Mesenchymal Stromal Cell on Liver Decellularised Extracellular Matrix for Tissue Engineering
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Porcine Liver Retrieval
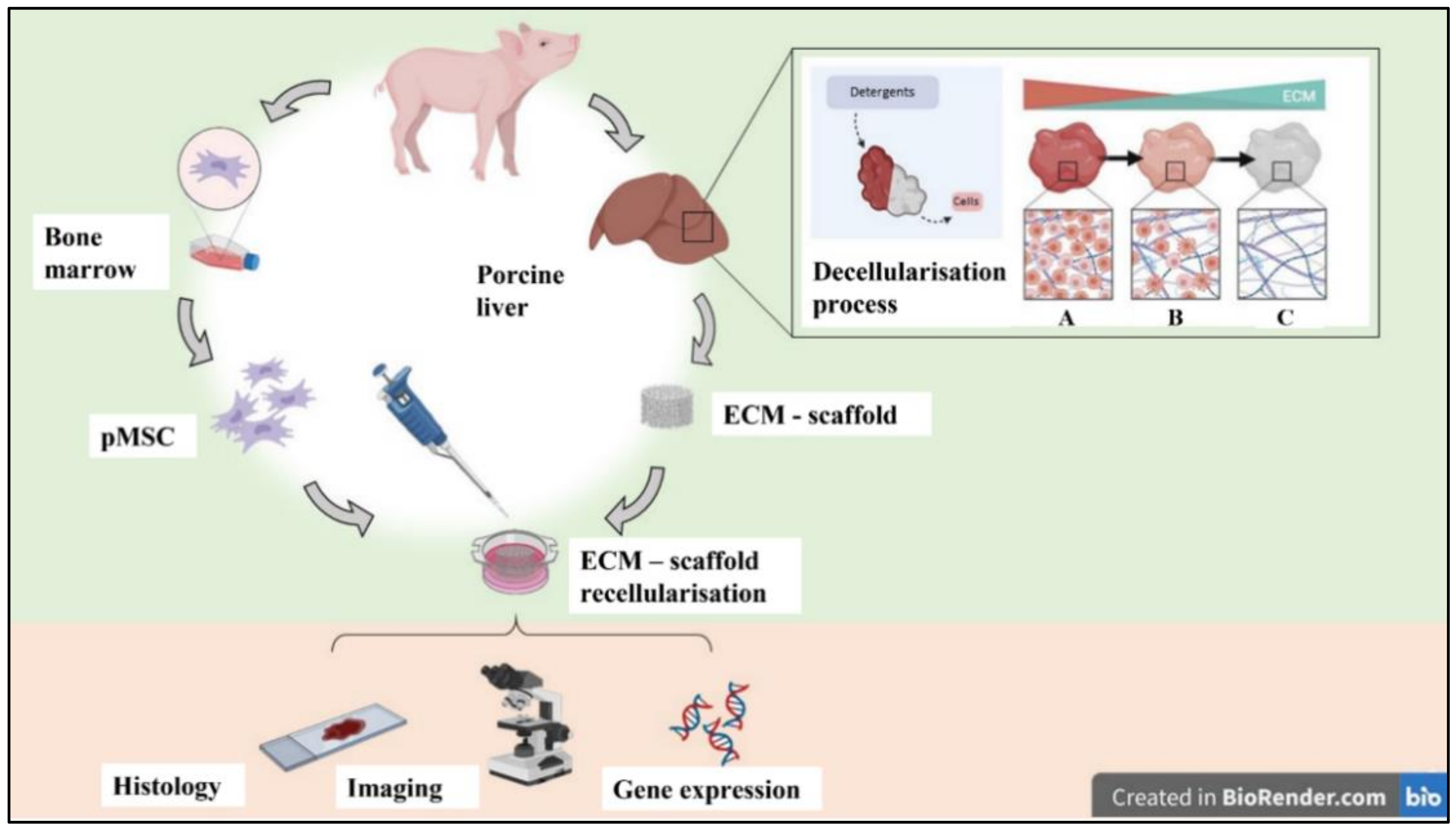
2.3. Liver Decellularisation Procedure
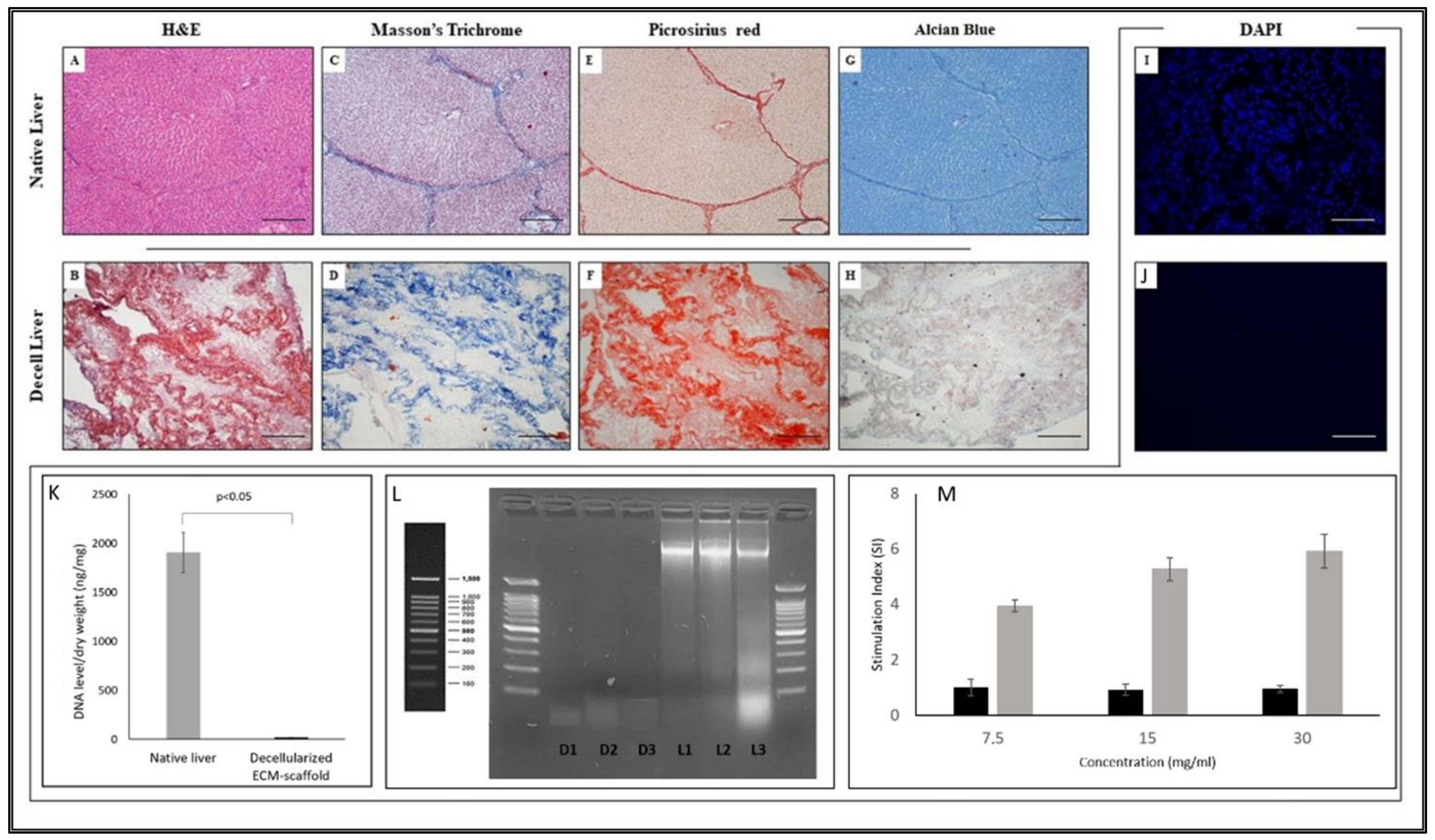
2.4. Histological Analysis and DAPI Staining
2.5. DNA Residuals Evaluation
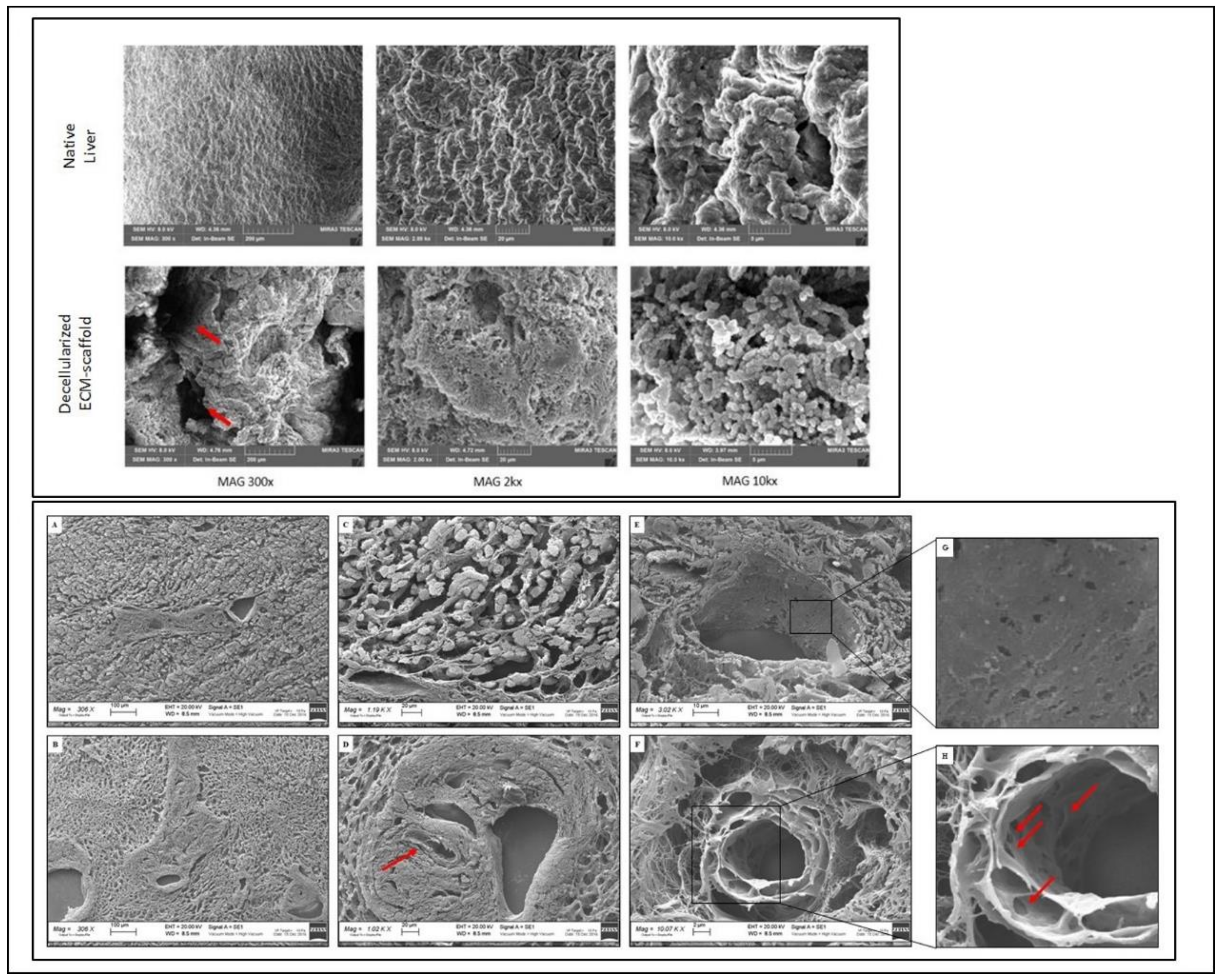
2.6. SEM Analysis
2.7. Scaffold Immunogenicity In Vitro Evaluation
2.8. Isolation and Expansion of pMSC
2.9. Characterisation of pMSC
2.10. Recellularisation
2.11. Metabolic Viability Based Assay(MTT)
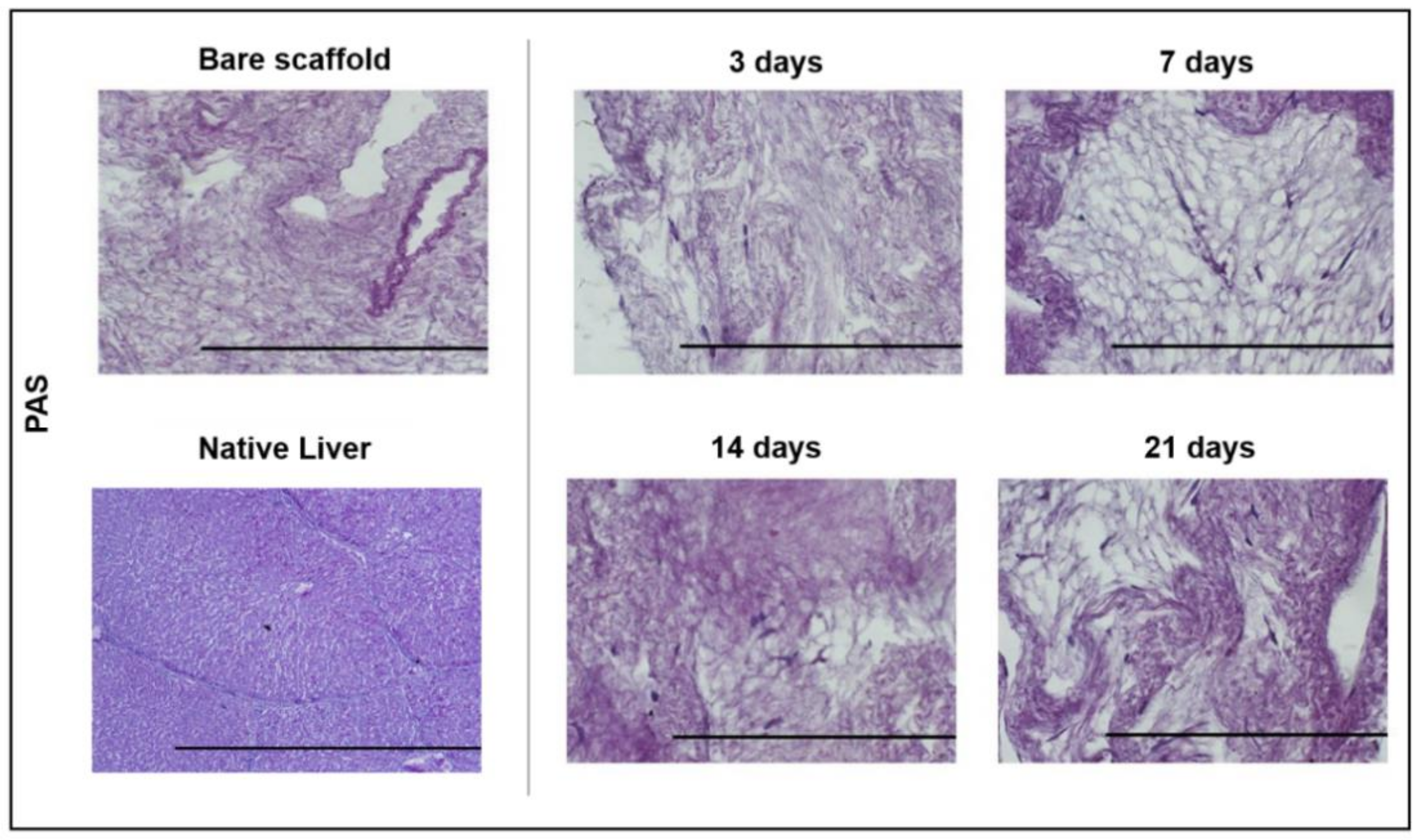
2.12. Periodic Acid Schiff (PAS) Staining
2.13. RNA Extraction, Retrotrascription and Real-Time PCR
2.14. Statistical Analysis
3. Results
3.1. Liver Decellularisation
3.2. 3D Architecture and Ultrastructure
3.3. Mesenchymal Stromal Cells Isolation and Characterisation
3.4. ECM Scaffold Recellularisation
Differentiation of pMSCs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Murray, C.J.L.; Lopez, A.D. Measuring the global burden of disease. N. Engl. J. Med. 2013, 369, 448–457. [Google Scholar] [CrossRef] [PubMed]
- Sarin, S.K.; Choudhury, A. Acute-on-chronic liver failure: Terminology, mechanisms and management. Nat. Rev. Gastro. Hepat. 2016, 13, 131–149. [Google Scholar] [CrossRef] [PubMed]
- Bernal, W.; Auzinger, G.; Dhawan, A.; Wendon, J. Seminar Acute liver failure. Lancet 2010, 376, 190–201. [Google Scholar] [CrossRef]
- Kwong, A.; Kim, W.R.; Lake, J.R.; Smith, J.M.; Schladt, D.P.; Skeans, M.A.; Noreen, S.M.; Foutz, J.; Miller, E.; Snyder, J.J.; et al. OPTN/SRTR 2018 Annual Data Report: Liver. Am. J. Transplant. 2020, 20, 193–299. [Google Scholar] [CrossRef] [PubMed]
- U.S. Department of Health and Human Services. OPTN/SRTR 2012 Annual Data Report—United States Organ Transplantation. Am. J. Transplant. 2014, 14, 1–192. [Google Scholar]
- Merion, R.M.; Pelletier, S.J.; Goodrich, N.; Englesbe, M.J.; Delmonico, F.L. Donation after cardiac death as a strategy to increase deceased donor liver availability. Ann. Surg. 2006, 244, 555–560. [Google Scholar] [CrossRef]
- Bilir, B.; Guinette, D.; Karrer, F.; Kumpe, D.A.; Krysl, J.; Stephens, J.; McGavran, L.; Ostrowska, A.; Durham, J. Hepatocyte transplantation in acute liver failure. Liver Transplant. 2000, 6, 32–40. [Google Scholar] [CrossRef]
- Nussler, A.; Konig, S.; Ott, M.; Sokal, E.; Christ, B.; Thasler, W.; Brulport, M.; Gabelein, G.; Schormann, W.; Schulze, M.; et al. Present status and perspectives of cell-based therapies for liver diseases. J. Hepatol. 2006, 45, 144–159. [Google Scholar] [CrossRef]
- O’Donoghue, K.; Fisk, N.M. Fetal stem cells. Best Pract. Res. Clin. Obstet. Gynaecol. 2004, 18, 853–875. [Google Scholar] [CrossRef]
- Nicolas, C.; Wang, Y.; Luebke-Wheeler, J.; Nyberg, S.L. Stem cell therapies for treatment of liver disease. Biomedicines 2016, 4, 2. [Google Scholar] [CrossRef]
- Ibars, E.P.; Cortes, M.; Tolosa, L.; Gómez-Lechón, M.J.; López, S.; Castell, J.V.; Mir, J. Hepatocyte transplantation program: Lessons learned and future strategies. World J. Gastroenterol. 2016, 22, 874–886. [Google Scholar] [CrossRef]
- Badylak, S.F.; Taylor, D.; Uygun, K. Whole Organ Tissue Engineering: Decellularization and Recellularization of Three-Dimensional Matrix Scaffolds. Annu. Rev. Biomed. Eng. 2010, 13, 27–53. [Google Scholar] [CrossRef]
- Baptista, P.M.; Orlando, G.; Mirmalek-Sani, S.H.; Siddiqui, M.; Atala, A.; Soker, S. Whole organ decellularization - A tool for bioscaffold fabrication and organ bioengineering. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 2009, 6526–6529. [Google Scholar]
- Uygun, B.E.; Soto-Gutierrez, A.; Yagi, H.; Izamis, M.L.; Guzzardi, M.A.; Shulman, C.; Milwid, J.; Kobayashi, N.; Tilles, A.; Berthiaume, F.; et al. Organ reengineering through development of a transplantable recellularized liver graft using decellularized liver matrix. Nat. Med. 2010, 16, 814–820. [Google Scholar] [CrossRef]
- Abazari, M.F.; Soleimanifar, F.; Enderami, S.E.; Nasiri, N.; Nejati, F.; Mousavi, S.A.; Soleimani, M.; Kiani, J.; Ghoraeian, P.; Kehtari, M. Decellularized amniotic membrane Scaffolds improve differentiation of iPSCs to functional hepatocyte-like cells. J. Cell Biochem. 2020, 121, 1169–1181. [Google Scholar] [CrossRef]
- Meyer, S.R.; Nagendran, J.; Desai, L.S.; Rayat, G.R.; Churchill, T.A.; Anderson, C.C.; Rajotte, R.V.; Lakey, J.R.; Ross, D.B. Decellularization reduces the immune response to aortic valve allografts in the rat. J. Thorac. Cardiovasc. Surg. 2005, 130, 469–476. [Google Scholar] [CrossRef]
- Sánchez-Romero, N.; Sainz-Arnal, P.; Pla-Palacín, I.; Dachary, P.R.; Almeida, H.; Pastor, C.; Soto, D.R.; Rodriguez, M.C.; Arbizu, E.O.; Martinez, L.B.; et al. The role of extracellular matrix on liver stem cell fate: A dynamic relationship in health and disease. Differentiation 2019, 106, 49–56. [Google Scholar] [CrossRef]
- Fitzpatrick, E.; Mitry, R.R.; Dhawan, A. Human hepatocyte transplantation: State of the art. J. Int. Med. 2009, 266, 339–357. [Google Scholar] [CrossRef]
- Lorvellec, M.; Scottoni, F.; Crowley, C.; Fiadeiro, R.; Maghsoudlou, P.; Pellegata, A.F.; Mazzacuva, F.; Gjinovci, A.; Lyne, A.M.; Zulini, J.; et al. Mouse decellularised liver scaffold improves human embryonic and induced pluripotent stem cells differentiation into hepatocyte-like cells. PLoS ONE 2017, 12(12), e0189586. [Google Scholar] [CrossRef]
- Wang, B.; Jakus, A.E.; Baptista, P.M.; Soker, S.; Soto-Gutierrez, A.; Abecassis, M.M.; Shah, R.N.; Wertheim, J.A. Functional Maturation of Induced Pluripotent Stem Cell Hepatocytes in Extracellular Matrix-A Comparative Analysis of Bioartificial Liver Microenvironments. Stem Cells Transl. Med. 2016, 5, 1257–1267. [Google Scholar] [CrossRef]
- Banas, A.; Yamamoto, Y.; Teratani, T.; Ochiya, T. Stem cell plasticity: Learning from hepatogenic differentiation strategies. Dev. Dynam. 2007, 236, 3228–3241. [Google Scholar] [CrossRef] [PubMed]
- Panta, W.; Imsoonthornruksa, S.; Yoisungnern, T.; Suksaweang, S.; Ketudat-Cairns, M.; Parnpai, R. Enhanced hepatogenic differentiation of human wharton’s jelly-derived mesenchymal stem cells by using three-step protocol. Int. J. Mol. Sci. 2019, 20, 3016. [Google Scholar] [CrossRef] [PubMed]
- Al Ghrbawy, N.M.; Afify, R.A.A.M.; Dyaa, N.; El Sayed, A.A. Differentiation of Bone Marrow: Derived Mesenchymal Stem Cells into Hepatocyte-like Cells. Indian J. Hematol. Blood Transfus. 2016, 32, 276–283. [Google Scholar] [CrossRef] [PubMed]
- Xie, P.Y.; Hu, X.J.; Guo, R.M.; Meng, X.C.; Pang, P.F.; Zhou, Z.Y.; Li, D.; Shan, H. Generation of functional hepatocyte-like cells from human bone marrow mesenchymal stem cells by overexpression of transcription factor HNF4α and FOXA2. Hepatob. Pancreat. Dis. Int. 2019, 18, 546–556. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Cui, L.; Zhou, X.; Yang, Q.; Wang, L.; Guo, G.; Hou, Y.; Cai, W.; Han, Z.; Shi, Y.; et al. Induction of hepatocyte-like cells from human umbilical cord-derived mesenchymal stem cells by defined microRNAs. J. Cell Mol. Med. 2017, 21, 881–893. [Google Scholar] [CrossRef]
- Najimi, M.; Khuu, D.N.; Lysy, P.A.; Jazouli, N.; Abarca, J.; Sempoux, C.; Sokal, E.M. Adult-derived human liver mesenchymal-like cells as a potential progenitor reservoir of hepatocytes? Cell Transplant. 2007, 16, 717–728. [Google Scholar] [CrossRef]
- Bari, E.; Di Silvestre, D.; Mastracci, L.; Grillo, F.; Grisoli, P.; Marrubini, G.; Nardini, M.; Mastrogiacomo, M.; Sorlini, M.; Rossi, R.; et al. GMP-compliant sponge-like dressing containing MSC lyo-secretome: Proteomic network of healing in a murine wound model. Eur. J. Pharm. Biopharm. 2020, 155, 37–48. [Google Scholar] [CrossRef]
- Comite, P.; Cobianchi, L.; Avanzini, M.A.; Zonta, S.; Mantelli, M.; Achille, V.; De Martino, M.; Cansolino, L.; Ferrari, C.; Alessiani, M.; et al. Isolation and Ex Vivo Expansion of Bone Marrow-Derived Porcine Mesenchymal Stromal Cells: Potential for Application in an Experimental Model of Solid Organ Transplantation in Large Animals. Transplant. Proc. 2010, 42, 1341–1343. [Google Scholar] [CrossRef]
- Crapo, P.M.; Gilbert, T.W.; Badylak, S.F. An overview of tissue and whole organ decellularization processes. Biomaterials 2011, 32, 3233–3243. [Google Scholar] [CrossRef]
- Croce, S.; Peloso, A.; Zoro, T.; Avanzini, M.A.; Cobianchi, L. A Hepatic Scaffold from Decellularized Liver Tissue: Food for Thought. Biomolecules 2019, 9, 813. [Google Scholar] [CrossRef]
- Li, Y.; Wu, Q.; Wang, Y.; Li, L.; Chen, F.; Shi, Y.; Bao, J.; Bu, H. Construction of bioengineered hepatic tissue derived from human umbilical cord mesenchymal stem cells via aggregation culture in porcine decellularized liver scaffolds. Xenotransplantation 2017, 24, 1–15. [Google Scholar] [CrossRef]
- Luo, Y.; Lou, C.; Zhang, S.; Zhu, Z.; Xing, Q.; Wang, P.; Liu, T.; Liu, H.; Li, C.; Shi, W.; et al. Three-dimensional hydrogel culture conditions promote the differentiation of human induced pluripotent stem cells into hepatocytes. Cytotherapy 2018, 20, 95–107. [Google Scholar] [CrossRef]
- Rajendran, D.; Hussain, A.; Yip, D.; Parekh, A.; Shrirao, A.; Cho, C.H. Long-term liver-specific functions of hepatocytes in electrospun chitosan nanofiber scaffolds coated with fibronectin. J. Biomed. Mater. Res. 2017, 105, 2119–2128. [Google Scholar] [CrossRef]
- Mazza, G.; Rombouts, K.; Rennie Hall, A.; Urbani, L.; Vinh Luong, T.; Al-Akkad, W.; Longato, L.; Brown, D.; Maghsoudlou, P.; Dhillon, A.P.; et al. Decellularized human liver as a natural 3D-scaffold for liver bioengineering and transplantation. Sci. Rep. 2015, 5, 1–15. [Google Scholar]
- Verstegen, M.M.A.; Willemse, J.; van den Hoek, S.; Kremers, G.J.; Luider, T.M.; van Huizen, N.A.; Willemssen, F.E.J.A.; Metselaar, H.J.; IJzermans, J.N.M.; van der Laan, L.J.W.; et al. Decellularization of Whole Human Liver Grafts Using Controlled Perfusion for Transplantable Organ Bioscaffolds. Stem Cells Dev. 2017, 26, 1304–1315. [Google Scholar] [CrossRef]
- Yagi, H.; Fukumitsu, K.; Fukuda, K.; Kitago, M.; Shinoda, M.; Obara, H.; Itano, O.; Kawachi, S.; Tanabe, M.; Coudriet, G.M.; et al. Human-Scale Whole-Organ Bioengineering for Liver Transplantation: A Regenerative Medicine Approach. Cell Transplant. 2013, 22, 231–242. [Google Scholar] [CrossRef]
- Moulisová, V.; Jiřík, M.; Schindler, C.; Červenková, L.; Pálek, R.; Rosendorf, J.; Arlt, J.; Bolek, L.; Šůsová, S.; Nietzsche, S.; et al. Novel morphological multi-scale evaluation system for quality assessment of decellularized liver scaffolds. J. Tissue Eng. 2020, 11, 2041731420921121. [Google Scholar] [CrossRef]
- Naeem, E.M.; Sajad, D.; Talaei-Khozani, T.; Khajeh, S.; Azarpira, N.; Alaei, S.; Tanideh, N.; Reza, T.M.; Razban, V. Decellularized liver transplant could be recellularized in rat partial hepatectomy model. J. Biomed. Mater. Res. A. 2019, 107, 2576–2588. [Google Scholar] [CrossRef]
- Peloso, A.; Ferrario, J.; Maiga, B.; Benzoni, I.; Bianco, C.; Citro, A.; Currao, M.; Malara, A.; Gaspari, A.; Balduini, A.; et al. Creation and implantation of acellular rat renal ECM-based scaffolds. Organogenesis 2015, 11, 58–74. [Google Scholar] [CrossRef]
- Liu, Z.Z.; Wong, M.L.; Griffiths, L.G. Effect of bovine pericardial extracellular matrix scaffold niche on seeded human mesenchymal stem cell function. Sci. Rep. 2016, 6, 1–12. [Google Scholar] [CrossRef]
- Zhang, J.; Zhao, X.; Liang, L.; Li, J.; Demirci, U.; Wang, S.Q. A decade of progress in liver regenerative medicine. Biomaterials 2018, 157, 161–176. [Google Scholar] [CrossRef] [PubMed]
- Keane, T.J.; Londono, R.; Turner, N.J.; Badylak, S.F. Consequences of ineffective decellularization of biologic scaffolds on the host response. Biomaterials 2012, 33, 1771–1781. [Google Scholar] [CrossRef] [PubMed]
- Alaby Pinheiro Faccioli, L.; Suhett Dias, G.; Hoff, V.; Lemos Dias, M.; Ferreira Pimentel, C.; Hochman-Mendez, C.; Braz Parente, D.; Labrunie, E.; Souza Mourão, P.; Rogério de Oliveira Salvalaggio, P.; et al. Optimizing the Decellularized Porcine Liver Scaffold Protocol. Cells Tissues Organs 2020, 11, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Brückner, S.; Tautenhahn, H.M.; Winkler, S.; Stock, P.; Dollinger, M.; Christ, B. A fat option for the pig: Hepatocytic differentiated mesenchymal stem cells for translational research. Exp. Cell Res. 2014, 321, 267–275. [Google Scholar] [CrossRef] [PubMed]
- Ghaedi, M.; Soleimani, M.; Shabani, I.; Duan, Y.; Lotfi, A.S. Hepatic differentiation from human mesenchymal stem cells on a novel nanofiber scaffold. Cell Mol. Biol. Lett. 2012, 17, 89–106. [Google Scholar] [CrossRef]
- Soto-Gutierrez, A.; Zhang, L.; Medberry, C.; Fukumitsu, K.; Faulk, D.; Jiang, H.; Reing, J.; Gramignoli, R.; Komori, J.; Ross, M.; et al. A Whole-Organ Regenerative Medicine Approach for Liver Replacement. Tissue Eng. Part C Methods 2011, 17, 677–686. [Google Scholar] [CrossRef]
- Bao, J.; Wu, Q.; Wang, Y.; Li, Y.; Li, L.; Chen, F.; Wu, X.; Xie, M.; Bu, H. Enhanced hepatic differentiation of rat bone marrow-derived mesenchymal stem cells in spheroidal aggregate culture on a decellularized liver scaffold. Int. J. Mol. Med. 2016, 38, 457–465. [Google Scholar] [CrossRef]
- Coronado, R.E.; Somaraki-Cormier, M.; Natesan, S.; Christy, R.J.; Ong, J.L.; Halff, G.A. Decellularization and Solubilization of Porcine Liver for Use as a Substrate for Porcine Hepatocyte Culture: Method Optimization and Comparison. Cell Transplant. 2017, 1840–1854. [Google Scholar] [CrossRef]
- Jaramillo, M.; Yeh, H.; Yarmush, M.L.; Uygun, B.E. Decellularized human liver extracellular matrix (hDLM)-mediated hepatic differentiation of human induced pluripotent stem cells (hIPSCs). J. Tissue Eng. Regen. Med. 2018, 12, e1962–e1973. [Google Scholar] [CrossRef]
- Kim, D.S.; Ryu, J.W.; Son, M.Y.; Oh, J.H.; Chung, K.S.; Lee, S.; Lee, J.J.; Ahn, J.H.; Min, J.S.; Ahn, J.; et al. A liver-specific gene expression panel predicts the differentiation status of in vitro hepatocyte models. Hepatology 2017, 66(5), 1662–1674. [Google Scholar] [CrossRef]
- Park, K.M.; Hussein, K.H.; Hong, S.H.; Ahn, C.; Yang, S.R.; Park, S.M.; Kweon, O.K.; Kim, B.M.; Woo, H.M. Decellularized Liver Extracellular Matrix as Promising Tools for Transplantable Bioengineered Liver Promotes Hepatic Lineage Commitments of Induced Pluripotent Stem Cells. Tissue Eng. 2016, 22, 449–460. [Google Scholar] [CrossRef]
- Ji, R.; Zhang, N.; You, N.; Li, Q.; Liu, W.; Jiang, N.; Liu, J.; Zhang, H.; Wang, D.; Tao, K.; et al. The differentiation of MSCs into functional hepatocyte-like cells in a liver biomatrix scaffold and their transplantation into liver-fibrotic mice. Biomaterials 2012, 33, 8995–9008. [Google Scholar] [CrossRef]
- Fráguas-Eggenschwiler, M.; Eggenschwiler, R.; Söllner, J.H.; Cortnumme, L.; Vondran, F.W.R.; Cantz, T.; Ott, M.; Niemann, H. Direct conversion of porcine primary fibroblasts into hepatocyte-like cells. Sci. Rep. 2021, 11, 9334. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Croce, S.; Cobianchi, L.; Zoro, T.; Dal Mas, F.; Icaro Cornaglia, A.; Lenta, E.; Acquafredda, G.; De Silvestri, A.; Avanzini, M.A.; Visai, L.; et al. Mesenchymal Stromal Cell on Liver Decellularised Extracellular Matrix for Tissue Engineering. Biomedicines 2022, 10, 2817. https://doi.org/10.3390/biomedicines10112817
Croce S, Cobianchi L, Zoro T, Dal Mas F, Icaro Cornaglia A, Lenta E, Acquafredda G, De Silvestri A, Avanzini MA, Visai L, et al. Mesenchymal Stromal Cell on Liver Decellularised Extracellular Matrix for Tissue Engineering. Biomedicines. 2022; 10(11):2817. https://doi.org/10.3390/biomedicines10112817
Chicago/Turabian StyleCroce, Stefania, Lorenzo Cobianchi, Tamara Zoro, Francesca Dal Mas, Antonia Icaro Cornaglia, Elisa Lenta, Gloria Acquafredda, Annalisa De Silvestri, Maria Antonietta Avanzini, Livia Visai, and et al. 2022. "Mesenchymal Stromal Cell on Liver Decellularised Extracellular Matrix for Tissue Engineering" Biomedicines 10, no. 11: 2817. https://doi.org/10.3390/biomedicines10112817
APA StyleCroce, S., Cobianchi, L., Zoro, T., Dal Mas, F., Icaro Cornaglia, A., Lenta, E., Acquafredda, G., De Silvestri, A., Avanzini, M. A., Visai, L., Brambilla, S., Bruni, G., Gravina, G. D., Pietrabissa, A., Ansaloni, L., & Peloso, A. (2022). Mesenchymal Stromal Cell on Liver Decellularised Extracellular Matrix for Tissue Engineering. Biomedicines, 10(11), 2817. https://doi.org/10.3390/biomedicines10112817







