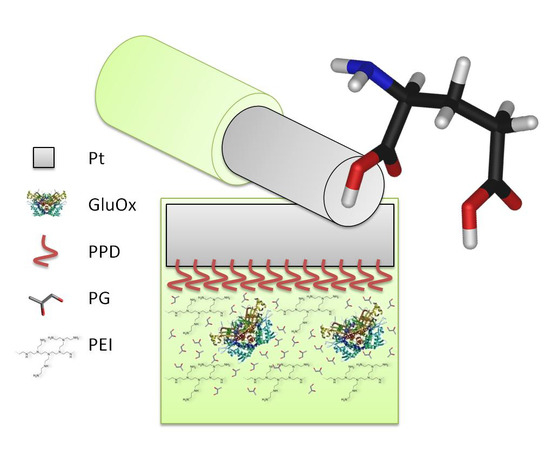
Propylene Glycol Stabilizes the Linear Response of Glutamate Biosensor: Potential Implications for In-Vivo Neurochemical Monitoring
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals and Reagents
2.2. Instrumentation and Software
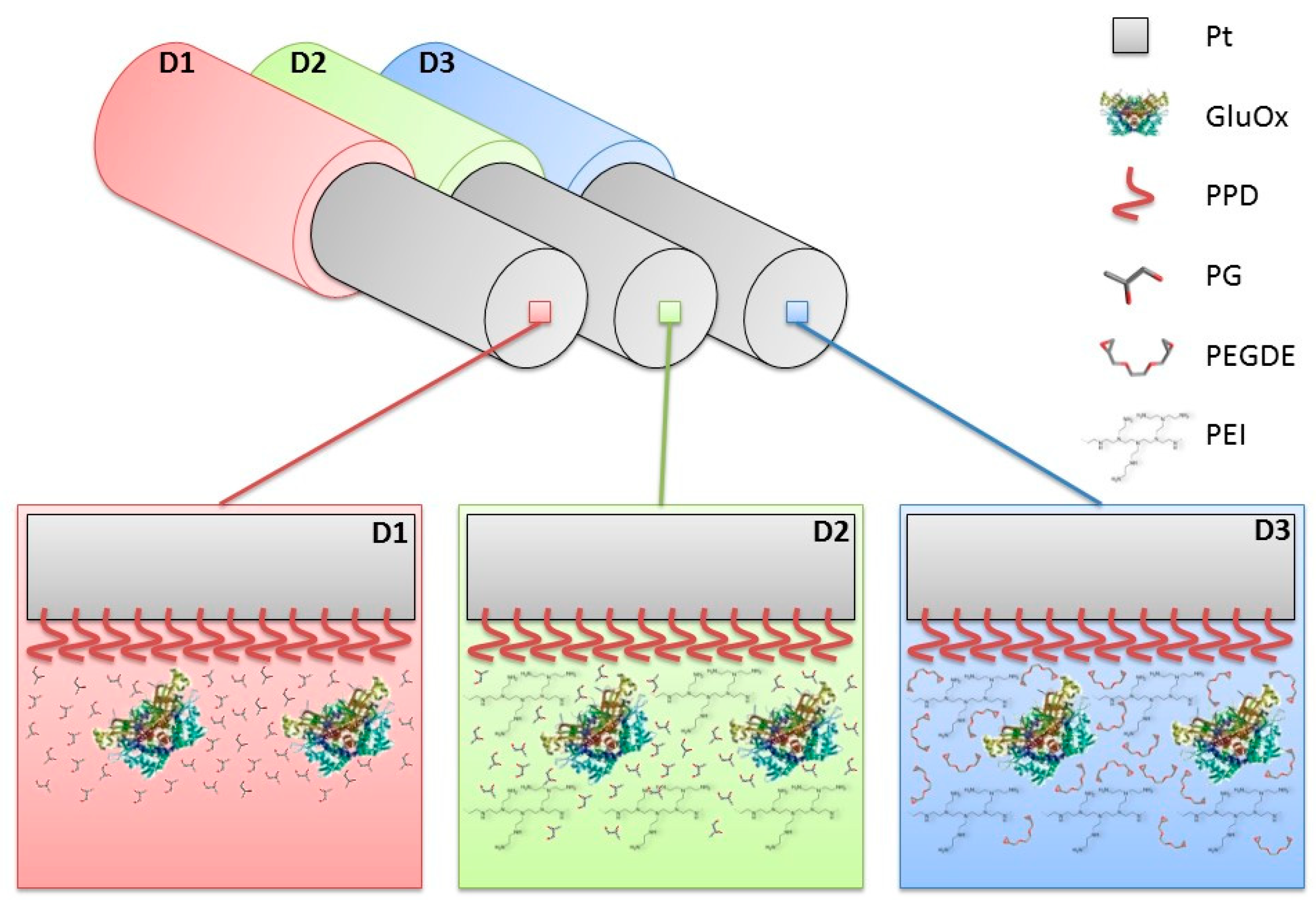
2.3. Biosensor Construction and Characterisation
2.4. Statistical Analysis
3. Results and Discussion
3.1. In Vitro Performances of Glutamate Biosensors
3.2. Effects of the Difference of Enzyme Concentration in Glutamate Biosensor Parameters
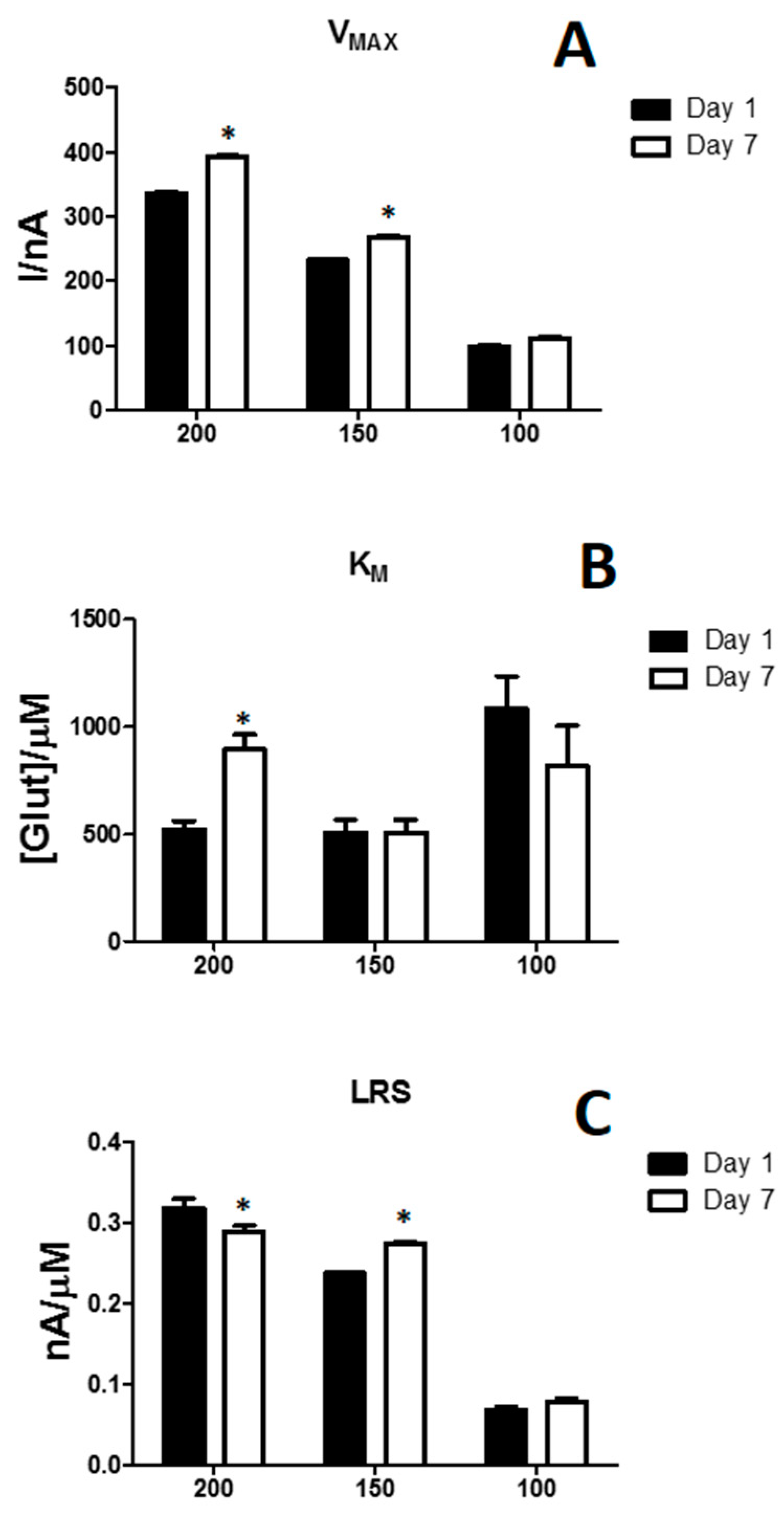
3.3. Effects of the Variation of Propylene Glycol in Glutamate Biosensor Performances
3.4. In Vitro Assessment of Continuous 48 h Glutamate Measurement in PBS
3.5. Effects of PG or PEGDE on Glutamate Biosensor Ageing
4. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Ford, R.; Quinn, S.J.; O’Neill, R.D. Characterization of Biosensors Based on Recombinant Glutamate Oxidase: Comparison of Crosslinking Agents in Terms of Enzyme Loading and Efficiency Parameters. Sensors 2016, 16, 1565. [Google Scholar] [CrossRef] [PubMed]
- Son, H.; Baek, J.H.; Go, B.S.; Jung, D.H.; Sontakke, S.B.; Chung, H.J.; Lee, D.H.; Roh, G.S.; Kang, S.S.; Cho, G.J.; et al. Glutamine has antidepressive effects through increments of glutamate and glutamine levels and glutamatergic activity in the medial prefrontal cortex. Neuropharmacology 2018, 143, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Sirca, D.; Vardeu, A.; Pinna, M.; Diana, M.; Enrico, P. A robust, state-of-the-art amperometric microbiosensor for glutamate detection. Biosens. Bioelectron. 2014, 61, 526–531. [Google Scholar] [CrossRef] [PubMed]
- Mikeladze, E.; Collins, A.; Sukhacheva, M.; Netrusov, A.; Csöregi, E. Characterization of a Glutamate Biosensor Based on a Novel Glutamate Oxidase Integrated into a Redox Hydrogel. Electroanalysis 2002, 14, 1052–1059. [Google Scholar] [CrossRef]
- Burmeister, J.J.; Pomerleau, F.; Palmer, M.; Day, B.K.; Huettl, P.; Gerhardt, G.A. Improved ceramic-based multisite microelectrode for rapid measurements of L-glutamate in the CNS. J. Neurosci. Methods 2002, 119, 163–171. [Google Scholar] [CrossRef]
- Rieben, N.; Cherouati, N.; Martinez, K.L. Glutamate Monitoring In Vitro and In Vivo: Recent Progress in the Field of Glutamate Biosensors. J. Nanosci. 2009, 1, 100–115. [Google Scholar] [CrossRef]
- Kang, Y.; Henchcliffe, C.; Verma, A.; Vallabhajosula, S.; He, B.; Kothari, P.J.; Pryor, K.O.; Mozley, P.D. 18F-FPEB PET/CT Shows mGluR5 Upregulation in Parkinson’s Disease. J. Neuroimag. 2018. [Google Scholar] [CrossRef] [PubMed]
- Jakaria, M.; Park, S.Y.; Haque, M.E.; Karthivashan, G.; Kim, I.S.; Ganesan, P.; Choi, D.K. Neurotoxic Agent-Induced Injury in Neurodegenerative Disease Model: Focus on Involvement of Glutamate Receptors. Front. Mol. Neurosci. 2018, 11, 307. [Google Scholar] [CrossRef] [PubMed]
- Müller Herde, A.; Schibli, R.; Weber, M.; Ametamey, S.M. Metabotropic glutamate receptor subtype 5 is altered in LPS-induced murine neuroinflammation model and in the brains of AD and ALS patients. Eur. J. Nucl. Med. Mol. Imaging 2018. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, L.; Tucci, S.; Guzman, N.; Paez, X. In vivo monitoring of glutamate in the brain by microdialysis and capillary electrophoresis with laser-induced fluorescence detection. J. Chromatogr. A 1993, 652, 393–398. [Google Scholar] [CrossRef]
- Globus, M.Y.; Busto, R.; Dietrich, W.D.; Martinez, E.; Valdes, I.; Ginsberg, M.D. Effect of ischemia on the in vivo release of striatal dopamine, glutamate, and gamma-aminobutyric acid studied by intracerebral microdialysis. J. Neurochem. 1988, 51, 1455–1464. [Google Scholar] [CrossRef] [PubMed]
- Sheng, W.; Hang, H.W.; Ruan, D.Y. In vivo microdialysis study of the relationship between lead-induced impairment of learning and neurotransmitter changes in the hippocampus. Environ. Toxicol. Pharmacol. 2005, 20, 233–240. [Google Scholar] [CrossRef] [PubMed]
- Kaul, S.; Faiman, M.D.; Lunte, C.E. Determination of GABA, glutamate and carbamathione in brain microdialysis samples by capillary electrophoresis with fluorescence detection. Electrophoresis 2011, 32, 284–291. [Google Scholar] [CrossRef] [PubMed]
- Murakami, G.; Nakamura, M.; Takita, M.; Ishida, Y.; Ueki, T.; Nakahara, D. Brain Rewarding Stimulation Reduces Extracellular Glutamate Through Glial Modulation in Medial Prefrontal Cortex of Rats. Neuropsychopharmacology 2015, 40, 2686–2695. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, R.D.; Lowry, J.P.; Rocchitta, G.; McMahon, C.P.; Serra, P.A. Designing sensitive and selective polymer/enzyme composite biosensors for brain monitoring in vivo. Trends Anal. Chem. 2008, 27, 78–88. [Google Scholar] [CrossRef]
- Hamdan, S.K.; Mohd Zain, A. In vivo Electrochemical Biosensor for Brain Glutamate Detection: A Mini Review. Malays. J. Med. Sci. 2014, 21, 12–26. [Google Scholar] [PubMed]
- Rocchitta, G.; Spanu, A.; Babudieri, S.; Latte, G.; Madeddu, G.; Galleri, G.; Nuvoli, S.; Bagella, P.; Demartis, M.I.; Fiore, V.; et al. Enzyme biosensors for biomedical applications: Strategies for safeguarding analytical performances in biological fluids. Sensors 2016, 16, 780. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Schellhorn, H.E. New developments and novel therapeutic perspectives for vitamin C. J. Nutr. 2007, 137, 2171–2184. [Google Scholar] [CrossRef] [PubMed]
- Rothwell, S.A.; Kinsella, M.E.; Zain, Z.M.; Serra, P.A.; Rocchitta, G.; Lowry, J.P.; O’Neill, R.D. Contributions by a novel edge effect to the permselectivity of an electrosynthesized polymer for microbiosensor applications. Anal. Chem. 2009, 81, 3911–3918. [Google Scholar] [CrossRef] [PubMed]
- Grieshaber, D.; MacKenzie, R.; Vörös, J.; Reimhult, E. Electrochemical Biosensors–Sensor Principles and Architectures. Sensors 2008, 8, 1400–1458. [Google Scholar] [CrossRef] [PubMed]
- Rathee, K.; Dhull, V.; Dhull, R.; Singh, S. Biosensors based on electrochemical lactate detection: A comprehensive review. Biochem. Biophys. Rep. 2015, 5, 35–54. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, R.D.; Chang, S.C.; Lowry, J.P.; McNeil, C.J. Comparisons of platinum, gold, palladium and glassy carbon as electrode materials in the design of biosensors for glutamate. Biosens. Bioelectron. 2004, 19, 1521–1528. [Google Scholar] [CrossRef] [PubMed]
- McMahon, C.P.; Rocchitta, G.; Serra, P.A.; Kirwan, S.M.; Lowry, J.P.; O’Neill, R.D. The efficiency of immobilised glutamate oxidase decreases with surface enzyme loading: An electrostatic effect, and reversal by a polycation significantly enhances biosensor sensitivity. Analyst 2006, 131, 68–72. [Google Scholar] [CrossRef] [PubMed]
- Rocchitta, G.; Secchi, O.; Alvau, M.D.; Migheli, R.; Calia, G.; Bazzu, G.; Farina, D.; Desole, M.S.; O’Neill, R.D.; Serra, P.A. Development and characterization of an implantable biosensor for telemetric monitoring of ethanol in the brain of freely moving rats. Anal. Chem. 2012, 84, 7072–7079. [Google Scholar] [CrossRef] [PubMed]
- McMahon, C.P.; Rocchitta, G.; Kirwan, S.M.; Killoran, S.J.; Serra, P.A.; Lowry, J.P.; O’Neill, R.D. Oxygen tolerance of an implantable polymer/enzyme composite glutamate biosensor displaying polycation-enhanced substrate sensitivity. Biosens. Bioelectron. 2007, 22, 1466–1473. [Google Scholar] [CrossRef] [PubMed]
- Pisoschi, A.M. Biosensors as Bio-Based Materials in Chemical Analysis: A Review. J. Biobased Mater. Bioenergy 2013, 7, 19–38. [Google Scholar] [CrossRef]
- Palmisano, F.; Zambonin, P.G.; Centonze, D. Amperometric biosensors based on electrosynthesised polymeric films. Fresenius J. Anal. Chem. 2000, 366, 586–601. [Google Scholar] [PubMed]
- Kirwan, S.M.; Rocchitta, G.; McMahon, C.P.; Craig, J.D.; Killoran, S.J.; O’Brien, K.B.; Serra, P.A.; Lowry, J.P.; O’Neill, R.D. Modifications of poly(o-phenylenediamine) permselective layer on pt-ir for biosensor application in neurochemical monitoring. Sensors 2007, 7, 420–437. [Google Scholar] [CrossRef]
- Rothwell, S.A.; Killoran, S.J.; O’Neill, R.D. Enzyme immobilization strategies and electropolymerization conditions to control sensitivity and selectivity parameters of a polymer-enzyme composite glucose biosensor. Sensors 2010, 10, 6439–6462. [Google Scholar] [CrossRef] [PubMed]
- Cosnier, S.; Holzinger, M. Electrosynthesized polymers for biosensing. Chem. Soc. Rev. 2011, 40, 2146–2156. [Google Scholar] [CrossRef] [PubMed]
- Pal, R.K.; Pradhan, S.; Narayanan, L.; Yadavalli, V.K. Micropatterned conductive polymer biosensors on flexible PDMS films. Sens. Actuator B Chem. 2018, 259, 498–504. [Google Scholar] [CrossRef]
- House, J.L.; Anderson, E.M.; Ward, W.K. Immobilization Techniques to Avoid Enzyme Loss from Oxidase-Based Biosensors: A One-Year Study. J. Diabetes Sci. Technol. 2007, 1, 18–27. [Google Scholar] [CrossRef] [PubMed]
- El Kaoutit, M.; Naranjo-Rodriguez, I.; Dominguez, M.; Hidalgo-Hidalgo de Cisneros, J.L. Overcoming the adverse effects of crosslinking in biosensors via addition of PEG: Improved sensing of hydrogen peroxide using immobilized peroxidase. Microchim. Acta. 2011, 175, 241–250. [Google Scholar] [CrossRef]
- Marques, M.E.; Mansur, A.A.P.; Mansur, H.S. Chemical functionalization of surfaces for building three-dimensional engineered biosensors. Appl. Surf. Sci. 2013, 275, 347–360. [Google Scholar] [CrossRef]
- Burmeister, J.J.; Davis, V.A.; Quintero, J.E.; Pomerleau, F.; Huettl, P.; Gerhardt, G.A. Glutaraldehyde cross-linked glutamate oxidase coated microelectrode arrays: Selectivity and resting levels of glutamate in the CNS. ACS Chem. Neurosci. 2013, 4, 721–728. [Google Scholar] [CrossRef] [PubMed]
- Vasylieva, N.; Barnych, B.; Meiller, A.; Maucler, C.; Pollegioni, L.; Lin, J.S.; Barbier, D.; Marinesco, S. Covalent enzyme immobilization by poly(ethylene glycol) diglycidyl ether (PEGDE) for microelectrode biosensor preparation. Biosens. Bioelectron. 2011, 26, 3993–4000. [Google Scholar] [CrossRef] [PubMed]
- Vasylieva, N.; Maucler, C.; Meiller, A.; Viscogliosi, H.; Lieutaud, T.; Barbier, D.; Marinesco, S. Immobilization method to preserve enzyme specificity in biosensors: consequences for brain glutamate detection. Anal. Chem. 2013, 85, 2507–2515. [Google Scholar] [CrossRef] [PubMed]
- Zigah, D.; Pellissier, M.; Fabre, B.; Barrière, F.; Hapiot, P. Covalent immobilization and SECM analysis in feedback mode of glucose oxidase on a modified oxidized silicon surface. J. Electroanal. Chem. 2009, 628, 144–147. [Google Scholar] [CrossRef]
- Gekko, K. Mechanism of polyol-induced protein stabilization: solubility of amino acids and diglycine in aqueous polyol solutions. J. Biochem. 1981, 90, 1633–1641. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Singh, J. Effect of polyols on the conformational stability and biological activity of a model protein lysozyme. AAPS Pharmscitech 2003, 4, E42. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.A. Additive effects of alcohols and polyols on thermostability of pepper leaf extracts. J. Amer. Soc. Hort. Sci. 2007, 132, 67–72. [Google Scholar]
- Liu, F.F.; Ji, L.; Zhang, L.; Dong, X.Y.; Sun, Y. Molecular basis for polyol-induced protein stability revealed by molecular dynamics simulations. J. Chem. Phys. 2010, 132, 225103. [Google Scholar] [CrossRef] [PubMed]
- Yusoff, M.; Hassan, B.N.; Ikhwanuddin, M.; Sheriff, S.M.; Hashim, F.; Mustafa, S.; Koh, I.C.C. Successful sperm cryopreservation of the brown-marbled grouper, Epinephelus fuscoguttatus using propylene glycol as cryoprotectant. Cryobiology 2018, 81, 168–173. [Google Scholar] [CrossRef] [PubMed]
- Calia, G.; Rocchitta, G.; Migheli, R.; Puggioni, G.; Spissu, Y.; Bazzu, G.; Mazzarello, V.; Lowry, J.P.; O’Neill, R.D.; Desole, M.S.; Serra, P.A. Biotelemetric monitoring of brain neurochemistry in conscious rats using microsensors and biosensors. Sensors 2009, 9, 2511–2523. [Google Scholar] [CrossRef] [PubMed]
- Ryan, M.R.; Lowry, J.P.; O’Neill, R.D. Biosensor for neurotransmitter L-glutamic acid designed for efficient use of L-glutamate oxidase and effective rejection of interference. Analyst 1997, 122, 1419–1424. [Google Scholar] [CrossRef] [PubMed]
- ICH Harmonised Tripartite Guideline Q2 (R1) (2005) Validation of Analytical Procedures: Text and Methodology. Available online: https://pacificbiolabs.com/wp-content/uploads/2017/12/Q2_R1__Guideline-4.pdf (accessed on 28 November 2018).
- Secchi, O.; Zinellu, M.; Spissu, Y.; Pirisinu, M.; Bazzu, G.; Migheli, R.; Desole, M.S.; O’Neill, R.D.; Serra, P.A.; Rocchitta, G. Further in-vitro characterization of an implantable biosensor for ethanol monitoring in the brain. Sensors 2013, 13, 9522–9535. [Google Scholar] [CrossRef] [PubMed]
- Qarawi, M.A. Role of polyols and surfactants in liquid protein formulations. World J. Pharm. Sci. 2017, 6, 190–196. [Google Scholar] [CrossRef]
- Panuszko, A.; Bruździak, P.; Kaczkowska, E.; Stangret, J. General Mechanism of Osmolytes’ Influence on Protein Stability Irrespective of the Type of Osmolyte Cosolvent. J. Phys. Chem. B 2016, 120, 11159–11169. [Google Scholar] [CrossRef] [PubMed]
- Timasheff, S.N. Protein-solvent preferential interactions, protein hydration, and the modulation of biochemical reactions by solvent components. Proc. Natl. Acad. Sci. USA. 2002, 99, 9721–9726. [Google Scholar] [CrossRef] [PubMed]
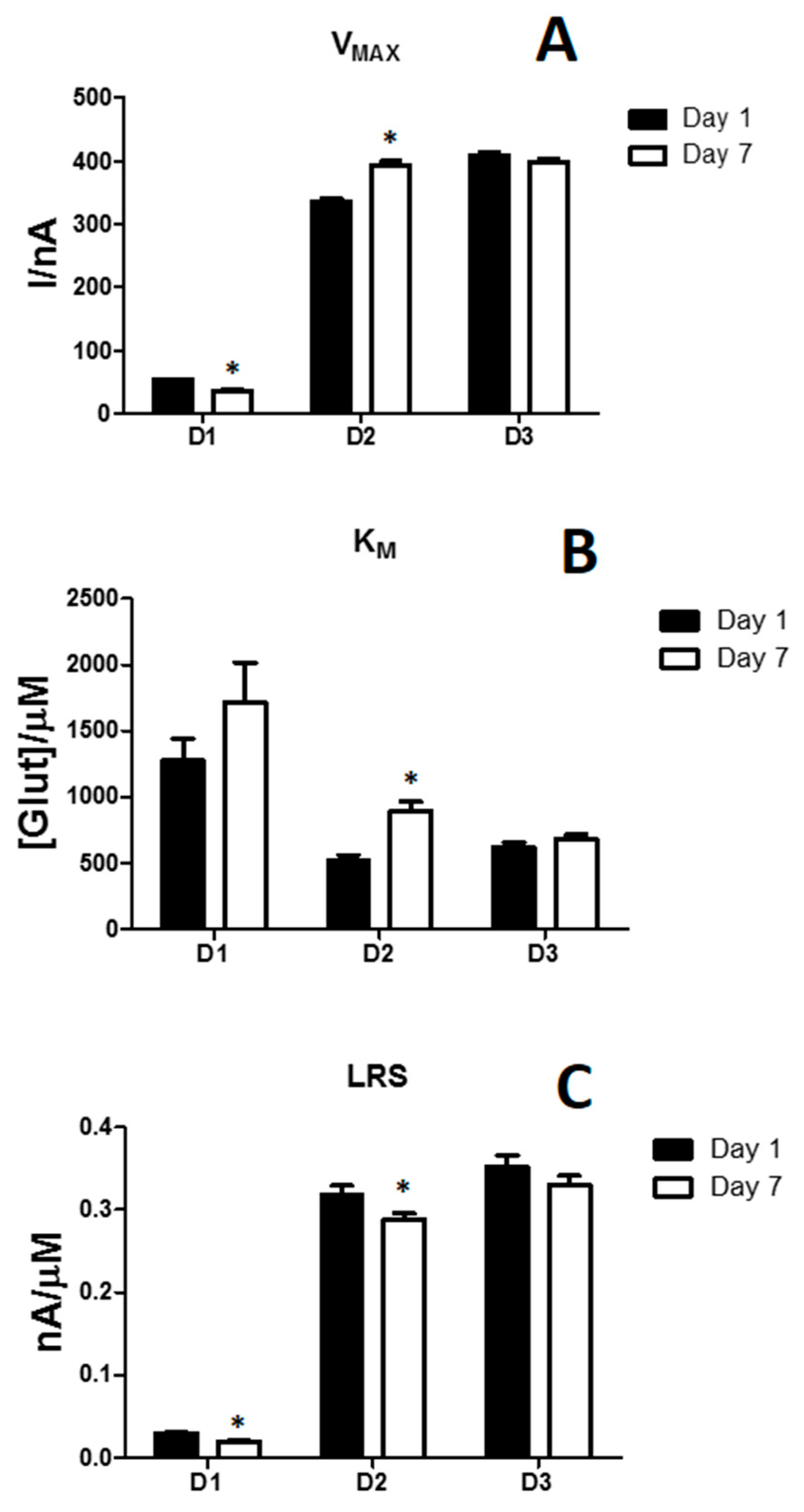
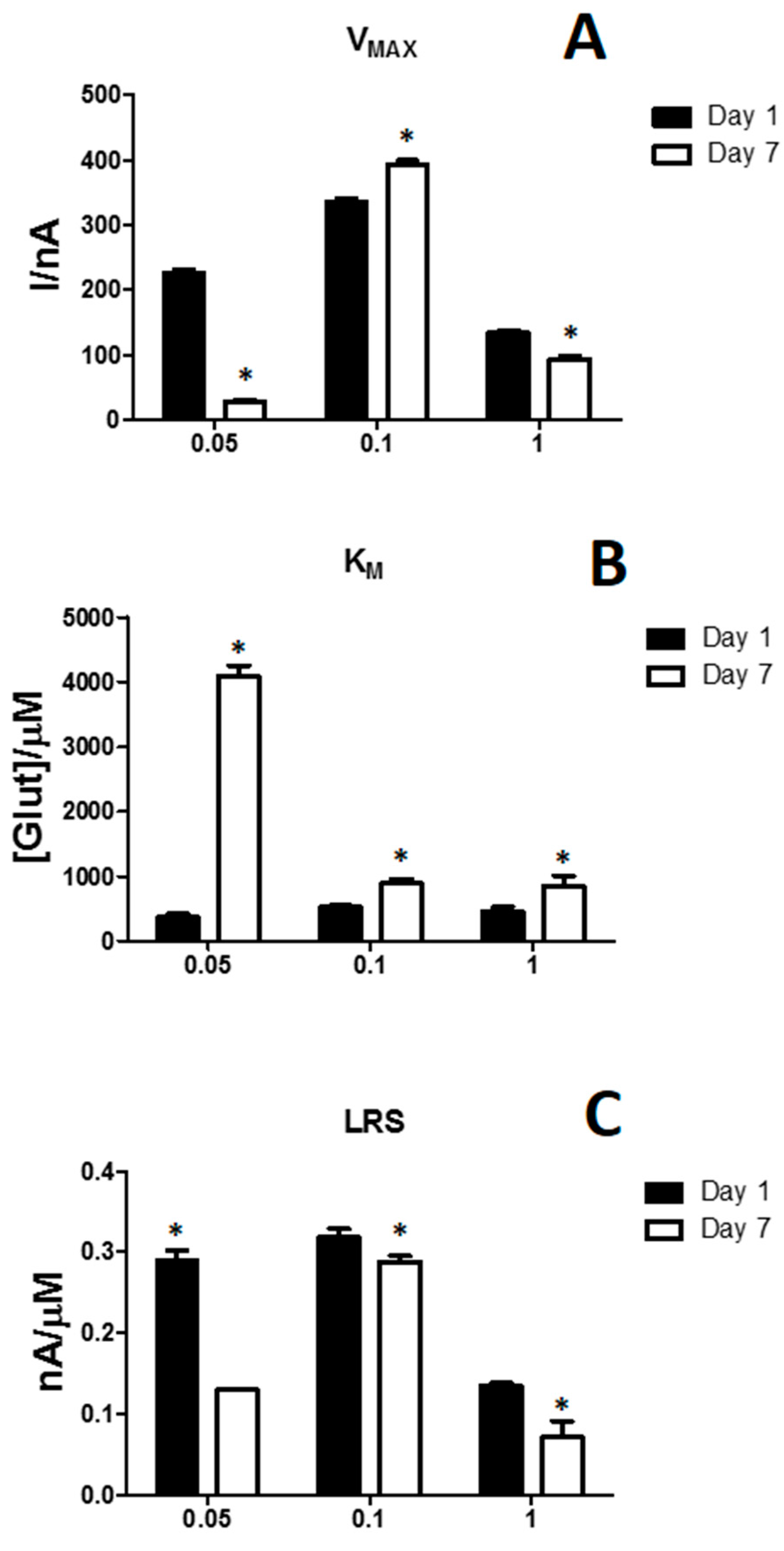
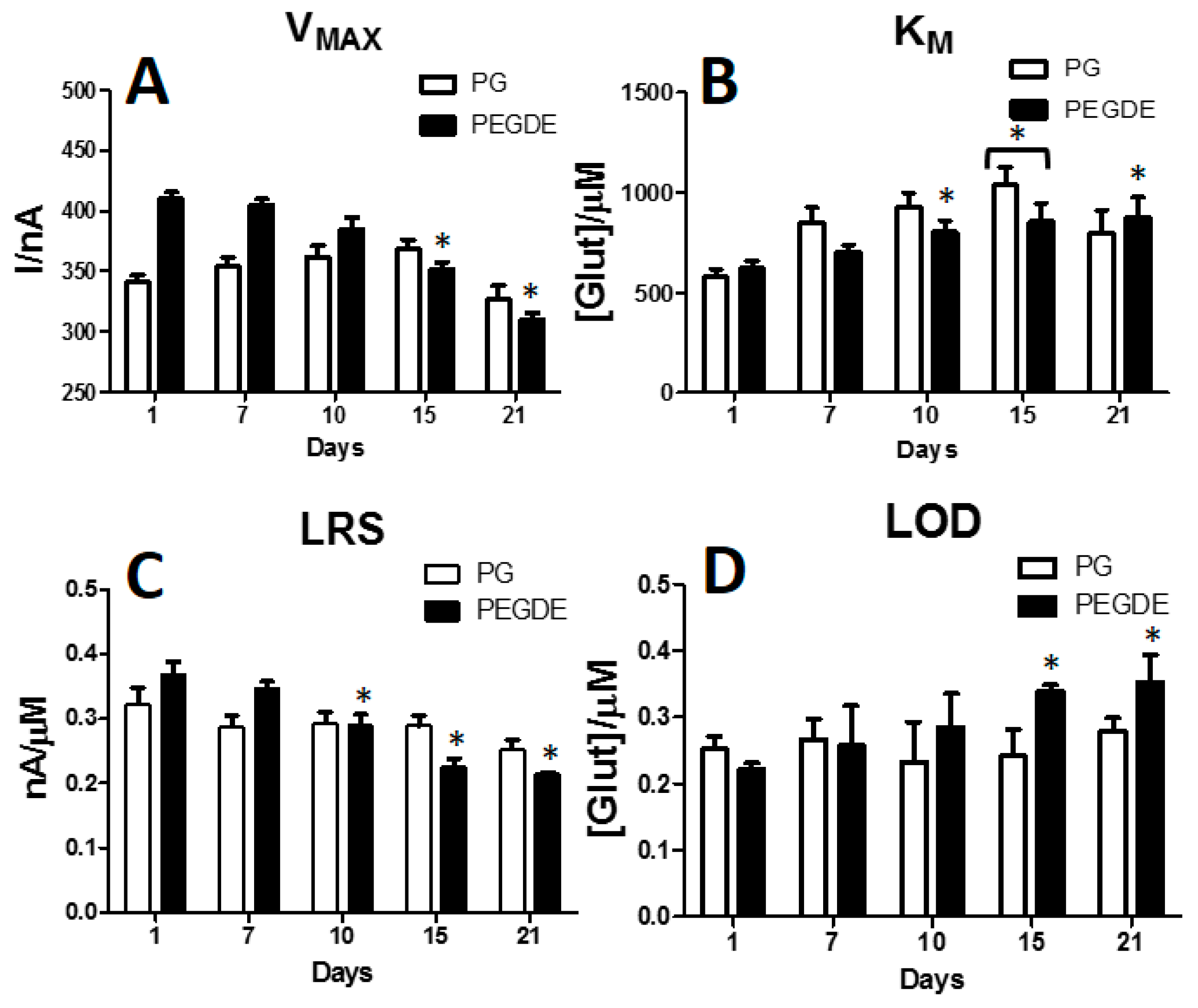
| Parameters | D1 | D2 | D3 |
|---|---|---|---|
| DAY 1 | |||
| VMAX (nA) | 52.71 ± 1.70 | 335.91 ± 5.03 | 409.73 ± 5.58 |
| KM [µM] | 1275.02 ± 163.81 | 524.53 ± 37.62 | 620.23 ± 37.65 |
| LRS (nA/µM) | 0.030 ± 0.001 | 0.318 ± 0.011 | 0.352 ± 0.014 |
| Ilim (nA) | 1.984 ± 0,056 | 1.450 ± 0.124 | 0.867 ± 0.074 |
| ΔI | −0.027 | 0.482 | 0.388 |
| LOD [µM] | 0.265 | 0.252 | 0.267 |
| DAY 7 | |||
| VMAX (nA) | 36.23 ± 1.67 | 392.16 ± 7.73 | 399.5 ± 4.88 |
| KM [µM] | 1717.12 ± 299.12 | 892.84 ± 73.55 | 680.34 ± 34.85 |
| LRS (nA/µM) | 0.020 ± 0.001 | 0.288 ± 0.008 | 0.330 ± 0.011 |
| Ilim(nA) | 26.072 ± 12.178 | 2.153 ± 0.430 | 4.127± 1.103 |
| ΔI | 15.103 | 1.061 | 1.848 |
| LOD[µM] | 0.270 | 0.221 | 0.257 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rocchitta, G.; Bacciu, A.; Arrigo, P.; Migheli, R.; Bazzu, G.; Serra, P.A. Propylene Glycol Stabilizes the Linear Response of Glutamate Biosensor: Potential Implications for In-Vivo Neurochemical Monitoring. Chemosensors 2018, 6, 58. https://doi.org/10.3390/chemosensors6040058
Rocchitta G, Bacciu A, Arrigo P, Migheli R, Bazzu G, Serra PA. Propylene Glycol Stabilizes the Linear Response of Glutamate Biosensor: Potential Implications for In-Vivo Neurochemical Monitoring. Chemosensors. 2018; 6(4):58. https://doi.org/10.3390/chemosensors6040058
Chicago/Turabian StyleRocchitta, Gaia, Andrea Bacciu, Paola Arrigo, Rossana Migheli, Gianfranco Bazzu, and Pier Andrea Serra. 2018. "Propylene Glycol Stabilizes the Linear Response of Glutamate Biosensor: Potential Implications for In-Vivo Neurochemical Monitoring" Chemosensors 6, no. 4: 58. https://doi.org/10.3390/chemosensors6040058
APA StyleRocchitta, G., Bacciu, A., Arrigo, P., Migheli, R., Bazzu, G., & Serra, P. A. (2018). Propylene Glycol Stabilizes the Linear Response of Glutamate Biosensor: Potential Implications for In-Vivo Neurochemical Monitoring. Chemosensors, 6(4), 58. https://doi.org/10.3390/chemosensors6040058