa demonstrate defective siderophore-mediated iron uptake, altered aerobic growth, and decreased superoxide dismutase and catalase activities. J. Bacteriol. 1996, 178, 3996–4003. [Google Scholar] [CrossRef] [PubMed]
- Neilands, J. Evolution of biological iron binding centers. In Structure and Bonding; Springer: Berlin/Heidelberg, Germany, 1972; Volume 11, pp. 145–170. [Google Scholar]
- Hurst, C.J. Neighborhoods and community involvement: No microbe is an island. In Manual of Environmental Microbiology, 3rd ed.; American Society of Microbiology: Washington, DC, USA, 2007; pp. 6–19. [Google Scholar]
- Behrens, S.; Kappler, A.; Obst, M. Linking environmental processes to the in situ functioning of microorganisms by high-resolution secondary ion mass spectrometry (NanoSIMS) and scanning transmission X-ray microscopy (STXM). Environ. Microbiol. 2012, 14, 2851–2869. [Google Scholar] [CrossRef] [PubMed]
- Tilman, D. The resource-ratio hypothesis of plant succession. Am. Nat. 1985, 125, 827–852. [Google Scholar] [CrossRef]
- Miller, T.E.; Burns, J.H.; Munguia, P.; Walters, E.L.; Kneitel, J.M.; Richards, P.M.; Mouquet, N.; Buckley, H.L. A critical review of twenty years’ use of the resource-ratio theory. Am. Nat. 2005, 165, 439–448. [Google Scholar] [PubMed]
- Litwin, C.M.; Calderwood, S.B. Role of iron in regulation of virulence genes. Clin. Microbiol. Rev. 1993, 6, 137–149. [Google Scholar] [CrossRef] [PubMed]
- Messenger, A.J.; Barclay, R. Bacteria, iron and pathogenicity. Biochem. Educ. 1983, 11, 54–63. [Google Scholar] [CrossRef]
- Drakesmith, H.; Prentice, A. Viral infection and iron metabolism. Nature Rev. Microbiol. 2008, 6, 541–552. [Google Scholar] [CrossRef] [PubMed]
- Skaar, E.P. The battle for iron between bacterial pathogens and their vertebrate hosts. PLoS Pathog. 2010, 6, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Ratledge, C.; Dover, L.G. Iron metabolism in pathogenic bacteria. Annu. Rev. Microbiol. 2000, 54, 881–941. [Google Scholar] [CrossRef] [PubMed]
- DePaola, A.; Ulaszek, J.; Kaysner, C.A.; Tenge, B.J.; Nordstrom, J.L.; Wells, J.; Puhr, N.; Gendel, S.M. Molecular, serological, and virulence characteristics of Vibrio parahaemolyticus isolated from environmental, food, and clinical sources in North America and Asia. Appl. Environ. Microbiol. 2003, 69, 3999–4005. [Google Scholar] [CrossRef] [PubMed]
- Stewart, J.R.; Gast, R.J.; Fujioka, R.S.; Solo-Gabriele, H.M.; Meschke, J.S.; Amaral-Zettler, L.A.; Del Castillo, E.; Polz, M.F.; Collier, T.K.; Strom, M.S. The coastal environment and human health: Microbial indicators, pathogens, sentinels and reservoirs. Environ. Health 2008, 7, S3. [Google Scholar] [CrossRef] [PubMed]
- Vigneault, B.; Percot, A.; Lafleur, M.; Campbell, P.G. Permeability changes in model and phytoplankton membranes in the presence of aquatic humic substances. Environ. Sci. Technol. 2000, 34, 3907–3913. [Google Scholar] [CrossRef]
- Tikhonov, V.; Yakushev, A.; Zavgorodnyaya, Y.A.; Byzov, B.; Demin, V. Effects of humic acids on the growth of bacteria. Eurasian Soil Sci. 2010, 43, 305–313. [Google Scholar] [CrossRef]
- Martinez, J.; Delgado-Iribarren, A.; Baquero, F. Mechanisms of iron acquisition and bacterial virulence. FEMS Microbiol. Lett. 1990, 75, 45–56. [Google Scholar] [CrossRef]
- Bullen, J. The significance of iron in infection. Rev. Infect. Dis. 1981, 3, 1127–1138. [Google Scholar] [CrossRef] [PubMed]
- Veyrier, F.J.; Cellier, M.F. Metal economy in host-microbe interactions. Front. Cell. Infect. Microbiol. 2015, 4, 190. [Google Scholar] [CrossRef] [PubMed]
- Nanotechproject.org. Available online: http://www.nanotechproject.org/ (accessed on 4 January 2017).
- Agrosource.net. Available online: www.agrosource.net/pdf/Crisis Declaration (accessed on 4 January 2017).

© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Current, K.M.; Dissanayake, N.M.; Obare, S.O. Effect of Iron Oxide Nanoparticles and Amoxicillin on Bacterial Growth in the Presence of Dissolved Organic Carbon. Biomedicines 2017, 5, 55. https://doi.org/10.3390/biomedicines5030055
Current KM, Dissanayake NM, Obare SO. Effect of Iron Oxide Nanoparticles and Amoxicillin on Bacterial Growth in the Presence of Dissolved Organic Carbon. Biomedicines. 2017; 5(3):55. https://doi.org/10.3390/biomedicines5030055
Chicago/Turabian StyleCurrent, Kelley M., Niluka M. Dissanayake, and Sherine O. Obare. 2017. "Effect of Iron Oxide Nanoparticles and Amoxicillin on Bacterial Growth in the Presence of Dissolved Organic Carbon" Biomedicines 5, no. 3: 55. https://doi.org/10.3390/biomedicines5030055
APA StyleCurrent, K. M., Dissanayake, N. M., & Obare, S. O. (2017). Effect of Iron Oxide Nanoparticles and Amoxicillin on Bacterial Growth in the Presence of Dissolved Organic Carbon. Biomedicines, 5(3), 55. https://doi.org/10.3390/biomedicines5030055




