Anti-Quorum Sensing Activities of Gliptins against Pseudomonas aeruginosa and Staphylococcus aureus
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains, Microbiological Media, and Chemicals
2.2. Detection of Minimum Inhibitory Concentrations (MICs) of Gliptins
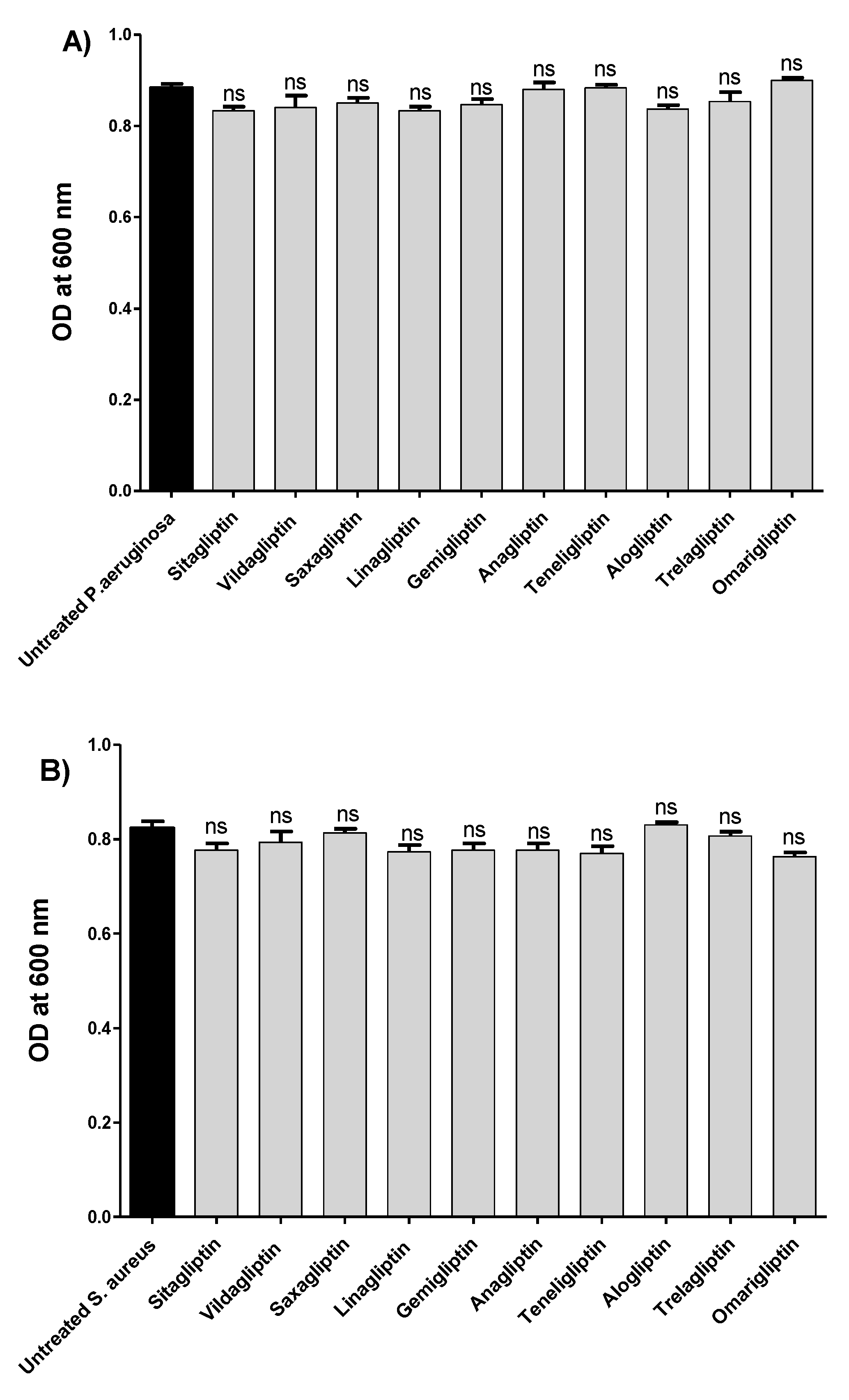
2.3. Assessment of Gliptins’ Effects on Bacterial Growth
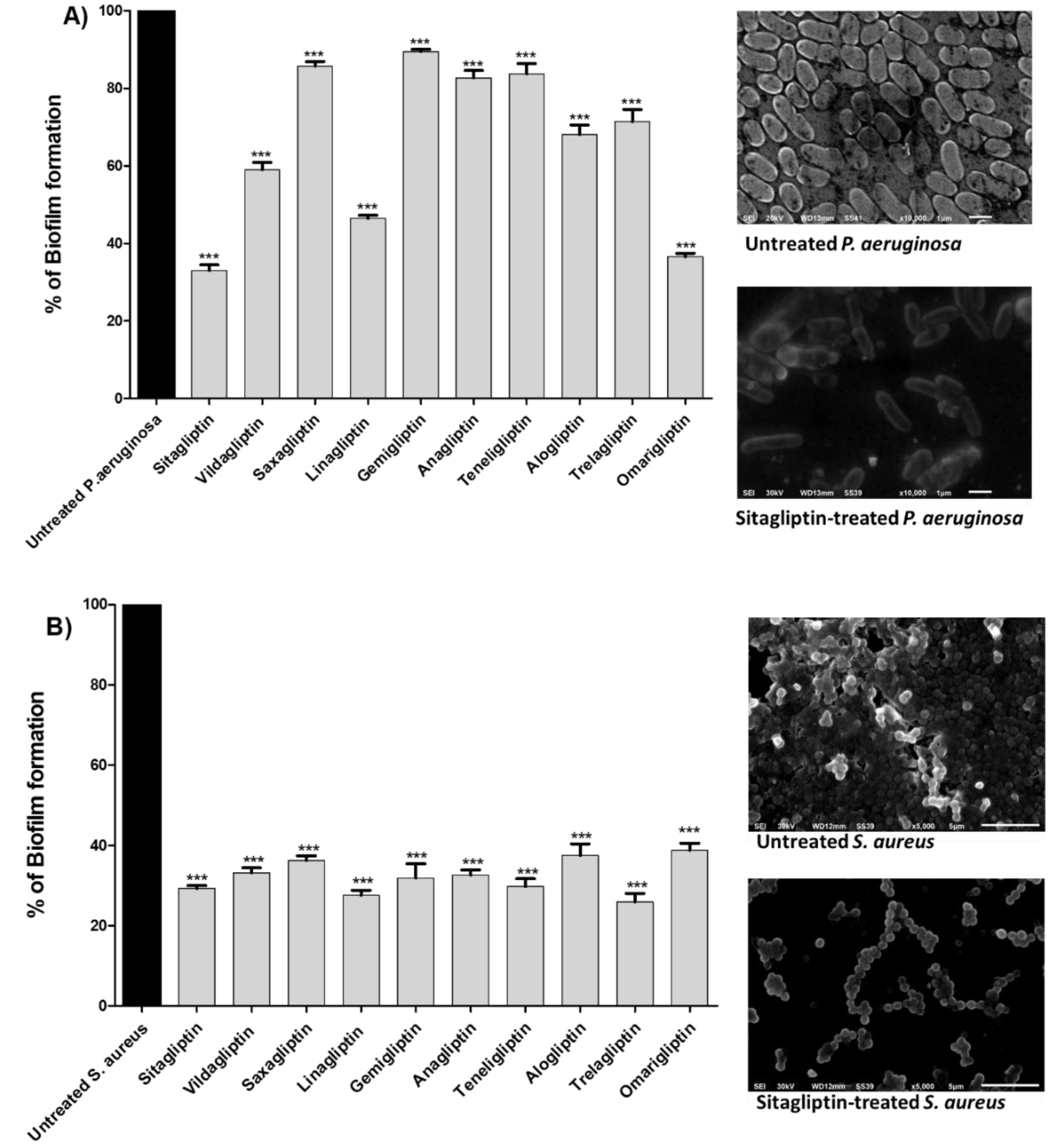
2.4. Evaluation of Antibiofilm Activities of Gliptins
2.5. Visualization of the Biofilm Formation Inhibition
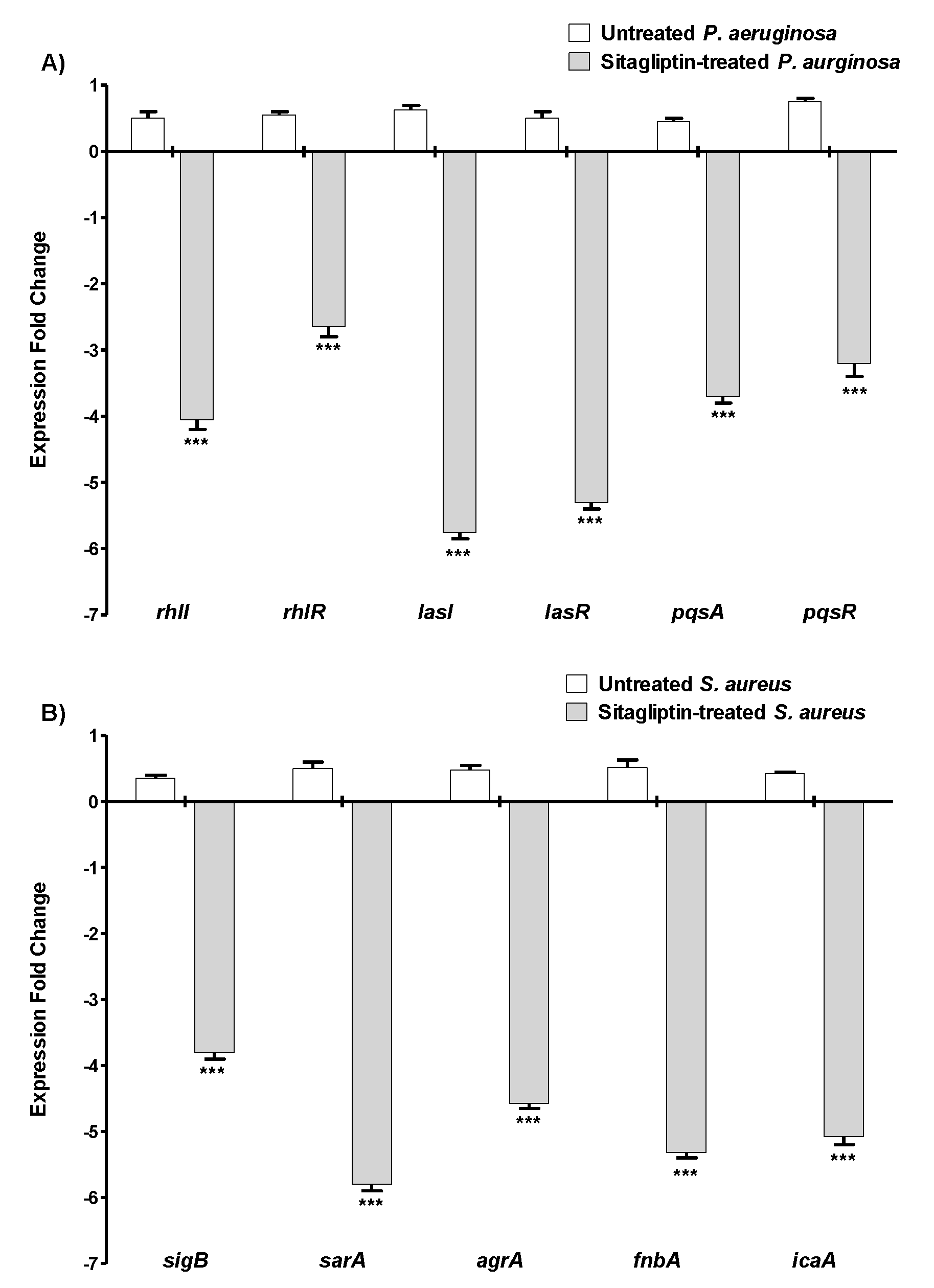
2.6. Quantification of QS- and Virulence-Encoding Gene Expression
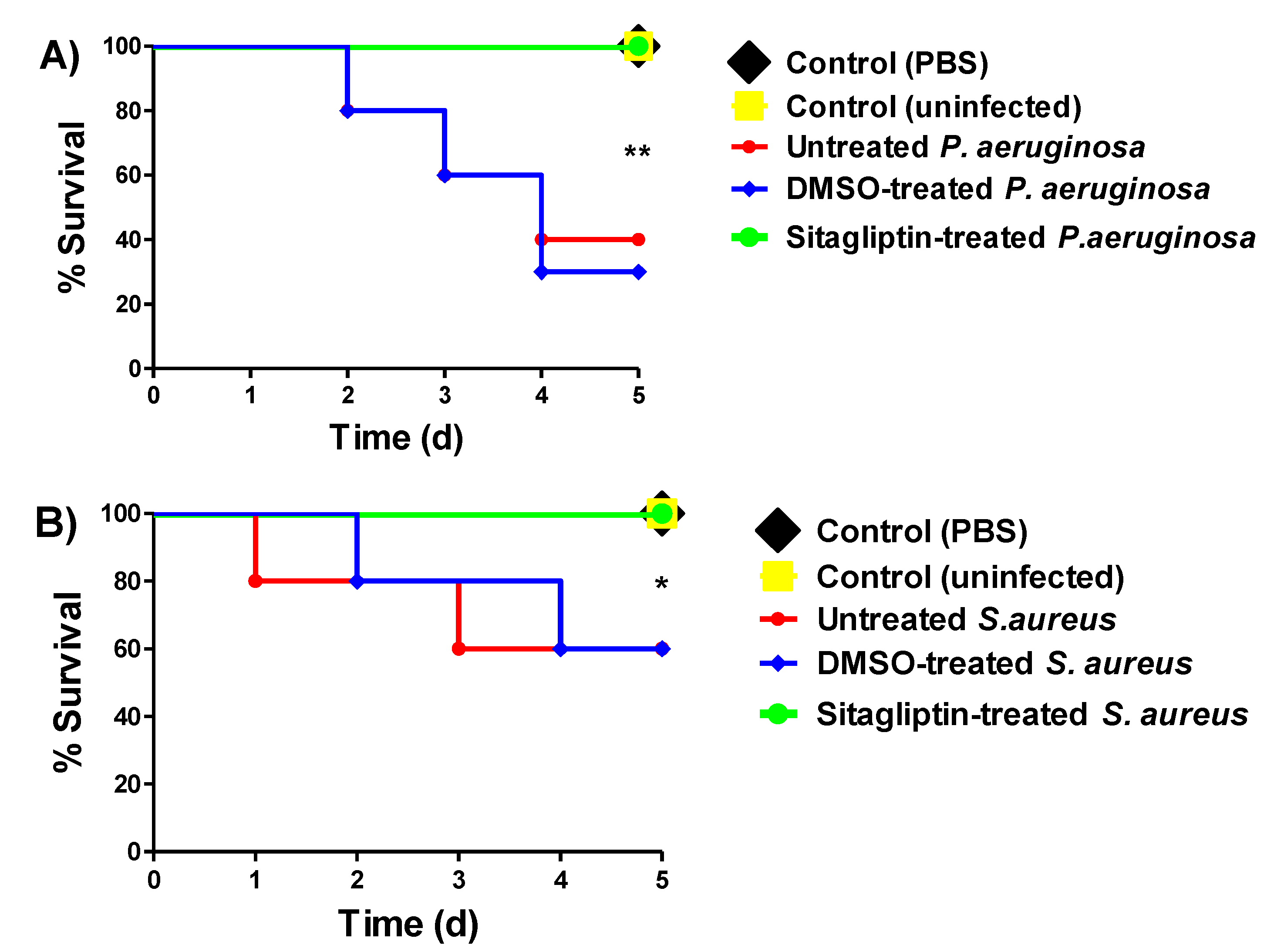
2.7. In Vivo Protection Assay
2.8. Ligand Construction, Target Preparation, and Docking Protocol
2.9. Statistical Analysis
3. Results and Discussion
3.1. Determination of Minimum Inhibitory Concentrations (MICs) of Gliptins
3.2. Antibiofilm Activities of Gliptins
3.3. In Vivo Antivirulence Activities of Gliptins
3.4. Gliptins’ Antivirulence Activities Could Be Due to Interference with QS
3.4.1. Gliptins Downregulate QS-Encoding Genes
3.4.2. Multi-Target Docking Analysis on QS Receptors
Docking Simulation on P. aeruginosa Virulence-Regulating Biotargets
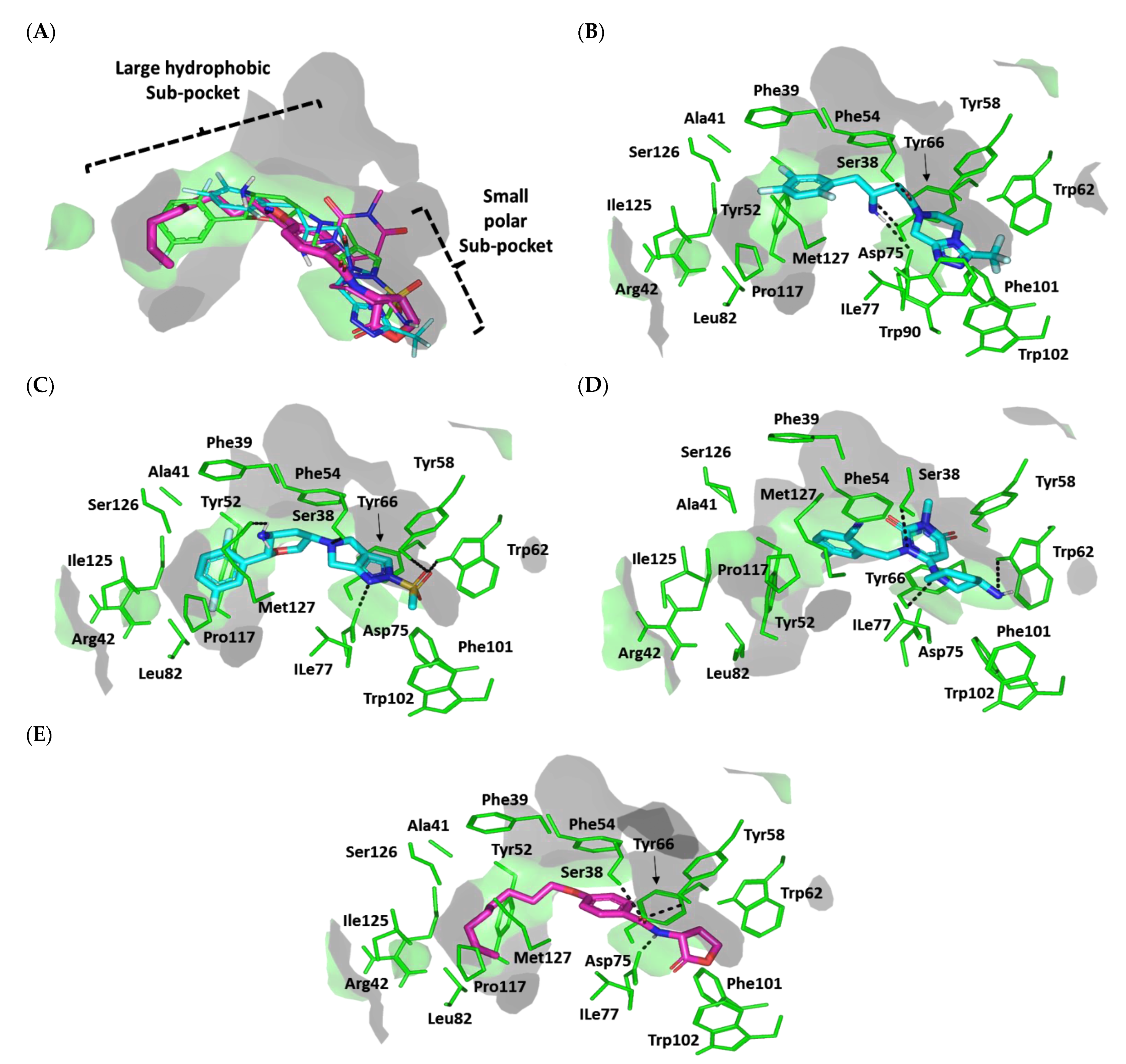
Docking Analysis at LasI-Type AHL Synthase Binding Site
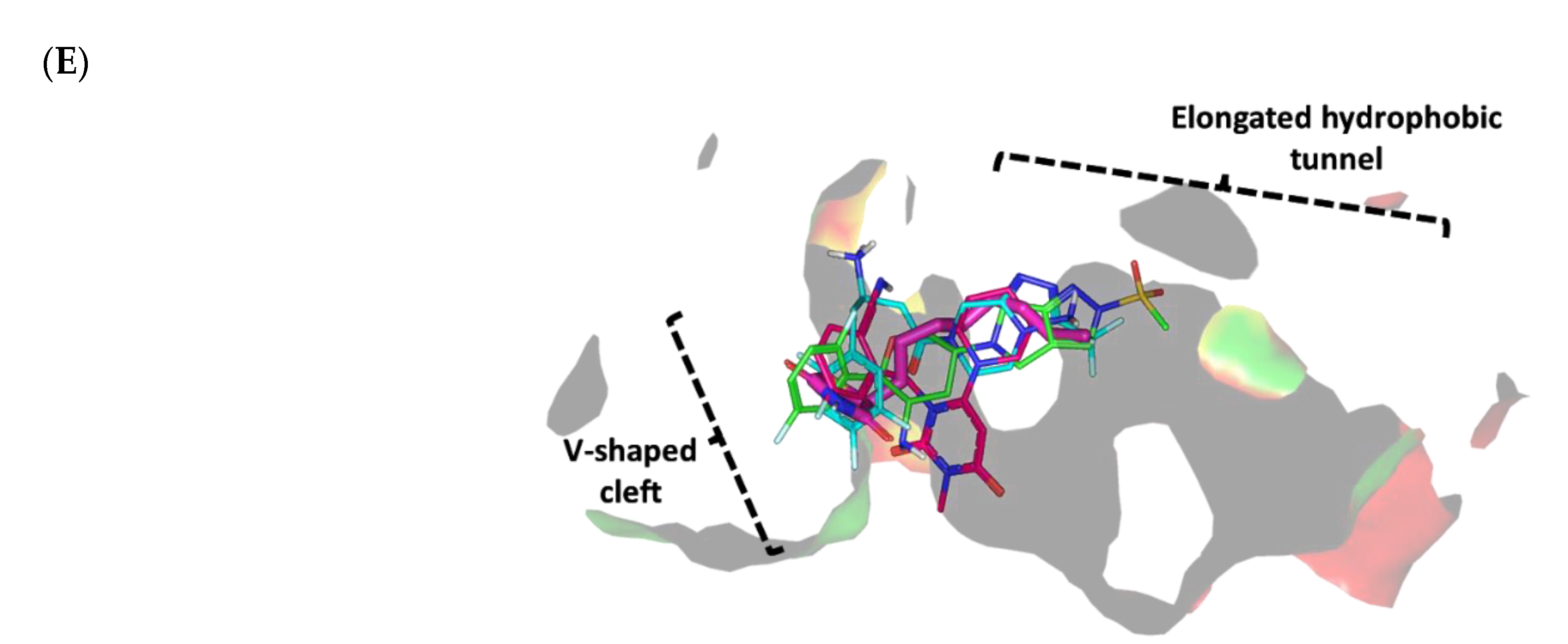
Docking Analysis at QscR-Type QS Protein Binding Site
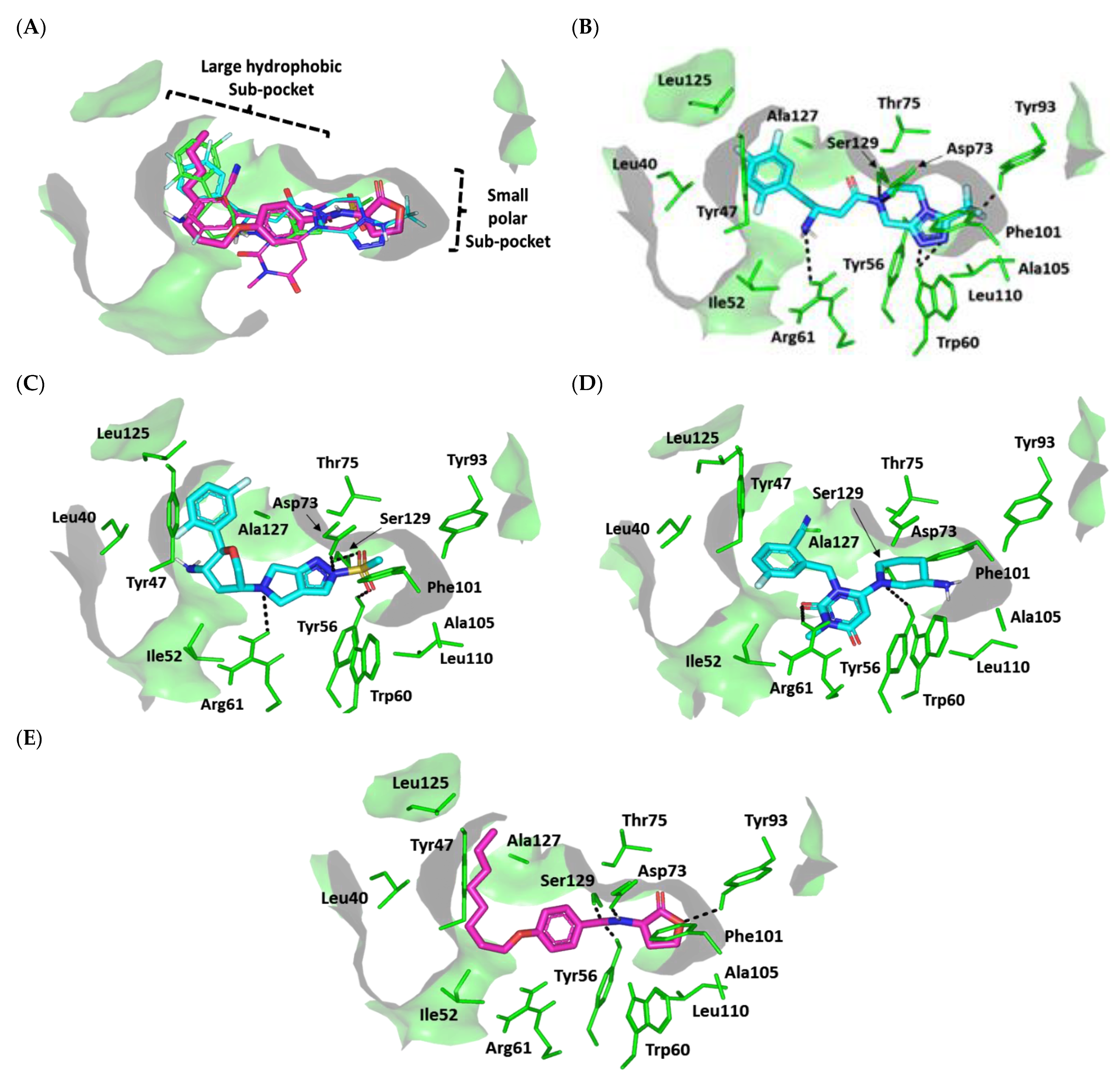
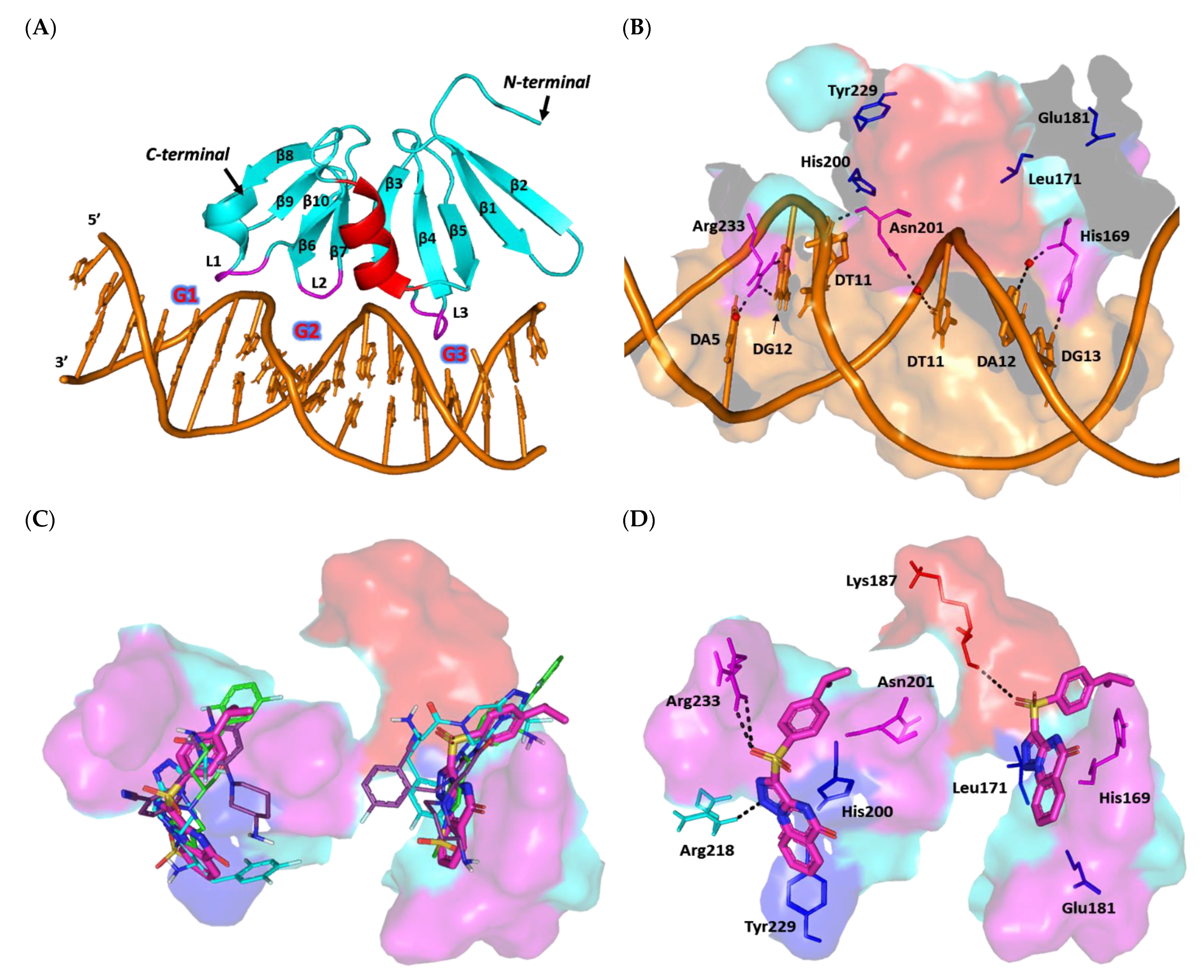
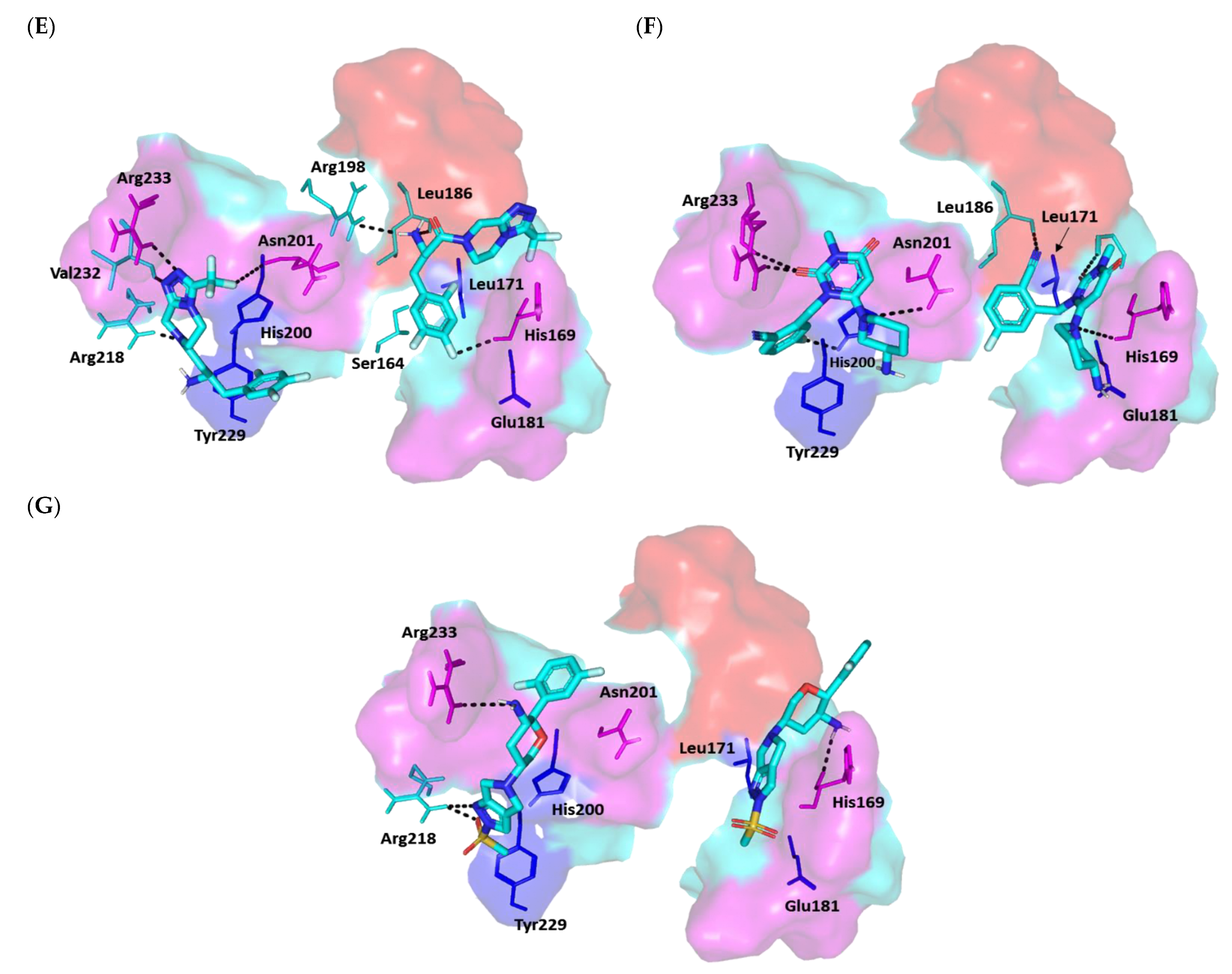
Docking Analysis at LasR-Type Quorum-Sensing Protein Binding Site
Docking Simulation on S. aureus Virulence-Regulating Biotargets
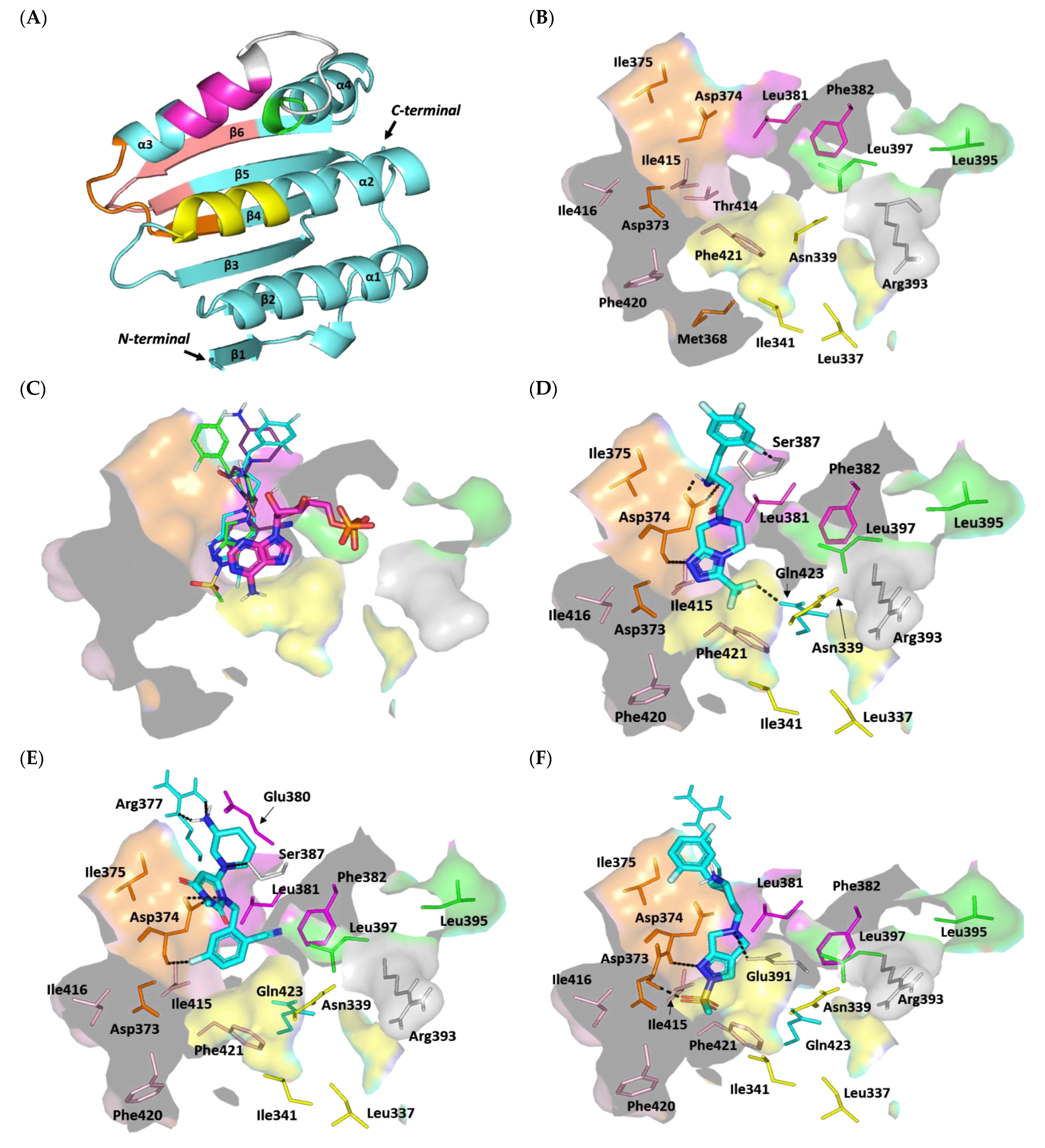
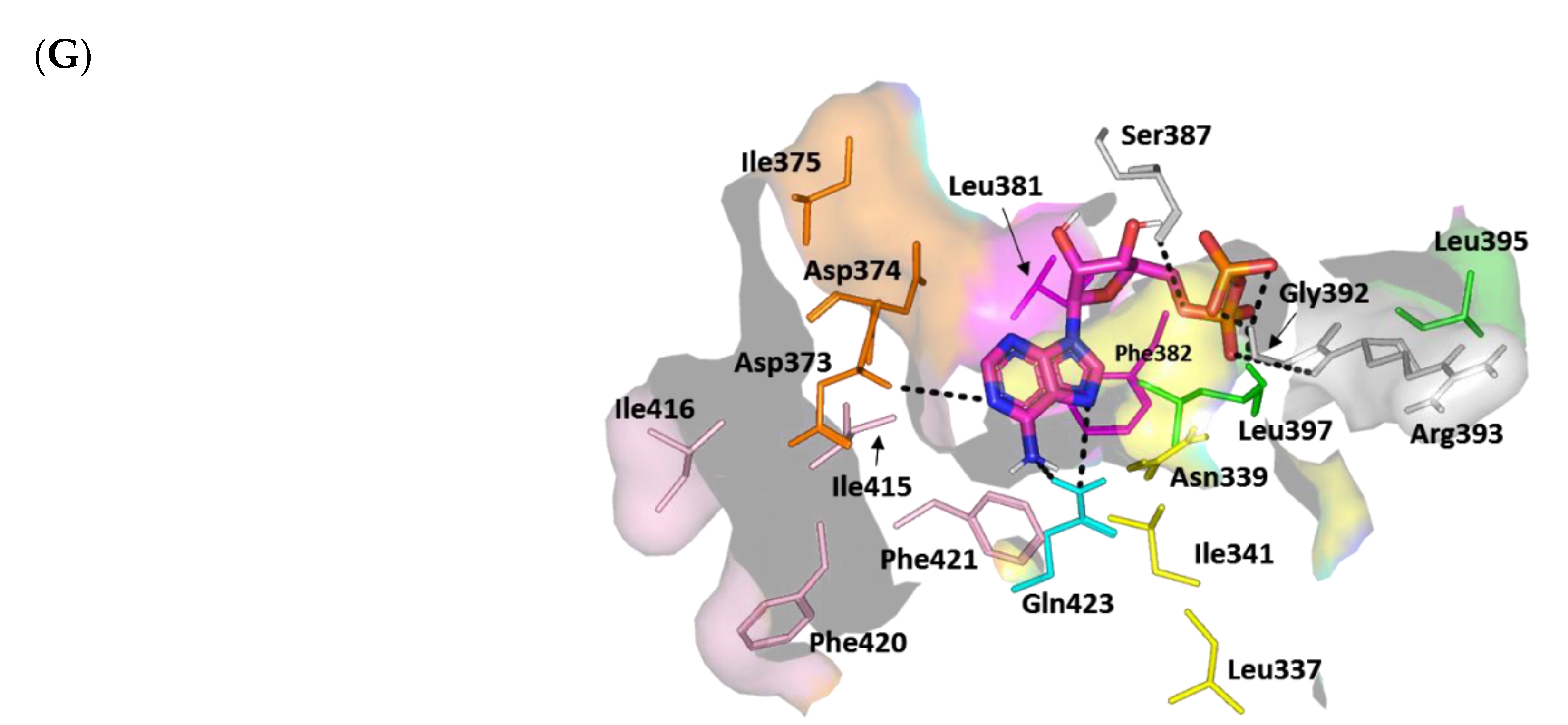
Docking Analysis at ArgC ATP-Binding Domain
Docking Analysis at ArgA Cytoplasmic Response Regulator
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Blair, J.M.; Webber, M.A.; Baylay, A.J.; Ogbolu, D.O.; Piddock, L.J. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 2015, 13, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Hoiby, N.; Bjarnsholt, T.; Givskov, M.; Molin, S.; Ciofu, O. Antibiotic resistance of bacterial biofilms. Int. J. Antimicrob. Agents 2010, 35, 322–332. [Google Scholar] [CrossRef] [PubMed]
- Almalki, A.J.; Ibrahim, T.S.; Taher, E.S.; Mohamed, M.F.A.; Youns, M.; Hegazy, W.A.H.; Al-Mahmoudy, A.M.M. Synthesis, Antimicrobial, Anti-Virulence and Anticancer Evaluation of New 5(4H)-Oxazolone-Based Sulfonamides. Molecules 2022, 27, 671. [Google Scholar] [CrossRef]
- Mohr, K.I. History of Antibiotics Research. Curr. Top. Microbiol. Immunol. 2016, 398, 237–272. [Google Scholar] [CrossRef] [PubMed]
- Cegelski, L.; Marshall, G.R.; Eldridge, G.R.; Hultgren, S.J. The biology and future prospects of antivirulence therapies. Nat. Rev. Microbiol. 2008, 6, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Rasko, D.A.; Sperandio, V. Anti-virulence strategies to combat bacteria-mediated disease. Nat. Rev. Drug Discov. 2010, 9, 117–128. [Google Scholar] [CrossRef] [PubMed]
- Abd El-Hamid, M.I.; Sewid, A.H.; Samir, M.; Hegazy, W.A.H.; Bahnass, M.M.; Mosbah, R.A.; Ghaith, D.M.; Khalifa, E.; Ramadan, H.; Alshareef, W.A.; et al. Clonal Diversity and Epidemiological Characteristics of ST239-MRSA Strains. Front. Cell. Infect. Microbiol. 2022, 12, 782045. [Google Scholar] [CrossRef]
- Muhlen, S.; Dersch, P. Anti-virulence Strategies to Target Bacterial Infections. Curr. Top. Microbiol. Immunol. 2016, 398, 147–183. [Google Scholar] [CrossRef]
- Kamaruzzaman, N.F.; Kendall, S.; Good, L. Targeting the hard to reach: Challenges and novel strategies in the treatment of intracellular bacterial infections. Br. J. Pharmacol. 2017, 174, 2225–2236. [Google Scholar] [CrossRef]
- Almalki, A.J.; Ibrahim, T.S.; Elhady, S.S.; Darwish, K.M.; Hegazy, W.A.H. Repurposing α-Adrenoreceptor Blockers as Promising Anti-Virulence Agents in Gram-Negative Bacteria. Antibiotics 2022, 11, 178. [Google Scholar] [CrossRef]
- Almalki, A.J.; Ibrahim, T.S.; Elhady, S.S.; Hegazy, W.A.H.; Darwish, K.M. Computational and Biological Evaluation of β-Adrenoreceptor Blockers as Promising Bacterial Anti-Virulence Agents. Pharmaceuticals 2022, 15, 110. [Google Scholar] [CrossRef] [PubMed]
- Rutherford, S.T.; Bassler, B.L. Bacterial quorum sensing: Its role in virulence and possibilities for its control. Cold Spring Harb. Perspect. Med. 2012, 2, a012427. [Google Scholar] [CrossRef] [PubMed]
- Hegazy, W.A.H.; Rajab, A.A.H.; Abu Lila, A.S.; Abbas, H.A. Anti-diabetics and antimicrobials: Harmony of mutual interplay. World J. Diabetes 2021, 12, 1832–1855. [Google Scholar] [CrossRef] [PubMed]
- Saqr, A.A.; Aldawsari, M.F.; Khafagy, E.-S.; Shaldam, M.A.; Hegazy, W.A.H.; Abbas, H.A. A Novel Use of Allopurinol as A Quorum-Sensing Inhibitor in Pseudomonas aeruginosa. Antibiotics 2021, 10, 1385. [Google Scholar] [CrossRef]
- Al Saqr, A.; Khafagy, E.S.; Alalaiwe, A.; Aldawsari, M.F.; Alshahrani, S.M.; Anwer, M.K.; Khan, S.; Lila, A.S.A.; Arab, H.H.; Hegazy, W.A.H. Synthesis of Gold Nanoparticles by Using Green Machinery: Characterization and In Vitro Toxicity. Nanomaterials 2021, 11, 808. [Google Scholar] [CrossRef]
- Aldawsari, M.F.; Khafagy, E.S.; Saqr, A.A.; Alalaiwe, A.; Abbas, H.A.; Shaldam, M.A.; Hegazy, W.A.H.; Goda, R.M. Tackling Virulence of Pseudomonas aeruginosa by the Natural Furanone Sotolon. Antibiotics 2021, 10, 871. [Google Scholar] [CrossRef]
- Hegazy, W.A.H.; Khayat, M.T.; Ibrahim, T.S.; Youns, M.; Mosbah, R.; Soliman, W.E. Repurposing of antidiabetics as Serratia marcescens virulence inhibitors. Braz. J. Microbiol. 2021, 52, 627–638. [Google Scholar] [CrossRef]
- Khayyat, A.N.; Abbas, H.A.; Khayat, M.T.; Shaldam, M.A.; Askoura, M.; Asfour, H.Z.; Khafagy, E.-S.; Abu Lila, A.S.; Allam, A.N.; Hegazy, W.A.H. Secnidazole Is a Promising Imidazole Mitigator of Serratia marcescens Virulence. Microorganisms 2021, 9, 2333. [Google Scholar] [CrossRef]
- Khayyat, A.N.; Hegazy, W.A.H.; Shaldam, M.A.; Mosbah, R.; Almalki, A.J.; Ibrahim, T.S.; Khayat, M.T.; Khafagy, E.S.; Soliman, W.E.; Abbas, H.A. Xylitol Inhibits Growth and Blocks Virulence in Serratia marcescens. Microorganisms 2021, 9, 1083. [Google Scholar] [CrossRef]
- Hentzer, M.; Wu, H.; Andersen, J.B.; Riedel, K.; Rasmussen, T.B.; Bagge, N.; Kumar, N.; Schembri, M.A.; Song, Z.; Kristoffersen, P.; et al. Attenuation of Pseudomonas aeruginosa virulence by quorum sensing inhibitors. EMBO J. 2003, 22, 3803–3815. [Google Scholar] [CrossRef]
- Askoura, M.; Youns, M.; Halim Hegazy, W.A. Investigating the influence of iron on Campylobacter jejuni transcriptome in response to acid stress. Microb. Pathog. 2020, 138, 103777. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Contreras, R. Is Quorum Sensing Interference a Viable Alternative to Treat Pseudomonas aeruginosa Infections? Front. Microbiol. 2016, 7, 1454. [Google Scholar] [CrossRef] [PubMed]
- Jiang, T.; Li, M. Quorum sensing inhibitors: A patent review. Expert Opin. Ther. Pat. 2013, 23, 867–894. [Google Scholar] [CrossRef]
- Hegazy, W.A.H.; Salem, I.M.; Alotaibi, H.F.; Khafagy, E.-S.; Ibrahim, D. Terazosin Interferes with Quorum Sensing and Type Three Secretion System and Diminishes the Bacterial Espionage to Mitigate the Salmonella Typhimurium Pathogenesis. Antibiotics 2022, 11, 465. [Google Scholar] [CrossRef] [PubMed]
- Hegazy, W.A.H.; Khayat, M.T.; Ibrahim, T.S.; Nassar, M.S.; Bakhrebah, M.A.; Abdulaal, W.H.; Alhakamy, N.A.; Bendary, M.M. Repurposing Anti-diabetic Drugs to Cripple Quorum Sensing in Pseudomonas aeruginosa. Microorganisms 2020, 8, 1285. [Google Scholar] [CrossRef] [PubMed]
- LaSarre, B.; Federle, M.J. Exploiting quorum sensing to confuse bacterial pathogens. Microbiol. Mol. Biol. Rev. 2013, 77, 73–111. [Google Scholar] [CrossRef]
- Wolska, K.I.; Grudniak, A.M.; Rudnicka, Z.; Markowska, K. Genetic control of bacterial biofilms. J. Appl. Genet. 2016, 57, 225–238. [Google Scholar] [CrossRef] [PubMed]
- Khayyat, A.N.; Abbas, H.A.; Mohamed, M.F.A.; Asfour, H.Z.; Khayat, M.T.; Ibrahim, T.S.; Youns, M.; Khafagy, E.-S.; Abu Lila, A.S.; Safo, M.K.; et al. Not Only Antimicrobial: Metronidazole Mitigates the Virulence of Proteus mirabilis Isolated from Macerated Diabetic Foot Ulcer. Appl. Sci. 2021, 11, 6847. [Google Scholar] [CrossRef]
- Abbas, H.A.; Hegazy, W.A.H. Targeting the virulence factors of Serratia marcescens by ambroxol. Roum. Arch. Microbiol. Immunol. 2017, 76, 27–32. [Google Scholar]
- Hegazy, W.A.H. Diclofenac inhibits virulence of Proteus mirabilis isolated from diabetic foot ulcer. Afr. J. Microbiol. Res. 2016, 10, 733–743. [Google Scholar] [CrossRef]
- Agha, K.A.; Abo-Dya, N.E.; Ibrahim, T.S.; Abdel-Aal, E.H.; Hegazy, W.A. Benzotriazole-Mediated Synthesis and Antibacterial Activity of Novel N-Acylcephalexins. Sci. Pharm. 2016, 84, 484–496. [Google Scholar] [CrossRef] [PubMed]
- Bendary, M.M.; Ibrahim, D.; Mosbah, R.A.; Mosallam, F.; Hegazy, W.A.H.; Awad, N.F.S.; Alshareef, W.A.; Alomar, S.Y.; Zaitone, S.A.; Abd El-Hamid, M.I. Thymol Nanoemulsion: A New Therapeutic Option for Extensively Drug Resistant Foodborne Pathogens. Antibiotics 2020, 10, 25. [Google Scholar] [CrossRef] [PubMed]
- Aldawsari, M.F.; Alalaiwe, A.; Khafagy, E.S.; Al Saqr, A.; Alshahrani, S.M.; Alsulays, B.B.; Alshehri, S.; Abu Lila, A.S.; Danish Rizvi, S.M.; Hegazy, W.A.H. Efficacy of SPG-ODN 1826 Nanovehicles in Inducing M1 Phenotype through TLR-9 Activation in Murine Alveolar J774A.1 Cells: Plausible Nano-Immunotherapy for Lung Carcinoma. Int. J. Mol. Sci. 2021, 22, 6833. [Google Scholar] [CrossRef] [PubMed]
- Hegazy, W.A.H.; Henaway, M. Hepatitis C virus pathogenesis: Serum IL-33 level indicates liver damage. Afr. J. Microbiol. Res. 2015, 9, 1386–1393. [Google Scholar] [CrossRef]
- Pushpakom, S.; Iorio, F.; Eyers, P.A.; Escott, K.J.; Hopper, S.; Wells, A.; Doig, A.; Guilliams, T.; Latimer, J.; McNamee, C.; et al. Drug repurposing: Progress, challenges and recommendations. Nat. Rev. Drug Discov. 2019, 18, 41–58. [Google Scholar] [CrossRef]
- Alandiyjany, M.N.; Abdelaziz, A.S.; Abdelfattah-Hassan, A.; Hegazy, W.A.H.; Hassan, A.A.; Elazab, S.T.; Mohamed, E.A.A.; El-Shetry, E.S.; Saleh, A.A.; ElSawy, N.A.; et al. Novel In Vivo Assessment of Antimicrobial Efficacy of Ciprofloxacin Loaded Mesoporous Silica Nanoparticles against Salmonella typhimurium Infection. Pharmaceuticals 2022, 15, 357. [Google Scholar] [CrossRef]
- El-Hamid, A.; Marwa, I.; El-Naenaeey, E.S.Y.; Hegazy, W.A.; Mosbah, R.A.; Nassar, M.S.; Bakhrebah, M.A.; Abdulaal, W.H.; Alhakamy, N.A.; Bendary, M.M. Promising Antibiofilm Agents: Recent Breakthrough against Biofilm Producing Methicillin-Resistant Staphylococcus aureus. Antibiotics 2020, 9, 667. [Google Scholar] [CrossRef]
- Vishwa, B.; Moin, A.; Gowda, D.V.; Rizvi, S.M.D.; Hegazy, W.A.H.; Abu Lila, A.S.; Khafagy, E.S.; Allam, A.N. Pulmonary Targeting of Inhalable Moxifloxacin Microspheres for Effective Management of Tuberculosis. Pharmaceutics 2021, 13, 79. [Google Scholar] [CrossRef]
- Abbas, H.A.; Hegazy, W.A.H. Repurposing anti-diabetic drug “Sitagliptin” as a novel virulence attenuating agent in Serratia marcescens. PLoS ONE 2020, 15, e0231625. [Google Scholar] [CrossRef]
- Alwattar, J.K.; Mneimneh, A.T.; Abla, K.K.; Mehanna, M.M.; Allam, A.N. Smart Stimuli-Responsive Liposomal Nanohybrid Systems: A Critical Review of Theranostic Behavior in Cancer. Pharmaceutics 2021, 13, 355. [Google Scholar] [CrossRef]
- Cao, H.; Lai, Y.; Bougouffa, S.; Xu, Z.; Yan, A. Comparative genome and transcriptome analysis reveals distinctive surface characteristics and unique physiological potentials of Pseudomonas aeruginosa ATCC 27853. BMC Genom. 2017, 18, 459. [Google Scholar] [CrossRef] [PubMed]
- Latimer, J.; Forbes, S.; McBain, A.J. Attenuated virulence and biofilm formation in Staphylococcus aureus following sublethal exposure to triclosan. Antimicrob. Agents Chemother. 2012, 56, 3092–3100. [Google Scholar] [CrossRef] [PubMed]
- Gowrishankar, S.; Poornima, B.; Pandian, S.K. Inhibitory efficacy of cyclo(L-leucyl-L-prolyl) from mangrove rhizosphere bacterium-Bacillus amyloliquefaciens (MMS-50) toward cariogenic properties of Streptococcus mutans. Res. Microbiol. 2014, 165, 278–289. [Google Scholar] [CrossRef] [PubMed]
- Youns, M.; Askoura, M.; Abbas, H.A.; Attia, G.H.; Khayyat, A.N.; Goda, R.M.; Almalki, A.J.; Khafagy, E.S.; Hegazy, W.A.H. Celastrol Modulates Multiple Signaling Pathways to Inhibit Proliferation of Pancreatic Cancer via DDIT3 and ATF3 Up-Regulation and RRM2 and MCM4 Down-Regulation. Onco Targets Ther. 2021, 14, 3849–3860. [Google Scholar] [CrossRef] [PubMed]
- Askoura, M.; Hegazy, W.A.H. Ciprofloxacin interferes with Salmonella Typhimurium intracellular survival and host virulence through repression of Salmonella pathogenicity island-2 (SPI-2) genes expression. Pathog. Dis. 2020, 78, ftaa011. [Google Scholar] [CrossRef]
- Askoura, M.; Abbas, H.A.; Al Sadoun, H.; Abdulaal, W.H.; Abu Lila, A.S.; Almansour, K.; Alshammari, F.; Khafagy, E.-S.; Ibrahim, T.S.; Hegazy, W.A.H. Elevated Levels of IL-33, IL-17 and IL-25 Indicate the Progression from Chronicity to Hepatocellular Carcinoma in Hepatitis C Virus Patients. Pathogens 2022, 11, 57. [Google Scholar] [CrossRef]
- Parai, D.; Banerjee, M.; Dey, P.; Chakraborty, A.; Islam, E.; Mukherjee, S.K. Effect of reserpine on Pseudomonas aeruginosa quorum sensing mediated virulence factors and biofilm formation. Biofouling 2018, 34, 320–334. [Google Scholar] [CrossRef]
- Gruber, J.D.; Chen, W.; Parnham, S.; Beauchesne, K.; Moeller, P.; Flume, P.A.; Zhang, Y.M. The role of 2,4-dihydroxyquinoline (DHQ) in Pseudomonas aeruginosa pathogenicity. PeerJ 2016, 4, e1495. [Google Scholar] [CrossRef]
- Antonic, V.; Stojadinovic, A.; Zhang, B.; Izadjoo, M.J.; Alavi, M. Pseudomonas aeruginosa induces pigment production and enhances virulence in a white phenotypic variant of Staphylococcus aureus. Infect. Drug Resist. 2013, 6, 175–186. [Google Scholar] [CrossRef]
- Lee, J.H.; Cho, H.S.; Kim, Y.; Kim, J.A.; Banskota, S.; Cho, M.H.; Lee, J. Indole and 7-benzyloxyindole attenuate the virulence of Staphylococcus aureus. Appl. Microbiol. Biotechnol. 2013, 97, 4543–4552. [Google Scholar] [CrossRef]
- Sambanthamoorthy, K.; Smeltzer, M.S.; Elasri, M.O. Identification and characterization of msa (SA1233), a gene involved in expression of SarA and several virulence factors in Staphylococcus aureus. Microbiology 2006, 152, 2559–2572. [Google Scholar] [CrossRef] [PubMed]
- Kot, B.; Sytykiewicz, H.; Sprawka, I. Expression of the Biofilm-Associated Genes in Methicillin-Resistant Staphylococcus aureus in Biofilm and Planktonic Conditions. Int. J. Mol. Sci. 2018, 19, 3487. [Google Scholar] [CrossRef] [PubMed]
- Askoura, M.; Almalki, A.J.; Lila, A.S.A.; Almansour, K.; Alshammari, F.; Khafagy, E.-S.; Ibrahim, T.S.; Hegazy, W.A.H. Alteration of Salmonella enterica Virulence and Host Pathogenesis through Targeting sdiA by Using the CRISPR-Cas9 System. Microorganisms 2021, 9, 2564. [Google Scholar] [CrossRef] [PubMed]
- Wadie, M.A.; Kishk, S.M.; Darwish, K.M.; Mostafa, S.M.; Elgawish, M.S. Simultaneous Determination of Losartan and Rosuvastatin in Rat Plasma Using Liquid Chromatography–Tandem Mass Spectrometric Technique for Application into Pharmacokinetic and Drug–Drug Interaction Studies. Chromatographia 2020, 83, 1477–1494. [Google Scholar] [CrossRef]
- Malebari, A.; Ibrahim, T.; Salem, I.; Salama, I.; Khayyat, A.; Mostafa, S.; El-Sabbagh, O.; Darwish, K. The Anticancer Activity for the Bumetanide-Based Analogs via Targeting the Tumor-Associated Membrane Bound Human Carbonic Anhydrase-IX Enzyme. Pharmaceuticals 2020, 13, 252. [Google Scholar] [CrossRef] [PubMed]
- El Raey, M.A.; El-Hagrassi, A.M.; Osman, A.F.; Darwish, K.M.; Emam, M. Acalypha wilkesiana flowers: Phenolic profiling, cytotoxic activity of their biosynthesized silver nanoparticles and molecular docking study for its constituents as Topoisomerase-I inhibitors. Biocatal. Agric. Biotechnol. 2019, 20, 101243. [Google Scholar] [CrossRef]
- Kitchen, D.B.; Decornez, H.; Furr, J.R.; Bajorath, J. Docking and scoring in virtual screening for drug discovery: Methods and applications. Nat. Rev. Drug Discov. 2004, 3, 935–949. [Google Scholar] [CrossRef] [PubMed]
- Wojciechowski, M.; Lesyng, B. Generalized Born Model: Analysis, Refinement, and Applications to Proteins. J. Phys. Chem. B 2004, 108, 18368–18376. [Google Scholar] [CrossRef]
- Labute, P. The generalized Born/volume integral implicit solvent model: Estimation of the free energy of hydration using London dispersion instead of atomic surface area. J. Comput. Chem. 2008, 29, 1693–1698. [Google Scholar] [CrossRef]
- The PyMOL Molecular Graphics System, 2.0.6; Schrödinger, LLC: New York, NY, USA, 2016; Impact, Schrödinger, LLC: New York, NY, USA, 2016; Prime, Schrödinger, LLC: New York, NY, USA, 2020.
- Hegazy, W.A.H.; Abbas, H.A. Evaluation of the role of SsaV ‘Salmonella pathogenicity island-2 dependent type III secretion system components on the virulence behavior of Salmonella enterica serovar Typhimurium. Afr. J. Biotechnol. 2017, 16, 718–726. [Google Scholar] [CrossRef]
- Gogoi, M.; Sharma, A.; Hazarika, N.K. Biofilm formation by bacterial isolates from patients on indwelling medical devices. Indian J. Med. Microbiol. 2015, 33, 319–320. [Google Scholar] [CrossRef] [PubMed]
- Ramanathan, S.; Ravindran, D.; Arunachalam, K.; Arumugam, V.R. Inhibition of quorum sensing-dependent biofilm and virulence genes expression in environmental pathogen Serratia marcescens by petroselinic acid. Antonie Leeuwenhoek 2018, 111, 501–515. [Google Scholar] [CrossRef] [PubMed]
- Abbas, H.A.; Shaldam, M.A.; Eldamasi, D. Curtailing Quorum Sensing in Pseudomonas aeruginosa by Sitagliptin. Curr. Microbiol. 2020, 77, 1051–1060. [Google Scholar] [CrossRef] [PubMed]
- Papenfort, K.; Bassler, B.L. Quorum sensing signal-response systems in Gram-negative bacteria. Nat. Rev. Microbiol. 2016, 14, 576–588. [Google Scholar] [CrossRef] [PubMed]
- Gould, T.A.; Schweizer, H.P.; Churchill, M.E. Structure of the Pseudomonas aeruginosa acyl-homoserinelactone synthase LasI. Mol. Microbiol. 2004, 53, 1135–1146. [Google Scholar] [CrossRef]
- Watson, W.T.; Minogue, T.D.; Val, D.L.; von Bodman, S.B.; Churchill, M.E. Structural basis and specificity of acyl-homoserine lactone signal production in bacterial quorum sensing. Mol. Cell 2002, 9, 685–694. [Google Scholar] [CrossRef]
- Wysoczynski-Horita, C.L.; Boursier, M.E.; Hill, R.; Hansen, K.; Blackwell, H.E.; Churchill, M.E.A. Mechanism of agonism and antagonism of the Pseudomonas aeruginosa quorum sensing regulator QscR with non-native ligands. Mol. Microbiol. 2018, 108, 240–257. [Google Scholar] [CrossRef]
- McCready, A.R.; Paczkowski, J.E.; Henke, B.R.; Bassler, B.L. Structural determinants driving homoserine lactone ligand selection in the Pseudomonas aeruginosa LasR quorum-sensing receptor. Proc. Natl. Acad. Sci. USA 2019, 116, 245–254. [Google Scholar] [CrossRef]
- Lidor, O.; Al-Quntar, A.; Pesci, E.C.; Steinberg, D. Mechanistic analysis of a synthetic inhibitor of the Pseudomonas aeruginosa LasI quorum-sensing signal synthase. Sci. Rep. 2015, 5, 16569. [Google Scholar] [CrossRef]
- Kontoyianni, M.; McClellan, L.M.; Sokol, G.S. Evaluation of Docking Performance: Comparative Data on Docking Algorithms. J. Med. Chem. 2004, 47, 558–565. [Google Scholar] [CrossRef]
- Moore, J.D.; Rossi, F.M.; Welsh, M.A.; Nyffeler, K.E.; Blackwell, H.E. A Comparative Analysis of Synthetic Quorum Sensing Modulators in Pseudomonas aeruginosa: New Insights into Mechanism, Active Efflux Susceptibility, Phenotypic Response, and Next-Generation Ligand Design. J. Am. Chem. Soc. 2015, 137, 14626–14639. [Google Scholar] [CrossRef] [PubMed]
- Lintz, M.J.; Oinuma, K.; Wysoczynski, C.L.; Greenberg, E.P.; Churchill, M.E. Crystal structure of QscR, a Pseudomonas aeruginosa quorum sensing signal receptor. Proc. Natl. Acad. Sci. USA 2011, 108, 15763–15768. [Google Scholar] [CrossRef] [PubMed]
- Tashiro, Y.; Yawata, Y.; Toyofuku, M.; Uchiyama, H.; Nomura, N. Interspecies interaction between Pseudomonas aeruginosa and other microorganisms. Microbes Environ. 2013, 28, 13–24. [Google Scholar] [CrossRef]
- Smalley, N.E.; An, D.; Parsek, M.R.; Chandler, J.R.; Dandekar, A.A. Quorum Sensing Protects Pseudomonas aeruginosa against Cheating by Other Species in a Laboratory Coculture Model. J. Bacteriol. 2015, 197, 3154–3159. [Google Scholar] [CrossRef] [PubMed]
- Gotoh, Y.; Eguchi, Y.; Watanabe, T.; Okamoto, S.; Doi, A.; Utsumi, R. Two-component signal transduction as potential drug targets in pathogenic bacteria. Curr. Opin. Microbiol. 2010, 13, 232–239. [Google Scholar] [CrossRef]
- Stephenson, K.; Hoch, J.A. Virulence- and antibiotic resistance-associated two-component signal transduction systems of Gram-positive pathogenic bacteria as targets for antimicrobial therapy. Pharmacol. Ther. 2002, 93, 293–305. [Google Scholar] [CrossRef]
- Stock, A.M.; Robinson, V.L.; Goudreau, P.N. Two-component signal transduction. Annu. Rev. Biochem. 2000, 69, 183–215. [Google Scholar] [CrossRef]
- Srivastava, S.K.; Rajasree, K.; Fasim, A.; Arakere, G.; Gopal, B. Influence of the AgrC-AgrA complex on the response time of Staphylococcus aureus quorum sensing. J. Bacteriol. 2014, 196, 2876–2888. [Google Scholar] [CrossRef]
- Thomason, P.; Kay, R. Eukaryotic signal transduction via histidine-aspartate phosphorelay. J. Cell Sci. 2000, 113 Pt 18, 3141–3150. [Google Scholar] [CrossRef]
- Stock, J.B.; Stock, A.M.; Mottonen, J.M. Signal transduction in bacteria. Nature 1990, 344, 395–400. [Google Scholar] [CrossRef]
- Bisicchia, P.; Bui, N.K.; Aldridge, C.; Vollmer, W.; Devine, K.M. Acquisition of VanB-type vancomycin resistance by Bacillus subtilis: The impact on gene expression, cell wall composition and morphology. Mol. Microbiol. 2011, 81, 157–178. [Google Scholar] [CrossRef]
- Dieppois, G.; Ducret, V.; Caille, O.; Perron, K. The transcriptional regulator CzcR modulates antibiotic resistance and quorum sensing in Pseudomonas aeruginosa. PLoS ONE 2012, 7, e38148. [Google Scholar] [CrossRef]
- Nguyen, H.T.; Wolff, K.A.; Cartabuke, R.H.; Ogwang, S.; Nguyen, L. A lipoprotein modulates activity of the MtrAB two-component system to provide intrinsic multidrug resistance, cytokinetic control and cell wall homeostasis in Mycobacterium. Mol. Microbiol. 2010, 76, 348–364. [Google Scholar] [CrossRef]
- Wilke, K.E.; Carlson, E.E. All signals lost. Sci. Transl. Med. 2013, 5, 203ps12. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Eguchi, Y.; Okada, T.; Minagawa, S.; Oshima, T.; Mori, H.; Yamamoto, K.; Ishihama, A.; Utsumi, R. Signal transduction cascade between EvgA/EvgS and PhoP/PhoQ two-component systems of Escherichia coli. J. Bacteriol. 2004, 186, 3006–3014. [Google Scholar] [CrossRef]
- Petrova, O.E.; Sauer, K. A novel signaling network essential for regulating Pseudomonas aeruginosa biofilm development. PLoS Pathog. 2009, 5, e1000668. [Google Scholar] [CrossRef]
- Xie, Q.; Zhao, A.; Jeffrey, P.D.; Kim, M.K.; Bassler, B.L.; Stone, H.A.; Novick, R.P.; Muir, T.W. Identification of a Molecular Latch that Regulates Staphylococcal Virulence. Cell Chem. Biol. 2019, 26, 548–558.e544. [Google Scholar] [CrossRef]
- Dutta, R.; Inouye, M. GHKL, an emergent ATPase/kinase superfamily. Trends Biochem. Sci. 2000, 25, 24–28. [Google Scholar] [CrossRef]
- Wilke, K.E.; Fihn, C.A.; Carlson, E.E. Screening serine/threonine and tyrosine kinase inhibitors for histidine kinase inhibition. Bioorg. Med. Chem. 2018, 26, 5322–5326. [Google Scholar] [CrossRef] [PubMed]
- Worthington, R.J.; Blackledge, M.S.; Melander, C. Small-molecule inhibition of bacterial two-component systems to combat antibiotic resistance and virulence. Future Med. Chem. 2013, 5, 1265–1284. [Google Scholar] [CrossRef]
- Huang, R.Z.; Zheng, L.K.; Liu, H.Y.; Pan, B.; Hu, J.; Zhu, T.; Wang, W.; Jiang, D.B.; Wu, Y.; Wu, Y.C.; et al. Thiazolidione derivatives targeting the histidine kinase YycG are effective against both planktonic and biofilm-associated Staphylococcus epidermidis. Acta Pharmacol. Sin. 2012, 33, 418–425. [Google Scholar] [CrossRef] [PubMed]
- Qin, Z.; Zhang, J.; Xu, B.; Chen, L.; Wu, Y.; Yang, X.; Shen, X.; Molin, S.; Danchin, A.; Jiang, H.; et al. Structure-based discovery of inhibitors of the YycG histidine kinase: New chemical leads to combat Staphylococcus epidermidis infections. BMC Microbiol. 2006, 6, 96. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Wang, F.; Niu, S.; Cao, J.; Wu, K.; Li, Y.; Yin, N.; Zhang, X.; Zhu, W.; Yin, Y. Discovery of novel inhibitors of Streptococcus pneumoniae based on the virtual screening with the homology-modeled structure of histidine kinase (VicK). BMC Microbiol. 2009, 9, 129. [Google Scholar] [CrossRef] [PubMed]
- Cai, X.; Zhang, J.; Chen, M.; Wu, Y.; Wang, X.; Chen, J.; Zhang, J.; Shen, X.; Qu, D.; Jiang, H. The effect of the potential PhoQ histidine kinase inhibitors on Shigella flexneri virulence. PLoS ONE 2011, 6, e23100. [Google Scholar] [CrossRef]
- Liu, H.; Zhao, D.; Chang, J.; Yan, L.; Zhao, F.; Wu, Y.; Xu, T.; Gong, T.; Chen, L.; He, N.; et al. Efficacy of novel antibacterial compounds targeting histidine kinase YycG protein. Appl. Microbiol. Biotechnol. 2014, 98, 6003–6013. [Google Scholar] [CrossRef] [PubMed]
- Eguchi, Y.; Kubo, N.; Matsunaga, H.; Igarashi, M.; Utsumi, R. Development of an antivirulence drug against Streptococcus mutans: Repression of biofilm formation, acid tolerance, and competence by a histidine kinase inhibitor, walkmycin C. Antimicrob. Agents Chemother. 2011, 55, 1475–1484. [Google Scholar] [CrossRef]
- Gilmour, R.; Foster, J.E.; Sheng, Q.; McClain, J.R.; Riley, A.; Sun, P.M.; Ng, W.L.; Yan, D.; Nicas, T.I.; Henry, K.; et al. New class of competitive inhibitor of bacterial histidine kinases. J. Bacteriol. 2005, 187, 8196–8200. [Google Scholar] [CrossRef]
- Sidote, D.J.; Barbieri, C.M.; Wu, T.; Stock, A.M. Structure of the Staphylococcus aureus AgrA LytTR domain bound to DNA reveals a beta fold with an unusual mode of binding. Structure 2008, 16, 727–735. [Google Scholar] [CrossRef]
- Rajasree, K.; Fasim, A.; Gopal, B. Conformational features of the Staphylococcus aureus AgrA-promoter interactions rationalize quorum-sensing triggered gene expression. Biochem. Biophys. Rep. 2016, 6, 124–134. [Google Scholar] [CrossRef]
- Nicod, S.S.; Weinzierl, R.O.; Burchell, L.; Escalera-Maurer, A.; James, E.H.; Wigneshweraraj, S. Systematic mutational analysis of the LytTR DNA binding domain of Staphylococcus aureus virulence gene transcription factor AgrA. Nucleic Acids Res. 2014, 42, 12523–12536. [Google Scholar] [CrossRef]
- Sully, E.K.; Malachowa, N.; Elmore, B.O.; Alexander, S.M.; Femling, J.K.; Gray, B.M.; DeLeo, F.R.; Otto, M.; Cheung, A.L.; Edwards, B.S.; et al. Selective chemical inhibition of agr quorum sensing in Staphylococcus aureus promotes host defense with minimal impact on resistance. PLoS Pathog. 2014, 10, e1004174. [Google Scholar] [CrossRef] [PubMed]
- Mahdally, N.H.; George, R.F.; Kashef, M.T.; Al-Ghobashy, M.; Murad, F.E.; Attia, A.S. Staquorsin: A Novel Staphylococcus aureus Agr-Mediated Quorum Sensing Inhibitor Impairing Virulence in vivo Without Notable Resistance Development. Front. Microbiol. 2021, 12, 700494. [Google Scholar] [CrossRef] [PubMed]
- Salam, A.M.; Quave, C.L. Targeting Virulence in Staphylococcus aureus by Chemical Inhibition of the Accessory Gene Regulator System In Vivo. mSphere 2018, 3, e00500-17. [Google Scholar] [CrossRef] [PubMed]
- Bezar, I.F.; Mashruwala, A.A.; Boyd, J.M.; Stock, A.M. Drug-like Fragments Inhibit agr-Mediated Virulence Expression in Staphylococcus aureus. Sci. Rep. 2019, 9, 6786. [Google Scholar] [CrossRef] [PubMed]
- Palaniappan, B.; Solomon, A.P.; Raj, C.D. Targeting AgrA quorum sensing regulator by bumetanide attenuates virulence in Staphylococcus aureus—A drug repurposing approach. Life Sci. 2021, 273, 119306. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.; Edelsbrunner, H.; Woodward, C. Anatomy of protein pockets and cavities: Measurement of binding site geometry and implications for ligand design. Protein Sci. Publ. Protein Soc. 1998, 7, 1884–1897. [Google Scholar] [CrossRef] [PubMed]
| Target Gene | Sequence (5′–3′) | Reference |
|---|---|---|
| lasI | For: CTACAGCCTGCAGAACGACA Rev: ATCTGGGTCTTGGCATTGAG | [16,25,47] |
| lasR | For: ACGCTCAAGTGGAAAATTGG Rev: GTAGATGGACGGTTCCCAGA | [16,25,47] |
| rhlI | For: CTCTCTGAATCGCTGGAAGG Rev: GACGTCCTTGAGCAGGTAGG | [16,25,47] |
| rhlR | For: AGGAATGACGGAGGCTTTTT Rev: CCCGTAGTTCTGCATCTGGT | [16,25,47] |
| pqsA | For: TTCTGTTCCGCCTCGATTTC Rev: AGTCGTTCAACGCCAGCAC | [16,25,48] |
| pqsR | For: AACCTGGAAATCGACCTGTG Rev: TGAAATCGTCGAGCAGTACG | [16,25,47] |
| rpoD | For: GGGCGAAGAAGGAAATGGTC Rev: CAGGTGGCGTAGGTGGAGAAC | [48] |
| SigB | For: AAGTGATTCGTAAGGACGTCT Rev: TCGATAACTATAACCAAAGCC T | [49,50] |
| SarA | For: TCTTGTTAATGCACAACAACGTAA Rev: TGT TTG CTT CAG TGA TTC GTT T | [51] |
| AgrA | For: GGAGTGATTTCAATGGCACA Rev: ATCCAT TTTACTAAGTCACCGATT | [51] |
| fnbA | For: AACTGCACAACCAGCAAATG Rev: TTGAGGTTGTGTCGTTTCCTT | [51] |
| icaA | For: CAATACTATTTCGGGTGTCTTCACTCT Rev: CAAGAAACTGCAATATCTTCGGTAATCAT | [52] |
| 16S rRNA | For: TGT CGT GAG ATG TTG GG Rev: CGA TTC CAG CTT CATGT | [50] |
| Tested Gliptin | P. aeruginosa | S. aureus |
|---|---|---|
| Sitagliptin | 10 mg/mL | 5 mg/mL |
| Vildagliptin | 10 mg/mL | 5 mg/mL |
| Saxagliptin | 2 mg/mL | 1 mg/mL |
| Linagliptin | 10 mg/mL | 5 mg/mL |
| Gemigliptin | 2 mg/mL | 1 mg/mL |
| Anagliptin | 2 mg/mL | 1 mg/mL |
| Teneligliptin | 2 mg/mL | 1 mg/mL |
| Alogliptin | 2 mg/mL | 1 mg/mL |
| Trelagliptin | 2 mg/mL | 1 mg/mL |
| Omarigliptin | 2 mg/mL | 1 mg/mL |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Khayat, M.T.; Abbas, H.A.; Ibrahim, T.S.; Khayyat, A.N.; Alharbi, M.; Darwish, K.M.; Elhady, S.S.; Khafagy, E.-S.; Safo, M.K.; Hegazy, W.A.H. Anti-Quorum Sensing Activities of Gliptins against Pseudomonas aeruginosa and Staphylococcus aureus. Biomedicines 2022, 10, 1169. https://doi.org/10.3390/biomedicines10051169
Khayat MT, Abbas HA, Ibrahim TS, Khayyat AN, Alharbi M, Darwish KM, Elhady SS, Khafagy E-S, Safo MK, Hegazy WAH. Anti-Quorum Sensing Activities of Gliptins against Pseudomonas aeruginosa and Staphylococcus aureus. Biomedicines. 2022; 10(5):1169. https://doi.org/10.3390/biomedicines10051169
Chicago/Turabian StyleKhayat, Maan T., Hisham A. Abbas, Tarek S. Ibrahim, Ahdab N. Khayyat, Majed Alharbi, Khaled M. Darwish, Sameh S. Elhady, El-Sayed Khafagy, Martin K. Safo, and Wael A. H. Hegazy. 2022. "Anti-Quorum Sensing Activities of Gliptins against Pseudomonas aeruginosa and Staphylococcus aureus" Biomedicines 10, no. 5: 1169. https://doi.org/10.3390/biomedicines10051169
APA StyleKhayat, M. T., Abbas, H. A., Ibrahim, T. S., Khayyat, A. N., Alharbi, M., Darwish, K. M., Elhady, S. S., Khafagy, E.-S., Safo, M. K., & Hegazy, W. A. H. (2022). Anti-Quorum Sensing Activities of Gliptins against Pseudomonas aeruginosa and Staphylococcus aureus. Biomedicines, 10(5), 1169. https://doi.org/10.3390/biomedicines10051169











