Fluorogenic Aptasensors with Small Molecules
Abstract
1. Introduction
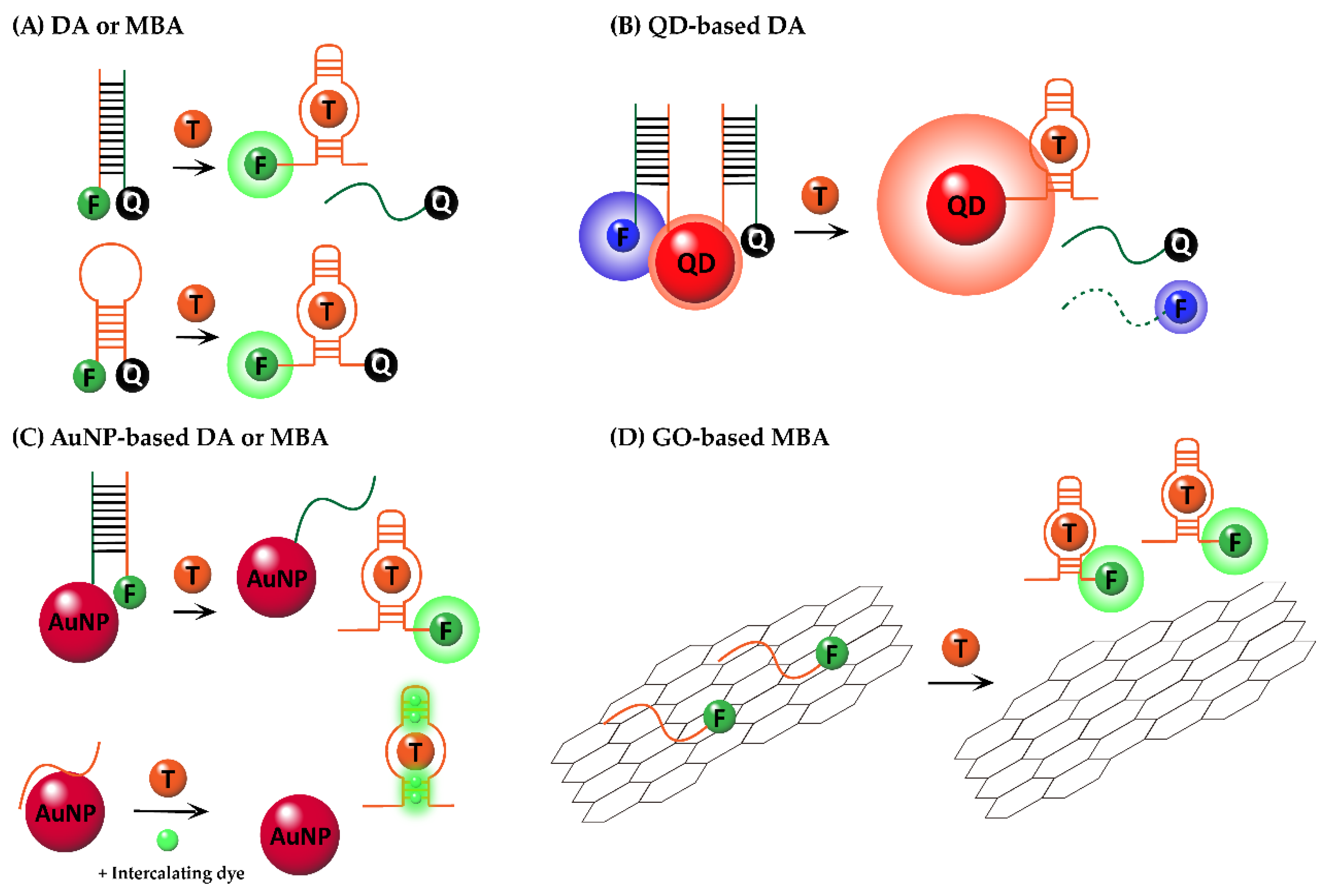
2. Duplex or Molecular Beacon-Type Fluorogenic Aptasensors with Small Molecules
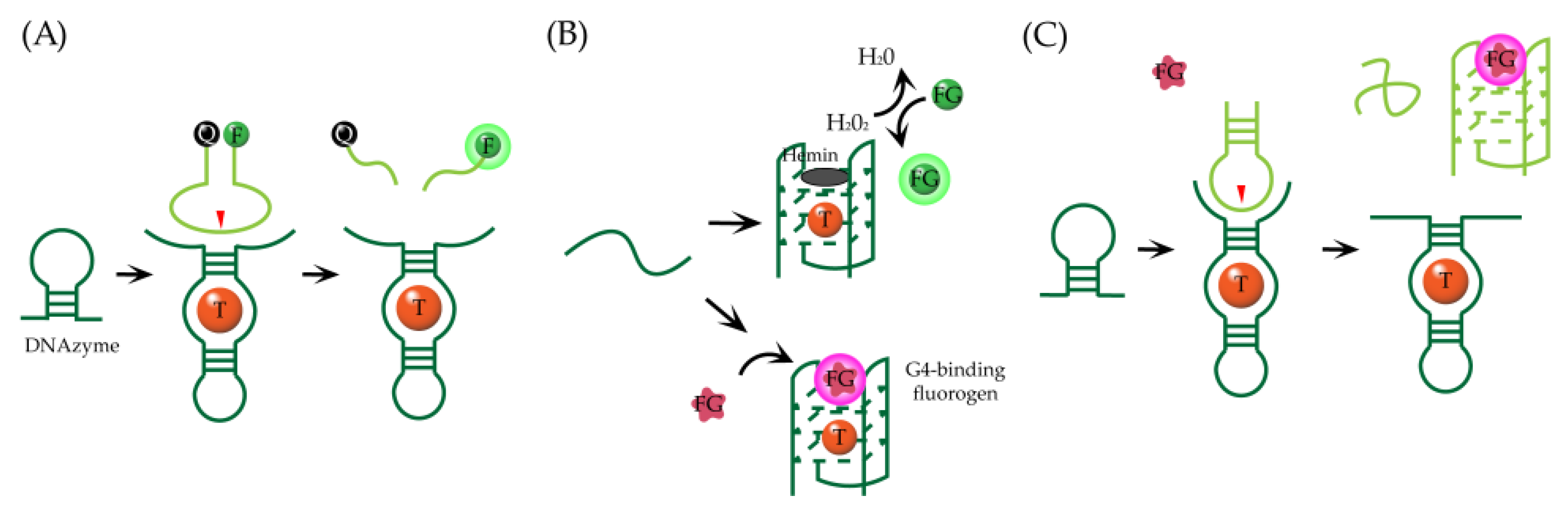
3. Fluorogenic Aptazymes with Small Molecules
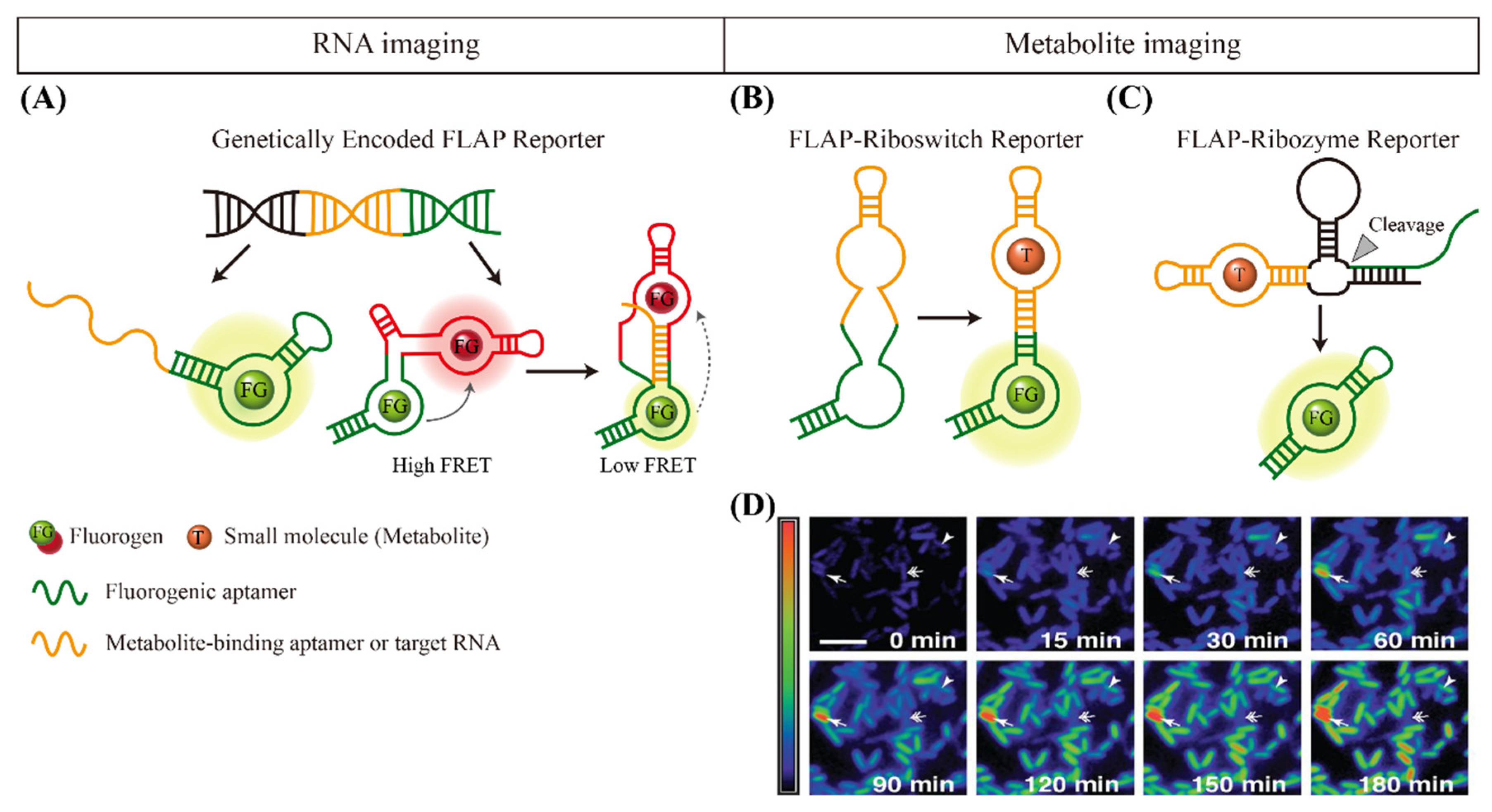
4. Fluorogen-Activating Aptamer Reporters for Imaging Small Molecules
| RNA Aptamer | Sequence (5′ to 3′) | Fluorogen | Ex/Em (nm) | Kd (nM) | Target | Ref. |
|---|---|---|---|---|---|---|
| Baby Spinach | GGU GAA GGA CGG GUC CAG UAG UUC GCU ACU GUU GAG UAG AGU GUG AGC UCC | DFHBI | 450/505 | ND | miRNA | [82] |
| Mango II | GGC ACG UAC GAA GGG ACG GUG CGG AGA GGA GAG UAC GUG C | TO1-Biotin | 492/561 | 1.8 | Small ncRNA | [83] |
| Mango III | GGC ACG UAC GAA GGA AGG AUU GGU AUG UGG UAU AUU CGU ACG UGC C | 15.0 | ||||
| Mango IV | GGC ACG UAC CGA GGG AGU GGU GAG GAU GAG GCG AGU ACG UGC | 10.4 | ||||
| Mango II | GGC ACG UAC GAA GGA GAG GAG AGG AAG AGG AGA GUA CGU GC | TO1-Biotin | 492/561 | 1.8 | Long ncRNA | [84] |
| Minimal Spinach | GGA UGU AAC UGA AUG AAA UGG UGA AGG ACG GGU CCA GUA GGC UGC UUC GGC AGC CUA CUU GUU GAG UAG AGU GUG AGC UCC GUA ACU AGU UAC AUC C | DFHBI-1T | 450/503 | 340 | Metabolite (SAM) | [85] |
| Minimal Broccoli | GGC CCG GAU AGC UCA GUC GGU AGA GCA GCG GAG ACG GUC GGG UCC AGA UAU UCG UAU CUG UCG AGU AGA GUG UGG GCU CCG CGG GUC CAG GGU UCA AGU CCC UGU UCG GGC GCC | DFHBI-1T | 450/503 | ND | Metabolite (SAM) | [85] |
| Minimal Mango | GGA UGC GUA ACC CUC AAG GAA CCC GCA AGC CAU CGG GAC UCA AGC CGC CGG UAC CUC CGA AGG GAC GGU GCG GAG AGG AGA GGG GGC ACU GGG CGG CUG UGU GAG AUU CUG CCA AAU AGA CAG CCG AA | YO3-biotin | 580/620 | 26 | Metabolite (SAM) | [85] |
| Corn | GAG GAA GGA GGU CUG AGG AGG UCA CU | DFHO | 470/505 | <1 | Metabolite (SAM) | [86] |
| Broccoli | GAG ACG GUC GGG UCC AGA UAU UCG UAU CUG UCG AGU AGA GUG UGG GCU C | DFHBI-1T | 470/503 | ND | Silver ion | [87] |
| Broccoli (modified) | GCG GAG ACG GUC GGG UCC AGA UAU UCG UAU CUG UCG AGU AGA GUG UGG GCU CCG | DFHBI-1T | 470/503 | ND | Metabolite (c-di-GMP, cAMP) | [88] |
5. Conclusions and Future Perspective
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- McKeague, M.; De Girolamo, A.; Valenzano, S.; Pascale, M.; Ruscito, A.; Velu, R.; Frost, N.R.; Hill, K.; Smith, M.; McConnell, E.M.; et al. Comprehensive analytical comparison of strategies used for small molecule aptamer evaluation. Anal. Chem. 2015, 87, 8608–8612. [Google Scholar] [CrossRef] [PubMed]
- Stojanovic, M.N.; Landry, D.W. Aptamer-based colorimetric probe for cocaine. J. Am. Chem. Soc. 2002, 124, 9678–9679. [Google Scholar] [CrossRef] [PubMed]
- Wei, H.; Li, B.; Li, J.; Wang, E.; Dong, S. Simple and sensitive aptamer-based colorimetric sensing of protein using unmodified gold nanoparticle probes. Chem. Commun. 2007, 36, 3735–3737. [Google Scholar] [CrossRef] [PubMed]
- Yildirim, N.; Long, F.; Gao, C.; He, M.; Shi, H.C.; Gu, A.Z. Aptamer-based optical biosensor for rapid and sensitive detection of 17beta-estradiol in water samples. Environ. Sci. Technol. 2012, 46, 3288–3294. [Google Scholar] [CrossRef]
- Hansen, J.A.; Wang, J.; Kawde, A.N.; Xiang, Y.; Gothelf, K.V.; Collins, G. Quantum-dot/aptamer-based ultrasensitive multi-analyte electrochemical biosensor. J. Am. Chem. Soc. 2006, 128, 2228–2229. [Google Scholar] [CrossRef]
- Wu, Z.S.; Guo, M.M.; Zhang, S.B.; Chen, C.R.; Jiang, J.H.; Shen, G.L.; Yu, R.Q. Reusable electrochemical sensing platform for highly sensitive detection of small molecules based on structure-switching signaling aptamers. Anal. Chem. 2007, 79, 2933–2939. [Google Scholar] [CrossRef]
- Gulbakan, B.; Barylyuk, K.; Schneider, P.; Pillong, M.; Schneider, G.; Zenobi, R. Native electrospray ionization mass spectrometry reveals multiple facets of aptamer-ligand interactions: From mechanism to binding constants. J. Am. Chem. Soc. 2018, 140, 7486–7497. [Google Scholar] [CrossRef]
- Nutiu, R.; Li, Y. Structure-switching signaling aptamers. J. Am. Chem. Soc. 2003, 125, 4771–4778. [Google Scholar] [CrossRef]
- Gomes de Castro, M.A.; Hobartner, C.; Opazo, F. Aptamers provide superior stainings of cellular receptors studied under super-resolution microscopy. PLoS ONE 2017, 12, e0173050. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.H.; Yang, H.H.; Zhu, C.L.; Chen, X.; Chen, G.N. A graphene platform for sensing biomolecules. Angew. Chem. Int. Ed. 2009, 48, 4785–4787. [Google Scholar] [CrossRef]
- Ozaki, H.; Nishihira, A.; Wakabayashi, M.; Kuwahara, M.; Sawai, H. Biomolecular sensor based on fluorescence-labeled aptamer. Bioorg. Med. Chem. Lett. 2006, 16, 4381–4384. [Google Scholar] [CrossRef]
- Karunanayake Mudiyanselage, A.P.; Wu, R.; Leon-Duque, M.A.; Ren, K.; You, M. “Second-generation” fluorogenic RNA-based sensors. Methods 2019, 161, 24–34. [Google Scholar] [CrossRef] [PubMed]
- Perez-Gonzalez, C.; Lafontaine, D.A.; Penedo, J.C. Fluorescence-based strategies to investigate the structure and dynamics of aptamer-ligand complexes. Front. Chem. 2016, 4, 33. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Cao, Z.; Lu, Y. Functional nucleic acid sensors. Chem. Rev. 2009, 109, 1948–1998. [Google Scholar] [CrossRef] [PubMed]
- Jo, H.; Ban, C. Aptamer–nanoparticle complexes as powerful diagnostic and therapeutic tools. Exp. Mol. Med. 2016, 48, e230. [Google Scholar] [CrossRef]
- Kim, S.E.; Ahn, K.Y.; Park, J.S.; Kim, K.R.; Lee, K.E.; Han, S.S.; Lee, J. Fluorescent ferritin nanoparticles and application to the aptamer sensor. Anal. Chem. 2011, 83, 5834–5843. [Google Scholar] [CrossRef]
- Ma, L.; Liu, J. Catalytic nucleic acids: Biochemistry, chemical biology, biosensors, and nanotechnology. Iscience 2020, 23, 100815. [Google Scholar] [CrossRef] [PubMed]
- Teller, C.; Shimron, S.; Willner, I. Aptamer-DNAzyme hairpins for amplified biosensing. Anal. Chem. 2009, 81, 9114–9119. [Google Scholar] [CrossRef]
- Yang, Y.; Huang, J.; Yang, X.; Quan, K.; Wang, H.; Ying, L.; Xie, N.; Ou, M.; Wang, K. Aptazyme-gold nanoparticle sensor for amplified molecular probing in living cells. Anal. Chem. 2016, 88, 5981–5987. [Google Scholar] [CrossRef]
- Ouellet, J. RNA fluorescence with light-up aptamers. Front. Chem. 2016, 4, 29. [Google Scholar] [CrossRef]
- Bouhedda, F.; Autour, A.; Ryckelynck, M. Light-Up RNA Aptamers and their cognate fluorogens: From their development to their applications. Int. J. Mol. Sci. 2018, 19, 44. [Google Scholar] [CrossRef]
- Kim, Y.S.; Raston, N.H.; Gu, M.B. Aptamer-based nanobiosensors. Biosens. Bioelectron. 2016, 76, 2–19. [Google Scholar]
- Wang, R.E.; Zhang, Y.; Cai, J.; Cai, W.; Gao, T. Aptamer-based fluorescent biosensors. Curr. Med. Chem. 2011, 18, 4175–4184. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Mahapatra, C.; Chen, H.; Peng, X.; Ramakrishna, S.; Nanda, H.S. Recent developments in fluorescent aptasensors for detection of antibiotics. Curr. Opin. Biomed. Eng. 2020, 13, 16–24. [Google Scholar] [CrossRef]
- Gedi, V.; Kim, Y.P. Detection and characterization of cancer cells and pathogenic bacteria using aptamer-based nano-conjugates. Sensors 2014, 14, 18302–18327. [Google Scholar] [CrossRef] [PubMed]
- Goud, K.Y.; Sharma, A.; Hayat, A.; Catanante, G.; Gobi, K.V.; Gurban, A.M.; Marty, J.L. Tetramethyl-6-carboxyrhodamine quenching-based aptasensing platform for aflatoxin B1: Analytical performance comparison of two aptamers. Anal. Biochem. 2016, 508, 19–24. [Google Scholar] [CrossRef]
- Sharma, A.; Catanante, G.; Hayat, A.; Istamboulie, G.; Ben Rejeb, I.; Bhand, S.; Marty, J.L. Development of structure switching aptamer assay for detection of aflatoxin M1 in milk sample. Talanta 2016, 158, 35–41. [Google Scholar] [CrossRef]
- Dwidar, M.; Yokobayashi, Y. Development of a histamine aptasensor for food safety monitoring. Sci. Rep. 2019, 9, 16659. [Google Scholar] [CrossRef]
- Wu, C.; Chen, T.; Han, D.; You, M.; Peng, L.; Cansiz, S.; Zhu, G.; Li, C.; Xiong, X.; Jimenez, E.; et al. Engineering of switchable aptamer micelle flares for molecular imaging in living cells. ACS Nano 2013, 7, 5724–5731. [Google Scholar] [CrossRef]
- Lin, J.H.; Tseng, W.B.; Lin, K.C.; Lee, C.Y.; Chandirasekar, S.; Tseng, W.L.; Hsieh, M.M. Oligonucleotide-based fluorescent probe for sensing of cyclic diadenylate monophosphate in Bacteria and diadenosine polyphosphates in human tears. ACS Sens. 2016, 1, 1132–1139. [Google Scholar] [CrossRef]
- Wang, G.F.; Zhu, Y.H.; Chen, L.; Zhang, X.J. Dual hairpin-like molecular beacon based on coralyne-adenosine interaction for sensing melamine in dairy products. Talanta 2014, 129, 398–403. [Google Scholar] [CrossRef]
- Shi, Y.J.; Chen, Y.J.; Hu, W.P.; Chang, L.S. Detection of Naja atra cardiotoxin using adenosine-based molecular beacon. Toxins 2017, 9, 24. [Google Scholar] [CrossRef]
- Zhang, C.Y.; Johnson, L.W. Single quantum-dot-based aptameric nanosensor for cocaine. Anal. Chem. 2009, 81, 3051–3055. [Google Scholar] [CrossRef]
- Long, F.; Shi, H.C.; Wang, H.C. Fluorescence resonance energy transfer based aptasensor for the sensitive and selective detection of 17 beta-estradiol using a quantum dot-bioconjugate as a nano-bioprobe. RSC Adv. 2014, 4, 2935–2941. [Google Scholar] [CrossRef]
- Sabet, F.S.; Hosseini, M.; Khabbaz, H.; Dadmehr, M.; Ganjali, M.R. FRET-based aptamer biosensor for selective and sensitive detection of aflatoxin B1 in peanut and rice. Food Chem. 2017, 220, 527–532. [Google Scholar] [CrossRef]
- Oh, E.; Hong, M.Y.; Lee, D.; Nam, S.H.; Yoon, H.C.; Kim, H.S. Inhibition assay of biomolecules based on fluorescence resonance energy transfer (FRET) between quantum dots and gold nanoparticles. J. Am. Chem. Soc. 2005, 127, 3270–3271. [Google Scholar] [CrossRef]
- Kim, Y.P.; Oh, Y.H.; Oh, E.; Ko, S.; Han, M.K.; Kim, H.S. Energy transfer-based multiplexed assay of proteases by using gold nanoparticle and quantum dot conjugates on a surface. Anal. Chem. 2008, 80, 4634–4641. [Google Scholar] [CrossRef]
- Zhang, J.Q.; Wang, Y.S.; Xue, J.H.; He, Y.; Yang, H.X.; Liang, J.; Shi, L.F.; Xiao, X.L. A gold nanoparticles-modified aptamer beacon for urinary adenosine detection based on structure-switching/fluorescence-“turning on” mechanism. J. Pharm. Biomed. Anal. 2012, 70, 362–368. [Google Scholar] [CrossRef]
- Zheng, D.; Seferos, D.S.; Giljohann, D.A.; Patel, P.C.; Mirkin, C.A. Aptamer nano-flares for molecular detection in living cells. Nano Lett. 2009, 9, 3258–3261. [Google Scholar] [CrossRef]
- Zhang, J.; Wang, L.; Zhang, H.; Boey, F.; Song, S.; Fan, C. Aptamer-based multicolor fluorescent gold nanoprobes for multiplex detection in homogeneous solution. Small 2010, 6, 201–204. [Google Scholar] [CrossRef]
- Lee, E.S.; Kim, G.B.; Ryu, S.H.; Kim, H.; Yoo, H.H.; Yoon, M.Y.; Lee, J.W.; Gye, M.C.; Kim, Y.P. Fluorescing aptamer-gold nanosensors for enhanced sensitivity to bisphenol A. Sens. Actuators B Chem. 2018, 260, 371–379. [Google Scholar] [CrossRef]
- He, Y.; Wang, Z.G.; Tang, H.W.; Pang, D.W. Low background signal platform for the detection of ATP: When a molecular aptamer beacon meets graphene oxide. Biosens. Bioelectron. 2011, 29, 76–81. [Google Scholar] [CrossRef]
- Zhao, H.M.; Gao, S.; Liu, M.; Chang, Y.Y.; Fan, X.F.; Quan, X. Fluorescent assay for oxytetracycline based on a long-chain aptamer assembled onto reduced graphene oxide. Microchim. Acta 2013, 180, 829–835. [Google Scholar] [CrossRef]
- Wang, Y.; Li, Z.; Hu, D.; Lin, C.T.; Li, J.; Lin, Y. Aptamer/graphene oxide nanocomplex for in situ molecular probing in living cells. J. Am. Chem. Soc. 2010, 132, 9274–9276. [Google Scholar] [CrossRef]
- Yi, M.; Yang, S.; Peng, Z.; Liu, C.; Li, J.; Zhong, W.; Yang, R.; Tan, W. Two-photon graphene oxide/aptamer nanosensing conjugate for in vitro or in vivo molecular probing. Anal. Chem. 2014, 86, 3548–3554. [Google Scholar] [CrossRef]
- Joyce, G.F. The antiquity of RNA-based evolution. Nature 2002, 418, 214–221. [Google Scholar] [CrossRef]
- Link, K.H.; Breaker, R.R. Engineering ligand-responsive gene-control elements: Lessons learned from natural riboswitches. Gene Ther. 2009, 16, 1189–1201. [Google Scholar] [CrossRef]
- Walter, J.G.; Stahl, F. Aptazymes: Expanding the specificity of natural catalytic nucleic acids by application of in vitro selected oligonucleotides. Adv. Biochem. Eng. Biotechnol. 2020, 170, 107–119. [Google Scholar]
- Roth, A.; Weinberg, Z.; Chen, A.G.Y.; Kim, P.B.; Ames, T.D.; Breaker, R.R. A widespread self-cleaving ribozyme class is revealed by bioinformatics. Nat. Chem. Biol. 2014, 10, 56–60. [Google Scholar] [CrossRef]
- Porta, H.; Lizardi, P.M. An allosteric hammerhead ribozyme. Bio/Technology 1995, 13, 161–164. [Google Scholar] [CrossRef]
- Tang, J.; Breaker, R.R. Rational design of allosteric ribozymes. Chem. Biol. 1997, 4, 453–459. [Google Scholar] [CrossRef]
- Lan, T.; Lu, Y. Metal ion-dependent DNAzymes and their applications as biosensors. Inter. Met. Ions Nucleic Acids 2012, 10, 217–248. [Google Scholar]
- Li, J.; Lu, Y. A highly sensitive and selective catalytic DNA biosensor for lead ions. J. Am. Chem. Soc. 2000, 122, 10466–10467. [Google Scholar] [CrossRef]
- Liu, J.; Brown, A.K.; Meng, X.; Cropek, D.M.; Istok, J.D.; Watson, D.B.; Lu, Y. A catalytic beacon sensor for uranium with parts-per-trillion sensitivity and millionfold selectivity. Proc. Natl. Acad. Sci. USA 2007, 104, 2056–2061. [Google Scholar] [CrossRef] [PubMed]
- Huang, P.-J.J.; Liu, J. An ultrasensitive light-up Cu2+ biosensor using a new DNAzyme cleaving a phosphorothioate-modified substrate. Anal. Chem. 2016, 88, 3341–3347. [Google Scholar] [CrossRef]
- Qi, L.; Zhao, Y.X.; Yuan, H.; Bai, K.; Zhao, Y.; Chen, F.; Dong, Y.H.; Wu, Y.Y. Amplified fluorescence detection of mercury(II) ions (Hg2+) using target-induced DNAzyme cascade with catalytic and molecular beacons. Analyst 2012, 137, 2799–2805. [Google Scholar] [CrossRef] [PubMed]
- Torabi, S.F.; Wu, P.W.; McGhee, C.E.; Chen, L.; Hwang, K.; Zheng, N.; Cheng, J.J.; Lu, Y. In vitro selection of a sodium-specific DNAzyme and its application in intracellular sensing. Proc. Natl. Acad. Sci. USA 2015, 112, 5903–5908. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.H.; Zhang, Y.P.; Ding, J.S.; Liu, J.W. In vitro selection in serum: RNA-cleaving DNAzymes for measuring Ca2+ and Mg2+. ACS Sens. 2016, 1, 600–606. [Google Scholar] [CrossRef]
- McGhee, C.E.; Loh, K.Y.; Lu, Y. DNAzyme sensors for detection of metal ions in the environment and imaging them in living cells. Curr. Opin. Biotech. 2017, 45, 191–201. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Chang, D.R.; Li, Y.F. Discovery and biosensing applications of diverse RNA-cleaving DNAzymes. Acc. Chem. Res. 2017, 50, 2273–2283. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.-M.; Zhang, X.-B.; Kong, R.-M.; Yang, B.; Tan, W. A ligation-triggered DNAzyme cascade for amplified fluorescence detection of biological small molecules with zero-background signal. J. Am. Chem. Soc. 2011, 133, 11686–11691. [Google Scholar] [CrossRef]
- Huang, J.; He, Y.; Yang, X.; Wang, K.; Quan, K.; Lin, X. Split aptazyme-based catalytic molecular beacons for amplified detection of adenosine. Analyst 2014, 139, 2994–2997. [Google Scholar] [CrossRef]
- Zhao, X.-H.; Kong, R.-M.; Zhang, X.-B.; Meng, H.-M.; Liu, W.-N.; Tan, W.; Shen, G.-L.; Yu, R.-Q. Graphene–DNAzyme based biosensor for amplified fluorescence “turn-on” detection of Pb2+ with a high selectivity. Anal. Chem. 2011, 83, 5062–5066. [Google Scholar] [CrossRef] [PubMed]
- Gao, F.; Wu, J.; Yao, Y.; Zhang, Y.; Liao, X.; Geng, D.; Tang, D. Proximity hybridization triggered strand displacement and DNAzyme assisted strand recycling for ATP fluorescence detection in vitro and imaging in living cells. RSC Adv. 2018, 8, 28161–28171. [Google Scholar] [CrossRef]
- Kosman, J.; Juskowiak, B. Bioanalytical application of peroxidase-mimicking DNAzymes: Status and challenges. Adv. Biochem. Eng. Biotechnol. 2020, 170, 59–84. [Google Scholar]
- Alizadeh, N.; Salimi, A.; Hallaj, R. Hemin/G-quadruplex horseradish peroxidase-mimicking DNAzyme: Principle and biosensing application. Adv. Biochem. Eng. Biotechnol. 2020, 170, 85–106. [Google Scholar]
- Li, B.L.; Du, Y.; Li, T.; Dong, S.J. Investigation of 3,3′,5,5′-tetramethylbenzidine as colorimetric substrate for a peroxidatic DNAzyme. Anal. Chim. Acta 2009, 651, 234–240. [Google Scholar] [CrossRef]
- Li, T.; Wang, E.; Dong, S. G-Quadruplex-based DNAzyme as a sensing platform for ultrasensitive colorimetric potassium detection. Chem. Commun. 2009, 5, 580–582. [Google Scholar] [CrossRef]
- Liang, G.; Man, Y.; Li, A.; Jin, X.X.; Pan, L.G.; Liu, X.H. Chemiluminescence assay for detection of 2-hydroxyfluorene using the G-quadruplex DNAzyme-H2O2-luminol system. Microchim. Acta 2018, 185, 54. [Google Scholar] [CrossRef]
- Xu, J.; Lee, E.S.; Gye, M.C.; Kim, Y.P. Rapid and sensitive determination of bisphenol A using aptamer and split DNAzyme. Chemosphere 2019, 228, 110–116. [Google Scholar] [CrossRef]
- Tian, R.; Zhang, B.; Zhao, M.; Zou, H.; Zhang, C.; Qi, Y.; Ma, Q. Fluorometric enhancement of the detection of H 2 O 2 using different organic substrates and a peroxidase-mimicking polyoxometalate. RSC Adv. 2019, 9, 12209–12217. [Google Scholar] [CrossRef]
- Nakayama, S.; Sintim, H.O. Biomolecule detection with peroxidase-mimicking DNAzymes; expanding detection modality with fluorogenic compounds. Mol. Biosyst. 2010, 6, 95–97. [Google Scholar] [CrossRef]
- Liu, Y.; Lai, P.D.; Wang, J.R.; Xing, X.W.; Xu, L. A superior G-quadruplex DNAzyme through functionalized modification of the hemin cofactor. Chem. Commun. 2020, 56, 2427–2430. [Google Scholar] [CrossRef] [PubMed]
- Fan, X.; Li, H.; Zhao, J.; Lin, F.; Zhang, L.; Zhang, Y.; Yao, S. A novel label-free fluorescent sensor for the detection of potassium ion based on DNAzyme. Talanta 2012, 89, 57–62. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.H.; Lee, E.S.; Lee, D.Y.; Kim, Y.P. Facile Determination of Sodium Ion and Osmolarity in Artificial Tears by Sequential DNAzymes. Sensors 2017, 17, 2840–2849. [Google Scholar] [CrossRef]
- Fu, T.; Ren, S.; Gong, L.; Meng, H.; Cui, L.; Kong, R.-M.; Zhang, X.-B.; Tan, W. A label-free DNAzyme fluorescence biosensor for amplified detection of Pb2+-based on cleavage-induced G-quadruplex formation. Talanta 2016, 147, 302–306. [Google Scholar] [CrossRef]
- Zhang, Z.; Sharon, E.; Freeman, R.; Liu, X.; Willner, I. Fluorescence detection of DNA, adenosine-5′-triphosphate (ATP), and telomerase activity by zinc (II)-protoporphyrin IX/G-quadruplex labels. Anal. Chem. 2012, 84, 4789–4797. [Google Scholar] [CrossRef]
- Swetha, P.; Fan, Z.; Wang, F.; Jiang, J.-H. Genetically encoded light-up RNA aptamers and their applications for imaging and biosensing. J. Mater. Chem. B 2020, 8, 3382–3392. [Google Scholar] [CrossRef]
- Bai, J.; Luo, Y.; Wang, X.; Li, S.; Luo, M.; Yin, M.; Zuo, Y.; Li, G.; Yao, J.; Yang, H.; et al. A protein-independent fluorescent RNA aptamer reporter system for plant genetic engineering. Nat. Commun. 2020, 11, 3847. [Google Scholar] [CrossRef]
- Paige, J.S.; Wu, K.Y.; Jaffrey, S.R. RNA mimics of green fluorescent protein. Science 2011, 333, 642–646. [Google Scholar] [CrossRef]
- Paige, J.S.; Nguyen-Duc, T.; Song, W.J.; Jaffrey, S.R. Fluorescence imaging of cellular metabolites with RNA. Science 2012, 335, 1194. [Google Scholar] [CrossRef] [PubMed]
- Soni, R.; Sharma, D.; Krishna, A.M.; Sathiri, J.; Sharma, A. A highly efficient Baby Spinach-based minimal modified sensor (BSMS) for nucleic acid analysis. Org. Biomol. Chem. 2019, 17, 7222–7227. [Google Scholar] [CrossRef]
- Autour, A.; Sunny, C.Y.J.; Adam, D.C.; Abdolahzadeh, A.; Galli, A.; Panchapakesan, S.S.S.; Rueda, D.; Ryckelynck, M.; Unrau, P.J. Fluorogenic RNA Mango aptamers for imaging small non-coding RNAs in mammalian cells. Nat. Commun. 2018, 9, 656. [Google Scholar] [CrossRef] [PubMed]
- Cawte, A.D.; Unrau, P.J.; Rueda, D.S. Live cell imaging of single RNA molecules with fluorogenic Mango II arrays. Nat. Commun. 2020, 11, 1283. [Google Scholar] [CrossRef] [PubMed]
- Jepsen, M.D.E.; Sparvath, S.M.; Nielsen, T.B.; Langvad, A.H.; Grossi, G.; Gothelf, K.V.; Andersen, E.S. Development of a genetically encodable FRET system using fluorescent RNA aptamers. Nat. Commun. 2018, 9, 18. [Google Scholar] [CrossRef]
- Kim, H.; Jaffrey, S.R. A fluorogenic RNA-based snsor aectivated by metabolite-induced RNA dimerization. Cell Chem. Biol. 2019, 26, 1725–1731. [Google Scholar] [CrossRef] [PubMed]
- Yu, Q.; Shi, J.; Mudiyanselage, A.; Wu, R.; Zhao, B.; Zhou, M.; You, M. Genetically encoded RNA-based sensors for intracellular imaging of silver ions. Chem. Commun. 2019, 55, 707–710. [Google Scholar] [CrossRef] [PubMed]
- You, M.; Litke, J.L.; Wu, R.; Jaffrey, S.R. Detection of low-abundance metabolites in live cells using an RNA integrator. Cell Chem. Biol. 2019, 26, 471–481. [Google Scholar] [CrossRef] [PubMed]
- Babendure, J.R.; Adams, S.R.; Tsien, R.Y. Aptamers switch on fluorescence of triphenylmethane dyes. J. Am. Chem. Soc. 2003, 125, 14716–14717. [Google Scholar] [CrossRef]
- Aw, S.S.; Tang, M.X.M.; Teo, Y.N.; Cohen, S.M. A conformation-induced fluorescence method for microRNA detection. Nucleic Acids Res. 2016, 44, e92. [Google Scholar] [CrossRef]
- Dolgosheina, E.V.; Jeng, S.C.Y.; Panchapakesan, S.S.S.; Cojocaru, R.; Chen, P.S.K.; Wilson, P.D.; Hawkins, N.; Wiggins, P.A.; Unrau, P.J. RNA Mango aptamer-fluorophore: A bright, high-affinity complex for RNA labeling and tracking. ACS Chem. Biol. 2014, 9, 2412–2420. [Google Scholar] [CrossRef] [PubMed]
- Song, W.J.; Filonov-, G.S.; Kim, H.; Hirsch, M.; Li, X.; Moon, J.D.; Jaffrey, S.R. Imaging RNA polymerase III transcription using a photostable RNA-fluorophore complex. Nat. Chem. Biol. 2017, 13, 1187–1194. [Google Scholar] [CrossRef]
- Filonov, G.S.; Moon, J.D.; Svensen, N.; Jaffrey, S.R. Broccoli: Rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 2014, 136, 16299–16308. [Google Scholar] [CrossRef] [PubMed]
- Nilaratanakul, V.; Hauer, D.A.; Griffin, D.E. Development of encoded Broccoli RNA aptamers for live cell imaging of alphavirus genomic and subgenomic RNAs. Sci. Rep. 2020, 10, 5233. [Google Scholar] [CrossRef]
- You, M.X.; Litke, J.L.; Jaffrey, S.R. Imaging metabolite dynamics in living cells using a Spinach-based riboswitch. Proc. Natl. Acad. Sci. USA 2015, 112, E2756–E2765. [Google Scholar] [CrossRef]
- Truong, J.; Hsieh, Y.F.; Truong, L.; Jia, G.; Hammond, M.C. Designing fluorescent biosensors using circular permutations of riboswitches. Methods 2018, 143, 102–109. [Google Scholar] [CrossRef]
- Auslander, S.; Fuchs, D.; Hurlemann, S.; Auslander, D.; Fussenegger, M. Engineering a ribozyme cleavage-induced split fluorescent aptamer complementation assay. Nucleic Acids Res. 2016, 44, e94. [Google Scholar] [CrossRef] [PubMed]
- Litke, J.L.; Jaffrey, S.R. Highly efficient expression of circular RNA aptamers in cells using autocatalytic transcripts. Nat. Biotechnol. 2019, 37, 667–675. [Google Scholar] [CrossRef]
- Tran, T.T.T.; Delgado, A.; Jeong, S. Organ-on-a-chip: The future of therapeutic aptamer research? BioChip J. 2021, 10. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, E.-S.; Lee, J.M.; Kim, H.-J.; Kim, Y.-P. Fluorogenic Aptasensors with Small Molecules. Chemosensors 2021, 9, 54. https://doi.org/10.3390/chemosensors9030054
Lee E-S, Lee JM, Kim H-J, Kim Y-P. Fluorogenic Aptasensors with Small Molecules. Chemosensors. 2021; 9(3):54. https://doi.org/10.3390/chemosensors9030054
Chicago/Turabian StyleLee, Eun-Song, Jeong Min Lee, Hea-Jin Kim, and Young-Pil Kim. 2021. "Fluorogenic Aptasensors with Small Molecules" Chemosensors 9, no. 3: 54. https://doi.org/10.3390/chemosensors9030054
APA StyleLee, E.-S., Lee, J. M., Kim, H.-J., & Kim, Y.-P. (2021). Fluorogenic Aptasensors with Small Molecules. Chemosensors, 9(3), 54. https://doi.org/10.3390/chemosensors9030054







