Artificial Intelligence: A Next-Level Approach in Confronting the COVID-19 Pandemic
Abstract
1. Introduction
2. Materials and Methods
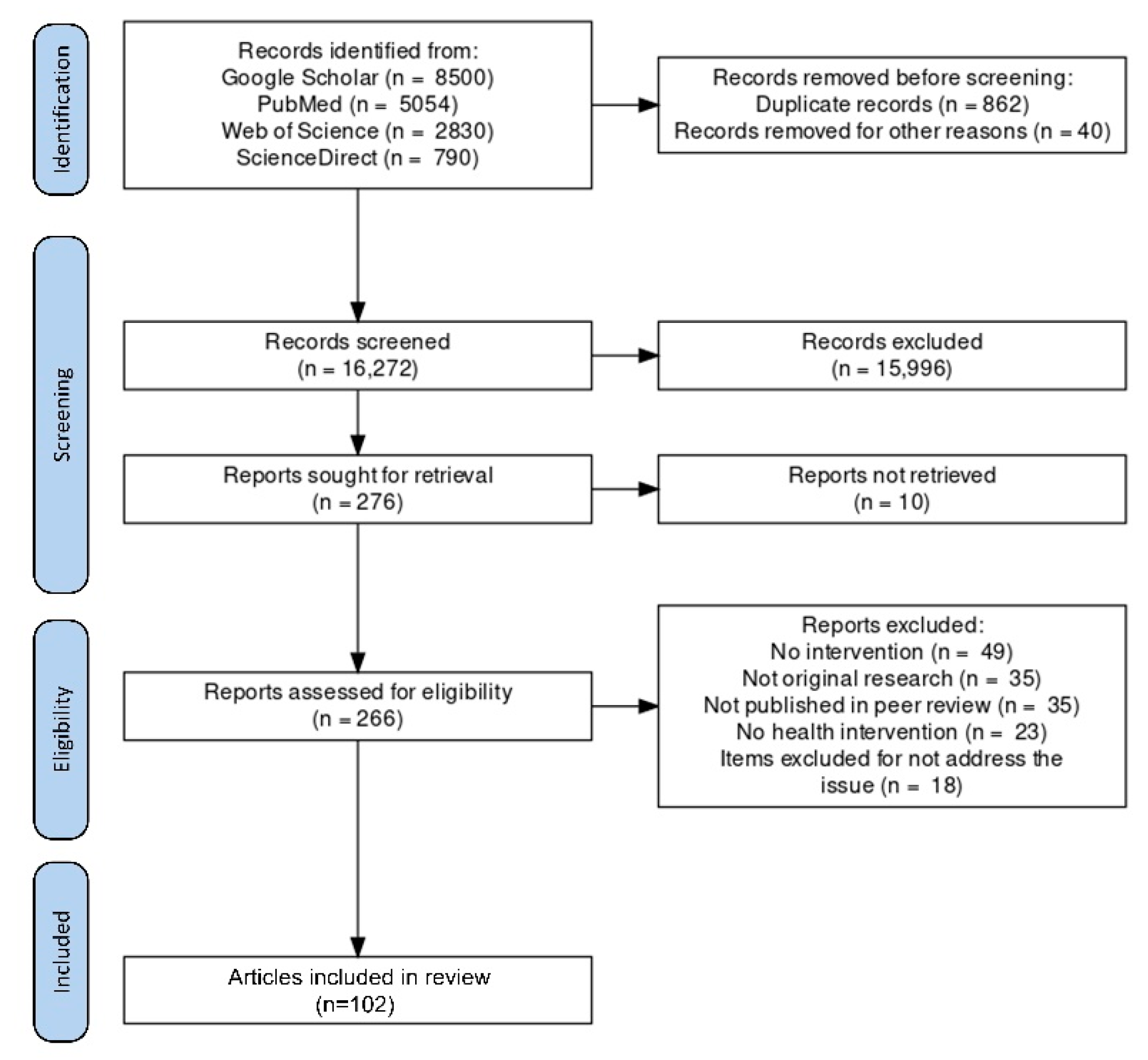
2.1. Search Strategy
2.2. Study Selection
3. Omicron
3.1. Secondary Infections
3.2. Mucormycosis
- Limitations
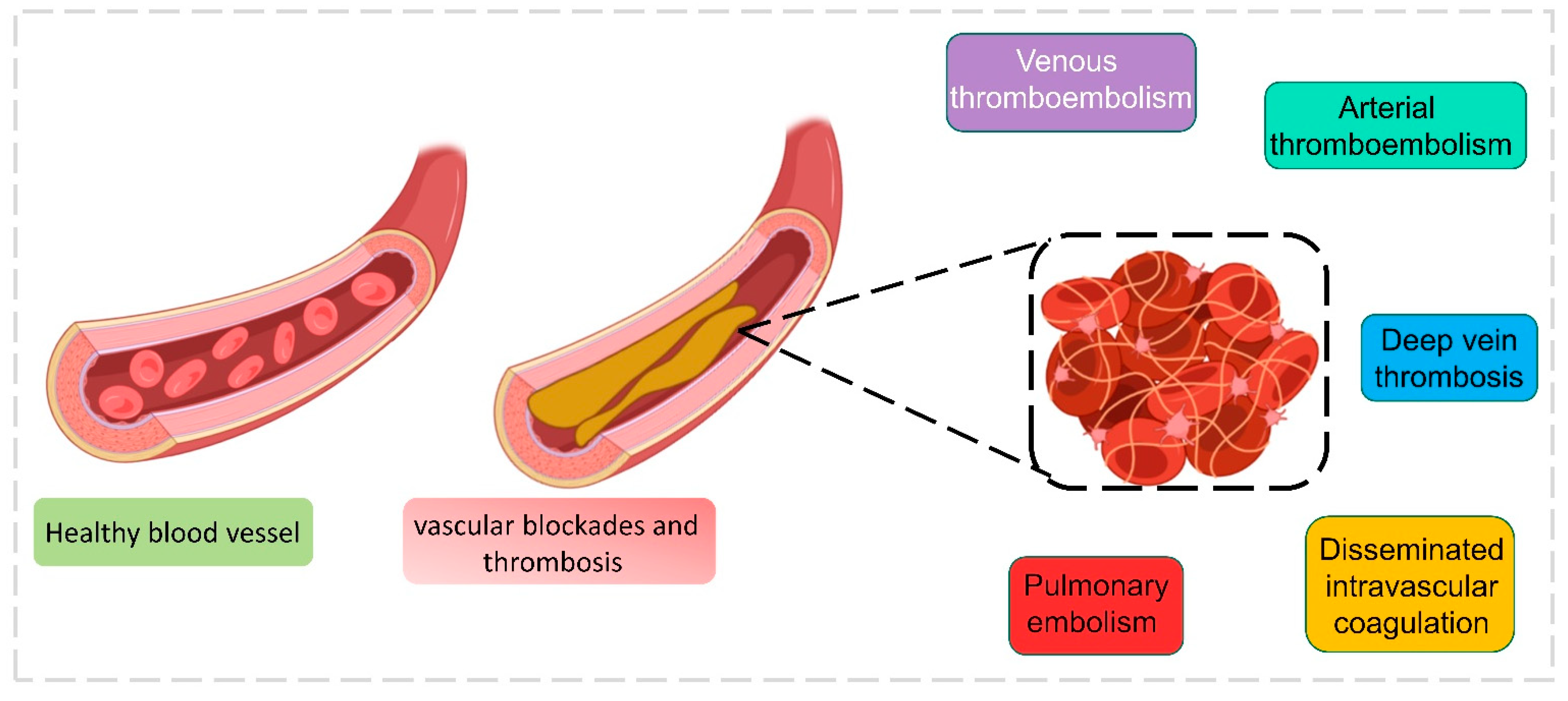
3.3. Blood Coagulation
- Limitations
3.4. Bacterial Infections in the Blood Stream and Urinary Samples of COVID-19 Patients
3.4.1. Blood Stream
- Limitations
3.4.2. Urinary Samples
3.5. Antibiotic Resistance: Improving Diagnosis Using ML
- Limitations
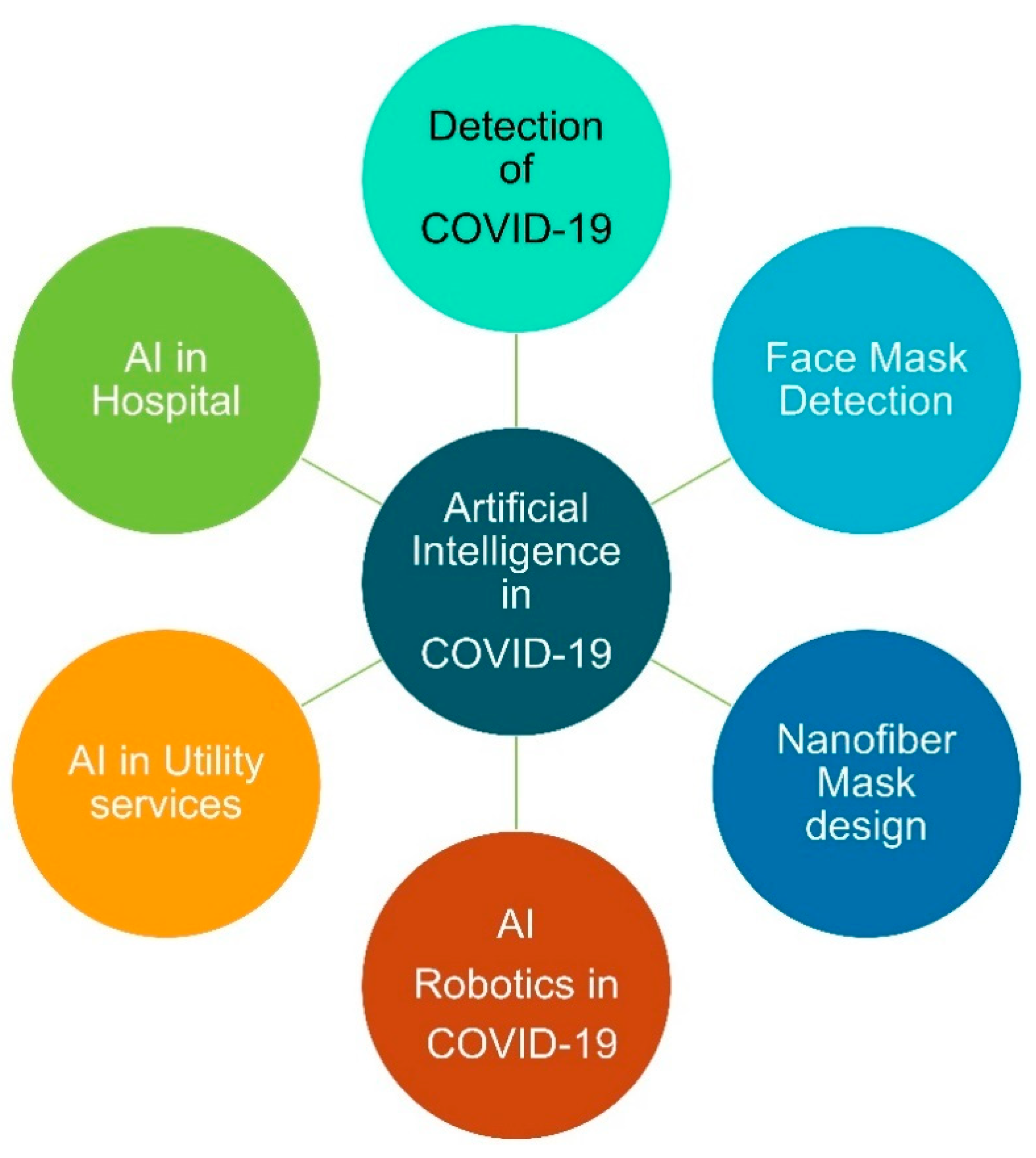
3.6. AI Detection of COVID-19 and Preventive Measures with AI
3.7. Face Mask Detection Systems Using AI
3.8. Nanofiber Masks Using AI
3.8.1. Machine Learning in Mask Production
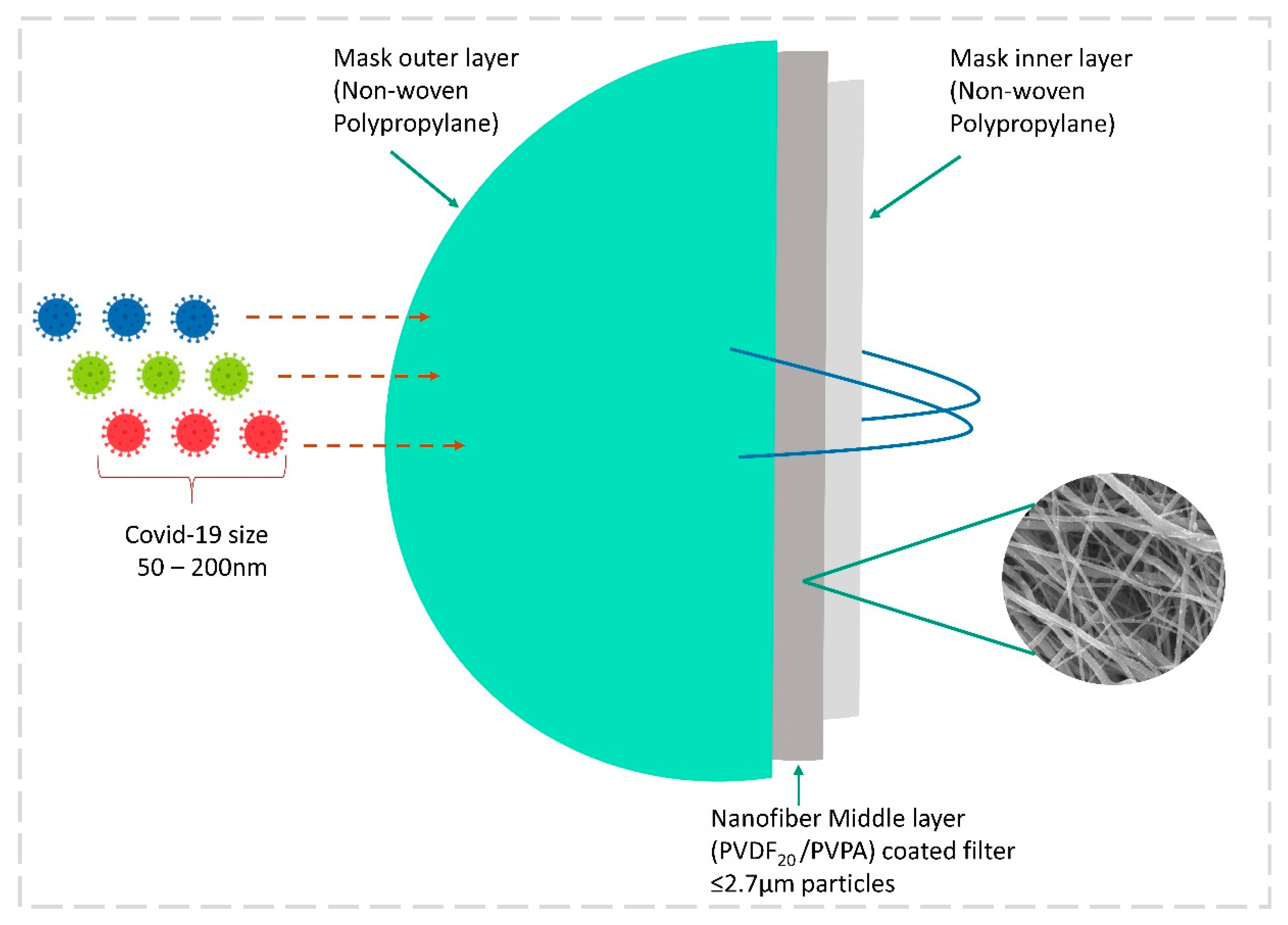
3.8.2. Electrospun Nanofiber Mask by Nanotechnology
3.9. Robotics against COVID-19
4. Impact of COVID-19 Pandemic in Public Life
4.1. AI in Utility Services
4.2. AI for Researchers
4.3. ML in Oil and Gas
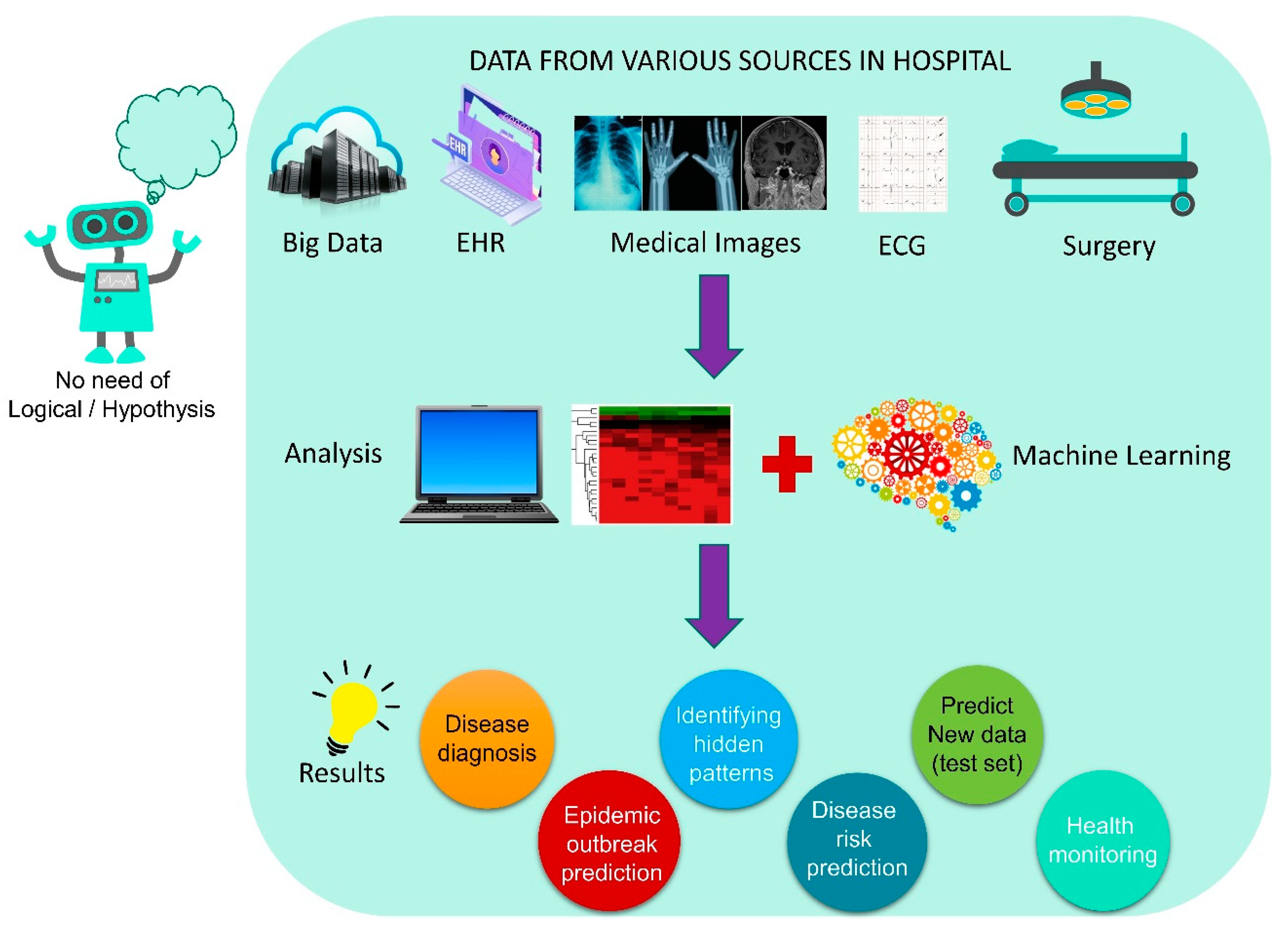
4.4. AI-Based Decision-Making in the Hospital
4.5. Telemonitoring during COVID-19
4.6. AI-Based Law in Various Countries
5. Discussion
- ➢
- A face recognition and attendance project is in the developmental stage, and a face mask recognition system must be included to detect a violation committed by a specific person in a closed office environment and send an alert to management.
- ➢
- The Center for Disease Control and Prevention (CDC) maintains a large database of “COVID-19 Science Update Database” published and pre-print databases which are easily accessible for any AI-based healthcare articles. Similarly, Lawrence Berkeley developed a COVID-19 literature search powered by advanced NLP algorithms, which uses AI to search attributes such as keyword, topic, title, year, and so on.
- ➢
- Another central data service for documents is CORD-19 and LitCovid, with a regular update on COVID-19 research outputs. These databases encourage the scientific community to access free articles, which helps in developing novel research ideas and outputs.
- ➢
- The promising results obtained from various drug makers’ companies’ AI-based technologies give hope to end the pandemic, but the real question is why these drug companies are having such difficulty sharing their data with low-income countries to end the pandemic. Until this happens, COVID-19 may repeatedly invade developed nations with various mutated stains.
- ➢
- Hospitals need to adopt recent AI technologies for analyzing radiographic images of patients with COVID-19, to reduce the burden on doctors. Telemonitoring and speech and text recognition using NLP will improve doctor–patient interactions in remote treatment methods.
- ➢
- AI offers doctors and health workers the ability to store and handle or share patient records very safely and securely through Cloud platforms such as Google Cloud AI, Amazon AI services, Microsoft Azure AI, IBM Watson Studio, ref. [199] H2O.ai, TensorFlow, DataRobot, Wipro Holmes AI and automation platform, Salesforce Einstein, Infosys Nia, and others that can be accessed through AI-infused technology.
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Origin of SARS-CoV-2. 26 March 2020. Available online: https://www.who.int/publications/i/item/origin-of-sars-cov-2 (accessed on 6 August 2022).
- World Health Organization. WHO Director-General’s Opening Remarks at the Media Briefing on COVID-19—11 March 2020. Available online: https://www.who.int/director-general/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020 (accessed on 6 August 2022).
- WHO. COVID-19 Weekly Epidemiological Update; World Health Organization: Geneva, Switzerland, 2021; pp. 1–23. [Google Scholar]
- Alhasan, M.; Hasaneen, M. Digital imaging, technologies and artificial intelligence applications during COVID-19 pandemic. Comput. Med. Imaging Graph. 2021, 91, 101933. [Google Scholar] [CrossRef] [PubMed]
- Anbalagan, S.; Arunprasanna, V.; Dinakaran, S.; Krishnan, M. Combinatory therapeutic approaches for common cold and SARS-CoV-2. Synergy 2020, 11, 100069. [Google Scholar] [CrossRef]
- Petrosillo, N.; Viceconte, G.; Ergonul, O.; Ippolito, G.; Petersen, E. COVID-19, SARS and MERS: Are they closely related? Clin. Microbiol. Infect. 2020, 26, 729–734. [Google Scholar] [CrossRef]
- Mallapaty, S. What the cruise-ship outbreaks reveal about COVID-19. Nature 2020, 580, 18. [Google Scholar] [CrossRef]
- Tuli, S.; Tuli, S.; Tuli, R.; Gill, S.S. Predicting the growth and trend of COVID-19 pandemic using machine learning and cloud computing. Internet Things 2020, 11, 100222. [Google Scholar] [CrossRef]
- Wang, H.; Jia, S.; Li, Z.; Duan, Y.; Tao, G.; Zhao, Z. A Comprehensive Review of Artificial Intelligence in Prevention and Treatment of COVID-19 Pandemic. Front. Genet. 2022, 13, 1–15. [Google Scholar] [CrossRef] [PubMed]
- National Institute of Health. Antiviral Agents, Including Antibody Products. Available online: https://www.covid19treatmentguidelines.nih.gov/ (accessed on 6 August 2022).
- Zahariadis, G.; Gooley, T.A.; Ryall, P.; Hutchinson, C.; Latchford, M.I.; Fearon, M.A.; Jamieson, F.B.; Richardson, S.; Kuschak, T.; Mederski, B. Risk of ruling out severe acute respiratory syndrome by ruling in another diagnosis: Variable incidence of atypical bacteria coinfection based on diagnostic assays. Can. Respir. J. 2006, 13, 17–22. [Google Scholar] [CrossRef]
- Arabi, Y.M.; Al-Omari, A.; Mandourah, Y.; Al-Hameed, F.; Sindi, A.A.; Alraddadi, B.; Shalhoub, S.; Almotairi, A.; Al Khatib, K.; Abdulmomen, A.; et al. Critically Ill Patients With the Middle East Respiratory Syndrome. Crit. Care Med. 2017, 45, 1683–1695. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Basera, P.; Anand, A.; Ozair, A. COVID-19-Associated Mucormycosis in a Tertiary Care Hospital in India: A Case Series. Cureus 2022, 13. [Google Scholar] [CrossRef]
- Aranjani, J.M.; Manuel, A.; Abdul Razack, H.I.; Mathew, S.T. COVID-19–associated mucormycosis: Evidence-based critical review of an emerging infection burden during the pandemic’s second wave in India. PLoS Negl. Trop. Dis. 2021, 15, e0009921. [Google Scholar] [CrossRef]
- Mahalaxmi, I.; Jayaramayya, K.; Venkatesan, D.; Subramaniam, M.D.; Renu, K.; Vijayakumar, P.; Narayanasamy, A.; Gopalakrishnan, A.V.; Kumar, N.S.; Sivaprakash, P.; et al. Mucormycosis: An opportunistic pathogen during COVID-19. Environ. Res. 2021, 201, 111643. [Google Scholar] [CrossRef] [PubMed]
- de Laat-Kremers, R.; De Jongh, R.; Ninivaggi, M.; Fiolet, A.; Fijnheer, R.; Remijn, J.; de Laat, B. Coagulation parameters predict COVID-19-related thrombosis in a neural network with a positive predictive value of 98%. Front. Immunol. 2022, 13, 977443. [Google Scholar] [CrossRef] [PubMed]
- Biswas, S.; Thakur, V.; Kaur, P.; Khan, A.; Kulshrestha, S.; Kumar, P. Blood clots in COVID-19 patients: Simplifying the curious mystery. Med. Hypotheses 2021, 146, 110371. [Google Scholar] [CrossRef]
- Palanisamy, N.; Vihari, N.; Meena, D.S.; Kumar, D.; Midha, N.; Tak, V.; Sharma, A.; Bohra, G.K.; Kothari, N.; Dutt, N.; et al. Clinical profile of bloodstream infections in COVID-19 patients: A retrospective cohort study. BMC Infect. Dis. 2021, 21, 933. [Google Scholar] [CrossRef]
- Kokkoris, S.; Papachatzakis, I.; Gavrielatou, E.; Ntaidou, T.; Ischaki, E.; Malachias, S.; Vrettou, C.; Nichlos, C.; Kanavou, A.; Zervakis, D.; et al. ICU-acquired bloodstream infections in critically ill patients with COVID-19. J. Hosp. Infect. 2021, 107, 95–97. [Google Scholar] [CrossRef]
- Cai, T.; Tascini, C.; Novelli, A.; Anceschi, U.; Bonkat, G.; Wagenlehner, F.; Bjerklund Johansen, T.E. The Management of Urinary Tract Infections during the COVID-19 Pandemic: What Do We Need to Know? Uro 2022, 2, 55–64. [Google Scholar] [CrossRef]
- Fakih, M.G.; Bufalino, A.; Sturm, L.; Huang, R.-H.; Ottenbacher, A.; Saake, K.; Winegar, A.; Fogel, R.; Cacchione, J. Coronavirus disease 2019 (COVID-19) pandemic, central-line-associated bloodstream infection (CLABSI), and catheter-associated urinary tract infection (CAUTI): The urgent need to refocus on hardwiring prevention efforts. Infect. Control Hosp. Epidemiol. 2022, 43, 26–31. [Google Scholar] [CrossRef]
- Díaz Pollán, B.; Guedez López, G.V.; García Clemente, P.M.; Jiménez González, M.; García Bujalance, S.; Gómez-Gil Mirá, M.R. Urinary Tract Infections in Hospitalized COVID-19 Patients, What’s Up, Doc? J. Clin. Med. 2022, 11, 1815. [Google Scholar] [CrossRef]
- Getahun, H.; Smith, I.; Trivedi, K.; Paulin, S.; Balkhy, H.H. Tackling antimicrobial resistance in the COVID-19 pandemic. Bull. World Health Organ. 2020, 98, 442–442A. [Google Scholar] [CrossRef] [PubMed]
- Lynch, C.; Mahida, N.; Gray, J. Antimicrobial stewardship: A COVID casualty? J. Hosp. Infect. 2020, 106, 401–403. [Google Scholar] [CrossRef] [PubMed]
- Karnoukh, K.I.; Drozdov, V.N.; Shikh, E.V.; Zhilina, S.V.; Lazareva, N.B. Etiology and Antimicrobial Resistance of Secondary Bacterial Infections in Patients Hospitalized with COVID-19: A Retrospective Analysis. Vestn. Ross. Akad. Meditsinskikh Nauk 2022, 77, 25–32. [Google Scholar] [CrossRef]
- Al Sulayyim, H.J.; Ismail, R.; Al Hamid, A.; Ghafar, N.A. Antibiotic Resistance during COVID-19: A Systematic Review. Int. J. Environ. Res. Public Health 2022, 19, 11931. [Google Scholar] [CrossRef] [PubMed]
- Ripa, M.; Galli, L.; Poli, A.; Oltolini, C.; Spagnuolo, V.; Mastrangelo, A.; Muccini, C.; Monti, G.; De Luca, G.; Landoni, G.; et al. Secondary infections in patients hospitalized with COVID-19: Incidence and predictive factors. Clin. Microbiol. Infect. 2021, 27, 451–457. [Google Scholar] [CrossRef]
- Miller, R.A. Medical Diagnostic Decision Support Systems--Past, Present, And Future: A Threaded Bibliography and Brief Commentary. J. Am. Med. Inform. Assoc. 1994, 1, 8–27. [Google Scholar] [CrossRef] [PubMed]
- Makridakis, S. The forthcoming Artificial Intelligence (AI) revolution: Its impact on society and firms. Futures 2017, 90, 46–60. [Google Scholar] [CrossRef]
- da Silva Neto, S.R.; Tabosa Oliveira, T.; Teixeira, I.V.; Aguiar de Oliveira, S.B.; Souza Sampaio, V.; Lynn, T.; Endo, P.T. Machine learning and deep learning techniques to support clinical diagnosis of arboviral diseases: A systematic review. PLoS Negl. Trop. Dis. 2022, 16, e0010061. [Google Scholar] [CrossRef]
- Sandhu, R.; Sood, S.K.; Kaur, G. An intelligent system for predicting and preventing MERS-CoV infection outbreak. J. Supercomput. 2016, 72, 3033–3056. [Google Scholar] [CrossRef] [PubMed]
- Colubri, A.; Silver, T.; Fradet, T.; Retzepi, K.; Fry, B.; Sabeti, P. Transforming Clinical Data into Actionable Prognosis Models: Machine-Learning Framework and Field-Deployable App to Predict Outcome of Ebola Patients. PLoS Negl. Trop. Dis. 2016, 10, e0004549. [Google Scholar] [CrossRef] [PubMed]
- Zimmerman, A.; Kalra, D. Usefulness of machine learning in COVID-19 for the detection and prognosis of cardiovascular complications. Rev. Cardiovasc. Med. 2020, 21, 345. [Google Scholar] [CrossRef]
- Kagiyama, N.; Shrestha, S.; Farjo, P.D.; Sengupta, P.P. Artificial Intelligence: Practical Primer for Clinical Research in Cardiovascular Disease. J. Am. Heart Assoc. 2019, 8, e012788. [Google Scholar] [CrossRef]
- Hirschberg, J.; Manning, C.D. Advances in natural language processing. Science 2015, 349, 261–266. [Google Scholar] [CrossRef]
- Russakovsky, O.; Deng, J.; Su, H.; Krause, J.; Satheesh, S.; Ma, S.; Huang, Z.; Karpathy, A.; Khosla, A.; Bernstein, M.; et al. ImageNet Large Scale Visual Recognition Challenge. Int. J. Comput. Vis. 2015, 115, 211–252. [Google Scholar] [CrossRef]
- Wehbe, R.M.; Sheng, J.; Dutta, S.; Chai, S.; Dravid, A.; Barutcu, S.; Wu, Y.; Cantrell, D.R.; Xiao, N.; Allen, B.D.; et al. DeepCOVID-XR: An artificial intelligence algorithm to detect COVID-19 on chest radiographs trained and tested on a large U.S. Clinical data set. Radiology 2021, 299, E167–E176. [Google Scholar] [CrossRef] [PubMed]
- Haleem, A.; Javaid, M.; Singh, R.P.; Suman, R. Applications of Artificial Intelligence (AI) for cardiology during COVID-19 pandemic. Sustain. Oper. Comput. 2021, 2, 71–78. [Google Scholar] [CrossRef]
- Gleichgerrcht, E.; Munsell, B.; Bhatia, S.; Vandergrift, W.A.; Rorden, C.; McDonald, C.; Edwards, J.; Kuzniecky, R.; Bonilha, L. Deep learning applied to whole-brain connectome to determine seizure control after epilepsy surgery. Epilepsia 2018, 59, 1643–1654. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Su, C.; Xiao, Y.; Wang, F. Artificial intelligence for COVID-19: Battling the pandemic with computational intelligence. Intell. Med. 2022, 2, 13–29. [Google Scholar] [CrossRef]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement. BMJ 2009, 339, 332–336. [Google Scholar] [CrossRef]
- Carrazco-Montalvo, A.; Armendáriz-Castillo, I.; Tello, C.L.; Morales, D.; Armas-Gonzalez, R.; Guizado-Herrera, D.; León-Sosa, A.; Ramos-Sarmiento, D.; Fuertes, B.; Patino, L.; et al. First detection of SARS-CoV-2 variant B.1.1.529 (Omicron) in Ecuador. New Microbes New Infect. 2022, 529, 100951. [Google Scholar] [CrossRef] [PubMed]
- Saxena, S.K.; Kumar, S.; Ansari, S.; Paweska, J.T.; Maurya, V.K.; Tripathi, A.K.; Abdel-Moneim, A.S. Characterization of the novel SARS-CoV-2 Omicron (B.1.1.529) variant of concern and its global perspective. J. Med. Virol. 2021, 94, 1738–1744. [Google Scholar] [CrossRef]
- Kupferschmidt, K. Where did ‘weird’ Omicron come from? Science 2021, 374, 1179. [Google Scholar] [CrossRef]
- World Health Organisation. World Helath Organisation Update on Omicron; World Health Organisation: Geneva, Switzerland, 2021. [Google Scholar]
- Alba, J.M.G.; Pérez-Martínez, Z.; Boga, J.A.; Rojo-Alba, S.; de Oña, J.G.; Alvarez-Argüelles, M.E.; Rodríguez, G.M.; Gonzalez, I.C.; González, I.H.; Coto, E.; et al. Emergence of New SARS-CoV2 Omicron Variants after the Change of Surveillance and Control Strategy. Microorganisms 2022, 10, 1954. [Google Scholar] [CrossRef] [PubMed]
- Lyngse, F.P.; Kirkeby, C.T.; Denwood, M.; Christiansen, L.E.; Mølbak, K.; Møller, C.H.; Skov, R.L.; Krause, T.G.; Rasmussen, M.; Sieber, R.N.; et al. Household transmission of SARS-CoV-2 Omicron variant of concern subvariants BA.1 and BA.2 in Denmark. Nat. Commun. 2022, 13, 5760. [Google Scholar] [CrossRef] [PubMed]
- Ma, K.; Chen, J. Omicron XE emerges as SARS-CoV-2 keeps evolving. Innovation 2022, 3, 100248. [Google Scholar] [CrossRef]
- Tegally, H.; Moir, M.; Everatt, J.; Giovanetti, M.; Scheepers, C.; Wilkinson, E.; Subramoney, K.; Makatini, Z.; Moyo, S.; Amoako, D.G.; et al. Emergence of SARS-CoV-2 Omicron lineages BA.4 and BA.5 in South Africa. Nat. Med. 2022, 28, 1785–1790. [Google Scholar] [CrossRef]
- Centers of Disease Control and Prevention. Science Brief: Omicron (B.1.1.529) Variant. Available online: https://www.cdc.gov/coronavirus/2019-ncov/science/science-briefs/scientific-brief-Omicron-variant.html#print (accessed on 3 February 2022).
- Gupta, A.; Pradhan, A.; Maurya, V.K.; Kumar, S.; Theengh, A.; Puri, B.; Saxena, S.K. Therapeutic approaches for SARS-CoV-2 infection. Methods 2021, 195, 29–43. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef]
- Walls, A.C.; Park, Y.-J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 183, 1735. [Google Scholar] [CrossRef]
- Chen, J.; Wang, R.; Gilby, N.B.; Wei, G.-W. Omicron Variant (B.1.1.529): Infectivity, Vaccine Breakthrough, and Antibody Resistance. J. Chem. Inf. Model. 2022, 62, 412–422. [Google Scholar] [CrossRef]
- Dawood, A.A. Increasing the frequency of omicron variant mutations boosts the immune response and may reduce the virus virulence. Microb. Pathog. 2022, 164, 105400. [Google Scholar] [CrossRef]
- Lai, C.C.; Wang, C.Y.; Hsueh, P.R. Co-infections among patients with COVID-19: The need for combination therapy with non-anti-SARS-CoV-2 agents? J. Microbiol. Immunol. Infect. 2020, 53, 505–512. [Google Scholar] [CrossRef]
- Esper, F.P.; Spahlinger, T.; Zhou, L. Rate and influence of respiratory virus co-infection on pandemic (H1N1) influenza disease. J. Infect. 2011, 63, 260–266. [Google Scholar] [CrossRef]
- Klein, E.Y.; Monteforte, B.; Gupta, A.; Jiang, W.; May, L.; Hsieh, Y.H.; Dugas, A. The frequency of influenza and bacterial coinfection: A systematic review and meta-analysis. Influenza Other Respi. Viruses 2016, 10, 394–403. [Google Scholar] [CrossRef] [PubMed]
- Santos, A.P.; Gonçalves, L.C.; Oliveira, A.C.C.; Queiroz, P.H.P.; Ito, C.R.M.; Santos, M.O.; Carneiro, L.C. Bacterial Co-Infection in Patients with COVID-19 Hospitalized (ICU and Not ICU): Review and Meta-Analysis. Antibiotics 2022, 11, 894. [Google Scholar] [CrossRef] [PubMed]
- Bengoechea, J.A.; Bamford, C.G.G. SARS-CoV-2, Bacterial Co-Infections, and Amr: The Deadly Trio in COVID-19? Juvenis Sci. 2020, 6, 42–50. [Google Scholar] [CrossRef]
- Jeong, W.; Keighley, C.; Wolfe, R.; Lee, W.L.; Slavin, M.A.; Kong, D.C.M.; Chen, S.C.A. The epidemiology and clinical manifestations of mucormycosis: A systematic review and meta-analysis of case reports. Clin. Microbiol. Infect. 2019, 25, 26–34. [Google Scholar] [CrossRef]
- John, T.M.; Jacob, C.N.; Kontoyiannis, D.P. When uncontrolled diabetes mellitus and severe COVID-19 converge: The perfect storm for mucormycosis. J. Fungi 2021, 7, 298. [Google Scholar] [CrossRef] [PubMed]
- Spellberg, B.; Edwards, J.; Ibrahim, A. Novel Perspectives on Mucormycosis: Pathophysiology, Presentation, and Management. Clin. Microbiol. Rev. 2005, 18, 556–569. [Google Scholar] [CrossRef] [PubMed]
- Syed-Abdul, S.; Babu, A.S.; Bellamkonda, R.S.; Itumalla, R.; Acharyulu, G.; Krishnamurthy, S.; Ramana, Y.V.S.; Mogilicharla, N.; Malwade, S.; Li, Y.-C. Using artificial intelligence-based models to predict the risk of mucormycosis among COVID-19 survivors: An experience from a public hospital in India. J. Infect. 2022, 84, 351–354. [Google Scholar] [CrossRef]
- Karthikeyan, S.; Ramkumar, G.; Aravindkumar, S.; Tamilselvi, M.; Ramesh, S.; Ranjith, A. A Novel Deep Learning-Based Black Fungus Disease Identification Using Modified Hybrid Learning Methodology. Contrast Media Mol. Imaging 2022, 2022, 4352730. [Google Scholar] [CrossRef]
- Subramaniam, S.; Scharrer, I. Procoagulant activity during viral infections. Front. Biosci. 2018, 23, 1060–1081. [Google Scholar] [CrossRef]
- Arachchillage, D.R.J.; Laffan, M. Abnormal coagulation parameters are associated with poor prognosis in patients with novel coronavirus pneumonia. J. Thromb. Haemost. 2020, 18, 1233–1234. [Google Scholar] [CrossRef] [PubMed]
- Kirchberger, I.; Berghaus, T.M.; von Scheidt, W.; Linseisen, J.; Meisinger, C. COVID-19 risk perceptions, worries and preventive behaviors in patients with previous pulmonary embolism. Thromb. Res. 2021, 202, 77–83. [Google Scholar] [CrossRef]
- Muñoz-Rivas, N.; Abad-Motos, A.; Mestre-Gómez, B.; Sierra-Hidalgo, F.; Cortina-Camarero, C.; Lorente-Ramos, R.M.; Torres-Rubio, P.; Arranz-García, P.; Franco-Moreno, A.I.; Gómez-Mariscal, E.; et al. Systemic thrombosis in a large cohort of COVID-19 patients despite thromboprophylaxis: A retrospective study. Thromb. Res. 2021, 199, 132–142. [Google Scholar] [CrossRef]
- Pancani, R.; Villari, L.; Foci, V.; Parri, G.; Barsotti, F.; Patrucco, F.; Malerba, M.; Vincenti, R.; Carrozzi, L.; Celi, A. Lower limb deep vein thrombosis in COVID-19 patients admitted to intermediate care respiratory units. Thromb. Res. 2021, 197, 44–47. [Google Scholar] [CrossRef] [PubMed]
- Heit, J.A.; Mohr, D.N.; Silverstein, M.D.; Petterson, T.M.; O’Fallon, W.M.; Melton, L.J. Predictors of recurrence after deep vein thrombosis and pulmonary embolism: A population-based cohort study. Arch. Intern. Med. 2002, 160, 761–768. [Google Scholar] [CrossRef]
- Aktaa, S.; Wu, J.; Nadarajah, R.; Rashid, M.; de Belder, M.; Deanfield, J.; Mamas, M.A.; Gale, C.P. Incidence and mortality due to thromboembolic events during the COVID-19 pandemic: Multi-sourced population-based health records cohort study. Thromb. Res. 2021, 202, 17–23. [Google Scholar] [CrossRef]
- Cui, S.; Chen, S.; Li, X.; Liu, S.; Wang, F. Prevalence of venous thromboembolism in patients with severe novel coronavirus pneumonia. J. Thromb. Haemost. 2020, 18, 1421–1424. [Google Scholar] [CrossRef]
- Malato, A.; Dentali, F.; Siragusa, S.; Fabbiano, F.; Kagoma, Y.; Boddi, M.; Gensini, G.F.; Peris, A.; Crowther, M.; Napolitano, M. The impact of deep vein thrombosis in critically ill patients: A meta-analysis of major clinical outcomes. Blood Transfus. 2015, 13, 559–568. [Google Scholar] [CrossRef]
- Bozzani, A.; Arici, V.; Tavazzi, G.; Franciscone, M.M.; Danesino, V.; Rota, M.; Rossini, R.; Sterpetti, A.V.; Ticozzelli, G.; Rumi, E.; et al. Acute arterial and deep venous thromboembolism in COVID-19 patients: Risk factors and personalized therapy. Surgery 2020, 168, 987–992. [Google Scholar] [CrossRef]
- Valle, C.; Bonaffini, P.A.; Dal Corso, M.; Mercanzin, E.; Franco, P.N.; Sonzogni, A.; Vacca, G.; Gianatti, A.; Sironi, S. Association between pulmonary embolism and COVID-19 severe pneumonia: Experience from two centers in the core of the infection Italian peak. Eur. J. Radiol. 2021, 137, 109613. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Gao, W.; Guo, W.; Guo, Y.; Shi, M.; Dong, G.; Ge, Q.; Zhu, J.; Lu, J. Prominent coagulation disorder is closely related to inflammatory response and could be as a prognostic indicator for ICU patients with COVID-19. J. Thromb. Thrombolysis 2020, 50, 825–832. [Google Scholar] [CrossRef]
- Teimury, A.; Khameneh, M.T.; Khaledi, E.M. Major coagulation disorders and parameters in COVID-19 patients. Eur. J. Med. Res. 2022, 27, 25. [Google Scholar] [CrossRef] [PubMed]
- Fang, K.; Dong, Z.; Chen, X.; Zhu, J.; Zhang, B.; You, J.; Xiao, Y.; Xia, W. Using machine learning to identify clotted specimens in coagulation testing. Clin. Chem. Lab. Med. 2021, 59, 1289–1297. [Google Scholar] [CrossRef]
- Lansbury, L.; Lim, B.; Baskaran, V.; Lim, W.S. Co-infections in people with COVID-19: A systematic review and meta-analysis. J. Infect. 2020, 81, 266–275. [Google Scholar] [CrossRef]
- Khatiwada, S.; Subedi, A. Lung microbiome and coronavirus disease 2019 (COVID-19): Possible link and implications. Hum. Microbiome J. 2020, 17, 100073. [Google Scholar] [CrossRef] [PubMed]
- Hoque, M.N.; Rahman, M.S.; Ahmed, R.; Hossain, M.S.; Islam, M.S.; Islam, T.; Hossain, M.A.; Siddiki, A.Z. Diversity and genomic determinants of the microbiomes associated with COVID-19 and non-COVID respiratory diseases. Gene Rep. 2021, 23, 101200. [Google Scholar] [CrossRef] [PubMed]
- Bonazzetti, C.; Morena, V.; Giacomelli, A.; Oreni, L.; Casalini, G.; Galimberti, L.R.; Bolis, M.; Rimoldi, M.; Ballone, E.; Colombo, R.; et al. Unexpectedly High Frequency of Enterococcal Bloodstream Infections in Coronavirus Disease 2019 Patients Admitted to an Italian ICU: An Observational Study. Crit. Care Med. 2021, 49, e31–e40. [Google Scholar] [CrossRef]
- Giacobbe, D.R.; Battaglini, D.; Ball, L.; Brunetti, I.; Bruzzone, B.; Codda, G.; Crea, F.; De Maria, A.; Dentone, C.; Di Biagio, A.; et al. Bloodstream infections in critically ill patients with COVID-19. Eur. J. Clin. Investig. 2020, 50, e13319. [Google Scholar] [CrossRef]
- Musuuza, J.S.; Watson, L.; Parmasad, V.; Putman-Buehler, N.; Christensen, L.; Safdar, N. Prevalence and outcomes of co-infection and superinfection with SARS-CoV-2 and other pathogens: A systematic review and metaanalysis. PLoS ONE 2021, 16, e0251170. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Guerra, B.A.; Gonzalez-Lara, M.F.; de-Leon-Cividanes, N.A.; Tamez-Torres, K.M.; Roman-Montes, C.M.; Rajme-Lopez, S.; Villalobos-Zapata, G.I.; Lopez-Garcia, N.I.; Martínez-Gamboa, A.; Sifuentes-Osornio, J.; et al. Antimicrobial resistance patterns and antibiotic use during hospital conversion in the COVID-19 pandemic. Antibiotics 2021, 10, 182. [Google Scholar] [CrossRef]
- Contou, D.; Claudinon, A.; Pajot, O.; Micaëlo, M.; Longuet Flandre, P.; Dubert, M.; Cally, R.; Logre, E.; Fraissé, M.; Mentec, H.; et al. Bacterial and viral co-infections in patients with severe SARS-CoV-2 pneumonia admitted to a French ICU. Ann. Intensive Care 2020, 10, 119. [Google Scholar] [CrossRef]
- Pourajam, S.; Kalantari, E.; Talebzadeh, H.; Mellali, H.; Sami, R.; Soltaninejad, F.; Amra, B.; Sajadi, M.; Alenaseri, M.; Kalantari, F.; et al. Secondary Bacterial Infection and Clinical Characteristics in Patients with COVID-19 Admitted to Two Intensive Care Units of an Academic Hospital in Iran During the First Wave of the Pandemic. Front. Cell. Infect. Microbiol. 2022, 12. [Google Scholar] [CrossRef]
- Pai, K.C.; Wang, M.S.; Chen, Y.F.; Tseng, C.H.; Liu, P.Y.; Chen, L.C.; Sheu, R.K.; Wu, C.L. An artificial intelligence approach to bloodstream infections prediction. J. Clin. Med. 2021, 10, 2901. [Google Scholar] [CrossRef] [PubMed]
- Zoabi, Y.; Kehat, O.; Lahav, D.; Weiss-Meilik, A.; Adler, A.; Shomron, N. Predicting bloodstream infection outcome using machine learning. Sci. Rep. 2021, 11, 20101. [Google Scholar] [CrossRef]
- Flores-Mireles, A.L.; Walker, J.N.; Caparon, M.; Hultgren, S.J. Urinary tract infections: Epidemiology, mechanisms of infection and treatment options. Nat. Rev. Microbiol. 2015, 13, 269–284. [Google Scholar] [CrossRef]
- Parish, A.; Holliday, K. Long-Term Care Acquired Urinary Tract Infections’ Antibiotic Resistance Patterns and Empiric Therapy: A Pilot Study. Geriatr. Nurs. 2012, 33, 473–478. [Google Scholar] [CrossRef] [PubMed]
- Hof, H. Candidurie! Was nun? Urologe 2017, 56, 172–179. [Google Scholar] [CrossRef]
- Bendala Estrada, A.D.; Calderón Parra, J.; Fernández Carracedo, E.; Muiño Míguez, A.; Ramos Martínez, A.; Muñez Rubio, E.; Rubio-Rivas, M.; Agudo, P.; Arnalich Fernández, F.; Estrada Perez, V.; et al. Inadequate use of antibiotics in the covid-19 era: Effectiveness of antibiotic therapy. BMC Infect. Dis. 2021, 21, 1144. [Google Scholar] [CrossRef] [PubMed]
- Bardi, T.; Pintado, V.; Gomez-Rojo, M.; Escudero-Sanchez, R.; Azzam Lopez, A.; Diez-Remesal, Y.; Martinez Castro, N.; Ruiz-Garbajosa, P.; Pestaña, D. Nosocomial infections associated to COVID-19 in the intensive care unit: Clinical characteristics and outcome. Eur. J. Clin. Microbiol. Infect. Dis. 2021, 40, 495–502. [Google Scholar] [CrossRef]
- Karaba, S.M.; Jones, G.; Helsel, T.; Smith, L.L.; Avery, R.; Dzintars, K.; Salinas, A.B.; Keller, S.C.; Townsend, J.L.; Klein, E.; et al. Prevalence of co-infection at the time of hospital admission in COVID-19 Patients, A multicenter study. Open Forum Infect. Dis. 2021, 8, ofaa578. [Google Scholar] [CrossRef]
- DeVoe, C.; Segal, M.R.; Wang, L.; Stanley, K.; Madera, S.; Fan, J.; Schouest, J.; Graham-Ojo, R.; Nichols, A.; Prasad, P.A.; et al. Increased rates of secondary bacterial infections, including Enterococcus bacteremia, in patients hospitalized with coronavirus disease 2019 (COVID-19). Infect. Control Hosp. Epidemiol. 2022, 43, 1416–1423. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.A.; Moore, C.L.; Cheung, K.H.; Brandt, C. Predicting urinary tract infections in the emergency department with machine learning. PLoS ONE 2018, 13, e0194085. [Google Scholar] [CrossRef]
- Burton, R.J.; Albur, M.; Eberl, M.; Cuff, S.M. Using artificial intelligence to reduce diagnostic workload without compromising detection of urinary tract infections. BMC Med. Inform. Decis. Mak. 2019, 19, 171. [Google Scholar] [CrossRef]
- Hsieh, C.C.; Lin, C.H.; Wang, W.Y.C.; Pauleen, D.J.; Chen, J.V. The outcome and implications of public precautionary measures in taiwan–declining respiratory disease cases in the COVID-19 pandemic. Int. J. Environ. Res. Public Health 2020, 17, 4877. [Google Scholar] [CrossRef] [PubMed]
- Zavala-Flores, E.; Salcedo-Matienzo, J. Medicación prehospitalaria en pacientes hospitalizados por COVID-19 en un hospital público de Lima-Perú. Acta Med. Peru. 2020, 37, 393–395. [Google Scholar] [CrossRef]
- Tiri, B.; Sensi, E.; Marsiliani, V.; Cantarini, M.; Priante, G.; Vernelli, C.; Martella, L.A.; Costantini, M.; Mariottini, A.; Andreani, P.; et al. Antimicrobial Stewardship Program, COVID-19, and Infection Control: Spread of Carbapenem-Resistant Klebsiella Pneumoniae Colonization in ICU COVID-19 Patients. What Did Not Work? J. Clin. Med. 2020, 9, 2744. [Google Scholar] [CrossRef]
- Fattorini, L.; Creti, R.; Palma, C.; Pantosti, A.; Unit of Antibiotic Resistance and Special Pathogens; Unit of Antibiotic Resistance and Special Pathogens. Bacterial coinfections in COVID-19: An underestimated adversary. Ann. Ist. Super. Sanita 2020, 56, 359–364. [Google Scholar] [CrossRef]
- Oonsivilai, M.; Mo, Y.; Luangasanatip, N.; Lubell, Y.; Miliya, T.; Tan, P.; Loeuk, L.; Turner, P.; Cooper, B.S. Using machine learning to guide targeted and locally-tailored empiric antibiotic prescribing in a children’s hospital in Cambodia [version 1; referees: 2 approved]. Wellcome Open Res. 2018, 3, 1–18. [Google Scholar] [CrossRef]
- Feretzakis, G.; Loupelis, E.; Sakagianni, A.; Kalles, D.; Martsoukou, M.; Lada, M.; Skarmoutsou, N.; Christopoulos, C.; Valakis, K.; Velentza, A.; et al. Using machine learning techniques to aid empirical antibiotic therapy decisions in the intensive care unit of a general hospital in Greece. Antibiotics 2020, 9, 50. [Google Scholar] [CrossRef]
- Martínez-Agüero, S.; Mora-Jiménez, I.; Lérida-García, J.; Álvarez-Rodríguez, J.; Soguero-Ruiz, C. Machine Learning Techniques to Identify Antimicrobial Resistance in the Intensive Care Unit. Entropy 2019, 21, 603. [Google Scholar] [CrossRef]
- Feretzakis, G.; Sakagianni, A.; Loupelis, E.; Kalles, D.; Skarmoutsou, N.; Martsoukou, M.; Christopoulos, C.; Lada, M.; Petropoulou, S.; Velentza, A.; et al. Machine learning for antibiotic resistance prediction: A prototype using off-the-shelf techniques and entry-level data to guide empiric antimicrobial therapy. Healthc. Inform. Res. 2021, 27, 214–221. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Zhao, C.; Yin, Y.; Chen, F.; Chen, H.; Wang, H. A Practical Approach for Predicting Antimicrobial Phenotype Resistance in Staphylococcus aureus through Machine Learning Analysis of Genome Data. Front. Microbiol. 2022, 13, 1–9. [Google Scholar] [CrossRef]
- Santerre, J.W.; Davis, J.J.; Xia, F.; Stevens, R. Machine Learning for Antimicrobial Resistance. arXiv 2016, arXiv:1607.01224. [Google Scholar]
- Feretzakis, G.; Loupelis, E.; Sakagianni, A.; Kalles, D.; Lada, M.; Christopoulos, C.; DImitrellos, E.; Martsoukou, M.; Skarmoutsou, N.; Petropoulou, S.; et al. Using machine learning algorithms to predict antimicrobial resistance and assist empirical treatment. Stud. Health Technol. Inform. 2020, 272, 75–78. [Google Scholar] [CrossRef]
- Lewin-Epstein, O.; Baruch, S.; Hadany, L.; Stein, G.Y.; Obolski, U. Predicting Antibiotic Resistance in Hospitalized Patients by Applying Machine Learning to Electronic Medical Records. Clin. Infect. Dis. 2021, 72, E848–E855. [Google Scholar] [CrossRef]
- Huang, T.S.; Lee, S.S.J.; Lee, C.C.; Chang, F.C. Detection of carbapenem-resistant Klebsiella pneumoniae on the basis of matrix-assisted laser desorption ionization time-of-flight mass spectrometry by using supervised machine learning approach. PLoS ONE 2020, 15, e0228459. [Google Scholar] [CrossRef]
- Sinha, A.; Rathi, M. COVID-19 prediction using AI analytics for South Korea. Appl. Intell. 2021, 51, 8579–8597. [Google Scholar] [CrossRef]
- Harmon, S.A.; Sanford, T.H.; Xu, S.; Turkbey, E.B.; Roth, H.; Xu, Z.; Yang, D.; Myronenko, A.; Anderson, V.; Amalou, A.; et al. Artificial intelligence for the detection of COVID-19 pneumonia on chest CT using multinational datasets. Nat. Commun. 2020, 11, 4080. [Google Scholar] [CrossRef]
- Buonsenso, D.; Pata, D.; Chiaretti, A. COVID-19 outbreak: Less stethoscope, more ultrasound. Lancet Respir. Med. 2020, 8, e27. [Google Scholar] [CrossRef]
- Born, J.; Wiedemann, N.; Cossio, M.; Buhre, C.; Brändle, G.; Leidermann, K.; Aujayeb, A.; Moor, M.; Rieck, B.; Borgwardt, K. Accelerating detection of lung pathologies with explainable ultrasound image analysis. Appl. Sci. 2021, 11, 672. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 1–9. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Chollet, F. Xception: Deep learning with depthwise separable convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 1251–1258. [Google Scholar]
- Zokaeinikoo, M.; Kazemian, P.; Mitra, P.; Kumara, S. AIDCOV: An Interpretable Artificial Intelligence Model for Detection of COVID-19 from Chest Radiography Images. ACM Trans. Manag. Inf. Syst. 2021, 12, 1–20. [Google Scholar] [CrossRef]
- Meng, H.; Xiong, R.; He, R.; Lin, W.; Hao, B.; Zhang, L.; Lu, Z.; Shen, X.; Fan, T.; Jiang, W.; et al. CT imaging and clinical course of asymptomatic cases with COVID-19 pneumonia at admission in Wuhan, China. J. Infect. 2020, 81, e33–e39. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Kaur, A.; Kumar, M. Face detection techniques: A review. Artif. Intell. Rev. 2019, 52, 927–948. [Google Scholar] [CrossRef]
- Niu, G.; Chen, Q. Learning an video frame-based face detection system for security fields. J. Vis. Commun. Image Represent. 2018, 55, 457–463. [Google Scholar] [CrossRef]
- Lalmuanawma, S.; Hussain, J.; Chhakchhuak, L. Applications of machine learning and artificial intelligence for COVID-19 (SARS-CoV-2) pandemic: A review. Chaos Solitons Fractals 2020, 139, 110059. [Google Scholar] [CrossRef]
- Sagayam, K.M. CNN-based Mask Detection System Using OpenCV and MobileNetV2. In Proceedings of the 2021 3rd International Conference on Signal Processing and Communication (ICPSC), Tamil Nadu, India, 13–14 May 2021; pp. 115–119. [Google Scholar]
- Loey, M.; Manogaran, G.; Taha, M.H.N.; Khalifa, N.E.M. Fighting against COVID-19: A novel deep learning model based on YOLO-v2 with ResNet-50 for medical face mask detection. Sustain. Cities Soc. 2021, 65, 102600. [Google Scholar] [CrossRef]
- Du, J. Understanding of object detection based on CNN family and YOLO. J. Phys. Conf. Ser. 2018, 1004, 12029. [Google Scholar] [CrossRef]
- Bhuiyan, M.R.; Khushbu, S.A.; Islam, M.S. A deep learning based assistive system to classify COVID-19 face mask for human safety with YOLOv3. In Proceedings of the 2020 11th International Conference on Computing, Communication and Networking Technologies (ICCCNT), Kharagpur, India, 1–3 July 2020; pp. 1–5. [Google Scholar]
- Degadwala, S.; Vyas, D.; Chakraborty, U.; Dider, A.R.; Biswas, H. Yolo-v4 Deep Learning Model for Medical Face Mask Detection. In Proceedings of the 2021 International Conference on Artificial Intelligence and Smart Systems (ICAIS), Coimbatore, India, 25–27 March 2021; pp. 209–213. [Google Scholar]
- Nagrath, P.; Jain, R.; Madan, A.; Arora, R.; Kataria, P.; Hemanth, J. SSDMNV2: A real time DNN-based face mask detection system using single shot multibox detector and MobileNetV2. Sustain. Cities Soc. 2021, 66, 102692. [Google Scholar] [CrossRef]
- Sethi, S.; Kathuria, M.; Kaushik, T. Face mask detection using deep learning: An approach to reduce risk of Coronavirus spread. J. Biomed. Inform. 2021, 120, 103848. [Google Scholar] [CrossRef]
- Varshini, B.; Yogesh, H.; Pasha, S.D.; Suhail, M.; Madhumitha, V.; Sasi, A. IoT-Enabled smart doors for monitoring body temperature and face mask detection. Glob. Transit. Proc. 2021, 2, 246–254. [Google Scholar] [CrossRef]
- Teboulbi, S.; Messaoud, S.; Hajjaji, M.A.; Mtibaa, A. Real-Time Implementation of AI-Based Face Mask Detection and Social Distancing Measuring System for COVID-19 Prevention. Sci. Program. 2021, 2021, 8340779. [Google Scholar] [CrossRef]
- Loey, M.; Manogaran, G.; Taha, M.H.N.; Khalifa, N.E.M. A hybrid deep transfer learning model with machine learning methods for face mask detection in the era of the COVID-19 pandemic. Measurement 2021, 167, 108288. [Google Scholar] [CrossRef] [PubMed]
- Jignesh Chowdary, G.; Punn, N.S.; Sonbhadra, S.K.; Agarwal, S. Face Mask Detection Using Transfer Learning of InceptionV3. In Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer: Berlin/Heidelberg, Germany, 2020; Volume 12581 LNCS, pp. 81–90. ISBN 9783030666644. [Google Scholar]
- Inamdar, M.; Mehendale, N. Real-Time Face Mask Identification Using Facemasknet Deep Learning Network. SSRN Electron. J. 2020. [Google Scholar] [CrossRef]
- Balasubramaniam, V. Facemask Detection Algorithm on COVID Community Spread Control using EfficientNet Algorithm. J. Soft Comput. Paradig. 2021, 3, 110–122. [Google Scholar] [CrossRef]
- Saravanan, T.M.; Karthiha, K.; Kavinkumar, R.; Gokul, S.; Mishra, J.P. A novel machine learning scheme for face mask detection using pretrained convolutional neural network. Mater. Today Proc. 2022, 58, 150–156. [Google Scholar] [CrossRef] [PubMed]
- Gupta, M.; Chaudhary, G.; Bansal, D.; Pandey, S. DTLMV2—A real-time deep transfer learning mask classifier for overcrowded spaces. Appl. Soft Comput. 2022, 127, 109313. [Google Scholar] [CrossRef] [PubMed]
- Ullah, N.; Javed, A.; Ali Ghazanfar, M.; Alsufyani, A.; Bourouis, S. A novel DeepMaskNet model for face mask detection and masked facial recognition. J. King Saud Univ.-Comput. Inf. Sci. 2022, 34, 9905–9914. [Google Scholar] [CrossRef]
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: A descriptive study. Lancet 2020, 395, 507–513. [Google Scholar] [CrossRef]
- Leow, Y.; Shi, J.K.; Liu, W.; Ni, X.P.; Yew, P.Y.M.; Liu, S.; Li, Z.; Xue, Y.; Kai, D.; Loh, X.J. Design and development of multilayer cotton masks via machine learning. Mater. Today Adv. 2021, 12, 100178. [Google Scholar] [CrossRef]
- Shin, J.; Jeong, S.; Kim, J.; Choi, Y.Y.; Choi, J.; Lee, J.G.; Kim, S.; Kim, M.; Rho, Y.; Hong, S.; et al. Dynamic Pore Modulation of Stretchable Electrospun Nanofiber Filter for Adaptive Machine Learned Respiratory Protection. ACS Nano 2021, 15, 15730–15740. [Google Scholar] [CrossRef] [PubMed]
- Abutaleb, A.; ArunPrasanna, V. Fabrication of biopolymer nanofibers from natural sources. Text. Res. J. 2021, 92, 004051752110550. [Google Scholar] [CrossRef]
- Qin, X.; Subianto, S. Electrospun nanofibers for filtration applications. In Electrospun Nanofibers; Elsevier: Amsterdam, The Netherlands, 2017; pp. 449–466. [Google Scholar]
- Shen, H.; Zhou, Z.; Wang, H.; Zhang, M.; Han, M.; Durkin, D.P.; Shuai, D.; Shen, Y. Development of Electrospun Nanofibrous Filters for Controlling Coronavirus Aerosols. Environ. Sci. Technol. Lett. 2021, 8, 545–550. [Google Scholar] [CrossRef]
- Ullah, S.; Ullah, A.; Lee, J.; Jeong, Y.; Hashmi, M.; Zhu, C.; Joo, K.I.; Cha, H.J.; Kim, I.S. Reusability Comparison of Melt-Blown vs Nanofiber Face Mask Filters for Use in the Coronavirus Pandemic. ACS Appl. Nano Mater. 2020, 3, 7231–7241. [Google Scholar] [CrossRef]
- Ishack, S.; Lipner, S.R. Applications of 3D Printing Technology to Address COVID-19–Related Supply Shortages. Am. J. Med. 2020, 133, 771–773. [Google Scholar] [CrossRef]
- Kumar, S.; Karmacharya, M.; Joshi, S.R.; Gulenko, O.; Park, J.; Kim, G.H.; Cho, Y.K. Photoactive Antiviral Face Mask with Self-Sterilization and Reusability. Nano Lett. 2021, 21, 337–343. [Google Scholar] [CrossRef]
- Shan, X.; Zhang, H.; Liu, C.; Yu, L.; Di, Y.; Zhang, X.; Dong, L.; Gan, Z. Reusable Self-Sterilization Masks Based on Electrothermal Graphene Filters. ACS Appl. Mater. Interfaces 2020, 12, 56579–56586. [Google Scholar] [CrossRef]
- Le, T.T.; Curry, E.J.; Vinikoor, T.; Das, R.; Liu, Y.; Sheets, D.; Tran, K.T.M.; Hawxhurst, C.J.; Stevens, J.F.; Hancock, J.N.; et al. Piezoelectric Nanofiber Membrane for Reusable, Stable, and Highly Functional Face Mask Filter with Long-Term Biodegradability. Adv. Funct. Mater. 2022, 32, 2113040. [Google Scholar] [CrossRef]
- Chaudhary, V.; Royal, A.; Chavali, M.; Yadav, S.K. Advancements in research and development to combat COVID-19 using nanotechnology. Nanotechnol. Environ. Eng. 2021, 6, 8. [Google Scholar] [CrossRef]
- El-Atab, N.; Qaiser, N.; Badghaish, H.; Shaikh, S.F.; Hussain, M.M.; Hussain, M.M. Flexible Nanoporous Template for the Design and Development of Reusable Anti-COVID-19 Hydrophobic Face Masks. ACS Nano 2020, 14, 7659–7665. [Google Scholar] [CrossRef]
- Kim, H.-D.; Kim, D.H.; Seo, P.W.; Bae, J. Design of Convolution Neural Network (CNN) Based Medicine Classifier for Nursing Robots. IEMEK J. Embed. Syst. Appl. 2021, 16, 187–193. [Google Scholar]
- Karabegović, I.; Husak, E.; Isić, S.; Karabegović, E.; Mahmić, M. Service Robots and Artificial Intelligence for Faster Diagnostics and Treatment in Medicine. In New Technologies, Development and Application IV, Proceedings of the International Conference “New Technologies, Development and Applications”, Sarajevo, Bosnia and Herzegovina, 24–26 June 2021; Springer: Cham, Switzerland, 2021; pp. 3–20. [Google Scholar]
- Shamout, M.; Ben-Abdallah, R.; Alshurideh, M.; Alzoubi, H.; Kurdi, B.; Hamadneh, S. A conceptual model for the adoption of autonomous robots in supply chain and logistics industry. Uncertain Supply Chain. Manag. 2022, 10, 577–592. [Google Scholar] [CrossRef]
- Ponce, P.; Mata, O.; Perez, E.; Lopez, J.R.; Molina, A.; McDaniel, T. S4 Features and Artificial Intelligence for Designing a Robot against COVID-19—Robocov. Futur. Internet 2022, 14, 22. [Google Scholar] [CrossRef]
- Suvarna, M.; Katragadda, A.; Sun, Z.; Choh, Y.B.; Chen, Q.; PS, P.; Wang, X. A machine learning framework to quantify and assess the impact of COVID-19 on the power sector: An Indian context. Adv. Appl. Energy 2022, 5, 100078. [Google Scholar] [CrossRef]
- Norouzi, N.; Zarazua de Rubens, G.; Choubanpishehzafar, S.; Enevoldsen, P. When pandemics impact economies and climate change: Exploring the impacts of COVID-19 on oil and electricity demand in China. Energy Res. Soc. Sci. 2020, 68, 101654. [Google Scholar] [CrossRef]
- Soni, S.; Roberts, K. An evaluation of two commercial deep learning-based information retrieval systems for COVID-19 literature. J. Am. Med. Informatics Assoc. 2021, 28, 132–137. [Google Scholar] [CrossRef]
- Ou, S.; He, X.; Ji, W.; Chen, W.; Sui, L.; Gan, Y.; Lu, Z.; Lin, Z.; Deng, S.; Przesmitzki, S.; et al. Machine learning model to project the impact of COVID-19 on US motor gasoline demand. Nat. Energy 2020, 5, 666–673. [Google Scholar] [CrossRef]
- Dias, R.D.; Shah, J.A.; Zenati, M.A. Artificial intelligence in cardiothoracic surgery. Minerva Cardioangiol. 2020, 68, 532–538. [Google Scholar] [CrossRef]
- Shabbir, A.; Shabbir, M.; Javed, A.R.; Rizwan, M.; Iwendi, C.; Chakraborty, C. Exploratory data analysis, classification, comparative analysis, case severity detection, and internet of things in COVID-19 telemonitoring for smart hospitals. J. Exp. Theor. Artif. Intell. 2022, 1–28. [Google Scholar] [CrossRef]
- Cingolani, M.; Scendoni, R.; Fedeli, P.; Cembrani, F. Artificial intelligence and digital medicine for integrated home care services in Italy: Opportunities and limits. Front. Public Health 2023, 10, 1095001. [Google Scholar] [CrossRef]
- Panicacci, S.; Donati, M.; Lubrano, A.; Vianello, A.; Ruiu, A.; Melani, L.; Tomei, A.; Fanucci, L. Telemonitoring in the Covid-19 Era: The Tuscany Region Experience. Healthcare 2021, 9, 516. [Google Scholar] [CrossRef]
- Artificial Intelligence and Machine Learning (AI/ML)-Enabled Medical Devices. Available online: https://www.fda.gov/medical-devices/software-medical-device-samd/artificial-intelligence-and-machine-learning-aiml-enabled-medical-devices (accessed on 14 November 2022).
- USA, Congress.Gov. No Vaccine Passports Act. Available online: https://www.congress.gov/bill/117th-congress/house-bill/2384?s=1&r=89 (accessed on 14 November 2022).
- Van Der Maarten, V. Data Responsibility V2.2–510 Global. Available online: https://centre.humdata.org/data-responsibility/ (accessed on 28 October 2022).
- UK General Data Protection Regulation (UK GDPR) and Data Protection Act (DPA). 2018. Available online: https://www.gov.uk/government/publications/nhs-covid-19-app-privacy-information/nhs-covid-19-app-privacy-notice#lawful-basis (accessed on 28 October 2022).
- General Data Protection Regulation. Available online: http://data.europa.eu/eli/reg/2016/679/oj (accessed on 28 October 2022).
- EU, Medical AI Tools. 2017/745 Medical Devices Regulations (MDR). Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:32017R0745 (accessed on 28 October 2022).
- EU, Medical AI Tools. The 2017/746 In Vitro Diagnostic Medical Devices Regulation (IVDR). Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=celex%3A32022R0112 (accessed on 28 October 2022).
- European Commission Proposal for a Regulation of the European Parliament and of the Council Laying Down Harmonised Rules on Artificial Intelligence (Artificial Intelligence Act) and Amending Certain Union Legislative Acts. Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/?qid=1623335154975&uri=CELEX%3A52021PC0206 (accessed on 30 October 2022).
- European Parliament Resolution of 16 February 2017 with Recommendations to the Commission on Civil Law Rules on Robotics (2015/2103(INL)). Available online: https://www.europarl.europa.eu/doceo/document/TA-8-2017-0051_EN.pdf (accessed on 30 October 2022).
- Government of Singapore Personal Data Protection Act 2012—Singapore Statutes Online. Available online: https://sso.agc.gov.sg/Act/PDPA2012?ProvIds=P1I-#pr4- (accessed on 30 October 2022).
- Therapeutic Goods Administration, Therapeutic Goods (Medical Devices) Regulations 2002. Available online: https://www.legislation.gov.au/Details/F2021C00217/Download (accessed on 30 October 2022).
- The State Council Notice of the State Council on Issuing the Development Plan for the New Generation of Artificial Intelligence. Available online: https://flia.org/wp-content/uploads/2017/07/A-New-Generation-of-Artificial-Intelligence-Development-Plan-1.pdf (accessed on 30 October 2022).
- Saudi Food and Drug Authority—Guidance on Software as a Medical Device, SFDA MDS-G23. Available online: https://www.sfda.gov.sa/sites/default/files/2021-04/SFDAArtificialIntelligenceEn.pdf (accessed on 30 October 2022).
- Gusev, A.V.; Morozov, S.P.; Kutichev, V.A.; Novitsky, R.E. Legal regulation of artificial intelligence software in healthcare in the Russian Federation. Med. Technol. Assess. Choice 2021, 1, 36. [Google Scholar] [CrossRef]
- Medical Devices Act—South Korea. Available online: https://elaw.klri.re.kr/eng_mobile/viewer.do?hseq=50798&type=sogan&key=31 (accessed on 14 November 2022).
- Singapore, Health Services Authority. Guidelines on Risk Classification of Standalone Medical Mobile Applications (SaMD) and Qualification of Clinical Decision Support Software (CDSS). Available online: https://www.hsa.gov.sg/announcements/regulatory-updates/consultation-on-regulatory-guidelines-for-classification-of-standalone-medical-mobile-applications-(samd)-and-qualification-of-clinical-decision-support-software-(cdss) (accessed on 14 November 2022).
- Standing Committee of the National People’s Congress Cybersecurity Law of the People’s Republic of China. Available online: http://www.xinhuanet.com//politics/2016-11/07/c_1119867015_2.htm (accessed on 30 October 2022).
- Gazette, G. Medical Device Act 2012 (ACT 737). Available online: www.federalgazette.agc.gov.my/outputaktap/20120209 (accessed on 14 November 2022).
- Abu Dhabi Department of Health Policy on Use of Artificial Intelligence (AI) in the Healthcare Sector of the Emirate of Abu Dhabi. Available online: https://www.google.com/url?sa=t&rct=j&q=&esrc=s&source=web&cd=&cad=rja&uact=8&ved=2ahUKEwi6iNbo0K77AhWjgv0HHdg5D-wQFnoECBAQAQ&url=https%3A%2F%2Fwww.doh.gov.ae%2F-%2Fmedia%2FE9C1470A575146B18015DEBE57E47F8D.ashx&usg=AOvVaw0TgyUjO4zetznNvkZiRKkt (accessed on 14 November 2022).
- House of Commons Digital Charter Implementation Act. Available online: https://www.parl.ca/DocumentViewer/en/43-2/bill/C-11/first-reading (accessed on 14 November 2022).
- Brazil. Law No 13, 709, of 14 August 2018 General Personal Data Protection Law (LGPD). Available online: http://www.planalto.gov.br/ccivil_03/_ato2015-2018/2018/lei/l13709.htm (accessed on 14 November 2022).
- Brazilian Artificial Intelligence Bill (Bill No. 21/2020). Available online: https://www.google.com/url?sa=t&rct=j&q=&esrc=s&source=web&cd=&cad=rja&uact=8&ved=2ahUKEwiY57269K_7AhVIh_0HHfGcDTQQFnoECAoQAQ&url=https%3A%2F%2Flegis.senado.leg.br%2Fsdleg-getter%2Fdocumento%2Fdownload%2Fa08e2a4b-da0c-4e58-8556-4e9f360e4c42&usg=AOvVaw15XW (accessed on 14 November 2022).
- Chatterjee, S.; Vardhan, B.; Singh, D.K.; Maitra, A.; Ojha, U.K. Should statins be considered for the management of mucormycosis in COVID-19? Diabetes Metab. Syndr. Clin. Res. Rev. 2021, 15, 102162. [Google Scholar] [CrossRef]
- Sujith, A.V.L.N.; Sajja, G.S.; Mahalakshmi, V.; Nuhmani, S.; Prasanalakshmi, B. Systematic review of smart health monitoring using deep learning and Artificial intelligence. Neurosci. Inform. 2022, 2, 100028. [Google Scholar] [CrossRef]
- Nguyen, D.C.; Ding, M.; Pathirana, P.N.; Seneviratne, A. Blockchain and AI-based solutions to combat coronavirus (COVID-19)-like epidemics: A survey. IEEE Access 2021, 9, 95730–95753. [Google Scholar] [CrossRef]
- Catalyst, N. What is telehealth? NEJM Catal. 2018, 4. Available online: https://catalyst.nejm.org/doi/full/10.1056/CAT.18.0268 (accessed on 14 November 2022).
- Zhang, J.; Hu, Q.; Wang, S.; Tao, J.; Gou, M. Digital light processing based three-dimensional printing for medical applications. Int. J. Bioprinting 2020, 6, 242. [Google Scholar] [CrossRef]
- Berber, B.; Aydin, C.; Kocabas, F.; Guney-Esken, G.; Yilancioglu, K.; Karadag-Alpaslan, M.; Caliseki, M.; Yuce, M.; Demir, S.; Tastan, C. Gene editing and RNAi approaches for COVID-19 diagnostics and therapeutics. Gene Ther. 2021, 28, 290–305. [Google Scholar] [CrossRef]
- Tang, T.-C.; An, B.; Huang, Y.; Vasikaran, S.; Wang, Y.; Jiang, X.; Lu, T.K.; Zhong, C. Materials design by synthetic biology. Nat. Rev. Mater. 2021, 6, 332–350. [Google Scholar] [CrossRef]
- Radivojević, T.; Costello, Z.; Workman, K.; Garcia Martin, H. A machine learning Automated Recommendation Tool for synthetic biology. Nat. Commun. 2020, 11, 4879. [Google Scholar] [CrossRef] [PubMed]
- Takebayashi, T.; Takahashi, K.; Okita, Y.; Kubo, H.; Hachisuka, K.; Domen, K. Impact of the robotic-assistance level on upper extremity function in stroke patients receiving adjunct robotic rehabilitation: Sub-analysis of a randomized clinical trial. J. Neuroeng. Rehabil. 2022, 19, 25. [Google Scholar] [CrossRef] [PubMed]
- Gambhir, S.; Malik, S.K.; Kumar, Y. Role of Soft Computing Approaches in HealthCare Domain: A Mini Review. J. Med. Syst. 2016, 40, 287. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, W.; Rasool, A.; Javed, A.R.; Baker, T.; Jalil, Z. Cyber Security in IoT-Based Cloud Computing: A Comprehensive Survey. Electronics 2021, 11, 16. [Google Scholar] [CrossRef]
- WHO. COVID19 Vaccine Tracker. Available online: https://covid19.trackvaccines.org/agency/who/ (accessed on 25 March 2022).



| Task | Input Data | Primary Models Used in the Study | Comparisons Models Used/Tested | Peformance/Accuracy | Comments/Limitations | References |
|---|---|---|---|---|---|---|
| Antibiotic resistance prediction in patients | 11,496 antimicrobial susceptability datasets from laboratory information system, internal medical ward, public hospital, Greece | stack ensemble (Microsoft Azure AutoML) | VotingEnsemble, MaxAbsScaler, LightGBM, Sparse Normalizer, XGBoost Classifier | AUCW is 0.822 and 0.850 Accuracy rate is 0.770 | Study uses 499 patients’ data. Data scientist is needed at this stage for pre-processing, feature selection, and final analysis. More data training needed to obtain more accurate results. | [107] |
| Prediction of antimicrobial resistance in Acinetobacter baumannii, Mycobacterium tuberculosis, and Streptococcus pneumoniae | K-mer representation of bacterial genomic data | random forest | Adaboost, logistic regression, deep learning | 80–92% | Lower data give less accuracy and higher number of data give elevated accuracy. Not possible to use by laboratories and hospitals having small datasets. | [109] |
| Predicting antibiotic resistance in hospitalized patients | 5590 instance datasets with different variables from general hospital laboratory, Greece | WEKA framework ML, run under Java platform | J48 algorithm, random forest, multinomial logistic regression, kNN algorithm, multilayer perceptron (MLP) | ROC area of 0.758 and accuracy of 75.8% | Low accuracy, limited dataset, and less clinical attributes. Increased dataset can give accurate results. | [110] |
| Prediction of antibiotic resistance in hospitalized patients | 16,000 antibiotic resistance tests of electronic medical record (EMR) | Ensemble of 3 models: Lasso logistic regression, neural networks, gradient boosted trees | Independent algorithms:Lasso logistic regression, neural networks, gradient boosted trees | Combined algorithm: 0.821, Xgb: 0.82, Lasso: 0.82, dnn: 0.803, auROC score was 0.8–0.88 | Bacterial details can increase AUROC score if included, additional information needed to improve accuracy such as antibiotics used prior to admission, microbiome composition, diet, and exercise. | [111] |
| Prediction of antimicrobial resistance from ICU patients in Pseudomonas and Enterococcus, Stenotrophomonas | Dataset of 32,997 collected from health information system of 2630 patients, University Hospital of Fuenlabrada, Spain | Logistic regression, K-nn, decision trees, random forest, multilayer perceptron | AMG: 82.2 ± 1.7, CAR: 79.6 ± 2.1, CF4: 74.9 ± 2.1, PAP: 77.1 ± 1.7, POL: 68.5 ± 7.0, QUI: 88.1 ± 1.6 | Accuracy differs based on various antibiotics and bacterial species; upgradation yet to undergo based on mechnical ventilation, bed; sepsis patients in ICU are to be considered. | [106] | |
| Prediction of Carbapenem-resistant Klebsiella pneumoniae | 46 Carbapenem-resistant Klebsiella pneumonia (CRKP) isolated from hospital patients | random forest | Logistic regression, Naïve Bayes, nearest neighbors, support vector machine | Accuracy: 97%, Carbapenem-resistant identification: 93% | Small sample size and limited data of CRKP. | [112] |
| Prediction of antimicrobial phenotype resistance of Staphylococcus aureus | K-mer representation of bacterial genomic data | Random forest, SVM, XGBoost | Among 10 anitibiotics used in the study, AUC was recorded between 82.02% for Linezolid and 96.13 for vancomycin. Cefoxitin registers AUC of 92.65% with sensitivity of 94% and major error 6.82% | Study uses 466 whole genome sequencing results to predict the antimicrobial resistance. | [108] |
| AI Models | Description of the Model | Accuracy | Image Dataset Sources | Reference |
|---|---|---|---|---|
| Hybrid deep transfer model | This model consists of SVM, decision tree, and accurate methods to detect face masks. | Achieved 99.64% on test | RMFD: real-world masked face dataset, SMFD: stimulated masked face dataset, random face in crowd | [135] |
| Inceptionv3 CNN | This module consists of 22 layers deep from GoogleNet to increase accuracy; this CNN detects persons without a mask. | Achieved 99.9% accuracy | Simulated masked face dataset was used in this study | [136] |
| Facemasknet model | This model uses DL model to identify masked face, properly masked face, and no mask face. | This model achieved 98.6% accuracy | Datasets used in this study are MFDD: masked face detection dataset, RWFCD: real-world face recognition dataset, SMFRD: simulated masked face recognition dataset | [137] |
| EfficientNet model | CNN-driven EfficientNet architecture is applied in this method and can be used for real-time detection of mask. | Achieved accuracy of 97.12% | Openly available face mask detection dataset | [138] |
| Deep learning model vgg16 | Trained with 2 datasets, it works well with medium and small datasets. | Accuracy of 96.50% was achieved | Two datasets with 1484 and 7200 images are used | [139] |
| DTLMV2 (deep transfer learning MobileNetV2) | This model uses a lightweight CNN which requires less computing power and is easily attached to computer vision and mobile system. | This model gains accuracy of 97.01% at validation data and 98% accuracy on training data | Crowd dataset with 7514 images is used | [140] |
| Deep Masknet framework | This model uses both face mask detection and masked facial recognition. | Obtained 100% accuracy on face mask detection and 93.33% of masked facial recognition | Mask detection and masked facial recognition (MDMFR) dataset | [141] |
| Type of Mask | Material Used | Filtering Size/Efficiency% | Characteristics | Reference |
|---|---|---|---|---|
| ML algorithm-based respirator | Elastic fiber membrane (EFM) | Up to 2.5 µm | Controlled mechanical stretching and relaxation of filter, response to slow and fast physical activity, cheaper material. | [144] |
| N95 | Electrostatic non-woven polypropylene | Up to 0.3 µm | High filtering efficiency of virus/bacteria, prevents air and droplets penetrating the edges. | [149] |
| Mask with nanofiber filter | Polyvinylidene fluorid (PVDF30) and (PVDF20) | ≤2.7 μm | Captures up to 99.9% of coronavirus aerosol. | [147] |
| Nanoparticles-coated non-woven surgical mask | Copper nanoparticles (CuNPs) | 99.37% | Photocatalytic and photothermal properties, reusable, self-cleaning ability, disposable. | [150] |
| Surgical mask with graphine | Melt-blown non-woven fabrics (MNF) and graphene layer with electric and thermal conductivity | 99.8% | Removal of virus/microbes by electrothermal method, high removal efficiency up to 10 cycles. | [151] |
| Nanofiber mask | Piezoelectric electrospun poly (l-lactic acid) (PLLA) nanofibers | >99% | Respiration electrifies the mask, long stable filter is humidity resistant, autoclavable, and degradable around 50 days. | [152] |
| N95 respirator nanofiber mask with copper nanoparticles | Nylon polymer fiber with copper nanoparticles | - | Antiviral and antimicrobial properties. | [153] |
| 3D-printed nanofiber-based mask | Combined silver and copper nanoparticles | 99.5% | Antimicrobial, high polarity, heat-insulating properties. | [153] |
| Nanoporous hard mask | Flexible silica fabricated with reactive ion etching process on polymeric membrane | Up to 5 nm | Reusable, high filtration efficiency, hydrophobic, antifouling, self-cleaning. | [154] |
| Country | Law/Regulation | Purpose of This Law | Date Effective |
|---|---|---|---|
| USA | No Vaccine Passports Act [168] | Relaxing the restrictions of forcing vaccine certificate | 4 August 2021 |
| The Netherlands (Red Cross) | 510 Data Responsibility Policy [169] | Data protection | 12 November 2018 |
| United Kingdom | Contact-tracing app (General Data Protection Regulation (UK GDPR) and Data Protection Act (DPA) 2018) [170] | Data protection and digital COVID tracking app | May 2020 |
| European Union | General Data Protection Regulation [171] | Data protection of public and health records | 27 April 2016 |
| European Union | Medical Devices Regulations 2017/745 (MDR) [172] | Protection of patients from medical device, protection of produced data using this device | 5 April 2017 |
| European Union | The 2017/746 In Vitro Diagnostic Medical Devices Regulation (IVDR) [173] | Protection of patient health and users, quality and safety of in vitro medical devices | 25 January 2022 |
| European Union | Regulation of the European Parliament and of the council laying down harmonized rules on AI (Artificial Intelligence ACT) and amending certain union legislative acts [174] | Facilitate and creating innovation in AI, creating trusted AI applications | 21 April 2021 |
| European Union | Civil Law Rules on Robotics [175] | Implementation of AI robotics | 16 February 2017 |
| Singapore | Personal Data Protection Act 2012 [176] | Data protection | 31 December 2021 |
| Australia | Therapeutic Goods (Medical Devices) Regulations 2002 [177] | Clinical decision support software | 25 February 2021 |
| China | Notice of the State Council Issuing the New Generation of Artificial Intelligence Development Plan. State Council Document. No. 35. 2017 [178] | Healthcare and management | 8 July 2017 |
| Kingdom of Saudi Arabia | Guidance on Software as a Medical Device/SFDA MDS-G23 [179] | AI- and BigData-based medical software to diagnose and predict patient conditions | 27 April 2021 |
| Russia | Development of AI in healthcare up to 2030, approved on 10 October 2019, No. 490 [180] | Software as medical device in healthcare | 10 October 2019 |
| South Korea | Medical Devices Act No. 15945, 11 December 2018 [181] | Software as medical device in healthcare | 11 December 2008 |
| Singapore | Standalone Medical Mobile Applications (SaMD) and Qualification of Clinical Decision Support Software (CDSS) [182] | Clinical decision support software | 19 July 2021 |
| China | Cybersecurity Law of the People’s Republic of China [183] | To preserve cyberspace sovereignty and national security | 7 November 2016 |
| Malaysia | Medical Device Act 737-2012 [184] | Medical device, software regulation in healthcare | 30 January 2012 |
| Emirate of Abu Dhabi | Artificial Intelligence (AI) in the Healthcare Sector of the Emirate of Abu Dhabi, Policy/AI/0.9, Version 0.9 [185] | Health system monitors, analysis, and public health observation | 30 April 2018 |
| Canada | Digital Charter Implementation Act, 2022 (Bill C-27) [186] | Protection of personal information, data, and health records, along with any serious direct cause to patients by AI | 16 June 2022 |
| Brazil | LGPD–General Personal Data Protection Law (Federal Law no. 13,709/2018) [187] | AI regulation in health sector of Brazil | 14 August 2018 |
| Brazil | Brazilian Artificial Intelligence Bill (Bill No. 21/2020) [188] | Development and applying of AI in various sectors of Brazil | 29 September 2021 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mahalakshmi, V.; Balobaid, A.; Kanisha, B.; Sasirekha, R.; Ramkumar Raja, M. Artificial Intelligence: A Next-Level Approach in Confronting the COVID-19 Pandemic. Healthcare 2023, 11, 854. https://doi.org/10.3390/healthcare11060854
Mahalakshmi V, Balobaid A, Kanisha B, Sasirekha R, Ramkumar Raja M. Artificial Intelligence: A Next-Level Approach in Confronting the COVID-19 Pandemic. Healthcare. 2023; 11(6):854. https://doi.org/10.3390/healthcare11060854
Chicago/Turabian StyleMahalakshmi, V., Awatef Balobaid, B. Kanisha, R. Sasirekha, and M. Ramkumar Raja. 2023. "Artificial Intelligence: A Next-Level Approach in Confronting the COVID-19 Pandemic" Healthcare 11, no. 6: 854. https://doi.org/10.3390/healthcare11060854
APA StyleMahalakshmi, V., Balobaid, A., Kanisha, B., Sasirekha, R., & Ramkumar Raja, M. (2023). Artificial Intelligence: A Next-Level Approach in Confronting the COVID-19 Pandemic. Healthcare, 11(6), 854. https://doi.org/10.3390/healthcare11060854






