Estimating the Prevalence of De Novo Monogenic Neurodevelopmental Disorders from Large Cohort Studies
Abstract
1. Introduction
2. Methods
2.1. Cohorts and Samples
2.2. Prevalence Estimation
2.3. Comparison to Previous Incidence Estimates
2.4. Comparison to Social Media Group Numbers for Top NDD Genes
3. Results
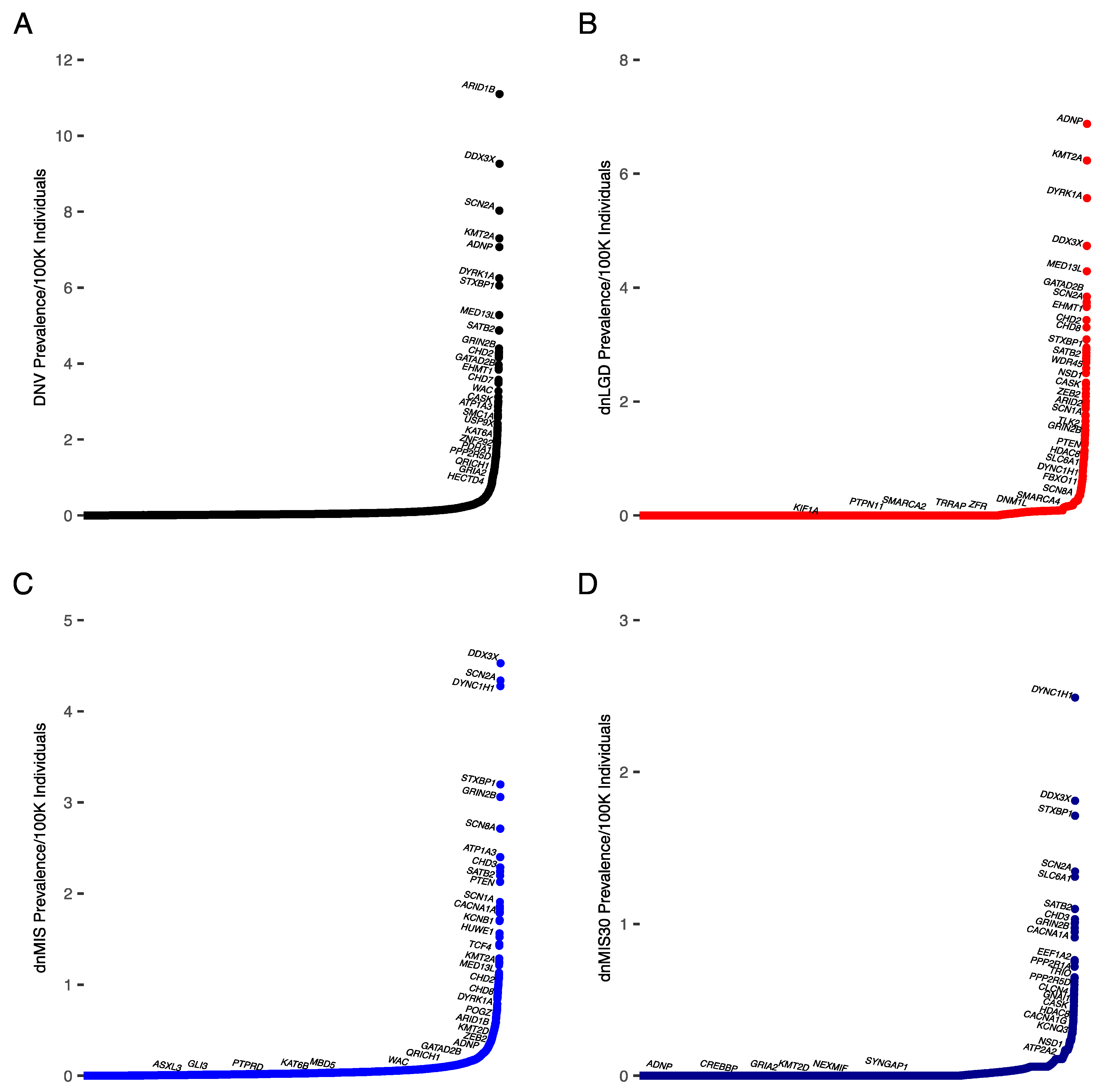
3.1. Prevalence Estimates for All NDDs
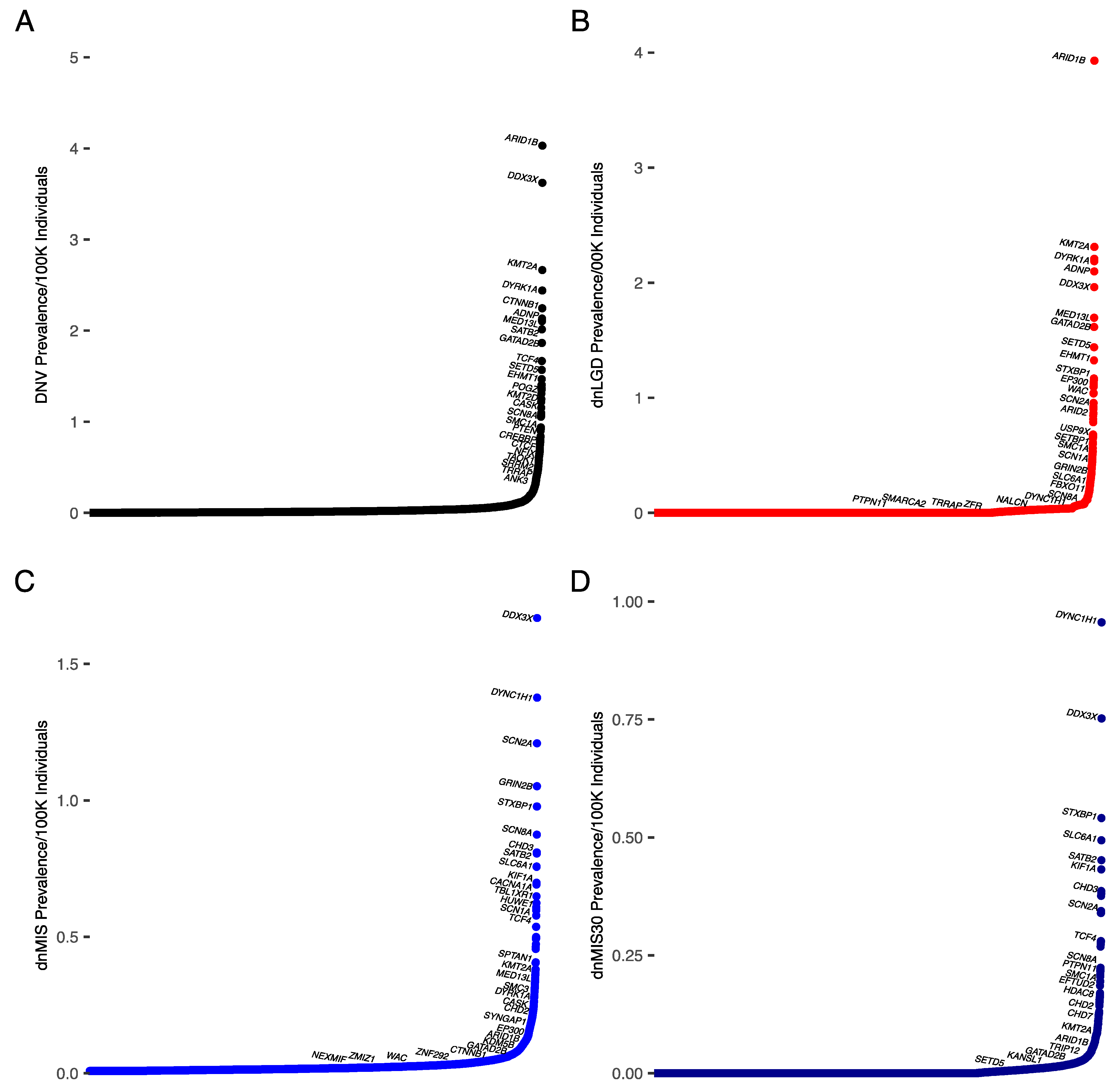
3.2. Prevalence Estimates for DD/ID
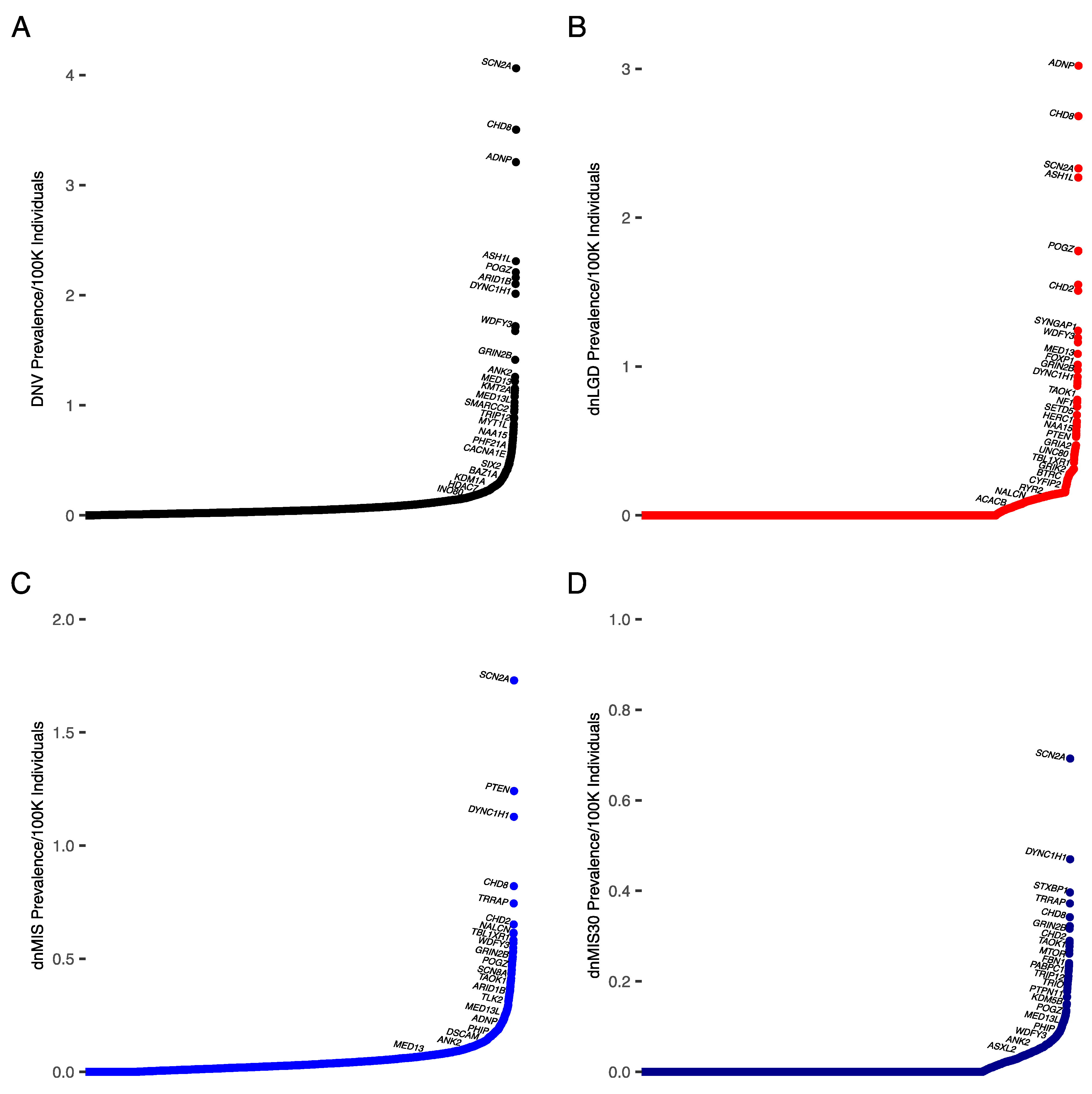
3.3. Prevalence Estimates for ASD
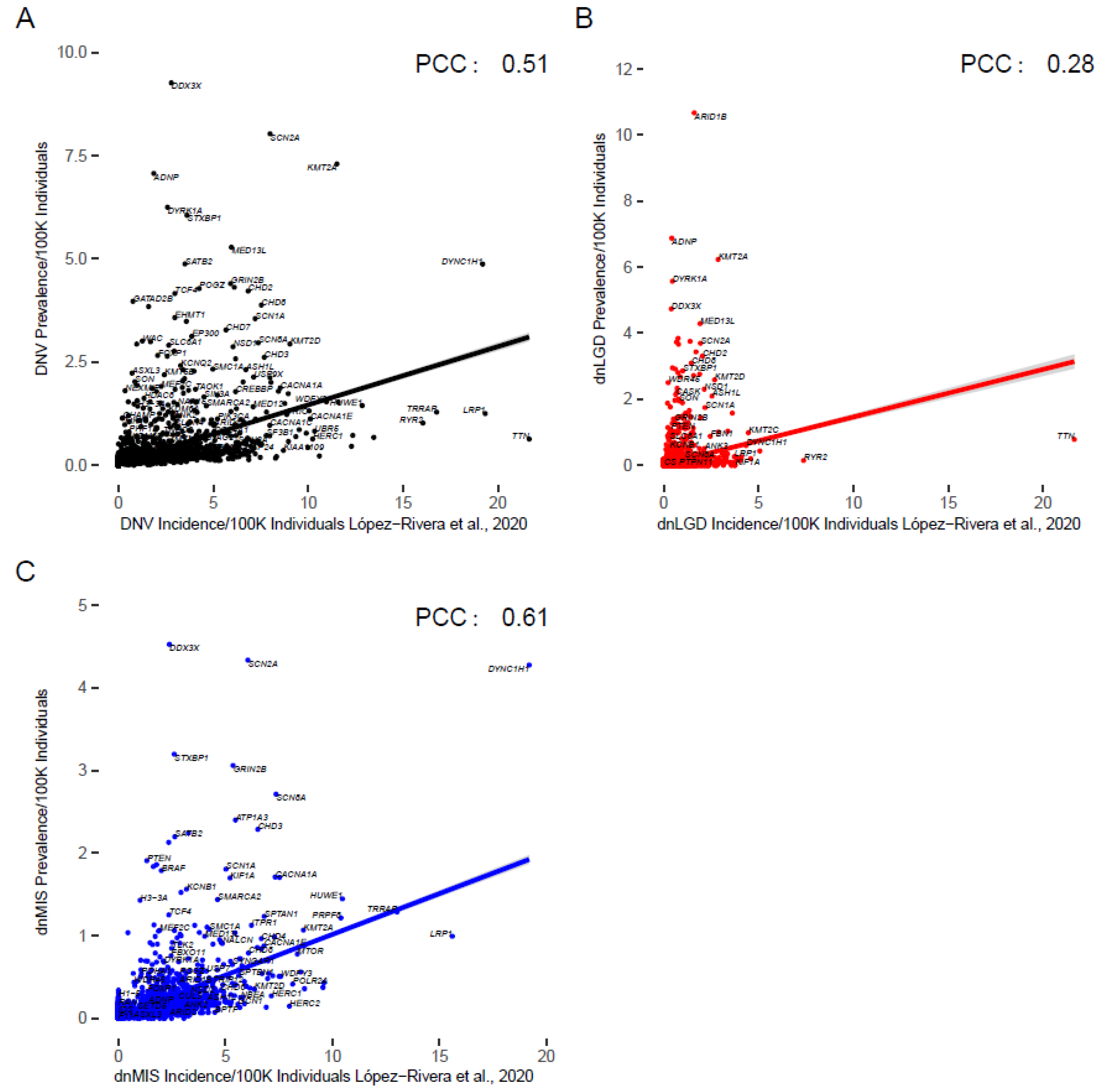
3.4. Comparison to Previous Estimates
3.5. Comparison to Social Media Groups
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rare Disease Act of 2002. Available online: https://www.congress.gov/107/plaws/publ280/PLAW-107publ280.pdf (accessed on 19 May 2022).
- Moliner, A.M.; Waligora, J. The European Union Policy in the Field of Rare Diseases. Adv. Exp. Med. Biol. 2017, 1031, 561–587. [Google Scholar] [CrossRef]
- Nguengang Wakap, S.; Lambert, D.M.; Olry, A.; Rodwell, C.; Gueydan, C.; Lanneau, V.; Murphy, D.; Le Cam, Y.; Rath, A. Estimating Cumulative Point Prevalence of Rare Diseases: Analysis of the Orphanet Database. Eur. J. Hum. Genet. 2020, 28, 165–173. Available online: https://www.nature.com/articles/s41431-019-0508-0 (accessed on 24 May 2022). [CrossRef] [PubMed]
- Zablotsky, B.; Black, L.I. Prevalence of Children Aged 3–17 Years with Developmental Disabilities, by Urbanicity: United States, 2015–2018. Natl. Health Stat. Report 2020, 139, 1–7. [Google Scholar]
- Iossifov, I.; O’Roak, B.J.; Sanders, S.J.; Ronemus, M.; Krumm, N.; Levy, D.; Stessman, H.A.; Witherspoon, K.T.; Vives, L.; Patterson, K.E.; et al. The Contribution of de Novo Coding Mutations to Autism Spectrum Disorder. Nature 2014, 515, 216–221. [Google Scholar] [CrossRef] [PubMed]
- Krumm, N.; Turner, T.N.; Baker, C.; Vives, L.; Mohajeri, K.; Witherspoon, K.; Raja, A.; Coe, B.P.; Stessman, H.A.; He, Z.-X.; et al. Excess of Rare, Inherited Truncating Mutations in Autism. Nat. Genet. 2015, 47, 582–588. [Google Scholar] [CrossRef]
- Coe, B.P.; Stessman, H.A.F.; Sulovari, A.; Geisheker, M.R.; Bakken, T.E.; Lake, A.M.; Dougherty, J.D.; Lein, E.S.; Hormozdiari, F.; Bernier, R.A.; et al. Neurodevelopmental Disease Genes Implicated by de Novo Mutation and Copy Number Variation Morbidity. Nat. Genet. 2019, 51, 106–116. [Google Scholar] [CrossRef] [PubMed]
- Kaplanis, J.; Samocha, K.E.; Wiel, L.; Zhang, Z.; Arvai, K.J.; Eberhardt, R.Y.; Gallone, G.; Lelieveld, S.H.; Martin, H.C.; McRae, J.F.; et al. Evidence for 28 genetic disorders discovered by combining healthcare and research data. Nature 2020, 586, 757–762. [Google Scholar] [CrossRef]
- Satterstrom, F.K.; Kosmicki, J.A.; Wang, J.; Breen, M.S.; De Rubeis, S.; An, J.-Y.; Peng, M.; Collins, R.; Grove, J.; Klei, L.; et al. Large-Scale Exome Sequencing Study Implicates Both Developmental and Functional Changes in the Neurobiology of Autism. Cell 2020, 180, 568–584.e23. [Google Scholar] [CrossRef]
- Titgemeyer, S.C.; Schaaf, C.P. Facebook Support Groups for Pediatric Rare Diseases: Cross-Sectional Study to Investigate Opportunities, Limitations, and Privacy Concerns. JMIR Pediatr. Parent. 2022, 5, e31411. [Google Scholar] [CrossRef]
- Gillentine, M.A.; Lupo, P.J.; Stankiewicz, P.; Schaaf, C.P. An Estimation of the Prevalence of Genomic Disorders Using Chromosomal Microarray Data. J. Hum. Genet. 2018, 63, 795–801. [Google Scholar] [CrossRef]
- Shourick, J.; Wack, M.; Jannot, A.-S. Assessing Rare Diseases Prevalence Using Literature Quantification. Orphanet J. Rare Dis. 2021, 16, 139. [Google Scholar] [CrossRef]
- Samocha, K.E.; Robinson, E.B.; Sanders, S.J.; Stevens, C.; Sabo, A.; McGrath, L.M.; Kosmicki, J.A.; Rehnström, K.; Mallick, S.; Kirby, A.; et al. A Framework for the Interpretation of de Novo Mutation in Human Disease. Nat. Genet. 2014, 46, 944–950. [Google Scholar] [CrossRef]
- López-Rivera, J.A.; Pérez-Palma, E.; Symonds, J.; Lindy, A.S.; McKnight, D.A.; Leu, C.; Zuberi, S.; Brunklaus, A.; Møller, R.S.; Lal, D. A Catalogue of New Incidence Estimates of Monogenic Neurodevelopmental Disorders Caused by de Novo Variants. Brain 2020, 143, 1099–1105. [Google Scholar] [CrossRef]
- Wang, T.; Hoekzema, K.; Vecchio, D.; Wu, H.; Sulovari, A.; Coe, B.P.; Gillentine, M.A.; Wilfert, A.B.; Perez-Jurado, L.A.; Kvarnung, M.; et al. Large-Scale Targeted Sequencing Identifies Risk Genes for Neurodevelopmental Disorders. Nat. Commun. 2020, 11, 4932. [Google Scholar] [CrossRef]
- Yuen, C.R.K.; Merico, D.; Bookman, M.; Howe, L.J.; Thiruvahindrapuram, B.; Patel, R.V.; Whitney, J.; Deflaux, N.; Bingham, J.; Wang, Z.; et al. Whole Genome Sequencing Resource Identifies 18 New Candidate Genes for Autism Spectrum Disorder. Nat. Neurosci. 2017, 20, 602–611. [Google Scholar] [CrossRef]
- De Rubeis, S.; He, X.; Goldberg, A.P.; Poultney, C.S.; Samocha, K.; Cicek, A.E.; Kou, Y.; Liu, L.; Fromer, M.; Walker, S.; et al. Synaptic, Transcriptional and Chromatin Genes Disrupted in Autism. Nature 2014, 515, 209–215. [Google Scholar] [CrossRef]
- McRae, J.F.; Clayton, S.; Fitzgerald, T.W.; Kaplanis, J.; Prigmore, E.; Rajan, D.; Sifrim, A.; Aitken, S.; Akawi, N.; Alvi, M. Deciphering Developmental Disorders Study Prevalence and Architecture of de Novo Mutations in Developmental Disorders. Nature 2017, 542, 433–438. [Google Scholar] [CrossRef]
- Epi4K Consortium; Epilepsy Phenome/Genome Project; Allen, A.S.; Berkovic, S.F.; Cossette, P.; Delanty, N.; Dlugos, D.; Eichler, E.E.; Epstein, M.P.; Glauser, T.; et al. De Novo Mutations in Epileptic Encephalopathies. Nature 2013, 501, 217–221. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Duyzend, M.H.; Coe, B.P.; Baker, C.; Hoekzema, K.; Gerdts, J.; Turner, T.N.; Zody, M.C.; Beighley, J.S.; Murali, S.C.; et al. Genome Sequencing Identifies Multiple Deleterious Variants in Autism Patients with More Severe Phenotypes. Genet. Med. 2019, 21, 1611–1620. [Google Scholar] [CrossRef] [PubMed]
- Lelieveld, S.H.; Reijnders, M.R.F.; Pfundt, R.; Yntema, H.G.; Kamsteeg, E.-J.; de Vries, P.; de Vries, B.B.A.; Willemsen, M.H.; Kleefstra, T.; Löhner, K.; et al. Meta-Analysis of 2,104 Trios Provides Support for 10 New Genes for Intellectual Disability. Nat. Neurosci. 2016, 19, 1194–1196. [Google Scholar] [CrossRef] [PubMed]
- Michaelson, J.J.; Shi, Y.; Gujral, M.; Zheng, H.; Malhotra, D.; Jin, X.; Jian, M.; Liu, G.; Greer, D.; Bhandari, A.; et al. Whole-Genome Sequencing in Autism Identifies Hot Spots for de Novo Germline Mutation. Cell 2012, 151, 1431–1442. [Google Scholar] [CrossRef] [PubMed]
- Rauch, A.; Wieczorek, D.; Graf, E.; Wieland, T.; Endele, S.; Schwarzmayr, T.; Albrecht, B.; Bartholdi, D.; Beygo, J.; Di Donato, N.; et al. Range of Genetic Mutations Associated with Severe Non-Syndromic Sporadic Intellectual Disability: An Exome Sequencing Study. Lancet 2012, 380, 1674–1682. [Google Scholar] [CrossRef]
- Takata, A.; Miyake, N.; Tsurusaki, Y.; Fukai, R.; Miyatake, S.; Koshimizu, E.; Kushima, I.; Okada, T.; Morikawa, M.; Uno, Y.; et al. Integrative Analyses of De Novo Mutations Provide Deeper Biological Insights into Autism Spectrum Disorder. Cell Rep. 2018, 22, 734–747. [Google Scholar] [CrossRef] [PubMed]
- Turner, T.N.; Coe, B.P.; Dickel, D.E.; Hoekzema, K.; Nelson, B.J.; Zody, M.C.; Kronenberg, Z.N.; Hormozdiari, F.; Raja, A.; Pennacchio, L.A.; et al. Genomic Patterns of De Novo Mutation in Simplex Autism. Cell 2017, 171, 710–722.e12. [Google Scholar] [CrossRef]
- Wang, T.; Kim, C.; Bakken, T.E.; Gillentine, M.A.; Henning, B.; Mao, Y.; Gilissen, C.; Consortium, T.S.; Nowakowski, T.J.; Eichler, E.E. Integrated Gene Analyses of de Novo Mutations from 46,612 Trios with Autism and Developmental Disorders. bioRxiv 2021. Available online: https://www.biorxiv.org/content/10.1101/2021.09.15.460398v1 (accessed on 6 May 2022).
- Buxbaum, J.D.; Bolshakova, N.; Brownfeld, J.M.; Anney, R.J.; Bender, P.; Bernier, R.; Cook, E.H.; Coon, H.; Cuccaro, M.; Freitag, C.M.; et al. The Autism Simplex Collection: An International, Expertly Phenotyped Autism Sample for Genetic and Phenotypic Analyses. Mol. Autism 2014, 5, 34. [Google Scholar] [CrossRef]
- Fischbach, G.D.; Lord, C. The Simons Simplex Collection: A Resource for Identification of Autism Genetic Risk Factors. Neuron 2010, 68, 192–195. [Google Scholar] [CrossRef]
- Feliciano, P.; Zhou, X.; Astrovskaya, I.; Turner, T.N.; Wang, T.; Brueggeman, L.; Barnard, R.; Hsieh, A.; Snyder, L.G.; Muzny, D.M.; et al. Exome Sequencing of 457 Autism Families Recruited Online Provides Evidence for Autism Risk Genes. NPJ Genom. Med. 2019, 4, 19. [Google Scholar] [CrossRef]
- Chen, R.; Davis, L.K.; Guter, S.; Wei, Q.; Jacob, S.; Potter, M.H.; Cox, N.J.; Cook, E.H.; Sutcliffe, J.S.; Li, B. Leveraging Blood Serotonin as an Endophenotype to Identify de Novo and Rare Variants Involved in Autism. Mol. Autism 2017, 8, 14. [Google Scholar] [CrossRef]
- Hashimoto, R.; Nakazawa, T.; Tsurusaki, Y.; Yasuda, Y.; Nagayasu, K.; Matsumura, K.; Kawashima, H.; Yamamori, H.; Fujimoto, M.; Ohi, K.; et al. Whole-Exome Sequencing and Neurite Outgrowth Analysis in Autism Spectrum Disorder. J. Hum. Genet. 2016, 61, 199–206. [Google Scholar] [CrossRef]
- Hamdan, F.F.; Srour, M.; Capo-Chichi, J.-M.; Daoud, H.; Nassif, C.; Patry, L.; Massicotte, C.; Ambalavanan, A.; Spiegelman, D.; Diallo, O.; et al. De Novo Mutations in Moderate or Severe Intellectual Disability. PLoS Genet. 2014, 10, e1004772. [Google Scholar] [CrossRef] [PubMed]
- Halvardson, J.; Zhao, J.J.; Zaghlool, A.; Wentzel, C.; Georgii-Hemming, P.; Månsson, E.; Ederth Sävmarker, H.; Brandberg, G.; Soussi Zander, C.; Thuresson, A.-C.; et al. Mutations in HECW2 Are Associated with Intellectual Disability and Epilepsy. J. Med. Genet. 2016, 53, 697–704. [Google Scholar] [CrossRef] [PubMed]
- Hamanaka, K.; Miyake, N.; Mizuguchi, T.; Miyatake, S.; Uchiyama, Y.; Tsuchida, N.; Sekiguchi, F.; Mitsuhashi, S.; Tsurusaki, Y.; Nakashima, M.; et al. Large-Scale Discovery of Novel Neurodevelopmental Disorder-Related Genes through a Unified Analysis of Single-Nucleotide and Copy Number Variants. Genome Med. 2022, 14, 40. [Google Scholar] [CrossRef]
- Chérot, E.; Keren, B.; Dubourg, C.; Carré, W.; Fradin, M.; Lavillaureix, A.; Afenjar, A.; Burglen, L.; Whalen, S.; Charles, P.; et al. Using Medical Exome Sequencing to Identify the Causes of Neurodevelopmental Disorders: Experience of 2 Clinical Units and 216 Patients. Clin. Genet. 2018, 93, 567–576. [Google Scholar] [CrossRef]
- Zhu, X.; Petrovski, S.; Xie, P.; Ruzzo, E.K.; Lu, Y.-F.; McSweeney, K.M.; Ben-Zeev, B.; Nissenkorn, A.; Anikster, Y.; Oz-Levi, D.; et al. Whole-Exome Sequencing in Undiagnosed Genetic Diseases: Interpreting 119 Trios. Genet. Med. 2015, 17, 774–781. [Google Scholar] [CrossRef]
- Helbig, K.L.; Farwell Hagman, K.D.; Shinde, D.N.; Mroske, C.; Powis, Z.; Li, S.; Tang, S.; Helbig, I. Diagnostic Exome Sequencing Provides a Molecular Diagnosis for a Significant Proportion of Patients with Epilepsy. Genet. Med. 2016, 18, 898–905. [Google Scholar] [CrossRef]
- Moreno-Ramos, O.A.; Olivares, A.M.; Haider, N.B.; de Autismo, L.C.; Lattig, M.C. Whole-Exome Sequencing in a South American Cohort Links ALDH1A3, FOXN1 and Retinoic Acid Regulation Pathways to Autism Spectrum Disorders. PLoS ONE 2015, 10, e0135927. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Deignan, J.L.; Dorrani, N.; Strom, S.P.; Kantarci, S.; Quintero-Rivera, F.; Das, K.; Toy, T.; Harry, B.; Yourshaw, M.; et al. Clinical Exome Sequencing for Genetic Identification of Rare Mendelian Disorders. JAMA 2014, 312, 1880–1887. [Google Scholar] [CrossRef] [PubMed]
- Tavassoli, T.; Kolevzon, A.; Wang, A.T.; Curchack-Lichtin, J.; Halpern, D.; Schwartz, L.; Soffes, S.; Bush, L.; Grodberg, D.; Cai, G.; et al. De Novo SCN2A Splice Site Mutation in a Boy with Autism Spectrum Disorder. BMC Med. Genet. 2014, 15, 35. [Google Scholar] [CrossRef]
- Werling, D.M.; Brand, H.; An, J.-Y.; Stone, M.R.; Zhu, L.; Glessner, J.T.; Collins, R.L.; Dong, S.; Layer, R.M.; Markenscoff-Papadimitriou, E.; et al. An Analytical Framework for Whole-Genome Sequence Association Studies and Its Implications for Autism Spectrum Disorder. Nat. Genet. 2018, 50, 727–736. [Google Scholar] [CrossRef]
- Veeramah, K.R.; O’Brien, J.E.; Meisler, M.H.; Cheng, X.; Dib-Hajj, S.D.; Waxman, S.G.; Talwar, D.; Girirajan, S.; Eichler, E.E.; Restifo, L.L.; et al. De Novo Pathogenic SCN8A Mutation Identified by Whole-Genome Sequencing of a Family Quartet Affected by Infantile Epileptic Encephalopathy and SUDEP. Am. J. Hum. Genet. 2012, 90, 502–510. [Google Scholar] [CrossRef] [PubMed]
- Veeramah, K.R.; Johnstone, L.; Karafet, T.M.; Wolf, D.; Sprissler, R.; Salogiannis, J.; Barth-Maron, A.; Greenberg, M.E.; Stuhlmann, T.; Weinert, S.; et al. Exome Sequencing Reveals New Causal Mutations in Children with Epileptic Encephalopathies. Epilepsia 2013, 54, 1270–1281. [Google Scholar] [CrossRef] [PubMed]
- Barcia, G.; Fleming, M.R.; Deligniere, A.; Gazula, V.-R.; Brown, M.R.; Langouet, M.; Chen, H.; Kronengold, J.; Abhyankar, A.; Cilio, R.; et al. De Novo Gain-of-Function KCNT1 Channel Mutations Cause Malignant Migrating Partial Seizures of Infancy. Nat. Genet. 2012, 44, 1255–1259. [Google Scholar] [CrossRef] [PubMed]
- de Ligt, J.; Willemsen, M.H.; van Bon, B.W.M.; Kleefstra, T.; Yntema, H.G.; Kroes, T.; Vulto-van Silfhout, A.T.; Koolen, D.A.; de Vries, P.; Gilissen, C.; et al. Diagnostic Exome Sequencing in Persons with Severe Intellectual Disability. N. Engl. J. Med. 2012, 367, 1921–1929. [Google Scholar] [CrossRef] [PubMed]
- Mignot, C.; Moutard, M.-L.; Rastetter, A.; Boutaud, L.; Heide, S.; Billette, T.; Doummar, D.; Garel, C.; Afenjar, A.; Jacquette, A.; et al. ARID1B Mutations Are the Major Genetic Cause of Corpus Callosum Anomalies in Patients with Intellectual Disability. Brain 2016, 139, e64. [Google Scholar] [CrossRef] [PubMed]
- Hoyer, J.; Ekici, A.B.; Endele, S.; Popp, B.; Zweier, C.; Wiesener, A.; Wohlleber, E.; Dufke, A.; Rossier, E.; Petsch, C.; et al. Haploinsufficiency of ARID1B, a Member of the SWI/SNF-A Chromatin-Remodeling Complex, Is a Frequent Cause of Intellectual Disability. Am. J. Hum. Genet. 2012, 90, 565–572. [Google Scholar] [CrossRef]
- Kleefstra, T.; de Leeuw, N. Kleefstra, T.; de Leeuw, N. Kleefstra Syndrome. In GeneReviews®; Adam, M.P., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Stamberger, H.; Nikanorova, M.; Willemsen, M.H.; Accorsi, P.; Angriman, M.; Baier, H.; Benkel-Herrenbrueck, I.; Benoit, V.; Budetta, M.; Caliebe, A.; et al. STXBP1 Encephalopathy: A Neurodevelopmental Disorder Including Epilepsy. Neurology 2016, 86, 954–962. [Google Scholar] [CrossRef]
- Issekutz, K.A.; Jr, J.M.G.; Prasad, C.; Smith, I.M.; Blake, K.D. An epidemiological analysis of CHARGE syndrome: Preliminary results from a Canadian study. Am. J. Med Genet. Part A 2005, 133A, 309–317. [Google Scholar] [CrossRef]
- Janssen, N.; Bergman, J.; Swertz, M.; Tranebjaerg, L.; Lodahl, M.; Schoots, J.; Hofstra, R.; van Ravenswaaij-Arts, C.M.A.; Hoefsloot, L.H. Mutation update on the CHD7 gene involved in CHARGE syndrome. Hum. Mutat. 2012, 33, 1149–1160. [Google Scholar] [CrossRef]
- Adam, M.P.; Hudgins, L.; Hannibal, M. Kabuki Syndrome. In GeneReviews®; Adam, M.P., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Tatton-Brown, K.; Cole. T/R.; Rahman, N. Sotos Syndrome. In GeneReviews®; Adam, M.P., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Adam, M.P.; Conta, J.; Bean, L.J.H. Mowat-Wilson Syndrome. In GeneReviews®; Adam, M.P., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Kaur, S.; Christodoulou, J. MECP2 Disorders. In GeneReviews®; Adam, M.P., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- A Koolen, D.; Morgan, A.; de Vries, B.B. Koolen-de Vries Syndrome. In GeneReviews®; Adam, M.P., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Koolen, A.D.; DDD Study; Pfundt, R.; Linda, K.; Beunders, G.; Veenstra-Knol, E.H.; Conta, J.H.; Fortuna, A.; Gillessen-Kaesbach, G.; Dugan, S.; et al. The Koolen-de Vries syndrome: A phenotypic comparison of patients with a 17q21.31 microdeletion versus a KANSL1 sequence variant. Eur. J. Hum. Genet. 2015, 24, 652–659. [Google Scholar] [CrossRef]
- Bayat, A.; Hjalgrim, H.; Møller, R.S. The incidence of SCN1A-related dravet syndrome in Denmark is 1:22,000: A population-based study from 2004 to 2009. Epilepsia 2015, 56. [Google Scholar] [CrossRef]
- Larsen, J.; Johannesen, K.M.; Ek, J.; Tang, S.; Marini, C.; Blichfeldt, S.; Kibaek, M.; von Spiczak, S.; Weckhuysen, S.; Frangu, M.; et al. The role of SLC2A1 mutations in myoclonic astatic epilepsy and absence epilepsy, and the estimated frequency of GLUT1 deficiency syndrome. Epilepsia 2015, 56, e203–e208. [Google Scholar] [CrossRef]
- Symonds, J.; Zuberi, S.M.; Stewart, K.; McLellan, A.; O‘Regan, M.; MacLeod, S.; Jollands, A.; Joss, S.; Kirkpatrick, M.; Brunklaus, A.; et al. Incidence and phenotypes of childhood-onset genetic epilepsies: A prospective population-based national cohort. Brain 2019, 142, 2303–2318. [Google Scholar] [CrossRef] [PubMed]
- Stevens, C.A. Rubinstein-Taybi Syndrome. In GeneReviews®; Adam, M.P., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Snijders Blok, L.; Madsen, E.; Juusola, J.; Gilissen, C.; Baralle, D.; Reijnders, M.R.F.; Venselaar, H.; Helsmoortel, C.; Cho, M.T.; Hoischen, A.; et al. Mutations in DDX3X Are a Common Cause of Unexplained Intellectual Disability with Gender-Specific Effects on Wnt Signaling. Am. J. Hum. Genet. 2015, 97, 343–352. [Google Scholar] [CrossRef] [PubMed]
- Amabile, S.; Jeffries, L.; McGrath, J.M.; Ji, W.; Spencer-Manzon, M.; Zhang, H.; Lakhani, S.A. DYNC1H1-Related Disorders: A Description of Four New Unrelated Patients and a Comprehensive Review of Previously Reported Variants. Am. J. Med. Genet. Part A 2020, 182, 2049–2057. [Google Scholar] [CrossRef] [PubMed]
- The Power of Being Counted—RARE-X. 2022. Available online: https://rare-x.org/case-studies/the-power-of-being-counted/ (accessed on 8 June 2022).
- van der Sluijs, P.J.; Jansen, S.; Vergano, S.A.; Adachi-Fukuda, M.; Alanay, Y.; AlKindy, A.; Baban, A.; Bayat, A.; Beck-Wödl, S.; Berry, K.; et al. The ARID1B Spectrum in 143 Patients: From Nonsyndromic Intellectual Disability to Coffin-Siris Syndrome. Genet. Med. 2019, 21, 1295–1307. [Google Scholar] [CrossRef]
- Vasko, A.; Drivas, T.G.; Schrier Vergano, S.A. Genotype-Phenotype Correlations in 208 Individuals with Coffin-Siris Syndrome. Genes 2021, 12, 937. [Google Scholar] [CrossRef]
- Schrier Vergano, S.; Santen, G.; Wieczorek, D.; Wollnik, B.; Matsumoto, N.; Deardorff, M.A. Coffin-Siris Syndrome. In GeneReviews®; Adam, M.P., Ardinger, H.H., Pagon, R.A., Wallace, S.E., Bean, L.J., Gripp, K.W., Mirzaa, G.M., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Lauer, E.; McCallion, P. Mortality of People with Intellectual and Developmental Disabilities from Select US State Disability Service Systems and Medical Claims Data. J. Appl. Res. Intellect. Disabil. 2015, 28, 394–405. [Google Scholar] [CrossRef]
- Srivastava, S.; Love-Nichols, J.A.; Dies, K.A.; Ledbetter, D.H.; Martin, C.L.; Chung, W.K.; Firth, H.V.; Frazier, T.; Hansen, R.L.; Prock, L.; et al. Meta-Analysis and Multidisciplinary Consensus Statement: Exome Sequencing Is a First-Tier Clinical Diagnostic Test for Individuals with Neurodevelopmental Disorders. Genet. Med. 2019, 21, 2413–2421. [Google Scholar] [CrossRef]
| Zablotsky and Black (2020) Prevalence % | 95% Confidence Interval | Current Study n | |
|---|---|---|---|
| All NDDs | 4.5% | 4–5% | 50,377 |
| DD/ID | 1.2% | 1.1–1.4% | 31,191 |
| ASD | 2.5% | 2.2–2.7% | 16,125 |
| Epilepsy | 0.8% | 0.7–0.9% | 1389 |
| NDDs (Prevalence/100,000; % in Cohort) | |||
|---|---|---|---|
| DNV | dnLGD | dnMIS | dnMIS30 |
| ARID1B (11.1; 0.3%) | ARID1B (10.7; 0.25%) | DDX3X (4.5; 0.14%) | DYNC1H1 (2.5; 0.09%) |
| DDX3X (9.3; 0.3%) | ADNP (6.9; 0.16%) | SCN2A (4.3; 0.17%) | DDX3X (1.8; 0.06%) |
| SCN2A (8; 0.3%) | KMT2A (6.2; 0.14%) | DYNC1H1 (4.3; 0.16%) | STXBP1 (1.7; 0.06%) |
| KMT2A (7.3; 0.22%) | DYRK1A (5.6; 0.13%) | STXBP1 (3.2; 0.11%) | SCN2A (1.3; 0.05%) |
| ADNP (7.1; 0.17%) | CTNNB1 (5.5, 0.13%) | GRIN2B (3.1; 0.13%) | SLC6A1 (1.3; 0.05%) |
| DYRK1A (6.2.; 0.18%) | DDX3X (4.7; 0.11%) | SCN8A (2.7; 0.09%) | SATB2 (1.1; 0.04%) |
| STXBP1 (6.1; 0.23%) | MED13L (4.3; 0.1%) | ATP1A3 (2.4; 0.08%) | CHD3 (1; 0.05%) |
| CTNNB1 (5.7; 0.13%) | GATAD2B (3.8; 0.09%) | CHD3 (2.3, 0.1%) | SMARCA4 (1; 0.04%) |
| MED13L (5.3; 0.18%) | POGZ (3.7; 0.09%) | KCNQ2 (2.2, 0.1%) | KIF1A (1; 0.05%) |
| SATB2 (4.9; 0.19%) | SETD5 (3.7; 0.1%) | SATB2 (2.2, 0.09%) | GRIN2B (1; 0.4%) |
| Gene/Syndrome | Current Cohort Estimate (All NDDs) | López-Rivera Estimate | Previous Estimates (Citation) |
|---|---|---|---|
| ARID1B/Coffin-Siris syndrome 1 | Most are due to LGD variants: 1/9009 | Most are due to LGD variants: 1/61,884 | 1/10,000–1/100,000 [47] |
| EHMT1/KMT2C/Kleefstra syndrome | LGD and MIS: EHMT1: 1/27,927 KMT2C: 1/90,759 | LGD and MIS: 1/33,686 LGD and MIS: 1/22,373 | At least 1/120,000 in those with NDDs [48] |
| STXBP1/STXBP1 encephalopathy | LGD and MIS: 1/16,516 | LGD and MIS: 1/27,664 | 1/91,862 [49] |
| CHD7/CHARGE syndrome | LGD and MIS: 1/30,513 | LGD and MIS: 1/17,642 | 1/8500–1/15,000 newborns [50,51] |
| KMT2D/KDM6A/Kabuki syndrome | LGD and MIS: KMT2D: 1/34,054 KDM6A: 1/77,272 | LGD and MIS: KMT2D: 1/11,061 KDM6A: 1/38,153 | 1/32,000–1/86,000 [52] |
| NSD1/Sotos syndrome | LGD and MIS: 1/34,843 | LGD and MIS: 1/16,552.5 | 1/14,000 [53] |
| ZEB2/Mowat-Wilson syndrome | LGD and MIS: 1/43,906 | LGD and MIS: 1/25,112 | 1/50,000–1/100,000 [54] |
| MECP2/Rett syndrome | LGD and MIS: 1/87,826 | LGD and MIS: 1/486,085 | 1/10,000–1/23,000 female births [55] |
| KANSL1/Koolen-de Vries syndrome syndrome | LGD and MIS: 1/65,049 | LGD and MIS: 1/59,802 | May be as frequent as deletion (1/55,000) [56,57] |
| SCN1A/Dravet syndrome | LGD and MIS: 1/28,161 | LGD and MIS: 1/13,877 | 1/22,000 incidence in Danish population [58] |
| SLC2A1/GLUT1 deficiency syndrome | MIS: 1/295,848 | MIS: 1/58,766.5 | 1/33,898–1/83,333 [59,60] |
| KCNQ2/KCNQ2 encephalopathy | MIS: 1/44,573 | MIS: 1/30,534 | 1/84,746 [60] |
| CREBBP/EP300/Rubinstein-Taybi syndrome | LGD and MIS: CREBBP: 1/56,126 EP300: 1/32,028 | LGD and MIS: CREBBP: 1/16,201 EP300: 1/25,862 | 1/100,000–1/125,000 [61] |
| DD (Prevalence/100,000; % in Cohort) | |||
|---|---|---|---|
| DNV | dnLGD | dnMIS | dnMIS30 |
| ARID1B (4; 0.38%) | ARID1B (3.9; 0.2) | DDX3X (1.7; 0.2%) | DYNC1H1 (1; 0.13%) |
| DDX3X (3.6; 0.37%) | KMT2A (2.3; 0.18%) | DYNC1H1 (1.4; 0.2%) | DDX3X (0.8; 0.09%) |
| KMT2A (2.7; 0.28%) | CTNNB1 (2.2; 0.18%) | SCN2A (1.2; 0.2%) | STXBP1 (0.54; 0.07%) |
| DYRK1A (2.4; 0.25%) | DYRK1A (2.19; 0.17%) | GRIN2B (1; 0.18%) | SLC6A1 (0.5; 0.07%) |
| CTNNB1 (2.2; 0.2%) | ADNP (2.1, 0.16%) | STXBP1 (0.96; 0.14%) | SATB2 (0.5; 0.07%) |
| ADNP (2.1; 0.19%) | DDX3X (1.96; 0.13%) | SCN8A (0.9; 0.12%) | KIF1A (0.4; 0.08%) |
| STXBP1 (2.1; 0.23%) | MED13L (1.7; 0.13%) | KCNQ2 (0.8; 0.15%) | CHD3 (0.4; 0.06%) |
| SCN2A (2.1; 0.28%) | GATAD2B (1.6; 0.12%) | CHD3 (0.8; 0.14%) | ATP1A3 (0.4; 0.04%) |
| MED13L (2; 0.24%) | SETD5 (1.4; 0.12%) | SATB2 (0.7; 0.12%) | CACNA1A (0.4; 0.07%) |
| SATB2 (1.9; 0.21%) | EHMT1 (1.3; 0.11%) | ATP1A3 (0.7; 0.1%) | SMARCA4 (0.3; 0.05%) |
| ASD (Prevalence/100,000; % in Cohort) | |||
|---|---|---|---|
| DNV | dnLGD | dnMIS | dnMIS30 |
| SCN2A (4.1; 0.22%) | ADNP (3, 0.12%) | SCN2A (1.7; 0.12%) | SCN2A (0.7; 0.05%) |
| CHD8 (3.5, 0.19%) | CHD8 (2.7; 0.11%) | PTEN (1.2; 0.07%) | DYNC1H1 (0.5; 0.03%) |
| ADNP (3.2; 0.16%) | SCN2A (2.3; 0.1%) | DYNC1H1 (1.1; 0.07%) | STXBP1 (0.4; 0.03%) |
| ASH1L (2.3; 0.1%) | ASH1L (2.3; 0.09%) | CHD8 (0.8; 0.07%) | TRRAP (0.4, 0.03%) |
| POGZ (2.2; 0.12%) | ARID1B (1.8; 0.07%) | TRRAP (0.7; 0.06%) | CHD8 (0.3; 0.03%) |
| CHD2 (2.2; 0.12%) | POGZ (1.8; 0.07%) | CHD2 (0.65; 0.06%) | GRIN2B (0.3; 0.03%) |
| ARID1B (2.1; 0.14%) | KMT5B (1.6; 0.06%) | NALCN (0.6; 0.06%) | CLCN4 (0.3; 0.02%) |
| DYNC1H1 (2; 0.11%) | CHD2 (1.5; 0.06%) | PABPC1 (0.6; 0.04%) | CHD2 (0.3; 0.02%) |
| WDFY3 (1.7; 0.1%) | SYNGAP1 (1.2; 0.05%) | CYFIP2 (0.58; 0.04%) | SLC6A1 (0.28; 0.02%) |
| KMT5B (1.7; 0.08%) | WDFY3 (1.2; 0.05%) | TBL1XR1 (0.56; 0.03%) | SMARCA4 (0.27; 0.02%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gillentine, M.A.; Wang, T.; Eichler, E.E. Estimating the Prevalence of De Novo Monogenic Neurodevelopmental Disorders from Large Cohort Studies. Biomedicines 2022, 10, 2865. https://doi.org/10.3390/biomedicines10112865
Gillentine MA, Wang T, Eichler EE. Estimating the Prevalence of De Novo Monogenic Neurodevelopmental Disorders from Large Cohort Studies. Biomedicines. 2022; 10(11):2865. https://doi.org/10.3390/biomedicines10112865
Chicago/Turabian StyleGillentine, Madelyn A., Tianyun Wang, and Evan E. Eichler. 2022. "Estimating the Prevalence of De Novo Monogenic Neurodevelopmental Disorders from Large Cohort Studies" Biomedicines 10, no. 11: 2865. https://doi.org/10.3390/biomedicines10112865
APA StyleGillentine, M. A., Wang, T., & Eichler, E. E. (2022). Estimating the Prevalence of De Novo Monogenic Neurodevelopmental Disorders from Large Cohort Studies. Biomedicines, 10(11), 2865. https://doi.org/10.3390/biomedicines10112865





