Towards the Improved Accuracy of Hepatitis E Diagnosis in Vulnerable and Target Groups: A Global Perspective on the Current State of Knowledge and the Implications for Practice
Abstract
1. Introduction
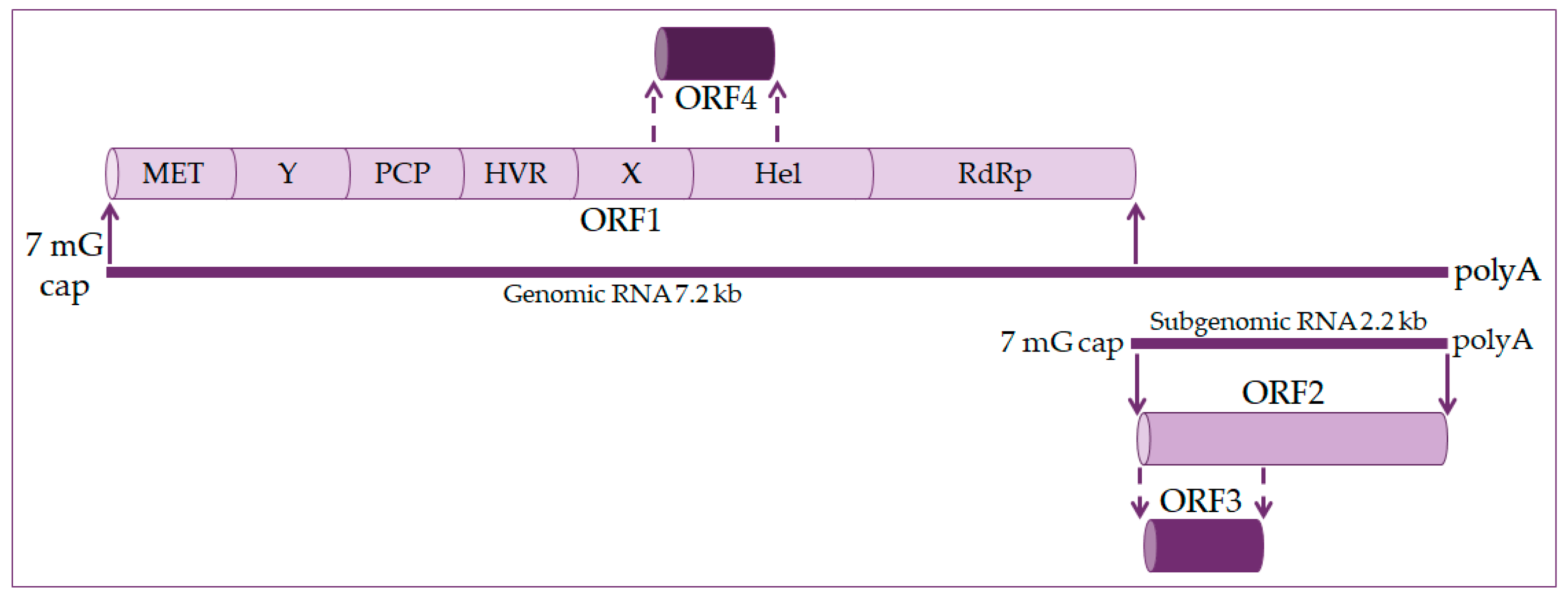
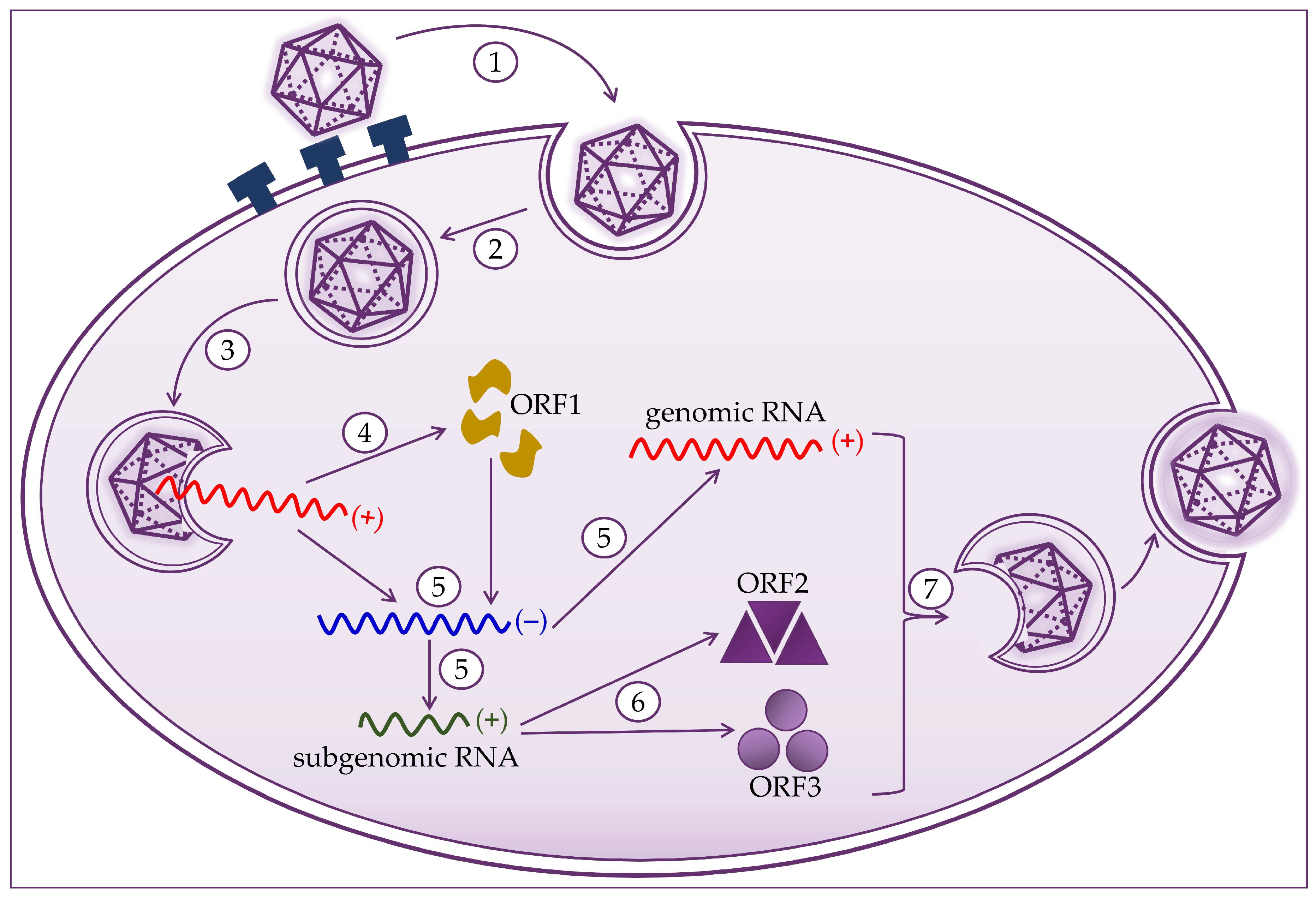
2. Molecular Biology of the Hepatitis E Virus
3. Epidemiology of HEV
4. HEV Infections in Pregnant Women
4.1. Clinical Presentation of Hepatitis E in Pregnant Women Depending on the HEV Genotype
4.2. Prevention Measures of HEV Infections in Pregnant Women
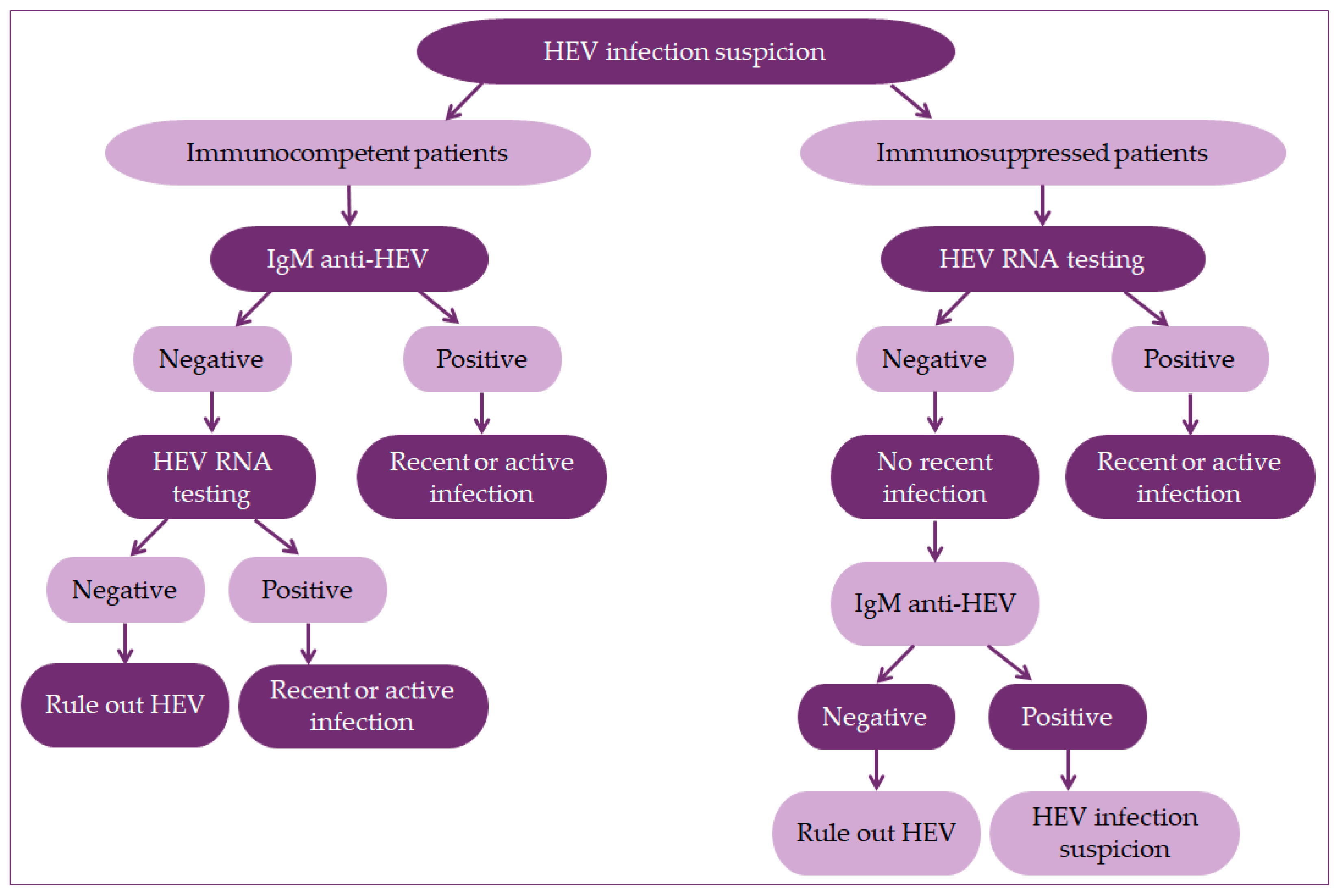
5. Laboratory Diagnostics of Hepatitis E Virus
5.1. Serological Tests
5.2. Nucleic Acid Tests
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- LeDesma, R.; Nimgaonkar, I.; Ploss, A. Hepatitis E Virus Replication. Viruses 2019, 11, 719. [Google Scholar] [CrossRef]
- Emerson, S.U.; Purcell, R.H. Hepatitis E virus. Rev. Med. Virol. 2003, 13, 145–154. [Google Scholar] [CrossRef]
- Lhomme, S.; Marion, O.; Abravanel, F.; Izopet, J.; Kamar, N. Clinical Manifestations, Pathogenesis and Treatment of Hepatitis E Virus Infections. J. Clin. Med. 2020, 9, 331. [Google Scholar] [CrossRef]
- Sridhar, S.; Teng, J.L.L.; Chiu, T.H.; Lau, S.K.P.; Woo, P.C.Y. Hepatitis E virus genotypes and evolution: Emergence of camel hepatitis E variants. Int. J. Mol. Sci. 2017, 18, 869. [Google Scholar] [CrossRef]
- Chaudhry, S.A.; Verma, N.; Koren, G. Hepatitis E infection during pregnancy. Can. Fam. Physician 2015, 61, 607–608. [Google Scholar]
- Fousekis, F.S.; Mitselos, I.V.; Christodoulou, D.K. Extrahepatic manifestations of hepatitis E virus: An overview. Clin. Mol. Hepatol. 2020, 26, 16–23. [Google Scholar] [CrossRef]
- Nair, V.P.; Anang, S.; Subramani, C.; Madhvi, A.; Bakshi, K.; Srivastava, A.; Shalimar; Nayak, B.; Kumar, C.T.R.; Surjit, M. Endoplasmic Reticulum Stress Induced Synthesis of a Novel Viral Factor Mediates Efficient Replication of Genotype-1 Hepatitis E Virus. PLoS Pathog. 2016, 12, e1005521. [Google Scholar] [CrossRef]
- Meng, X.J. Recent advances in Hepatitis E Virus. J. Viral Hepat. 2010, 17, 153–161. [Google Scholar] [CrossRef]
- Goel, A.; Aggarwal, R. Hepatitis E: Epidemiology, Clinical Course, Prevention, and Treatment. Gastroenterol. Clin. N. Am. 2020, 49, 315–330. [Google Scholar] [CrossRef]
- Yin, X.; Feng, Z. Hepatitis E Virus Entry. Viruses 2019, 11, 883. [Google Scholar] [CrossRef]
- Wu, C.; Wu, X.; Xia, J. Hepatitis E virus infection during pregnancy. Virol. J. 2020, 17, 1–11. [Google Scholar] [CrossRef]
- Al-Sadeq, D.W.; Majdalawieh, A.F.; Mesleh, A.G.; Abdalla, O.M.; Nasrallah, G.K. Laboratory challenges in the diagnosis of hepatitis E virus. J. Med. Microbiol. 2018, 67, 466–480. [Google Scholar] [CrossRef]
- Pallerla, S.R.; Harms, D.; Johne, R.; Todt, D.; Steinmann, E.; Schemmerer, M.; Wenzel, J.J.; Hofmann, J.; Shih, J.W.K.; Wedemeyer, H.; et al. Hepatitis E Virus Infection: Circulation, Molecular Epidemiology, and Impact on Global Health. Pathogens 2020, 9, 856. [Google Scholar] [CrossRef]
- Aslan, A.T.; Balaban, H.Y. Hepatitis E virus: Epidemiology, diagnosis, clinical manifestations, and treatment. World J. Gastroenterol. 2020, 26, 5543–5560. [Google Scholar] [CrossRef]
- Lhomme, S.; Legrand-Abravanel, F.; Kamar, N.; Izopet, J. Screening, diagnosis and risks associated with Hepatitis E virus infection. Expert Rev. Anti-Infect. Ther. 2019, 17, 403–418. [Google Scholar] [CrossRef]
- Khudyakov, Y.; Kamili, S. Serological diagnostics of hepatitis E virus infection. Virus Res. 2011, 161, 84–92. [Google Scholar] [CrossRef]
- Kar, P.; Karna, R. A Review of the Diagnosis and Management of Hepatitis E. Curr. Treat. Options Infect. Dis. 2020, 12, 310–320. [Google Scholar] [CrossRef]
- Nan, Y.; Zhang, Y.J. Molecular biology and infection of hepatitis E virus. Front. Microbiol. 2016, 7, 1419. [Google Scholar]
- Oechslin, N.; Moradpour, D.; Gouttenoire, J. On the Host Side of the Hepatitis E Virus Life Cycle. Cells 2020, 9, 1294. [Google Scholar] [CrossRef]
- Kenney, S.P.; Meng, X.J. Hepatitis E virus genome structure and replication strategy. Cold Spring Harb. Perspect. Med. 2019, 9, a031724. [Google Scholar] [CrossRef]
- Himmelsbach, K.; Bender, D.; Hildt, E. Life cycle and morphogenesis of the hepatitis E virus. Emerg. Microbes Infect. 2018, 7, 196. [Google Scholar] [CrossRef] [PubMed]
- Debing, Y.; Moradpour, D.; Neyts, J.; Gouttenoire, J. Update on hepatitis E virology: Implications for clinical practice. J. Hepatol. 2016, 65, 200–212. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, T.; Nasir, S.; Musa, T.H.; AlRyalat, S.A.S.; Khan, M.; Hui, J. Epidemiology, diagnosis, vaccines, and bibliometric analysis of the 100 top-cited studies on Hepatitis E virus. Hum. Vaccines Immunother. 2020, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Panda, S.K.; Varma, S.P.K. Hepatitis E: Molecular Virology and Pathogenesis. J. Clin. Exp. Hepatol. 2013, 3, 114–124. [Google Scholar] [CrossRef]
- Ju, X.; Ding, Q. Hepatitis E virus assembly and release. Viruses 2019, 11, 539. [Google Scholar] [CrossRef]
- Thakur, V.; Ratho, R.K.; Kumar, S.; Saxena, S.K.; Bora, I.; Thakur, P. Viral Hepatitis E and Chronicity: A Growing Public Health Concern. Front. Microbiol. 2020, 11. [Google Scholar] [CrossRef]
- Sayed, I.M.; Vercouter, A.S.; Abdelwahab, S.F.; Vercauteren, K.; Meuleman, P. Is hepatitis E virus an emerging problem in industrialized countries? Hepatology 2015, 62, 1883–1892. [Google Scholar] [CrossRef]
- World Health Organization Hepatitis, E. Available online: https://www.who.int/news-room/fact-sheets/detail/hepatitis-e (accessed on 17 December 2020).
- Teshale, E. Travel-Related Infectious Diseases. In Yellow Book; Brunette, G., Nemhauser, J., Eds.; Oxford University Press: New York, NY, USA, 2019. [Google Scholar]
- Horvatits, T.; Schulze zur Wiesch, J.; Lütgehetmann, M.; Lohse, A.W.; Pischke, S. The Clinical Perspective on Hepatitis E. Viruses 2019, 11, 617. [Google Scholar] [CrossRef]
- Denner, J. Hepatitis e virus (HEV)—The future. Viruses 2019, 11, 251. [Google Scholar] [CrossRef]
- Clemente-Casares, P.; Ramos-Romero, C.; Ramirez-Gonzalez, E.; Mas, A. Hepatitis E virus in industrialized countries: The silent threat. Biomed. Res. Int. 2016, 2016. [Google Scholar] [CrossRef]
- Primadharsini, P.P.; Nagashima, S.; Okamoto, H. Genetic variability and evolution of hepatitis E virus. Viruses 2019, 11, 456. [Google Scholar] [CrossRef] [PubMed]
- Gupta, E.; Agarwala, P. Hepatitis E virus infection: An old virus with a new story! Indian J. Med. Microbiol. 2018, 36, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Hakim, M.S.; Wang, W.; Bramer, W.M.; Geng, J.; Huang, F.; de Man, R.A.; Peppelenbosch, M.P.; Pan, Q. The global burden of hepatitis E outbreaks: A systematic review. Liver Int. 2017, 37, 19–31. [Google Scholar] [CrossRef]
- Teshale, E.H.; Hu, D.J.; Holmberg, S.D. The two faces of hepatitis E virus. Clin. Infect. Dis. 2010, 51, 328–334. [Google Scholar] [CrossRef] [PubMed]
- Narayanan, S.; Abutaleb, A.; Sherman, K.E.; Kottilil, S. Clinical features and determinants of chronicity in hepatitis E virus infection. J. Viral Hepat. 2019, 26, 414–421. [Google Scholar] [CrossRef]
- Sooryanarain, H.; Meng, X.J. Hepatitis E virus: Reasons for emergence in humans. Curr. Opin. Virol. 2019, 34, 10–17. [Google Scholar] [CrossRef]
- Blasco-Perrin, H.; Abravanel, F.; Blasco-Baque, V.; Péron, J.M. Hepatitis E, the neglected one. Liver Int. 2016, 36, 130–134. [Google Scholar] [CrossRef]
- Carratalà, A.; Joost, S. Population density and water balance influence the global occurrence of hepatitis E epidemics. Sci. Rep. 2019, 9. [Google Scholar] [CrossRef]
- Wallace, S.J.; Crossan, C.; Hussaini, S.H.; Dalton, H.R. Hepatitis E: A largely underestimated, emerging threat. Br. J. Hosp. Med. 2019, 80, 399–404. [Google Scholar] [CrossRef]
- Wedemeyer, H.; Pischke, S.; Manns, M.P. Pathogenesis and treatment of hepatitis E virus infection. Gastroenterology 2012, 142. [Google Scholar] [CrossRef]
- Sclair, S.N.; Schiff, E.R. An update on the hepatitis E virus. Curr. Gastroenterol. Rep. 2013, 15. [Google Scholar] [CrossRef] [PubMed]
- Takova, K.; Koynarski, T.; Minkov, I.; Ivanova, Z.; Toneva, V.; Zahmanova, G. Increasing Hepatitis E Virus Seroprevalence in Domestic Pigs and Wild Boar in Bulgaria. Animals 2020, 10, 1521. [Google Scholar] [CrossRef]
- Khuroo, M.S.; Khuroo, M.S.; Khuroo, N.S. Hepatitis E: Discovery, global impact, control and cure. World J. Gastroenterol. 2016, 22, 7030–7045. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, S.; Wu, J.; Jiang, Y.; Zhang, H.; Li, S.; Liu, H.; Yang, C.; Tang, H.; Guo, N.; et al. Hepatitis E virus infection in acute non-traumatic neuropathy: A large prospective case-control study in China. EBioMedicine 2018, 36, 122–130. [Google Scholar] [CrossRef] [PubMed]
- Cong, W.; Sui, J.C.; Zhang, X.Y.; Qian, A.D.; Chen, J.; Zhu, X.Q. Seroprevalence of hepatitis E virus among pregnant women and control subjects in China. J. Med. Virol. 2015, 87, 446–450. [Google Scholar] [CrossRef]
- Ricci, A.; Allende, A.; Bolton, D.; Chemaly, M.; Davies, R.; Fernandez Escamez, P.S.; Herman, L.; Koutsoumanis, K.; Lindqvist, R.; Nørrung, B.; et al. Public health risks associated with hepatitis E virus (HEV) as a food-borne pathogen. EFSA J. 2017, 15. [Google Scholar] [CrossRef]
- Ijaz, S.; Vyse, A.J.; Morgan, D.; Pebody, R.G.; Tedder, R.S.; Brown, D. Indigenous hepatitis E virus infection in England: More common than it seems. J. Clin. Virol. 2009, 44, 272–276. [Google Scholar] [CrossRef] [PubMed]
- Webb, G.W.; Dalton, H.R. Hepatitis E: An underestimated emerging threat. Ther. Adv. Infect. Dis. 2019, 6. [Google Scholar] [CrossRef]
- Đaković Rode, O.; Jemeršić, L.; Vince, A. Hepatitis E in Croatia-Guidelines for Diagnosis and Treatment. Liječnički Vjesn. 2016, 138, 289–296. [Google Scholar]
- Jemeršić, L.; Keros, T.; Maltar, L.; Barbić, L.; Čavlek, T.V.; Jeličić, P.; Rode, O.Đ.; Prpić, J. Differences in hepatitis E virus (HEV) presence in naturally infected seropositive domestic pigs and wild boars—An indication of wild boars having an important role in HEV epidemiology. Vet. Arh. 2017, 87, 651–663. [Google Scholar] [CrossRef]
- Mrzljak, A.; Dinjar-Kujundzic, P.; Jemersic, L.; Prpic, J.; Barbic, L.; Savic, V.; Stevanovic, V.; Vilibic-Cavlek, T. Epidemiology of hepatitis e in South-East Europe in the “one Health” concept. World J. Gastroenterol. 2019, 25, 3168–3182. [Google Scholar] [CrossRef] [PubMed]
- Jemeršić, L.; Prpić, J.; Brnić, D.; Keros, T.; Pandak, N.; Rode, O.Đ. Genetic diversity of hepatitis E virus (HEV) strains derived from humans, swine and wild boars in Croatia from 2010 to 2017. BMC Infect. Dis. 2019, 19, 269. [Google Scholar] [CrossRef] [PubMed]
- Nelson, K.E.; Labrique, A.B.; Kmush, B.L. Epidemiology of genotype 1 and 2 hepatitis E virus infections. Cold Spring Harb. Perspect. Med. 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Abebe, M.; Ali, I.; Ayele, S.; Overbo, J.; Aseffa, A.; Mihret, A. Seroprevalence and risk factors of Hepatitis E Virus infection among pregnant women in Addis Ababa, Ethiopia. PLoS ONE 2017, 12, e0180078. [Google Scholar] [CrossRef]
- Kmush, B.L.; Nelson, K.E.; Labrique, A.B. Risk factors for hepatitis E virus infection and disease. Expert Rev. Anti-Infect. Ther. 2015, 13, 41–53. [Google Scholar] [CrossRef] [PubMed]
- Bergløv, A.; Hallager, S.; Weis, N. Hepatitis E during pregnancy: Maternal and foetal case-fatality rates and adverse outcomes—A systematic review. J. Viral Hepat. 2019, 26, 1240–1248. [Google Scholar] [CrossRef]
- El-Mokhtar, M.A.; Othman, E.R.; Khashbah, M.Y.; Ismael, A.; Ghaliony, M.A.; Seddik, M.I.; Sayed, I.M. Evidence of the Extrahepatic Replication of Hepatitis E Virus in Human Endometrial Stromal Cells. Pathogens 2020, 9, 295. [Google Scholar] [CrossRef]
- Wedemeyer, H.; Rybczynska, J.; Pischke, S.; Krawczynski, K. Immunopathogenesis of hepatitis E virus infection. Semin. Liver Dis. 2013, 33, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Xin, S.; Xiao, L. Clinical manifestations of hepatitis E. Adv. Exp. Med. Biol. 2016, 948, 175–189. [Google Scholar] [CrossRef]
- Yu, W.; Hao, X.; Li, Y.; Yang, C.; Li, Y.; He, Z.; Huang, F. Vertical transmission of hepatitis E virus in pregnant rhesus macaques. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Racicot, K.; Mor, G. Risks associated with viral infections during pregnancy. J. Clin. Invest. 2017, 127, 1591–1599. [Google Scholar] [CrossRef] [PubMed]
- Sookoian, S. Liver disease during pregnancy: Acute viral hepatitis. Ann. Hepatol. 2006, 5, 231–236. [Google Scholar] [CrossRef]
- Abravanel, F.; Lhomme, S.; Dubois, M.; Peron, J.M.; Alric, L.; Kamar, N.; Izopet, J. Hepatitis E virus. Med. Mal. Infect. 2013, 43, 263–270. [Google Scholar] [CrossRef] [PubMed]
- Cornish, E.F.; Filipovic, I.; Åsenius, F.; Williams, D.J.; McDonnell, T. Innate Immune Responses to Acute Viral Infection During Pregnancy. Front. Immunol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Gouilly, J.; Chen, Q.; Siewiera, J.; Cartron, G.; Levy, C.; Dubois, M.; Al-Daccak, R.; Izopet, J.; Jabrane-Ferrat, N.; El Costa, H. Genotype specific pathogenicity of hepatitis E virus at the human maternal-fetal interface. Nat. Commun. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Seth, A.; Sherman, K.E. Hepatitis E: What We Think We Know. Clin. Liver Dis. 2020, 15, S37–S44. [Google Scholar] [CrossRef]
- Aggarwal, R.A. Hepatitis E: Clinical presentation in disease-endemic areas and diagnosis. Semin. Liver Dis. 2013, 33, 30–40. [Google Scholar] [CrossRef]
- Shalimar; Acharya, S.K. Hepatitis E and Acute Liver Failure in Pregnancy. J. Clin. Exp. Hepatol. 2013, 3, 213–224. [Google Scholar] [CrossRef]
- Julin, C.H.; Hjortaas, K.; Dembinski, J.L.; Sandbu, S.; Øverbø, J.; Stene-Johansen, K.; Dudman, S. Hepatitis E in Pregnant Women and the Potential Use of HEV Vaccine to Prevent Maternal Infection and Mortality. Curr. Trop. Med. Rep. 2019, 6, 197–204. [Google Scholar] [CrossRef]
- Navaneethan, U.; Al Mohajer, M.; Shata, M.T. Hepatitis E and pregnancy: Understanding the pathogenesis. Liver Int. 2008, 28, 1190–1199. [Google Scholar] [CrossRef]
- Rac, M.W.F.; Sheffield, J.S. Prevention and management of viral hepatitis in pregnancy. Obstet. Gynecol. Clin. N. Am. 2014, 41, 573–592. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, R.; Naik, S. Epidemiology of hepatitis E: Current status. J. Gastroenterol. Hepatol. 2009, 24, 1484–1493. [Google Scholar] [CrossRef]
- Terrault, N.A.; Levy, M.T.; Cheung, K.W.; Jourdain, G. Viral hepatitis and pregnancy. Nat. Rev. Gastroenterol. Hepatol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Hao, X.; Li, Y.; Long, F.; He, Q.; Huang, F.; Yu, W. Successful establishment of hepatitis E virus infection in pregnant BALB/c mice. Viruses 2019, 11, 451. [Google Scholar] [CrossRef]
- Li, Y.; Huang, X.; Zhang, Z.; Li, S.; Zhang, J.; Xia, N.; Zhao, Q. Prophylactic Hepatitis E Vaccines: Antigenic Analysis and Serological Evaluation. Viruses 2020, 12, 109. [Google Scholar] [CrossRef] [PubMed]
- Chibber, R.M.; Usmani, M.A.; Al-Sibai, M.H. Should HEV infected mothers breast feed? Arch. Gynecol. Obstet. 2004, 270, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Aggarwal, R.; Jameel, S. Hepatitis E. Hepatology 2011, 54, 2218–2226. [Google Scholar] [CrossRef]
- Krain, L.J.; Atwell, J.E.; Nelson, K.E.; Labrique, A.B. Fetal and neonatal health consequences of vertically transmitted hepatitis E virus infection. Am. J. Trop. Med. Hyg. 2014, 90, 365–370. [Google Scholar] [CrossRef][Green Version]
- Kar, P.; Sengupta, A. A guide to the management of hepatitis E infection during pregnancy. Expert Rev. Gastroenterol. Hepatol. 2019, 13, 205–211. [Google Scholar] [CrossRef]
- Webb, G.W.; Kelly, S.; Dalton, H.R. Hepatitis A and Hepatitis E: Clinical and Epidemiological Features, Diagnosis, Treatment, and Prevention. Clin. Microbiol. Newsl. 2020, 42, 171–179. [Google Scholar] [CrossRef]
- Taherkhani, R.; Farshadpour, F. Epidemiology of hepatitis E in pregnant women and children in Iran: A general overview. J. Clin. Transl. Hepatol. 2016, 4, 269–276. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Dalton, H.R.; Kamar, N.; Baylis, S.A.; Moradpour, D.; Wedemeyer, H.; Negro, F.; European Association for the Study of the Liver. EASL Clinical Practice Guidelines on hepatitis E virus infection. J. Hepatol. 2018, 68, 1256–1271. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Wang, Y. Laboratory diagnosis of HEV infection. In Hepatitis E Virus. Advances in Experimental Medicine and Biology; Wang, Y., Ed.; Springer: Dordrecht, The Netherlands, 2016; Volume 948, pp. 191–209. [Google Scholar]
- Candido, A.; Taffon, S.; Chionne, P.; Pisani, G.; Madonna, E.; Dettori, S.; Hamza, A.; Valdarchi, C.; Bruni, R.; Ciccaglione, A.R. Diagnosis of HEV infection by serological and real-time PCR assays: A study on acute non-A-C hepatitis collected from 2004 to 2010 in Italy. BMC Res. Notes 2012, 5, 297. [Google Scholar] [CrossRef] [PubMed]
- Marrone, G.; Biolato, M.; Mercurio, G.; Capobianchi, M.R.; Garbuglia, A.R.; Liguori, A.; Vassallo, G.; Gasbarrini, A.; Miele, L.; Grieco, A. Acute HEV hepatitis: Clinical and laboratory diagnosis. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 764–770. [Google Scholar] [CrossRef]
- Myint, K.S.A.; Endy, T.P.; Gibbons, R.V.; Laras, K.; Mammen, M.P.; Sedyaningsih, E.R.; Seriwatana, J.; Glass, J.S.; Narupiti, S.; Corwin, A.L. Evaluation of diagnostic assays for hepatitis E virus in outbreak settings. J. Clin. Microbiol. 2006, 44, 1581–1583. [Google Scholar] [CrossRef]
- Aggarwal, R. Diagnosis of hepatitis E. Nat. Rev. Gastroenterol. Hepatol. 2013, 10, 24–33. [Google Scholar] [CrossRef]
- Doting, M.H.E.; Weel, J.; Niesters, H.G.M.; Riezebos-Brilman, A.; Brandenburg, A. The added value of hepatitis E diagnostics in determining causes of hepatitis in routine diagnostic settings in the Netherlands. Clin. Microbiol. Infect. 2017, 23, 667–671. [Google Scholar] [CrossRef][Green Version]
- Sampedro, A.; Casanovas, I.; Ceballos, J.; Rodriguez-Granger, J.; Cobo, F.; Navarro, J.M. Comparative evaluation of two immunoassays for serological diagnosis of hepatitis E. J. Med. Virol. 2020, 92, 260–262. [Google Scholar] [CrossRef]
- Pérez-Gracia, M.T.; Suay-García, B.; Mateos-Lindemann, M.L. Hepatitis E and pregnancy: Current state. Rev. Med. Virol. 2017, 27, e1929. [Google Scholar] [CrossRef]
- Larrue, H.; Abravanel, F.; Péron, J. Hepatitis E, what’s the real issue? Liver Int. 2020, 40, 43–47. [Google Scholar] [CrossRef]
- Prasidthrathsint, K.; Stapleton, J.T. Laboratory Diagnosis and Monitoring of Viral Hepatitis. Gastroenterol. Clin. N. Am. 2019, 48, 259–279. [Google Scholar] [CrossRef] [PubMed]
- Mirazo, S.; Ramos, N.; Mainardi, V.; Arbiza, J.; Gerona, S. Transmission, diagnosis, and management of hepatitis E: An update. Hepatic Med. Evid. Res. 2014, 6, 45. [Google Scholar] [CrossRef] [PubMed]
- Nicot, F.; Cazabat, M.; Lhomme, S.; Marion, O.; Sauné, K.; Chiabrando, J.; Dubois, M.; Kamar, N.; Abravanel, F.; Izopet, J. Quantification of HEV RNA by droplet digital PCR. Viruses 2016, 8, 233. [Google Scholar] [CrossRef]
- Lan, X.; Yang, B.; Bao, Y.L.; Xiang, P.Y.; Xue, R.L.; Ji, X.L. Reverse transcription-loop-mediated isothermal amplification assay for rapid detection of hepatitis E virus. J. Clin. Microbiol. 2009, 47, 2304–2306. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control. Options for National Testing and Surveillance for Hepatitis E virus in the EU/EEA—Operational Guidance; ECDC: Stockholm, Sweden, 2019. [Google Scholar]
- Rivero Juárez, A. Consensus Document of the Diagnosis, Management, and Prevention of Infection with the Hepatitis E virus: Study Group for Viral Hepatitis (GEHEP) of the Spanish Society of Infectious Diseases and Clinical Microbiology (SEIMC). 2018. Available online: http://www.seimc.org (accessed on 2 December 2020).
- Anastasiou, O.E.; Thodou, V.; Berger, A.; Wedemeyer, H.; Ciesek, S. Comprehensive evaluation of hepatitis E serology and molecular testing in a large cohort. Pathogens 2020, 9, 137. [Google Scholar] [CrossRef]
- Desai, A.N. Hepatitis E. JAMA 2020, 323, 1862. [Google Scholar] [CrossRef]
- Bi, H.; Yang, R.; Wu, C.; Xia, J. Hepatitis E Virus and Blood Transfusion Safety. Epidemiol. Infect. 2020, 148, e158. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhang, X.; Zhu, F.; Jin, H.; Wang, B. A preliminary cost-effectiveness analysis of hepatitis E vaccination among pregnant women in epidemic regions. Hum. Vaccin. Immunother. 2016, 12, 2003–2009. [Google Scholar] [CrossRef][Green Version]
- Ankcorn, M.J.; Tedder, S.R.; Cairns, J.; Sandmann, F.G. Cost-Effectiveness Analysis of Screening for Persistent Hepatitis E Virus Infection in Solid Organ Transplant Patients in the United Kingdom: A Model-Based Economic Evaluation. Value Health 2020, 23, 309–318. [Google Scholar] [CrossRef]
| Genus | Orthohepevirus |
| Family | Hepeviridae |
| Species | A, B, C, D |
| Genotypes | 1,2,3,4,5,6,7,8 |
| Virion | 32–34 nm, icosahedral |
| Envelope | quasi-enveloped |
| Genome | single-stranded, positive-sense, linear RNA |
| Genome size | 7.2 kb |
| Sensitivity | Inactivation at temperatures above 70 °C |
| HEV Genotype | Host | Endemic Regions |
|---|---|---|
| 1 | Human | Asia, North Africa, Mexico, Middle East |
| 2 | Human | Asia, North Africa, Mexico, Middle East |
| 3 | Human, rabbit, pig, mongoose | Worldwide |
| 4 | Human, domestic pig, and wild boar | South and East Asia, the United States, Europe, China, and Japan |
| 5 | Wild boar | Japan |
| 6 | Wild boar | Japan |
| 7 | Camel | Dubai |
| 8 | Camel | Dubai, western China |
| Method | Type of Molecule Detected | Advantages | Disadvantages | |
|---|---|---|---|---|
| Serological tests | Enzyme immunoassay (EIA) | IgM and/or IgG anti-HEV antibodies | Quick, easy to perform, less expensive than a molecular test, high sensitivity, low specificity, epidemiological surveys | Discrepancies in the results, vary in sensitivity, specificity, false-positive reaction through cross-reactive antibodies |
| Viral capsid antigen and phosphoprotein | Technically less demanding than PCR | Antigens disappear quickly with seroconversion, low sensitivity, the poor performance of polypeptides, limited application, less sensitivity than real-time PCR | ||
| Rapid immunochromatographic assays | IgM anti-HEV antibodies | Sensitivity comparable with PCR, higher than ELISA, fast, simple, point of care testing | Low accuracy in detecting other genotypes than genotype 3 | |
| Nucleic acid tests | Reversion transcription and PCR (RT-PCR) | ORF2 region | High sensitivity and specificity, identification of HEV genotypes | Low levels of HEV RNA before symptoms, a brief period of viremia |
| Nested PCR | ORF2 region | High specificity and sensitivity for genotype 1 | The complicated procedure, strict requirements, contamination | |
| Real-time PCR (qPCR) | ORF2 and ORF3 region | High specificity, high sensitivity, more sensitive than nPCR, simple operation, quantification | Trained staff, test facilities, high cost in developing countries, polymorphisms in primer site results with false-negative results | |
| Loop-mediated isothermal amplification assay (LAMP) | ORF3 region | Fast, no special equipment, high sensitivity, and efficiency | Limited experience, works for genotypes 1 and 4, no data for other genotypes | |
| RNA sequencing | ORF1 and ORF2 region | Accurate genotyping | Variability, lack of standardization, high cost, polymorphisms in primer site results in false-negative results |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Talapko, J.; Meštrović, T.; Pustijanac, E.; Škrlec, I. Towards the Improved Accuracy of Hepatitis E Diagnosis in Vulnerable and Target Groups: A Global Perspective on the Current State of Knowledge and the Implications for Practice. Healthcare 2021, 9, 133. https://doi.org/10.3390/healthcare9020133
Talapko J, Meštrović T, Pustijanac E, Škrlec I. Towards the Improved Accuracy of Hepatitis E Diagnosis in Vulnerable and Target Groups: A Global Perspective on the Current State of Knowledge and the Implications for Practice. Healthcare. 2021; 9(2):133. https://doi.org/10.3390/healthcare9020133
Chicago/Turabian StyleTalapko, Jasminka, Tomislav Meštrović, Emina Pustijanac, and Ivana Škrlec. 2021. "Towards the Improved Accuracy of Hepatitis E Diagnosis in Vulnerable and Target Groups: A Global Perspective on the Current State of Knowledge and the Implications for Practice" Healthcare 9, no. 2: 133. https://doi.org/10.3390/healthcare9020133
APA StyleTalapko, J., Meštrović, T., Pustijanac, E., & Škrlec, I. (2021). Towards the Improved Accuracy of Hepatitis E Diagnosis in Vulnerable and Target Groups: A Global Perspective on the Current State of Knowledge and the Implications for Practice. Healthcare, 9(2), 133. https://doi.org/10.3390/healthcare9020133








