Predictive Performance of Machine Learning Models for Heart Failure Readmission: A Systematic Review
Abstract
1. Introduction
2. Materials and Methods
2.1. Protocol and Registration
2.2. Eligibility Criteria
2.3. Information Sources
2.4. Search Strategy
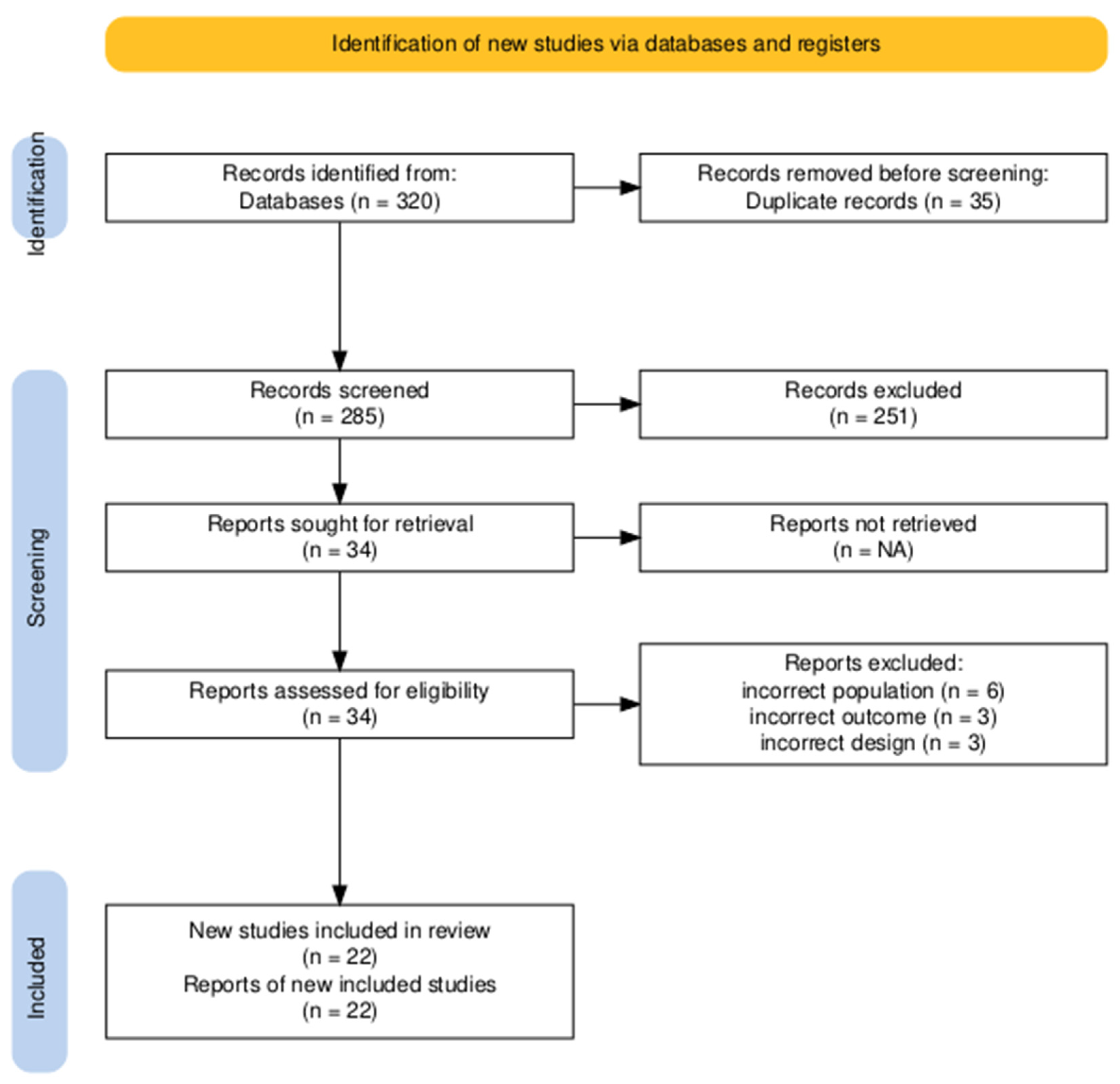
2.5. Selection Process
2.6. Data Collection Process
2.7. Data Items
2.8. Study Risk of Bias Assessment
2.9. Reporting Bias Assessment
3. Results
3.1. Study Characteristics
3.2. Synthesis of Findings
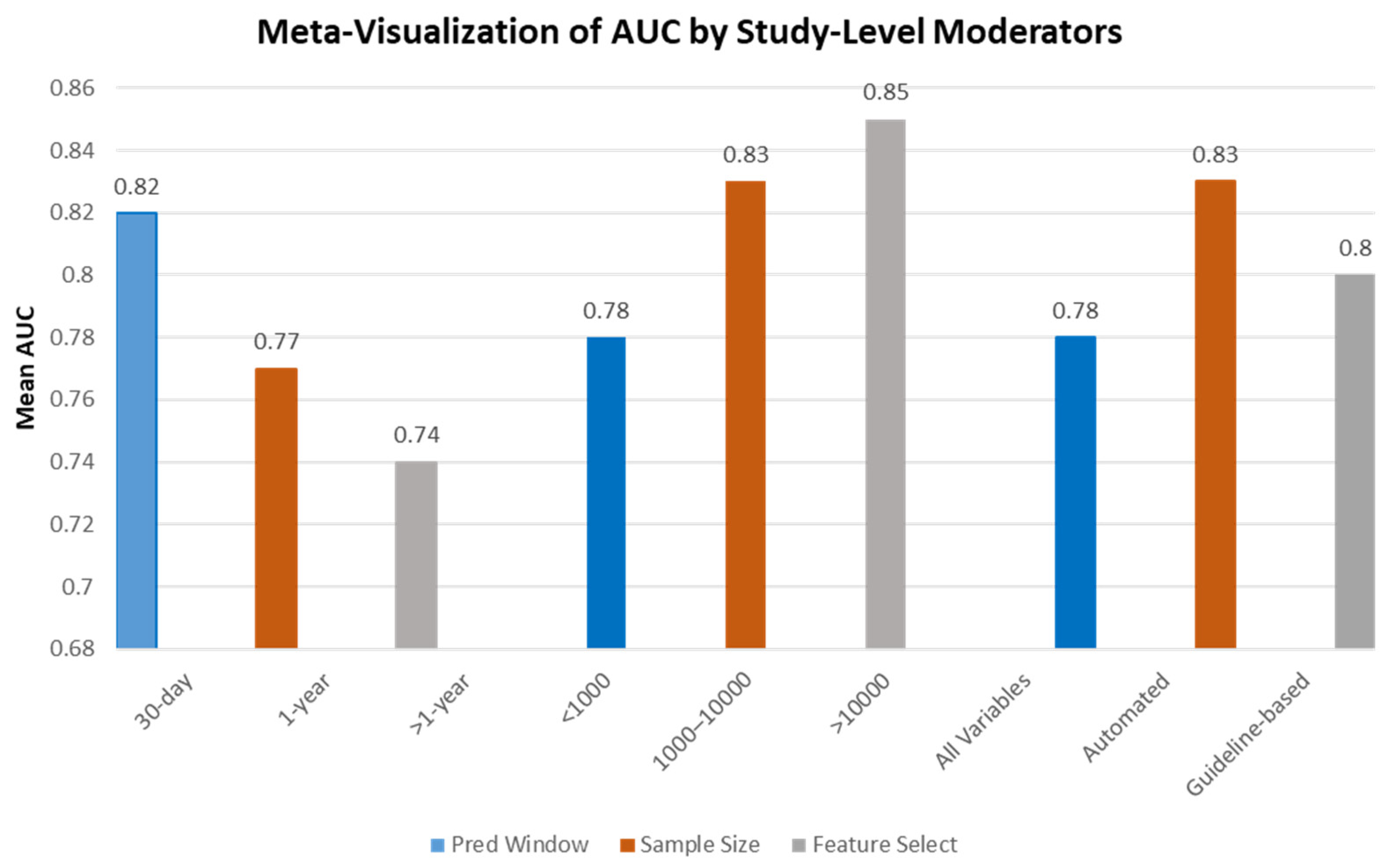
3.3. Subgroup Analysis of Model Performance
4. Discussion
4.1. Strengths of ML in Predicting HF Readmissions
4.2. Effectiveness of Supervised Learning Methods
4.3. Expanding Unsupervised Learning Potential
5. Analysis of Machine Learning Approaches
5.1. Algorithm Types and Prediction Windows
5.2. Strengths and Limitations
5.3. Additional Performance Metrics
6. Methodological Considerations
6.1. Integration Recommendation for Methodological Considerations
6.2. Factors Contributing to Discrepancies in ML Model Performance
7. Clinical Context
7.1. Prediction Windows and the Complexity of HF Readmissions
7.2. Patient Demographics and Risk Factors
7.3. Addressing Geographic Considerations
8. Translational Implications and Implementation Considerations
8.1. Implications for Healthcare Organizations
8.2. Ethical Considerations in ML Implementation
8.3. Infrastructure and Clinical Integration Challenges
8.4. Implementation Roadmap for Clinical Integration
8.5. Barriers to Clinical Adoption: Technical vs. Sociocultural Perspectives
8.6. Methodological Recommendations for Future Research
8.7. Limitations and Future Directions
9. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Shahim, B.; Kapelios, C.J.; Savarese, G.; Lund, L.H. Global Public Health Burden of Heart Failure: An Updated Review. Card. Fail. Rev. 2023, 9, e11. [Google Scholar] [CrossRef] [PubMed]
- Savarese, G.; Lund, L.H. Global Public Health Burden of Heart Failure. Card. Fail. Rev. 2017, 3, 7. [Google Scholar] [CrossRef] [PubMed]
- McLaren, D.P.; Jones, R.; Plotnik, R.; Zareba, W.; McIntosh, S.; Alexis, J.; Chen, L.; Block, R.; Lowenstein, C.J.; Kutyifa, V. Prior hospital admission predicts thirty-day hospital readmission for heart failure patients. Cardiol. J. 2016, 23, 155–162. [Google Scholar] [CrossRef]
- Krumholz, H.M.; Chaudhry, S.I.; Spertus, J.A.; Mattera, J.A.; Hodshon, B.; Herrin, J. Do Non-Clinical Factors Improve Prediction of Readmission Risk? JACC Heart Fail. 2016, 4, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Keeney, T.; Jette, D.U.; Cabral, H.; Jette, A.M. Frailty and Function in Heart Failure: Predictors of 30-Day Hospital Readmission? J. Geriatr. Phys. Ther. 2021, 44, 101–107. [Google Scholar] [CrossRef]
- Lv, Q.; Zhang, X.; Wang, Y.; Xu, X.; He, Y.; Liu, J.; Chang, H.; Zhao, Y.; Zang, X. Multi-trajectories of symptoms and their associations with unplanned 30-day hospital readmission among patients with heart failure: A longitudinal study. Eur. J. Cardiovasc. Nurs. 2024, 23, 737–745. [Google Scholar] [CrossRef]
- Frizzell, J.D.; Liang, L.; Schulte, P.J.; Yancy, C.W.; Heidenreich, P.A.; Hernandez, A.F.; Bhatt, D.L.; Fonarow, G.C.; Laskey, W.K. Prediction of 30-Day All-Cause Readmissions in Patients Hospitalized for Heart Failure: Comparison of Machine Learning and Other Statistical Approaches. JAMA Cardiol. 2017, 2, 204–209. [Google Scholar] [CrossRef]
- Ahmad, T.; Lund, L.H.; Rao, P.; Ghosh, R.; Warier, P.; Vaccaro, B.; Dahlström, U.; O’Connor, C.M.; Felker, G.M.; Desai, N.R. Machine Learning Methods Improve Prognostication, Identify Clinically Distinct Phenotypes, and Detect Heterogeneity in Response to Therapy in a Large Cohort of Heart Failure Patients. J. Am. Heart Assoc. 2018, 7, e008081. [Google Scholar] [CrossRef]
- Sarker, I.H. Machine Learning: Algorithms, Real-World Applications and Research Directions. SN Comput. Sci. 2021, 2, 160. [Google Scholar] [CrossRef]
- Mortazavi, B.J.; Downing, N.S.; Bucholz, E.M.; Dharmarajan, K.; Manhapra, A.; Li, S.-X.; Negahban, S.N.; Krumholz, H.M. Analysis of Machine Learning Techniques for Heart Failure Readmissions. Circ. Cardiovasc. Qual. Outcomes 2016, 9, 629–640. [Google Scholar] [CrossRef]
- Jing, L.; Ulloa Cerna, A.E.; Good, C.W.; Sauers, N.M.; Schneider, G.; Hartzel, D.N.; Leader, J.B.; Kirchner, H.L.; Hu, Y.; Riviello, D.M.; et al. A Machine Learning Approach to Management of Heart Failure Populations. JACC Heart Fail. 2020, 8, 578–587. [Google Scholar] [CrossRef]
- Yu, M.-Y.; Son, Y.-J. Machine learning–based 30-day readmission prediction models for patients with heart failure: A systematic review. Eur. J. Cardiovasc. Nurs. 2024, 23, 711–719. [Google Scholar] [CrossRef] [PubMed]
- Jahangiri, S.; Abdollahi, M.; Rashedi, E.; Azadeh-Fard, N. A machine learning model to predict heart failure readmission: Toward optimal feature set. Front. Artif. Intell. 2024, 7, 1363226. [Google Scholar] [CrossRef]
- Hernandez, L.M.; Blazer, D.G. The Impact of Social and Cultural Environment on Health; National Academies Press: Washington, DC, USA, 2020. Available online: https://www.ncbi.nlm.nih.gov/books/NBK19924/ (accessed on 24 January 2024).
- Dawkins, B.; Renwick, C.; Ensor, T.; Shinkins, B.; Jayne, D.; Meads, D. What Factors Affect Patients’ Ability to Access healthcare? An Overview of Systematic Reviews. Trop. Med. Int. Health 2021, 26, 1177–1188. [Google Scholar] [CrossRef] [PubMed]
- Pavlou, M.; Ambler, G.; Seaman, S.R.; Guttmann, O.; Elliott, P.; King, M.; Omar, R.Z. How to develop a more accurate risk prediction model when there are few events. BMJ 2015, 351, h3868. [Google Scholar] [CrossRef] [PubMed]
- Stødle, K.; Flage, R.; Guikema, S.D.; Aven, T. Data-driven predictive modeling in risk assessment: Challenges and directions for proper uncertainty representation. Risk Anal. 2023, 43, 2644–2658. [Google Scholar] [CrossRef]
- Kleiner Shochat, M.; Fudim, M.; Shotan, A.; Blondheim, D.S.; Kazatsker, M.; Dahan, I.; Asif, A.; Rozenman, Y.; Kleiner, I.; Weinstein, J.M.; et al. Prediction of readmissions and mortality in patients with heart failure: Lessons from the IMPEDANCE-HF extended trial. ESC Heart Fail. 2018, 5, 788–799. [Google Scholar] [CrossRef]
- Hu, Y.; Ma, F.; Hu, M.; Shi, B.; Pan, D.; Ren, J. Development and validation of a machine learning model to predict the risk of readmission within one year in HFpEF patients: Short title: Prediction of HFpEF readmission. Int. J. Med. Inform. 2024, 194, 105703. [Google Scholar] [CrossRef]
- Yordanov, T.R.; Lopes, R.R.; Ravelli, A.C.; Vis, M.; Houterman, S.; Marquering, H.; Abu-Hanna, A. An integrated approach to geographic validation helped scrutinize prediction model performance and its variability. J. Clin. Epidemiol. 2023, 157, 13–21. [Google Scholar] [CrossRef]
- PRISMA. Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA). 2020. Available online: https://www.prisma-statement.org/ (accessed on 24 January 2024).
- Moons, K.G.M.; de Groot, J.A.H.; Bouwmeester, W.; Vergouwe, Y.; Mallett, S.; Altman, D.G.; Reitsma, J.B.; Collins, G.S. Critical Appraisal and Data Extraction for Systematic Reviews of Prediction Modelling Studies: The CHARMS Checklist. PLoS Med. 2014, 11, e1001744. [Google Scholar] [CrossRef]
- Allam, A.; Nagy, M.; Thoma, G.; Krauthammer, M. Neural networks versus Logistic regression for 30 days all-cause readmission prediction. Sci. Rep. 2019, 9, 9277. [Google Scholar] [CrossRef]
- Angraal, S.; Mortazavi, B.J.; Gupta, A.; Khera, R.; Ahmad, T.; Desai, N.R.; Jacoby, D.L.; Masoudi, F.A.; Spertus, J.A.; Krumholz, H.M. Machine Learning Prediction of Mortality and Hospitalization in Heart Failure with Preserved Ejection Fraction. JACC Heart Fail. 2020, 8, 12–21. [Google Scholar] [CrossRef]
- Golas, S.B.; Shibahara, T.; Agboola, S.; Otaki, H.; Sato, J.; Nakae, T.; Hisamitsu, T.; Kojima, G.; Felsted, J.; Kakarmath, S.; et al. A machine learning model to predict the risk of 30-day readmissions in patients with heart failure: A retrospective analysis of electronic medical records data. BMC Med. Inform. Decis. Mak. 2018, 18, 44. [Google Scholar] [CrossRef]
- Jiang, W.; Siddiqui, S.; Barnes, S.; Barouch, L.A.; Korley, F.; Martinez, D.A.; Toerper, M.; Cabral, S.; Hamrock, E.; Levin, S. Readmission risk trajectories for patients with heart failure using a dynamic prediction approach: Retrospective study. JMIR Med. Inform. 2019, 7, e1475. [Google Scholar] [CrossRef] [PubMed]
- Mahajan, S.M.; Mahajan, A.S.; King, R.; Negahban, S. Predicting Risk of 30-Day Readmissions Using Two Emerging Machine Learning Methods. Stud. Health Technol. Inform. 2018, 250, 250–255. Available online: https://pubmed.ncbi.nlm.nih.gov/29857454/ (accessed on 24 January 2024). [PubMed]
- Mahajan, S.M.; Ghani, R. Using ensemble machine learning methods for predicting risk of readmission for heart failure. In MEDINFO 2019: Health and Wellbeing e-Networks for All; IOS Press: Amsterdam, The Netherlands, 2019; pp. 243–247. [Google Scholar] [CrossRef]
- Pishgar, M.; Harford, S.; Theis, J.; Galanter, W.; Rodríguez-Fernández, J.M.; Chaisson, L.H.; Zhang, Y.; Trotter, A.; Kochendorfer, K.M.; Boppana, A.; et al. A process mining-deep learning approach to predict survival in a cohort of hospitalized COVID-19 patients. BMC Med. Inform. Decis. Mak. 2022, 22, 194. [Google Scholar] [CrossRef] [PubMed]
- Desai, R.J.; Wang, S.V.; Vaduganathan, M.; Evers, T.; Schneeweiss, S. Comparison of machine learning methods with traditional models for use of administrative claims with electronic medical records to predict heart failure outcomes. JAMA Netw. Open 2020, 3, e1918962. [Google Scholar] [CrossRef]
- Shameer, K.; Johnson, K.W.; Yahi, A.; Miotto, R.; Li, L.I.; Ricks, D.; Jebakaran, J.; Kovatch, P.; Sengupta, P.P.; Gelijns, S.; et al. Predictive modeling of hospital readmission rates using electronic medical record-wide machine learning: A case-study using Mount Sinai heart failure cohort. Pac. Symp. Biocomput. 2017, 22, 276–287. [Google Scholar] [CrossRef]
- Tukpah, A.M.C.; Cawi, E.; Wolf, L.; Nehorai, A.; Cummings-Vaughn, L. Development of an Institution-Specific Readmission Risk Prediction Model for Real-time Prediction and Patient-Centered Interventions. J. Gen. Intern. Med. 2021, 36, 3910–3912. [Google Scholar] [CrossRef]
- Turgeman, L.; May, J.H. A mixed-ensemble model for hospital readmission. Artif. Intell. Med. 2016, 72, 72–82. [Google Scholar] [CrossRef]
- Yu, S.; Farooq, F.; Van Esbroeck, A.; Fung, G.; Anand, V.; Krishnapuram, B. Predicting readmission risk with institution-specific prediction models. Artif. Intell. Med. 2015, 65, 89–96. [Google Scholar] [CrossRef]
- Sarijaloo, F.; Park, J.; Zhong, X.; Wokhlu, A. Predicting 90-day acute heart failure readmission and death using machine learning-supported decision analysis. Clin. Cardiol. 2021, 44, 230–237. [Google Scholar] [CrossRef]
- Lorenzoni, G.; Sabato, S.S.; Lanera, C.; Bottigliengo, D.; Minto, C.; Ocagli, H.; De Paolis, P.; Gregori, D.; Iliceto, S.; Pisanò, F. Comparison of Machine Learning Techniques for Prediction of Hospitalization in Heart Failure Patients. J. Clin. Med. 2019, 8, 1298. [Google Scholar] [CrossRef] [PubMed]
- Friz, P.H.; Esposito, V.; Marano, G.; Primitz, L.; Bovio, A.; Delgrossi, G.; Bombelli, M.; Grignaffini, G.; Monza, G.; Boracchi, P. Machine learning and LACE index for predicting 30-day readmissions after heart failure hospitalization in elderly patients. Intern. Emerg. Med. 2022, 17, 1727–1737. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Dong, W.; Wang, J.; Lu, X.; Kaymak, U.; Huang, Z. Interpretable clinical prediction via attention-based neural network. BMC Med. Inform. Decis. Mak. 2020, 20, 131. [Google Scholar] [CrossRef] [PubMed]
- Lv, H.; Yang, X.; Wang, B.; Wang, S.; Du, X.; Tan, Q.; Hao, Z.; Xia, Y. Machine learning–driven models to predict prognostic outcomes in patients hospitalized with heart failure using electronic health records: Retrospective study. J. Med. Internet Res. 2021, 23, e24996. [Google Scholar] [CrossRef]
- Bat-Erdene, B.I.; Zheng, H.; Son, S.H.; Lee, J.Y. Deep learning-based prediction of heart failure rehospitalization during 6, 12, 24-month follow-ups in patients with acute myocardial infarction. Health Inform. J. 2022, 28, 14604582221101529. [Google Scholar] [CrossRef]
- Awan, S.E.; Bennamoun, M.; Sohel, F.; Sanfilippo, F.M.; Chow, B.J.; Dwivedi, G. Feature selection and transformation by machine learning reduce variable numbers and improve prediction for heart failure readmission or death. PLoS ONE 2019, 14, e0218760. [Google Scholar] [CrossRef]
- Sharma, V.; Kulkarni, V.; Mcalister, F.; Eurich, D.; Keshwani, S.; Simpson, S.H.; Voaklander, D.; Samanani, S. Predicting 30-Day Readmissions in Patients with Heart Failure Using Administrative Data: A Machine Learning Approach. J. Card. Fail. 2021, 28, 710–722. [Google Scholar] [CrossRef]
- Huang, Y.; Talwar, A.; Chatterjee, S.; Aparasu, R.R. Application of machine learning in predicting hospital readmissions: A scoping review of the literature. BMC Med. Res. Methodol. 2021, 21, 96. [Google Scholar] [CrossRef]
- Sabouri, M.; Rajabi, A.; Hajianfar, G.; Gharibi, O.; Mohebi, M.; Avval, A.; Naderi, N.; Shiri, I. Machine learning based readmission and mortality prediction in heart failure patients. Sci. Rep. 2023, 13, 18671. [Google Scholar] [CrossRef]
- Hidayaturrohman, Q.A.; Hanada, E. Predictive Analytics in Heart Failure Risk, Readmission, and Mortality Prediction: A Review. Cureus 2024, 16, 11. [Google Scholar] [CrossRef]
- Flores, A.M.; Schuler, A.; Eberhard, A.V.; Olin, J.W.; Cooke, J.P.; Leeper, N.J.; Shah, N.H.; Ross, E.G. Unsupervised Learning for Automated Detection of Coronary Artery Disease Subgroups. J. Am. Heart Assoc. 2021, 10, e021976. [Google Scholar] [CrossRef] [PubMed]
- Bednarski, B.; Williams, M.C.; Pieszko, K.; Miller, R.J.H.; Huang, C.; Kwiecinski, J.; Sharir, T.; Di Carli, M.; Fish, M.B.; Ruddy, T.D.; et al. Unsupervised machine learning improves risk stratification of patients with visual normal SPECT myocardial perfusion imaging assessments. Eur. Heart J. 2022, 43 (Suppl. 2), ehac544.300. [Google Scholar] [CrossRef]
- Bell-Navas, A.; Villalba-Orero, M.; Lara-Pezzi, E.; Garicano-Mena, J.; Clainche, S.L. Heart Failure Prediction using Modal Decomposition and Masked Autoencoders for Scarce Echocardiography Databases. arXiv 2025, arXiv:2504.07606. [Google Scholar] [CrossRef]
- Wideqvist, M.; Cui, X.; Magnusson, C.; Schaufelberger, M.; Fu, M. Hospital readmissions of patients with heart failure from real world: Timing and associated risk factors. ESC Heart Fail. 2021, 8, 1388–1397. [Google Scholar] [CrossRef] [PubMed]
- Lam, C.S.P.; Piña, I.L.; Zheng, Y.; Bonderman, D.; Pouleur, A.-C.; Saldarriaga, C.; Pieske, B.; Blaustein, R.O.; Nkulikiyinka, R.; Westerhout, C.M.; et al. Age, Sex, and Outcomes in Heart Failure with Reduced EF: Insights from the VICTORIA Trial. JACC Heart Fail. 2023, 11, 1246–1257. [Google Scholar] [CrossRef]
- Cersosimo, A.; Zito, E.; Pierucci, N.; Matteucci, A.; La Fazia, V.M. A Talk with ChatGPT: The Role of Artificial Intelligence in Shaping the Future of Cardiology and Electrophysiology. J. Pers. Med. 2025, 15, 205. [Google Scholar] [CrossRef]
| Study/Country | Number of Patients | % of Gender | Average Age (yrs) | Algorithm | AUC | Accuracy | Precision | Readmission Days |
|---|---|---|---|---|---|---|---|---|
| Allam et al. [23], USA | 272,778 | 49 females | 73 | SLA | 0.64 | NR | NR | 30 days |
| Angraal et al. [24], USA | 1767 | 50 females | 72 | SLA | 0.76 | NR | NR | 3-year |
| Frizzell et al. [7], USA | 238,581 | 54.5 females | 80 | SLA | 0.62 | NR | NR | 30 days |
| Golas et al. [25], USA | 28,031 | 53 males | 65 | SLA | 0.70 | NR | NR | 30 days |
| Jiang et al. [26], USA | 534 | 64 females | 75 | ULA | 0.73 | NR | NR | 30 days |
| Mahajan et al. [27], USA | 1778 | 97.6 males | 72 | ULA | 0.72 | NR | NR | 30 days |
| Mahajana et al. [28], USA | 36,245 | NA | NA | ELT | 0.70 | NR | NR | 30 days |
| Pishgar et al. [29], USA | 38,597 | 46.3 females | 70 | SLA | 0.93 | 0.84 | 0.89 | 30 days |
| Desai et al. [30], USA | 9502 | 45 female | 78 | SLA | 0.76 | NR | NR | 1-year |
| Shameer et al. [31], USA | 1068 | NA | NA | SLA | 0.78 | NR | NR | 30 days |
| Tukpah et al. [32], USA | 965 | NA | NA | SLA | 0.69 | 0.78 | 0.58 | 30 days |
| Turgeman & May [33], USA | 965 | NA | 79 | SLA | NR | 0.85 | NR | 30 days |
| Yu et al. [34], USA | 20,588 | NA | 65 | SLA | 0.65 | NR | NR | 30 days |
| Sarijaloo et al. [35], USA | 2441 | NA | 65 | SLA | 0.75 | 0.75 | NR | 90 days |
| Mortazavi et al. [10], USA | 1653 | NA | NA | SLA | 0.67 | NR | NR | 180 days |
| Lorenzoni et al. [36], Italy | 380 | 60 females | 73 | SLA | 0.81 | 0.81 | NR | 1-year |
| Friz et al. [37], Italy | 3079 | 55.3 females | 81 | SLA | 0.74 | 0.60 | 0.70 | 30 days |
| Chen et al. [38], China | 736 | NA | 72 | SLA | 0.67 | 0.67 | 0.71 | 1-year |
| Lv et al. [39], China | 13,602 | 52 females | 72 | SLA | 0.81 | 0.77 | 0.76 | 1-year |
| Bat-Erdene et al. [40], Korea | 11,011 | NA | NA | SLA | 0.99 | 0.99 | 0.98 | 1-year |
| Awan et al. [41], Australia | 10,757 | 49 males | 81 | SLA | 0.62 | 48.42 | 0.70 | 30 days |
| Sharma et al. [42], Canada | 9845 | 56 males | 71 | SLA | 0.65 | NR | NR | 30 days |
| Study Name | Participant Selection | Predictor Assessment | Outcome Assessment | Model Development | Analysis |
|---|---|---|---|---|---|
| Allam et al. [23] | L | L | L | L | L |
| Awan et al. [41] | M | L | L | L | L |
| Frizzell et al. [7] | L | L | L | L | L |
| Golas et al. [25] | M | L | L | L | L |
| Jiang et al. [26] | L | L | L | L | L |
| Mahajan et al. [27] | L | L | L | L | L |
| Mahajana et al. [28] | H | L | L | L | L |
| Pishgar et al. [29] | L | L | L | L | L |
| Polo Friz et al. [37] | M | L | L | L | L |
| Shameer et al. [31] | L | L | L | L | L |
| Sharma et al. [42] | L | L | L | L | L |
| Turgeman & May [33] | L | L | L | L | L |
| Yu et al. [34] | L | L | L | L | L |
| Sarijaloo et al. [35] | L | L | L | L | L |
| Mortazavi et al. [10] | L | L | L | L | L |
| Bat-Erdene et al. [40] | L | L | L | L | L |
| Chen et al. [38] | L | L | L | L | L |
| Study | Prediction Window | Sample Size | Key Findings | Strengths | Limitations | Population Diversity (% Non-White) |
|---|---|---|---|---|---|---|
| Allam et al. [23] | 30 days | Large | Deep Learning (Neural Network), Traditional ML (Logistic Regression) | Neural networks showed slightly better performance than logistic regression for 30-day readmission risk. | Large dataset; comparison of deep learning and traditional approaches. | Homogeneous cohort; dependent on billing codes. |
| Frizzell et al. [7] | 30 days | Large multicenter | Traditional ML (Logistic Regression, Random Forest, Gradient Boosting) | Ensemble methods did not substantially outperform logistic regression, with AUC ~0.62–0.72. | Multicenter design; rigorous validation. | U.S.-centric data; moderate discrimination. |
| Angraal et al. [24] | 3 years | Medium | Ensemble (Random Forest) | Random forest achieved reasonable predictive power (AUC 0.76) for long-term (3-year) readmission risk. | Emphasis on long-term outcomes; advanced ML pipeline. | Relatively small/less diverse sample; limited generalizability. |
| Jiang et al. [26] | Dynamic (varied) | Not Reported (NR) | Unsupervised ML (Clustering: k-means) | Identified risk trajectories and clusters (e.g., “rapid decompensators”); segmentation associated with markedly different readmission risks. | Novel dynamic prediction; insight into patient heterogeneity. | Unsupervised results are harder to translate into protocols; lack of clinical actionability. |
| Golas et al. [25] | 30 days | 11,510 patients, 27,334 admissions | Traditional ML (Random Forest, Logistic Regression, SVM, Gradient Boosting) | Random forest and logistic regression had similar AUCs (0.76), supporting use of EHR data for prediction. | Large EHR dataset; real-time application design. | Limited external validation; single institution setting. |
| Shameer et al. [31] | 30 days | Large | Traditional ML (Elastic Net Logistic Regression) | AUC 0.72; demonstrated EHR-wide ML is feasible and valuable for readmission prediction. | Comprehensive variable set; relevant to clinical workflows. | Single-center design; focus on billing code predictors. |
| Bat-Erdene et al. [40] | 6, 12, 24 months | Moderate | Deep Learning | Deep learning outperformed traditional approaches for 6–24-month readmission prediction. | Extended follow-up window; leveraged advanced neural networks. | Lacked clinical interpretability; smaller dataset. |
| Chen et al. [38] | 1 year | Not Reported (NR) | Deep Learning (Attention-based Neural Network) | Attention mechanisms improved interpretability and prediction with AUC 0.82. | Introduced model interpretability; highlighted features via attention weights. | Lacked comparison to other ML approaches; cohort size NR. |
| Lv et al. [39] | Dynamic | Not Reported (NR) | Unsupervised ML (Clustering for trajectory patterns) | High timing prediction (89% accuracy) through symptom trajectory clustering. | Focus on dynamic, interpretable trajectories; novel approach. | Hard to translate unsupervised findings into actionable clinical tools; sample size NR. |
| Sarijaloo et al. [35] | 90 days | Moderate | Ensemble (Random Forest, Gradient Boosting) | ML models improved prediction of 90-day readmission and death versus clinical risk models. | Included robust clinical and administrative data. | Model complexity limits bedside application. |
| Study | Key Strength | Critical Limitation | Clinical Limitations |
|---|---|---|---|
| Huang et al. [43] | Comprehensive scoping review of 42 studies | U.S.-centered sample (82% of included studies; no quality assessment of primary studies | Limited generalizability to non-Western healthcare systems |
| Angraal et al. [24] | Long-term (3-year prediction capability) | Homogeneous cohort (72% White participants); no SGLT2 inhibitor data | Underestimates risk in Asian/younger populations |
| Shameer et al. [31] | Health Electronic records (HER)-wide feature engineering | Single-center design; reliance on billing codes over clinical narratives | May miss social determinants affecting readmission |
| Allam et.al. [23] | Comparison of neural networks vs. logistic regression | Limited to 30-day readmission prediction | Provide insight on algorithm selection for short-term risk assessment |
| Frizzell et al. [7] | Multicenter study design | Focus on traditional statistical approaches | Establishes baseline for comparing ML to conventional methods |
| Jiang et al. [26] | Novel unsupervised approach for dynamic risk trajectories | Complex implementation in clinical settings | Offers new perspective on evolving readmission risk over time |
| Study | ML Algorithm(s) Used | Prediction Window | AUC | Key Features Used |
|---|---|---|---|---|
| Allam et al. [23] | Neural Network, Logistic Regression | 30 days | 0.64 | Billing codes, labs |
| Frizzell et al. [7] | Random Forest, Gradient Boosting, Logistic Regression | 30 days | 0.62–0.72 | EHR, demographics |
| Golas et al. [25] | Random Forest, Logistic Regression, SVM, Gradient Boosting | 30 days | 0.76 | EHR, demographic, clinical, admission data |
| Chen et al. [38] | Deep Learning (Attention-based Neural Network) | 1 year | 0.82 | EHR, text |
| Jiang et al. [26] | Unsupervised k-Means Clustering | Dynamic | 0.73 | Trajectory patterns |
| Shameer et al. [31] | Elastic Net Logistic Regression | 30 days | 0.72 | EHR-wide features, billing codes |
| Bat-Erdene et al. [40] | Deep Learning | 6, 12, 24 months | 0.80–0.85 | Epidemiologic, labs, admission/discharge data |
| Sarijaloo et al. [35] | Random Forest, Gradient Boosting | 90 days | 0.76 | Clinical, administrative, labs |
| Lv et al. [39] | Unsupervised Clustering for Trajectory Patterns | Dynamic | Not Reported | Symptom trajectories |
| Angraal et al. [24] | Random Forest (Ensemble) | 3 years | 0.76 | Demographic, clinical variables |
| Polo Friz et al. [37] | Supervised ML (Random Forest, SVM, Logistic Regression) | 30 days | ~0.69 | LACE index, administrative, clinical |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alnomasy, N.; Pangket, P.; Mostoles, R., Jr.; Alrashedi, H.; Pasay-an, E.; Cho, H.; Alsayed, S.; Gonzales, A.; Alharbi, A.A.M.; Alatawi, N.A.H.; et al. Predictive Performance of Machine Learning Models for Heart Failure Readmission: A Systematic Review. Biomedicines 2025, 13, 2111. https://doi.org/10.3390/biomedicines13092111
Alnomasy N, Pangket P, Mostoles R Jr., Alrashedi H, Pasay-an E, Cho H, Alsayed S, Gonzales A, Alharbi AAM, Alatawi NAH, et al. Predictive Performance of Machine Learning Models for Heart Failure Readmission: A Systematic Review. Biomedicines. 2025; 13(9):2111. https://doi.org/10.3390/biomedicines13092111
Chicago/Turabian StyleAlnomasy, Nader, Petelyne Pangket, Romeo Mostoles, Jr., Habib Alrashedi, Eddieson Pasay-an, Hwayoung Cho, Sharifah Alsayed, Analita Gonzales, Amal A. Mohammad Alharbi, Nuha Ayad H. Alatawi, and et al. 2025. "Predictive Performance of Machine Learning Models for Heart Failure Readmission: A Systematic Review" Biomedicines 13, no. 9: 2111. https://doi.org/10.3390/biomedicines13092111
APA StyleAlnomasy, N., Pangket, P., Mostoles, R., Jr., Alrashedi, H., Pasay-an, E., Cho, H., Alsayed, S., Gonzales, A., Alharbi, A. A. M., Alatawi, N. A. H., Torres, S., Abudawood, K., & Alamoudi, F. A. (2025). Predictive Performance of Machine Learning Models for Heart Failure Readmission: A Systematic Review. Biomedicines, 13(9), 2111. https://doi.org/10.3390/biomedicines13092111








