osr1 Maintains Renal Progenitors and Regulates Podocyte Development by Promoting wnt2ba via the Antagonism of hand2
Abstract
1. Introduction
2. Materials and Methods
2.1. Creation and Maintenance of Zebrafish Lines
2.2. Live Imaging and Dextran Injections
2.3. WISH, FISH, IF, Sectioning and Image Acquisition
2.4. Genotyping
2.5. Morpholinos and RT-PCR
2.6. Statistics and Measurements
3. Results
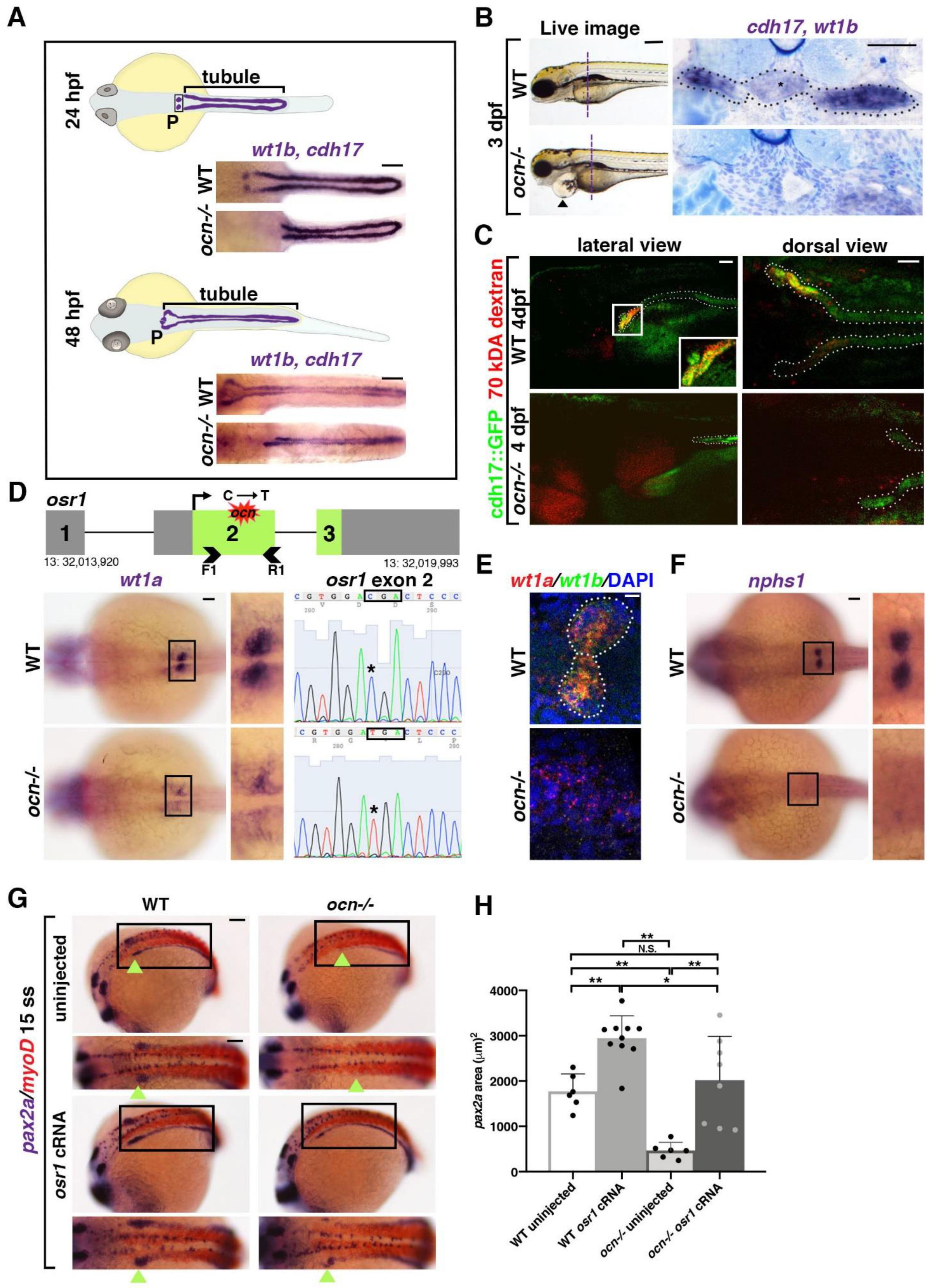
3.1. ocn Encodes a Premature Stop Codon in osr1 and Mutants Exhibits Defective Podocyte and Pronephric Tubule Development
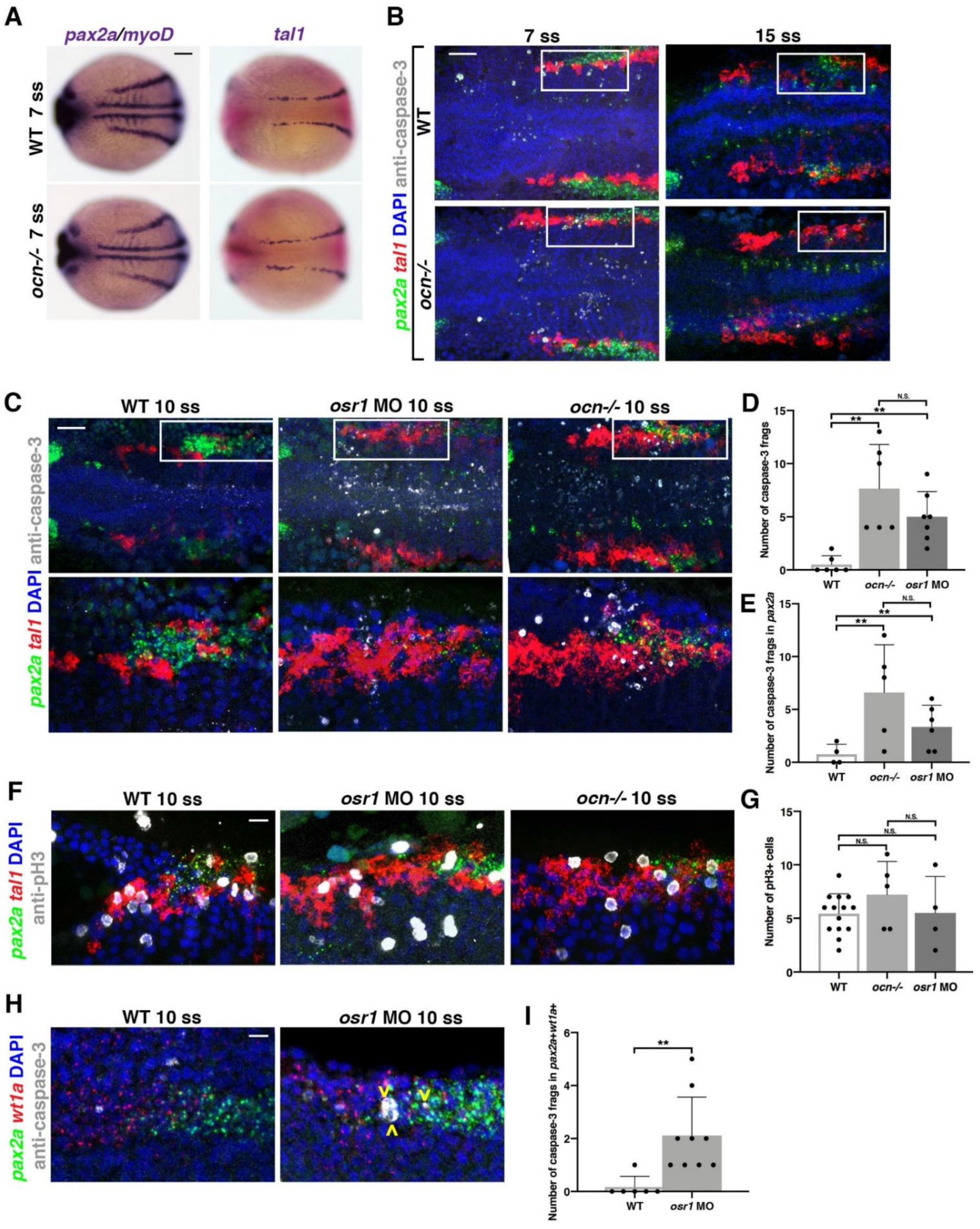
3.2. Kidney Progenitors Are Specified in osr1 Deficient Animals, but Subsequently Undergo Apoptosis
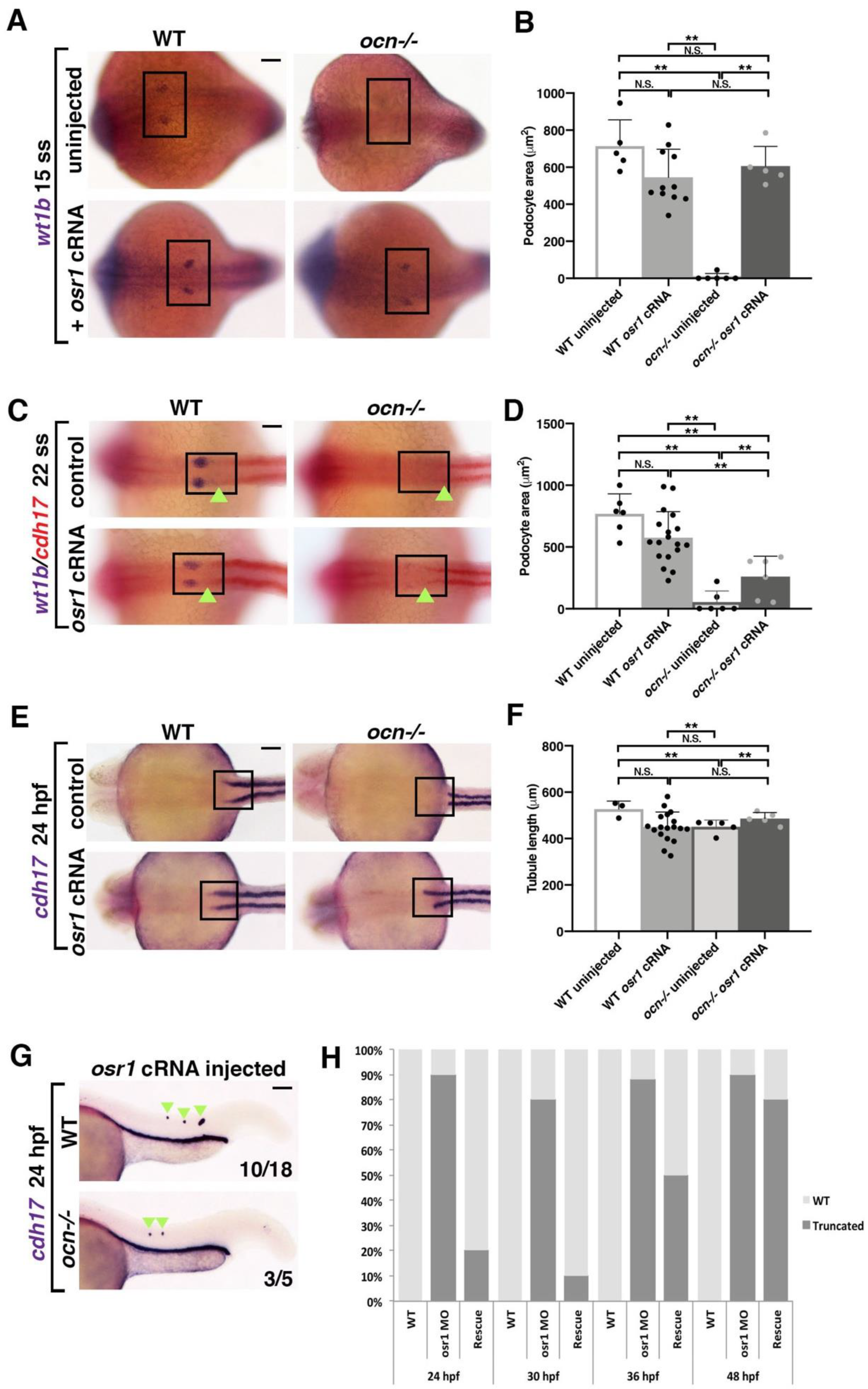
3.3. Ectopic osr1 Is Transiently Sufficient to Rescue Renal Progenitors
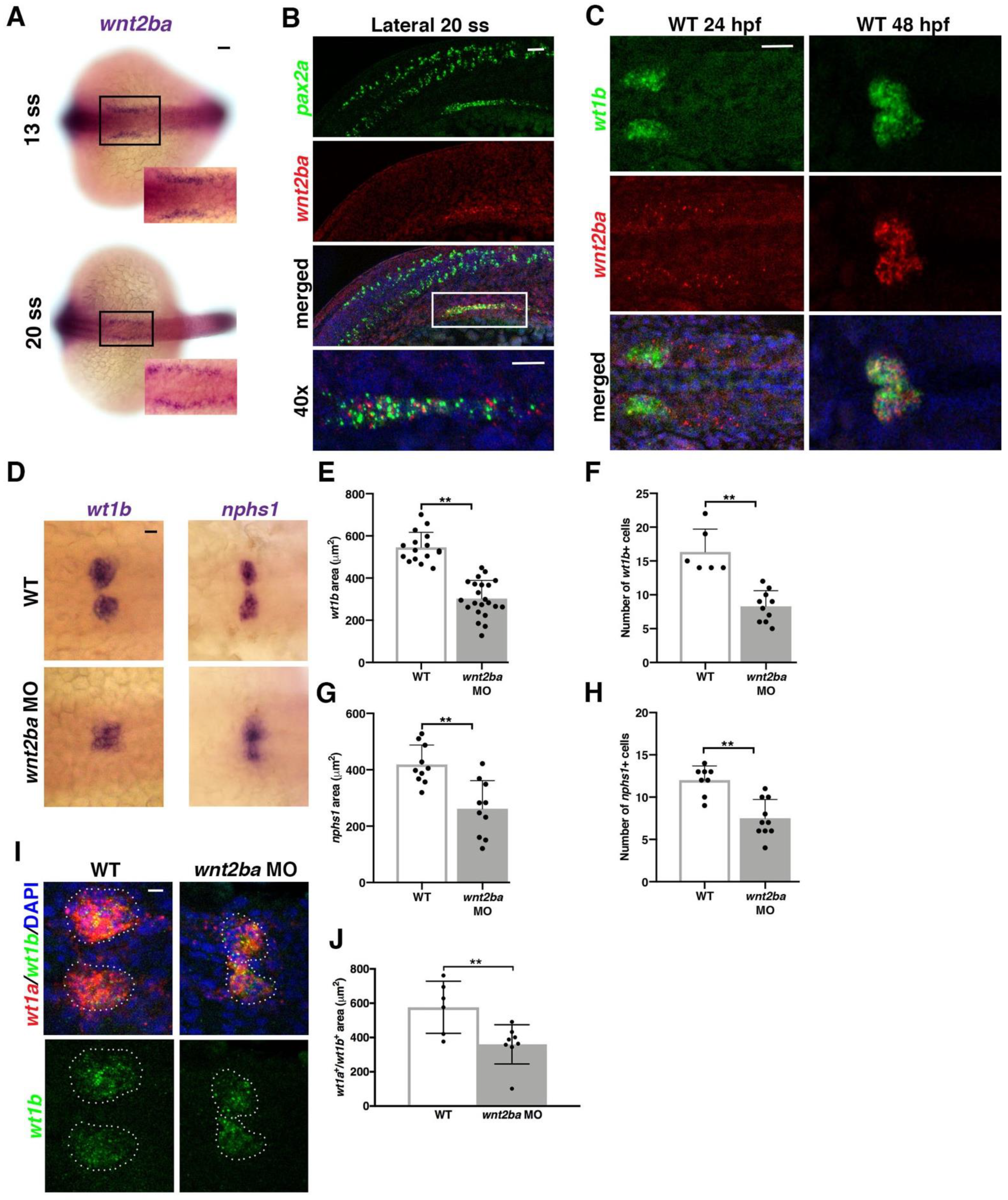
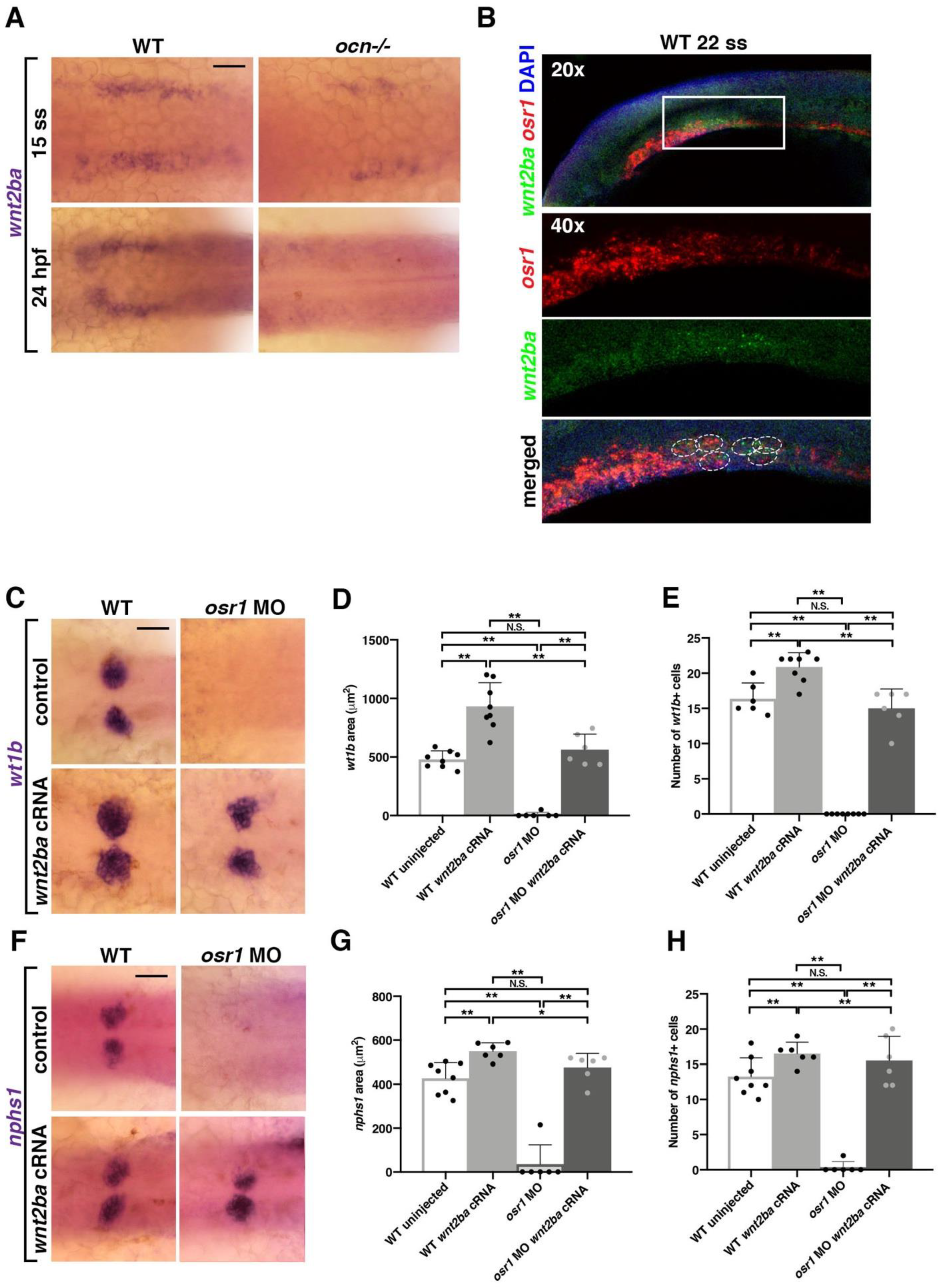
3.4. wnt2ba Is a Novel Podocyte Marker and Regulator
3.5. osr1 Promotes wnt2ba in the Podocyte Developmental Pathway
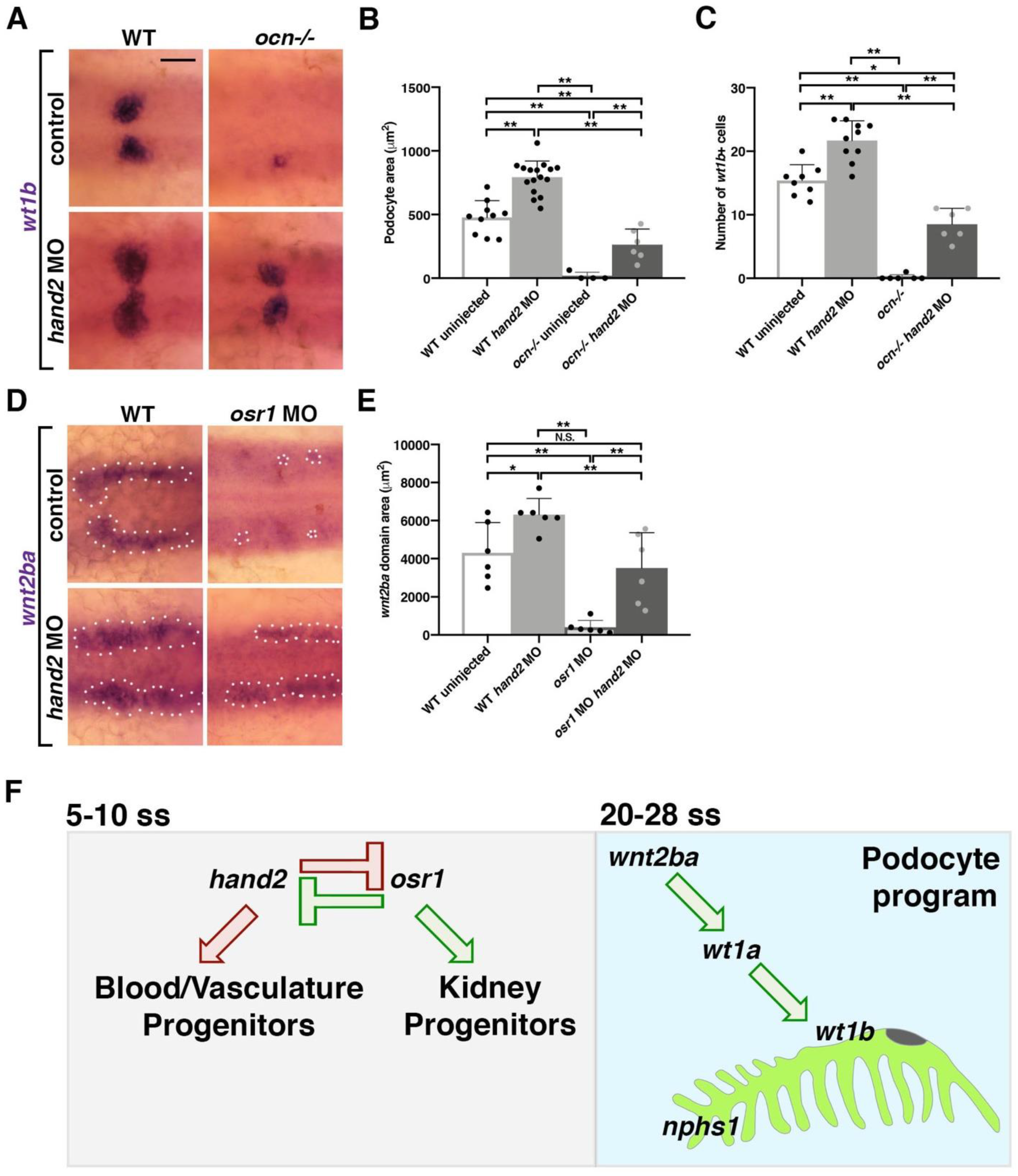
3.6. hand2 Suppresses Podocyte Development by Restricting wnt2ba Expression and Podocyte Development
4. Discussion
4.1. Osr1 Acts to Promote Podocytes
4.2. Osr1 Is Needed for Kidney Cell Maintenance
4.3. Wnt2ba Is a Novel Regulator of Podocyte Development
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Ichimura, K.; Kakuta, S.; Kawasaki, Y.; Miyaki, T.; Nonami, T.; Miyazaki, N.; Nakao, T.; Enomoto, S.; Arai, S.; Koike, M.; et al. Morphological process of podocyte development revealed by block-face scanning electron microscopy. J. Cell Sci. 2017, 130, 132–142. [Google Scholar] [CrossRef] [PubMed]
- Grahammer, F. New structural insights into podocyte biology. Cell Tissue Res. 2017, 369, 5–10. [Google Scholar] [CrossRef] [PubMed]
- Pavenstädt, H.; Kriz, W.; Kretzler, M. Cell biology of the glomerular podocyte. Physiol Rev. 2003, 83, 253–307. [Google Scholar] [CrossRef]
- Garg, P. A review of podocyte biology. Am. J. Nephrol. 2018, 47, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Zhuo, J.L.; Li, X.C. 2013. Proximal nephron. Compr. Physiol. 2013, 3, 1079. [Google Scholar] [PubMed]
- Wiggins, R.C. The spectrum of podocytopathies: A unifying view of glomerular diseases. Kidney Int. 2007, 71, 1205–1214. [Google Scholar] [CrossRef]
- Romagnani, P.; Remuzzi, G.; Glassock, R.; Levin, A.; Jager, K.J.; Tonelli, M.; Massy, Z.; Wanner, C.; Anders, H. Chronic kidney disease. Nat. Rev. Dis. Primers 2017, 3, 17088. [Google Scholar] [CrossRef]
- Reiser, J.; Sever, S. Podocyte biology and pathogenesis of kidney disease. Annu. Rev. Med. 2013, 64, 357–366. [Google Scholar] [CrossRef]
- Gerlach, G.F.; Wingert, R.A. Kidney organogenesis in the zebrafish: Insights into vertebrate nephrogenesis and regeneration. Wiley Interdiscip. Rev. Dev. Biol. 2013, 2, 559–585. [Google Scholar] [CrossRef]
- Perens, E.A.; Garavito-Aguilar, Z.V.; Guio-Vega, G.P.; Peña, K.T.; Schindler, Y.L.; Yelon, D. Hand2 inhibits kidney specification while promoting vein formation within the posterior mesoderm. eLife 2016, 5, e19941. [Google Scholar] [CrossRef]
- Little, M.H.; McMahon, A.P. Mammalian kidney development: Principles, progress, and projections. Cold Spring Harb. Perspect. Biol. 2012, 4, a008300. [Google Scholar] [CrossRef] [PubMed]
- McMahon, A.P. Development of the Mammalian Kidney. Curr. Top. Dev. Biol. 2016, 117, 31. [Google Scholar] [PubMed]
- Luyckx, V.A.; Shukha, K.; Brenner, B.M. Low nephron number and its clinical consequences. Rambam Maimonides Med. J. 2011, 2, e0061. [Google Scholar] [CrossRef] [PubMed]
- Little, M.H. Growing kidney tissue from stem cells: How far from "party trick" to medical application? Cell Stem Cell 2016, 18, 695–698. [Google Scholar] [CrossRef] [PubMed]
- Drummond, B.E.; Wingert, R.A. Insights into kidney stem cell development and regeneration using zebrafish. World J. Stem Cells 2016, 8, 22–31. [Google Scholar] [CrossRef]
- Drummond, I.A.; Davidson, A.J. Zebrafish kidney development. Methods Cell Biol. 2016, 134, 391–429. [Google Scholar]
- Diep, C.Q.; Ma, D.; Deo, R.C.; Holm, T.M.; Naylor, R.W.; Arora, N.; Wingert, R.A.; Bollig, F.; Djordjevic, G.; Lichman, B.; et al. Identification of adult nephron progenitors capable of kidney regeneration in zebrafish. Nature 2011, 470, 95–100. [Google Scholar] [CrossRef]
- Naylor, R.W.; Przepiorski, A.; Ren, Q. HNF1B is essential for nephron segmentation during nephrogenesis. J. Am. Soc. Nephrol. 2013, 24, 77–87. [Google Scholar] [CrossRef]
- Hsu, H.; Lin, G.; Chung, B. Parallel early development of zebrafish interrenal glands and pronephros: Differential control by Wt1 and Ff1b. Development 2003, 130, 2107–2116. [Google Scholar] [CrossRef]
- Bollig, F.; Mehringer, R.; Perner, B.; Hartung, C.; Schäfer, M.; Schartl, M.; Volff, J.; Winkler, C.; Englert, C. Identification and comparative expression analysis of a second Wt1 gene in zebrafish. Dev. Dyn. 2006, 235, 554. [Google Scholar] [CrossRef]
- O’Brien, L.L.; Grimaldi, M.; Kostun, Z.; Wingert, R.A.; Selleck, R.; Davidson, A.J. Wt1a, Foxc1a, and the Notch Mediator Rbpj physically interact and regulate the formation of podocytes in zebrafish. Dev. Biol. 2011, 358, 318–330. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Chen, Z.; Zeng, C.; Wang, L.; Xu, F.; Hou, Q.; Liu, Z. Ultrastructural characterization of the pronephric glomerulus development in zebrafish. J. Morphol. 2016, 277, 1104–1112. [Google Scholar] [CrossRef] [PubMed]
- Wingert, R.A.; Selleck, R.; Yu, J.; Song, H.; Chen, Z.; Song, A.; Zhou, Y.; Thisse, B.; Thisse, C.; McMahon, A.P.; et al. The cdx genes and retinoic acid control the positioning and segmentation of the zebrafish pronephros. PLoS Genet. 2007, 3, 1922–1938. [Google Scholar] [CrossRef] [PubMed]
- Wingert, R.A.; Davidson, A.J. Zebrafish nephrogenesis involves dynamic spatiotemporal expression changes in renal progenitors and essential signals from retinoic acid and irx3b. Dev. Dyn. 2011, 240, 2011–2027. [Google Scholar] [CrossRef] [PubMed]
- Desgrange, A.; Cereghini, S. Nephron patterning: Lessons from Xenopus, zebrafish, and mouse studies. Cells 2015, 4, 483–499. [Google Scholar] [CrossRef]
- Mugford, J.W.; Sipilä, P.; McMahon, J.A.; McMahon, A.P. Osr1 expression demarcates a multi-potent population of intermediate mesoderm that undergoes progressive restriction to an Osr1-dependent nephron progenitor compartment within the mammalian kidney. Dev. Biol. 2008, 324, 88–98. [Google Scholar] [CrossRef]
- James, R.G.; Kamei, C.N.; Wang, Q.; Jiang, R.; Schultheiss, T.M. Odd-Skipped related 1 is required for development of the metanephric kidney and regulates formation and differentiation of kidney precursor cells. Development 2006, 133, 2995–3004. [Google Scholar] [CrossRef]
- Wang, Q.; Lan, Y.; Cho, E.; Maltby, K.M.; Jiang, R. Odd-Skipped Related 1 (Odd 1) is an essential regulator of heart and urogenital development. Dev. Biol. 2005, 288, 582–594. [Google Scholar] [CrossRef]
- Tena, J.J.; Neto, A.; de la Calle-Mustienes, E.; Bras-Pereira, C.; Casares, F.; Gómez-Skarmeta, J.L. Odd-Skipped genes encode repressors that control kidney development. Dev. Biol. 2007, 301, 518–531. [Google Scholar] [CrossRef]
- Mudumana, S.P.; Hentschel, D.; Liu, Y.; Vasilyev, A.; Drummond, I.A. Odd skipped related1 reveals a novel role for endoderm in regulating kidney versus vascular cell fate. Development 2008, 135, 3355–3367. [Google Scholar] [CrossRef]
- Tomar, R.; Mudumana, S.P.; Pathak, N.; Hukriede, N.A.; Drummond, I.A. Osr1 is Required for Podocyte Development Downstream of Wt1a. J. Am. Soc. Neph. 2014, 25, 2539–2545. [Google Scholar] [CrossRef] [PubMed]
- Neto, A.; Mercader, N.; Gómez-Skarmeta, J.L. The Osr1 and Osr2 genes act in the pronephric anlage downstream of retinoic acid signaling and upstream of Wnt2b to maintain pectoral fin development. Development 2012, 139, 301–311. [Google Scholar] [CrossRef] [PubMed]
- Perens, E.A.; Diaz, J.T.; Quesnel, A.; Crump, J.G.; Yelon, D. osr1 couples intermediate mesoderm cell fate with temporal dynamics of vessel progenitor cell differentiation. Development 2021, 148, dev198408. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Iglesias, D.; Eliopoulos, N.; El Kares, R.; Chu, L.; Romagnani, P.; Goodyer, P. A variant OSR1 allele which disturbs OSR1 mRNA expression in renal progenitor cells is associated with reduction of newborn kidney size and function. Hum. Mol. Genet. 2011, 20, 4167–4174. [Google Scholar] [CrossRef] [PubMed]
- Kroeger, P.T.; Poureetezadi, S.J.; McKee, R.; Jou, J.; Miceli, R.; Wingert, R.A. Production of haploid zebrafish embryos by in vitro fertilization. J. Vis. Exp. 2014, 89, 51708. [Google Scholar] [CrossRef]
- Chambers, B.E.; Gerlach, G.F.; Clark, E.G.; Chen, K.H.; Levesque, A.E.; Leshchiner, I.; Goessling, W.; Wingert, R.A. Tfap2a is a novel gatekeeper of nephron differentiation during kidney development. Development 2019, 146, dev172387. [Google Scholar] [CrossRef]
- Kroeger, P.T.; Drummond, B.E.; Miceli, R.; McKernan, M.; Gerlach, G.F.; Marra, A.N.; Fox, A.; McCampbell, K.K.; Leshchiner, I.; Rodriguez-Mari, A.; et al. The zebrafish kidney mutant zeppelin reveals that brca2/fancd1 is essential for pronephros development. Dev. Biol. 2017, 428, 148–163. [Google Scholar] [CrossRef]
- Li, Y.; Cheng, C.N.; Verdun, V.A.; Wingert, R.A. Zebrafish nephrogenesis is regulated by interactions between retinoic acid, mecom, and Notch signaling. Dev. Biol. 2014, 386, 111–122. [Google Scholar] [CrossRef]
- Cheng, C.N.; Li, Y.; Marra, A.N.; Verdun, V.; Wingert, R.A. Flat mount preparation for observation and analysis of zebrafish embryo specimens stained by whole mount in situ hybridization. J. Vis. Exp. 2014, 89, 51604. [Google Scholar] [CrossRef]
- Cheng, C.N.; Wingert, R.A. Nephron proximal tubule patterning and corpuscles of Stannius formation are regulated by the sim1a transcription factor and retinoic acid in zebrafish. Dev. Biol. 2015, 399, 100–116. [Google Scholar] [CrossRef]
- Marra, A.N.; Ulrich, M.; White, A.; Springer, M.; Wingert, R.A. Visualizing multiciliated cells in the zebrafish through a combined protocol of whole mount fluorescent in situ hybridization and immunofluorescence. J. Vis. Exp. 2017, 129, 56261. [Google Scholar] [CrossRef] [PubMed]
- Marra, A.N.; Chambers, B.E.; Chambers, J.M.; Drummond, B.E.; Adeeb, B.D.; Wesselman, H.M.; Morales, E.E.; Handa, N.; Pettini, T.; Ronshaugen, M.; et al. Visualizing gene expression during zebrafish pronephros development and regeneration. Methods Cell. Biol. 2019, 154, 183–215. [Google Scholar] [PubMed]
- Wakahara, T.; Kusu, N.; Yamauchi, H.; Kimura, I.; Konishi, M.; Miyake, A.; Itoh, N. Fibin, a novel secreted lateral plate mesoderm signal, is essential for pectoral fin bud initiation in zebrafish. Dev. Biol. 2007, 303, 527–535. [Google Scholar] [CrossRef] [PubMed]
- Maves, L.; Tyler, A.; Moens, C.B.; Tapscott, S.J. Pbx acts with Hand2 in early myocardial differentiation. Dev. Biol. 2009, 333, 409–418. [Google Scholar] [CrossRef]
- Gerlach, G.F.; Wingert, R.A. Zebrafish pronephros tubulogenesis and epithelial identity maintenance are reliant on the polarity proteins prkc iota and zeta. Dev. Biol. 2014, 396, 183–200. [Google Scholar] [CrossRef]
- Leshchiner, I.; Alexa, K.; Kelsey, P.; Adzhubei, I.; Austin-Tse, C.A.; Cooney, J.D.; Anderson, H.; King, M.J.; Stottmann, R.W.; Garnaas, M.K.; et al. Mutation mapping and identification by whole-genome sequencing. Genome Res. 2012, 22, 1541–1548. [Google Scholar] [CrossRef]
- Ryan, S.; Willer, J.; Marjoram, L.; Bagwell, J.; Mankiewicz, J.; Leshchiner, I.; Goessling, W.; Bagnat, M.; Katsanis, N. Rapid identification of kidney cyst mutations by whole exome sequencing in zebrafish. Development 2013, 140, 4445–4451. [Google Scholar] [CrossRef]
- Swartz, M.E.; Sheehan-Rooney, K.; Dixon, M.J.; Eberhart, J.K. Examination of a palatogenic gene program in zebrafish. Dev. Dyn. 2011, 240, 2204–2220. [Google Scholar] [CrossRef]
- Liu, H.; Lan, Y.; Xu, J.; Chang, C.; Brugmann, S.A.; Jiang, R. Odd-Skipped related-1 controls neural crest chondrogenesis during tongue development. Proc. Natl. Acad. Sci. USA 2013, 110, 18555–18560. [Google Scholar] [CrossRef]
- Terashima, A.V.; Mudumana, S.P.; Drummond, I.A. Odd skipped related 1 is a negative feedback regulator of nodal-induced endoderm development. Dev. Dyn. 2014, 243, 1571–1580. [Google Scholar] [CrossRef]
- Gering, M.; Rodaway, A.R.; Göttgens, B.; Patient, R.K.; Green, A.R. The SCL gene specifies haemangioblast development from early mesoderm. EMBO J. 1998, 17, 4029–4045. [Google Scholar] [CrossRef] [PubMed]
- Liao, E.C.; Paw, B.H.; Oates, A.C.; Pratt, S.J.; Postlethwait, J.H.; Zon, L.I. SCL/Tal-1 transcription factor acts downstream of cloche to specify hematopoietic and vascular progenitors in zebrafish. Genes Dev. 1998, 12, 621–626. [Google Scholar] [CrossRef] [PubMed]
- Kroeger, P.T.; Wingert, R.A. Using zebrafish to study podocyte genesis during kidney development and regeneration. Genesis 2014, 52, 771–792. [Google Scholar] [CrossRef] [PubMed]
- Perner, B.; Englert, C.; Bollig, F. The Wilms tumor genes Wt1a and Wt1b control different steps during formation of the zebrafish pronephros. Dev. Biol. 2007, 309, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Liu, H.; Chai, H.; Lan, Y.; Jiang, R. Osr1 interacts synergistically with Wt1 to regulate kidney organogenesis. PLoS ONE 2016, 11, e0159597. [Google Scholar] [CrossRef]
- Lin, Y.; Liu, A.; Zhang, S.; Ruusunen, T.; Kreidberg, J.A.; Peltoketo, H.; Drummond, I.; Vainio, S. Induction of ureter branching as a response to Wnt-2b signaling during early kidney organogenesis. Dev. Dyn. 2001, 222, 26–39. [Google Scholar] [CrossRef]
- Iglesias, D.M.; Hueber, P.; Chu, L.; Campbell, R.; Patenaude, A.; Dziarmaga, A.J.; Quinlan, J.; Mohamed, O.; Dufort, D.; Goodyer, P.R. canonical WNT signaling during kidney development. Am. J. Physiol. Ren. Physiol. 2007, 293, 494. [Google Scholar] [CrossRef]
- Combes, A.N.; Combes, A.N.; Zappia, L.; Er, P.X.; Oshlack, A.; Little, M.H. Single-cell analysis reveals congruence between kidney organoids and human fetal kidney. Genome Med. 2019, 11, 3. [Google Scholar] [CrossRef]
- Goss, A.M.; Tian, Y.; Tsukiyama, T.; Cohen, E.D.; Zhou, D.; Lu, M.M.; Yamaguchi, T.P.; Morrisey, E.E. Wnt2/2b and Β-Catenin signaling are necessary and sufficient to specify lung progenitors in the foregut. Dev. Cell 2009, 17, 290–298. [Google Scholar] [CrossRef]
- Rankin, S.A.; Gallas, A.L.; Neto, A.; Gómez-Skarmeta, J.L.; Zorn, A.M. Suppression of Bmp4 signaling by the zinc-finger repressors Osr1 and Osr2 is required for Wnt/Β-Catenin-mediated lung specification in Xenopus. Development 2012, 139, 3010–3020. [Google Scholar] [CrossRef]
- Lyons, J.P.; Miller, R.K.; Zhou, X.; Weidinger, G.; Deroo, T.; Denayer, T.; Park, J.; Ji, H.; Hong, J.Y.; Li, A.; et al. Requirement of Wnt/Β-Catenin signaling in pronephric kidney development. Mech. Dev. 2009, 126, 142–159. [Google Scholar] [CrossRef] [PubMed]
- Verkade, H.; Heath, J.K. Wnt signaling mediates diverse developmental processes in zebrafish. Methods Mol. Biol. 2008, 469, 225. [Google Scholar] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Drummond, B.E.; Chambers, B.E.; Wesselman, H.M.; Gibson, S.; Arceri, L.; Ulrich, M.N.; Gerlach, G.F.; Kroeger, P.T.; Leshchiner, I.; Goessling, W.; et al. osr1 Maintains Renal Progenitors and Regulates Podocyte Development by Promoting wnt2ba via the Antagonism of hand2. Biomedicines 2022, 10, 2868. https://doi.org/10.3390/biomedicines10112868
Drummond BE, Chambers BE, Wesselman HM, Gibson S, Arceri L, Ulrich MN, Gerlach GF, Kroeger PT, Leshchiner I, Goessling W, et al. osr1 Maintains Renal Progenitors and Regulates Podocyte Development by Promoting wnt2ba via the Antagonism of hand2. Biomedicines. 2022; 10(11):2868. https://doi.org/10.3390/biomedicines10112868
Chicago/Turabian StyleDrummond, Bridgette E., Brooke E. Chambers, Hannah M. Wesselman, Shannon Gibson, Liana Arceri, Marisa N. Ulrich, Gary F. Gerlach, Paul T. Kroeger, Ignaty Leshchiner, Wolfram Goessling, and et al. 2022. "osr1 Maintains Renal Progenitors and Regulates Podocyte Development by Promoting wnt2ba via the Antagonism of hand2" Biomedicines 10, no. 11: 2868. https://doi.org/10.3390/biomedicines10112868
APA StyleDrummond, B. E., Chambers, B. E., Wesselman, H. M., Gibson, S., Arceri, L., Ulrich, M. N., Gerlach, G. F., Kroeger, P. T., Leshchiner, I., Goessling, W., & Wingert, R. A. (2022). osr1 Maintains Renal Progenitors and Regulates Podocyte Development by Promoting wnt2ba via the Antagonism of hand2. Biomedicines, 10(11), 2868. https://doi.org/10.3390/biomedicines10112868






