A Two-Stage De-Identification Process for Privacy-Preserving Medical Image Analysis
Abstract
:1. Introduction
2. Related Work
2.1. DICOM Header De-Identification Techniques
2.2. Pixel-Level De-Identification in DICOM Images
3. Material and Methods
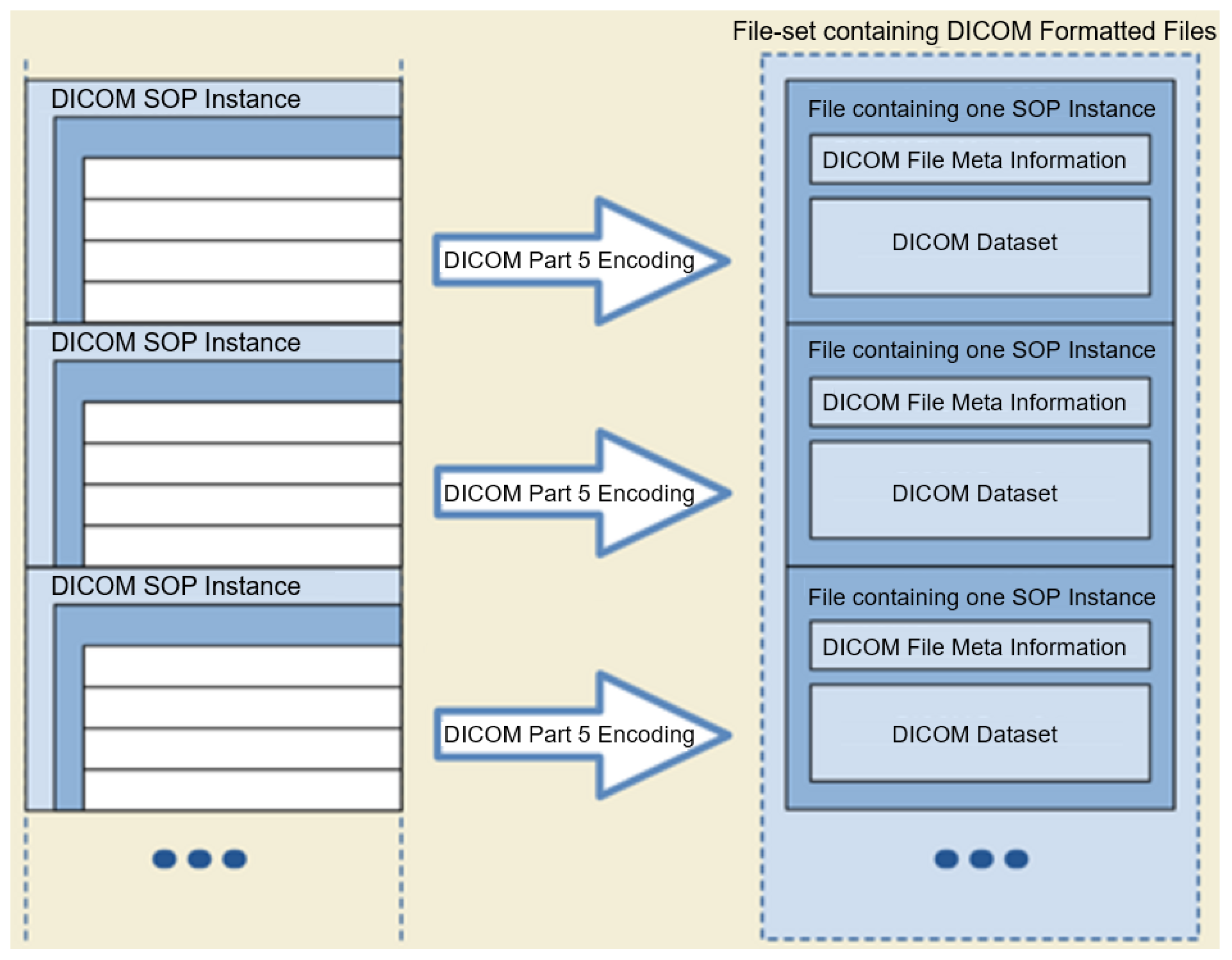
- DICOM (Digital Imaging and Communication On Medicine): is an international standard in medical domain and covers all imaging modalities and organs [9]. DICOM is supported by almost all vendors of clinical imaging systems.
- ANALYZE: This format is widely used in the functional neuroimaging field [31]. Each data item consists of two files: (1) a file with contains the actual data in a binary format with the filename extension .img, and (2) a header file with filename extension .hdr and it holds information about the data, such as voxel size and the number of voxels in each dimension. This format is progressively replaced by NIfTI.
- NIfTI (Neuroimaging Informatics Technology Initiative): This is a format for neuroimaging developed in the early 2000s by the DFWG (Data Format Working Group) to improve the ANALYZE format [32]. The initial version of NIfTI, that is NIfTI-1, combines header and image sections. Extensions of NIfTI provide flexibility to store additional information such as experimental design and data acquisition details. More recently, the Brain Imaging Data Structure (BIDS) (Brain Imaging Data Structure, https://bids.neuroimaging.io/, accessed on: 4 April 2022) format is rapidly replacing NIfTI.
3.1. DICOM Image Processing
- Tag: Each DICOM element in a DICOM file has a tag, which uniquely defines that element. A DICOM tag consists of 2 main parts: (1) Group number (2) Element number. For instance, an element with the Tag (0028, 0010) belongs to the Image group 0028 (i.e., attributes that describe the image and its properties). The element number 0010 corresponds to the number of rows in the image. Similarly, under the same group, element number 0011 refers to the number of columns in an image. In the case of the dataset used in this paper, both the number of rows and columns are 512.
- Value Representation (VR): A VR is a two-character code that defines an element’s type. There are a variety of data types in DICOM. For instance, UI refers to a Unique Identifier data type, and SH refers to a Short String data type.
- Value Length: This shows the length of a DICOM element. For example, the length of the value of the Patient Sex element (i.e., ‘F’ for female, ‘M’ for male, and ‘O’ for other) is 2 (note the trailing space which is mandated by the VR for this element).
- Value Field: This holds the actual value of the element. Values with VRs, consisted of character strings (except for the UI VRs), need to be padded with a space character when necessary to achieve even length. On the other hand, values with a VR of UI should be padded with a single trailing NULL character when necessary to achieve even length. Finally, values with a VR of OB (i.e., Other Byte String) also need to be padded with a trailing NULL to ensure even length. During de-identification or any type of value manipulation, these rules need to be followed.
- Type 1: Attributes under this category must be present and have a valid value (depending on the VR of the element).
- Type 2: Attributes under this category must be present, however, they may contain the value of “unknown”, or simply a zero-length value (unlike Type 1 attributes).
- Type 3: Attributes under this type are optional. This means that they may or may not be included in the DICOM header and if present, their value could be of zero length. Removing such attributes is not a violation of the DICOM standard.
- Type 1C: The letter C stands for Conditional. Attributes under this category cannot be simply removed as their existence is conditional. If the condition is met, then the attribute shall be treated as a Type 1 attribute (i.e., their existence is required and cannot have a zero-length value). If the condition is not met, these attributes will not be included in the dataset. They cannot be simply de-identified without investigating the condition, since it is indeed a DICOM protocol violation if the specified conditions are met and a Type 1C attribute is not included.
- Type 2C: Similar to Type 1C attributes, the existence of the attributes under this category is conditional. If the specified conditions are met, then the attribute is treated as a Type 2 attribute. If the specified condition is not met, the attribute may not be included in the dataset. Such an attribute cannot be simply de-identified without investigating the required conditions that dictate their existence, and it is indeed a DICOM protocol violation if the specified conditions are met and a Type 2C attribute is not included due to removal.
3.2. DICOM De-Identification Considerations
- Clear: The attribute will remain; however, its value will be cleared to an empty string of length 0 (the VR of the attribute must be consulted).
- Overwrite: The attribute will remain; however, its value will be overwritten with a word such as ‘CONFIDENTIAL’ or a randomly assigned value (the VR of the attribute must be consulted).
- Remove: The entire attribute will be removed from the DICOM file (the attribute type must be consulted).
- UID/ID: DICOM uses UIDs to make possible the communication between multiple application entities over a PACS system. The DICOM standard uses certain UID’s so that a sender and a receiver node in a PACS system can encode/decode the transferred DICOM files (i.e., they can talk DICOM). For instance, the sender can use a certain value for the Transfer Syntax UID to inform the receiver of the compression algorithm used to compress the data by the sender, so that the receiver can successfully decompress the data upon receiving it. Such UIDs take publicly available standard values. There is a variety of services available in DICOM. For instance, the standard SOP Classes and their UID classes for Storage services can be found in section B.5 of the DICOM standard (Standard SOP Classes, https://dicom.nema.org/dicom/2013/output/chtml/part04/sect_B.5.html, accessed on: 4 April 2022). As mentioned in the previous section, some of the UIDs and IDs are also used to maintain links between the patient, study, series, and DICOM instances.Among all of the different UIDs in the collected DICOM dataset, it has been confirmed that it is only the instance UIDs that bear a potential risk of disclosing the patient’s identity. Because other types of UID within this dataset (e.g., SOP Class UIDs, Transfer Syntax UID, Irradiation Event UID, Frame of Reference UID, etc.) are either about the method of data transmission over a PACS network, method of data storage, or simply keeping track of the frame of reference in a given series. For instance, the SOP Class UID definition (globally known and unique) contains the rules and semantics which may restrict the use of certain services for the current data at hand. For example, the SOP Class UID of 1.2.840.10008.5.1.4.1.1.2 refers to the CT Image Storage service, and the SOP Class UID of 1.2.840.10008.1.1 refers to Verification SOP Class (i.e., verification of basic connectivity between a sender and receiver over a PACS network). For a given DICOM file, these Class UIDs have nothing to do with the data within the file and only describe the type of service that is required for that file.While a Class UID is similar to a governing template with no actual patient-related data, Instance UIDs have a direct connection with the actual data within a DICOM file. For instance, for a given DICOM file, Study Instance UID and Series Instance UID can directly and uniquely point to which exact study, and within that study, to which series that particular DICOM file belongs.Patient ID and Study ID need to be de-identified to protect the identity of the patient. It is important to note that in the dataset used in this paper, it was found that Class UIDs (as opposed to Instance UIDs) were not de-identified by the export process withing the hospital since they do not disclose anything about the patient’s identity. On the other hand, all Instance UIDs were de-identified by the export process.
- Dates/Times: There are many date and time attributes in a DICOM header and it is customary to replace their values with some randomly generated date and time strings.
- Sensitive names: There are names such as; Station Name, Patient Name, Physician’s Name, Radiologist’s Name, Institution Name/Address, that require de-identification.
- Sensitive numbers: There are certain numbers that one might need to be de-identified. One such attribute, Accession Number, is a RIS (Radiological Information System) generated number that identifies the order for the Study. This value could be used to reconstruct the identity of the patient [16].
4. De-Idenfication Process
- Different regulations and conditions appear in different authorities such as GDPR and HIPAA, complicating the development of international standards for de-identification in medical imaging data [37].
- De-identification techniques are usually employed in the preparation stages for data transfer or sharing [38]. In cases where the patient withdraws their consent, it detaches data governance from data ownership (hindering the right to be forgotten, GDPR article 17 [39]). In addition to this, if the legislation changes, the new data that will be generated from the original data should not include those patients data. In this case, the sharing protocol should be clear of ambiguity and defined to satisfy the data owners requirements.
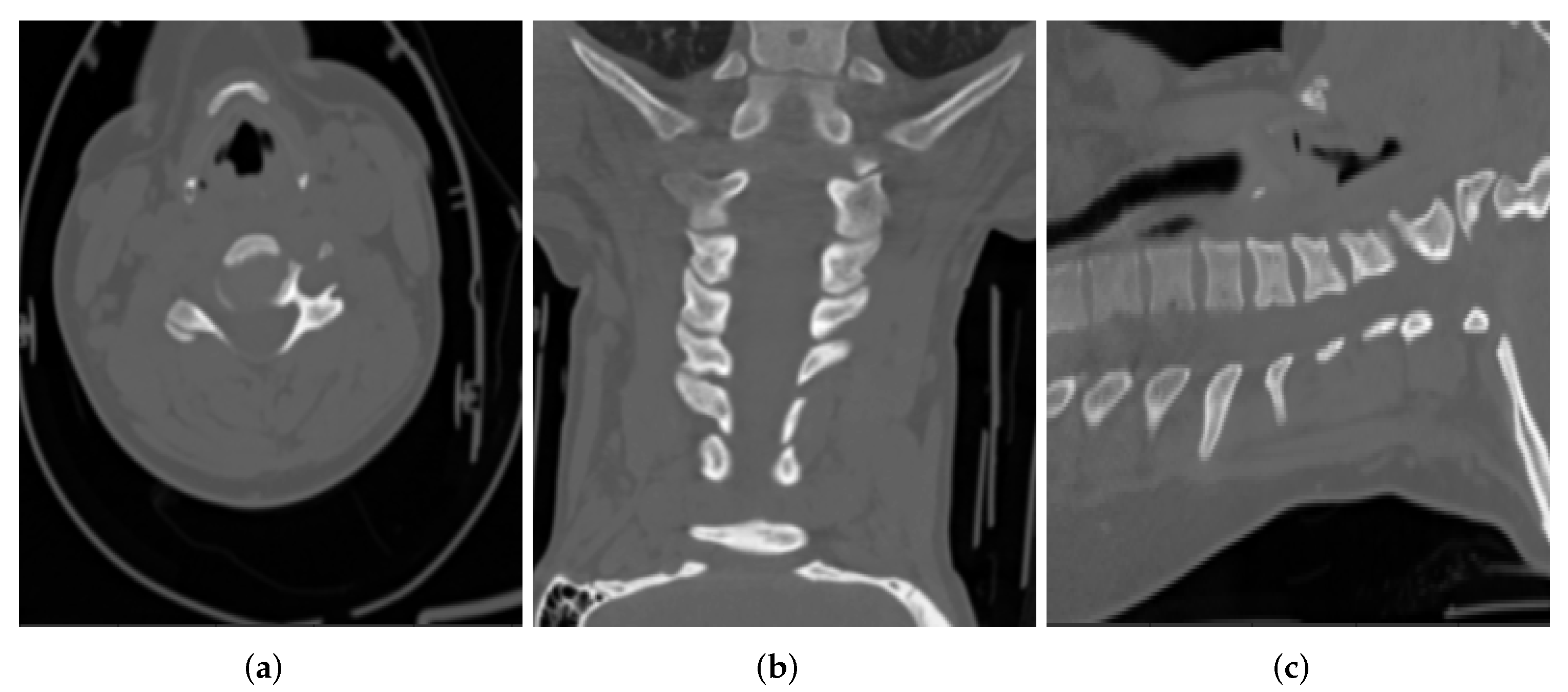
- Requirements of the de-identification process vary according to the type of datasets [40]. For example, it is harder to link a radiographic image of a section of the leg back to an individual, compared to a computed tomography scan of their head, where the contours of the face can be reconstructed directly from the image.
4.1. Stage 1: De-Identification at Source (NIMIS)
- The following Instance UIDs are changed while preserving the relationship in the DICOM data model:
- –
- Media Storage SOP Instance UID: Overwritten
- –
- SOP Instance UID: Overwritten
- –
- Study Instance UID: Overwritten
- –
- Series Instance UID: Overwritten
- Station Name: Overwritten as CONFIDENTIAL
- Source Application Entity Title: Overwritten as CONFIDENTIAL
- The following IDs have been overwritten:
- –
- Patient ID: Overwritten as !T000 for all patients.
- –
- Study ID: Overwritten as CONFIDENTIAL
- –
- Scheduled Procedure Step ID: Overwritten as CONFIDENTIAL
- Sensitive numbers:
- –
- Accession Number: Overwritten as CONFIDENTIAL
- Obvious Patient information attributes:
- –
- Patient’s Name: Overwritten as CONFIDENTIAL
- –
- Patient’s Birth Date: Cleared
- –
- Patient’s Sex: Cleared
- –
- Patient’s Age: Cleared
- Dates and Times:
- –
- Study Date: Overwritten with a random date
- –
- Series Date: Overwritten with a random date
- –
- Acquisition Date: Overwritten with a random date
- –
- Content Date: Overwritten with a random date
- –
- Acquisition DateTime: Overwritten with a random date and time
- –
- Study Time: Overwritten with a random time
- –
- Series Time: Overwritten with a random time
- –
- Acquisition Time: Overwritten with a random time
- –
- Content Time: Overwritten with a random time
4.2. Stage 2: Validation and Further De-Identification of DICOM
- Private attributes:
- –
- Private Creator: Removed
- –
- [Anonymization Status]: Removed
- Sequential attributes:
- –
- Request Attributes Sequence: Removed because one of its sub-attributes (Requested Procedure ID) is determined as NOT SAFE by the clinical experts. It would be worthwhile to note that the reason that the Requested Procedure ID attribute is not directly removed is that it is a Type 1C attribute, whose existence is conditioned over the existence of the Request Attributes Sequence, which is a Type 3 attribute and can be safely removed.
- Dates and Times:
- –
- Date of Last Calibration: Overwritten with a random date
- –
- Time of Last Calibration: Overwritten with a random time
- Names:
- –
- Institution Name: Cleared
- –
- Institution Address: Cleared
- –
- Referring Physician’s Name: Cleared
- Comments:
- –
- Image Comments: Cleared
- UIDs:
- –
- Irradiation Event UID: Removed
- –
- Private Information Creator UID (and Private Information as a result of this): Removed
5. Discussion
- It is advisable and often required to de-identify data at the source site (e.g., the hospital where the data are generated) before moving it off-site.
- Apart from DICOM header de-identification, there is a need for handling and validating the de-identification of burned-in pixel annotations as well as the facial features, where possible. In particular, it should be noted that in certain applications of Artificial Intelligence (AI) in radiation therapy, radiography and head and neck scans, the facial feature information might be necessary. For the current dataset, it has been confirmed that there are no facial features recorded in the DICOM images.
- There is a need to actively evaluate the tools’ compliance with DICOM standards by validating conformance to DICOM Application-Level Confidentiality Profile Attributes (ALCPA) (DICOM Application-Level Confidentiality Profile Attributes: https://dicom.nema.org/dicom/2013/output/chtml/part15/chapter_E.html#table_E.1-1, accessed on: 4 April 2022.).
- One should define concrete privacy preservation requirements for the specific use case and data at hand.
- One should ensure traceability and compliance audit by keeping a record of software, version, affected data portion, results, etc. for every de-identification event.
- Fine-grained de-identification details: It is essential for the credibility of any DICOM de-identification tools to provide a detailed explanation of the employed methods for DICOM de-identification. However, the existing literature lacks fine-grained de-identification details, which makes it difficult for the readers to elicit the best practices for de-identifying DICOM files. In this paper, an exhaustive attribute-level two-stage DICOM de-identification process has been described in a step by step fashion. In addition, the best practices of DICOM header de-identification have been described in order to ensure compatibility with the DICOM standard. The considerations outlined for DICOM de-identification in this paper such as the importance of considering attribute type and value representation will provide a guideline for clinical experts and researchers towards developing tools that ensure patients’ privacy.
- Conforming to the DICOM standard: Although most of the existing techniques claim to comply with the DICOM standard, the lack of sufficient details regarding their proposed de-identification methods, makes it difficult to ensure their DICOM standard compatibility. As a result, one can only hope that these approaches will abide by the DICOM standard and can be safely deployed in a DICOM-driven PACS network. However, this work has provided a highly detailed de-identification process in order to ensure the reader that the proposed method fully conforms to the DICOM standard and is deployable in a PACS network.
- Validation with expert radiologists: It is essential to consult and validate the de-identification of PII in DICOM file headers with expert clinical radiologists to ensure that the de-identification process will not jeopardise the diagnosis process by the radiologists. The validation of the de-identification methods by a domain expert can further guarantee the deployability of such methods in a real-life production environment. During the development of the proposed two-stage de-identification approach, multiple workshops have been held with expert radiologists to validate DICOM compatibility and as well as the integrity of the de-identified DICOM files. To the best of our knowledge, the state-of-the-art de-identification approaches do not incorporate this step. In the future work, it is hoped that consultation and validation with domain experts will become common practice in the future, especially in medical applications where the proposed methods have a direct impact on patient’s well-being.
- Open source: In order to ensure reproducibility, it is crucial to always share the underlying code for the developed methods, and this becomes even more important in medical applications. The de-identification tool developed in this paper is open-source and publicly available on [36].
- Deployment in production environments: Not all of the de-identification tools have been developed and tested by deploying them in an actual production environment. The proposed method has now been rigorously tested and fully deployed in the Medical Imaging Ireland (Med-I) platform.
- When it comes to DICOM de-identification, one must determine whether de-identifying the DICOM header attributes would suffice or a pixel-level de-identification is also required.
- Upon de-identifying any DICOM header attributes, one should constantly prevent violating the DICOM standard by checking the (1) type and (2) value representation of the attributes to be de-identified. Any violation could result in DICOM incompatibility and potential issues in a PACS network during the deployment phase.
- When developing DICOM de-identification tools, one should always be aware of the fact that, depending on the scanner machine, different DICOM attributes might be used in the generated DICOM files. As a result, it is a good practice to ensure that the developed de-identification method is both standalone and independent of the imaging modality in order to ensure the generalisability of these methods across multiple machines.
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ML | Machine Learning |
| AI | Artificial Intelligence |
| ALCPA | Application-Level Confidentiality Profile Attributes |
| HSE | Health Service Executive |
| PACS | Picture Archiving and Communication System |
| NIMIS | National Integrated Medical Imaging System |
| DICOM | Digital Imaging and Communications in Medicine |
| CT | Computed Tomography |
| PII | Personally Identifiable Information |
| PHI | Protected Health Information |
| GDPR | General Data Protection Regulation |
| UID | Unique Identification Number |
| NIfTI | Neuroimaging Informatics Technology Initiative |
| MINC | Medical Imaging NetCDF (Network Common Data Form) |
| MRI | Magnetic Resonance Imaging |
Appendix A. Example DICOM Header before and after De-Identification by the Clinical Team
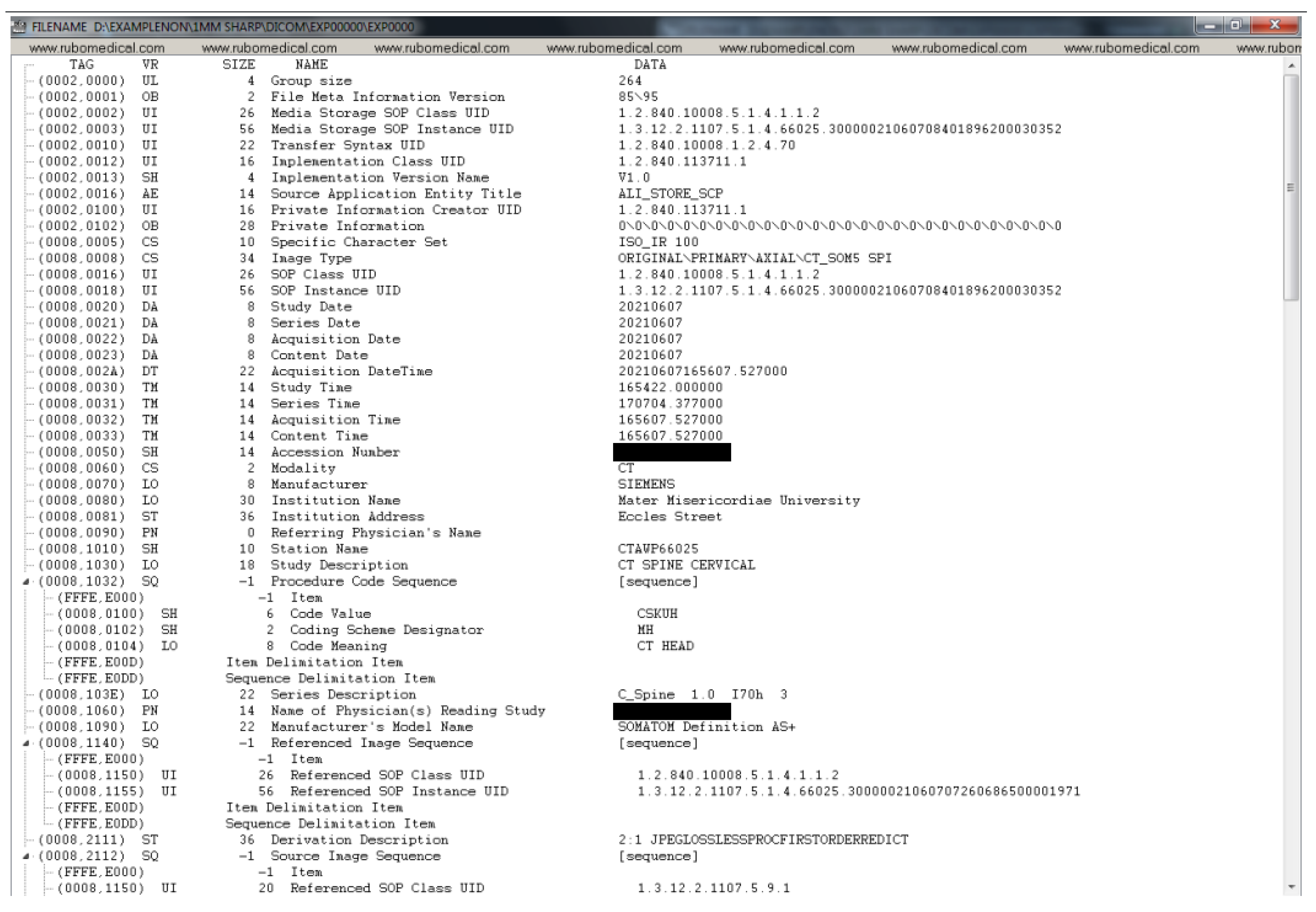
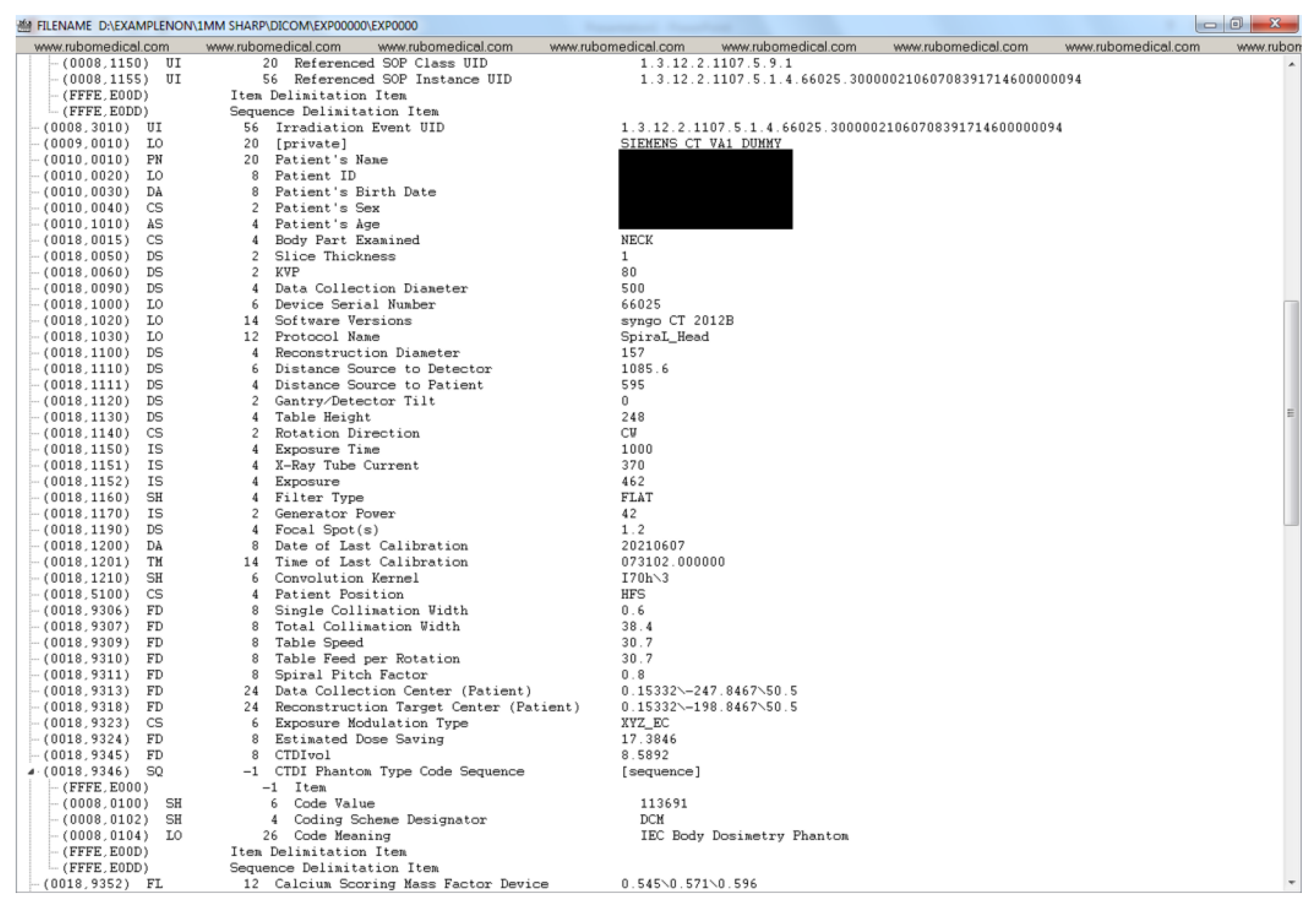
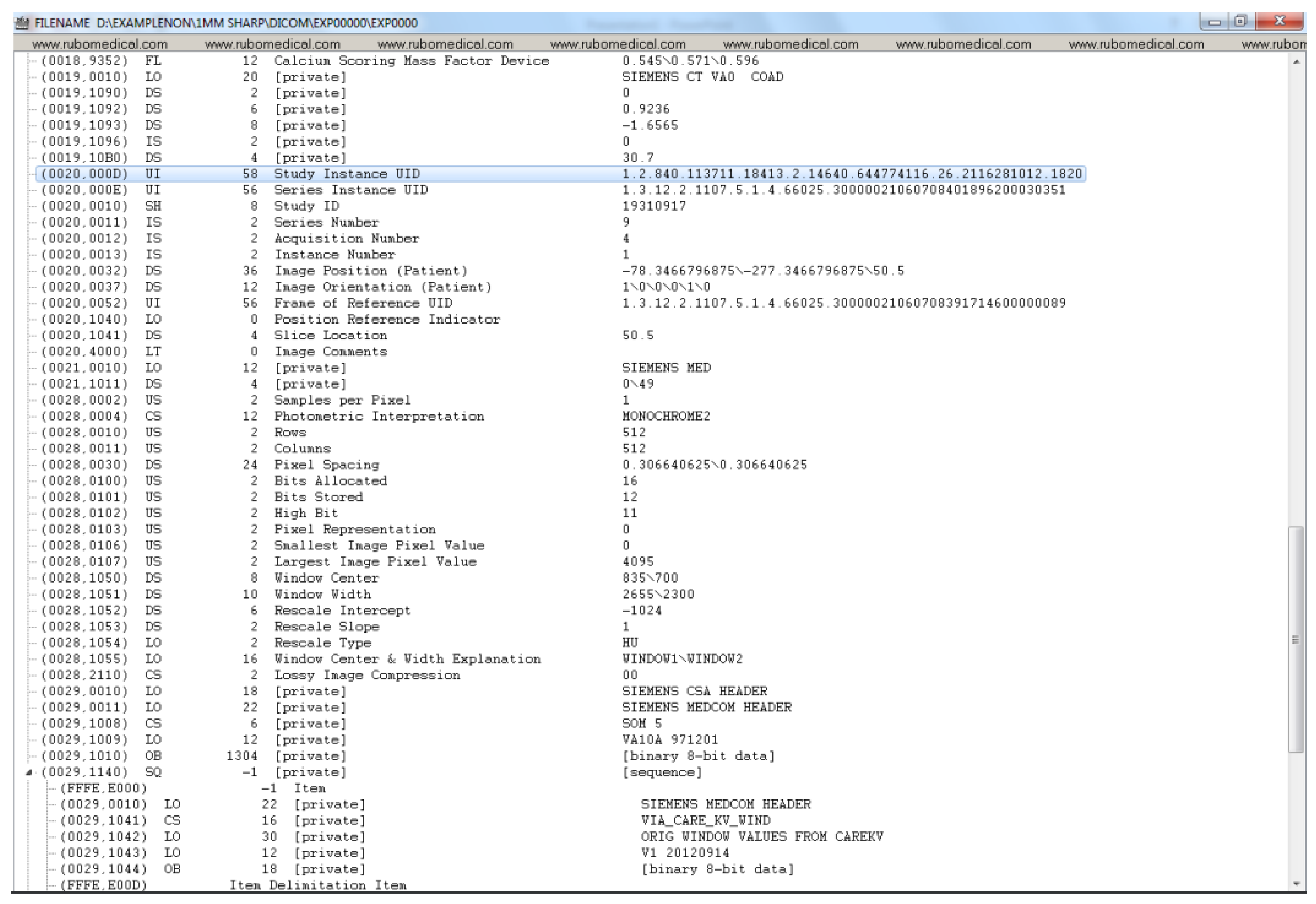
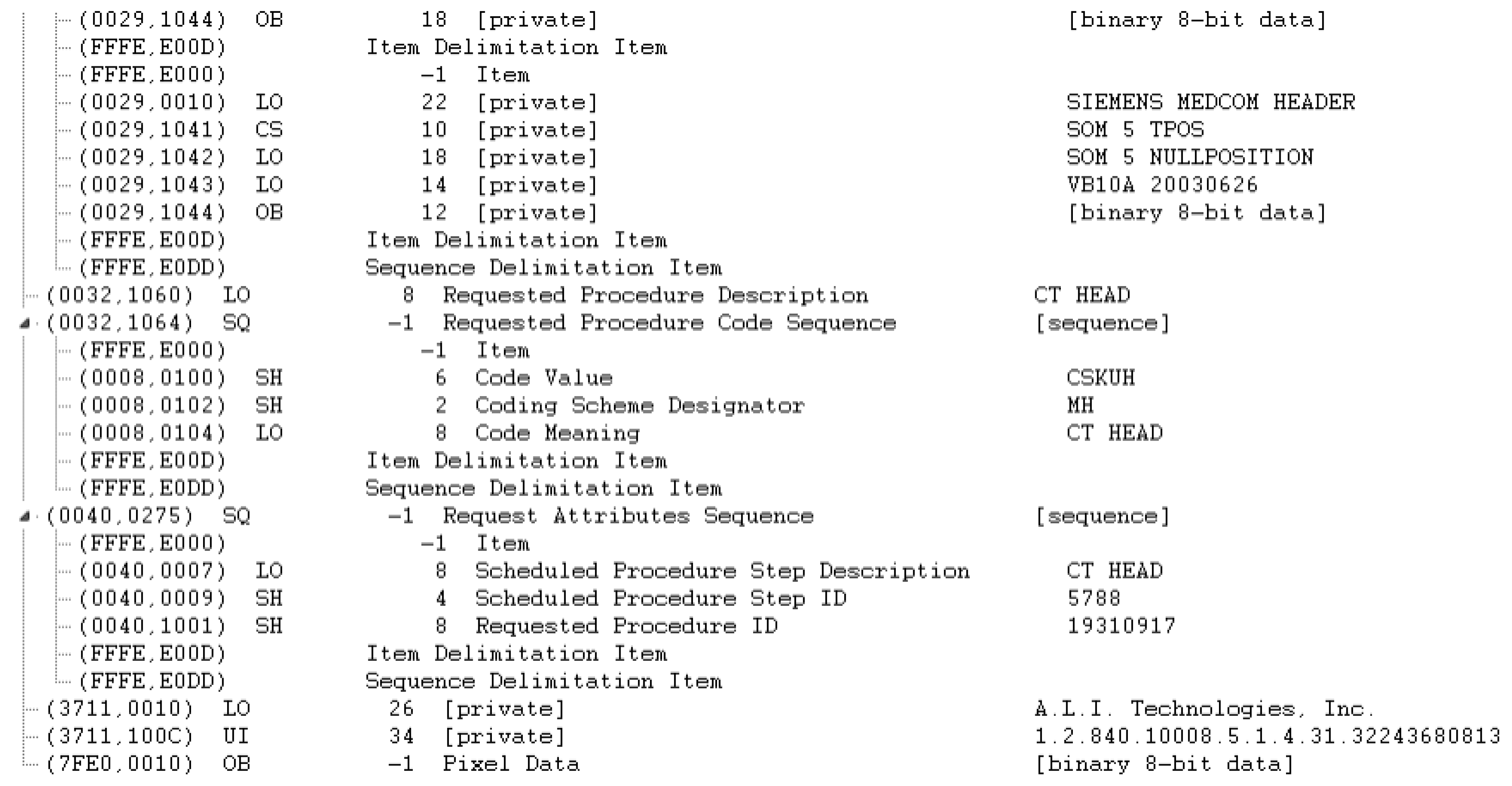
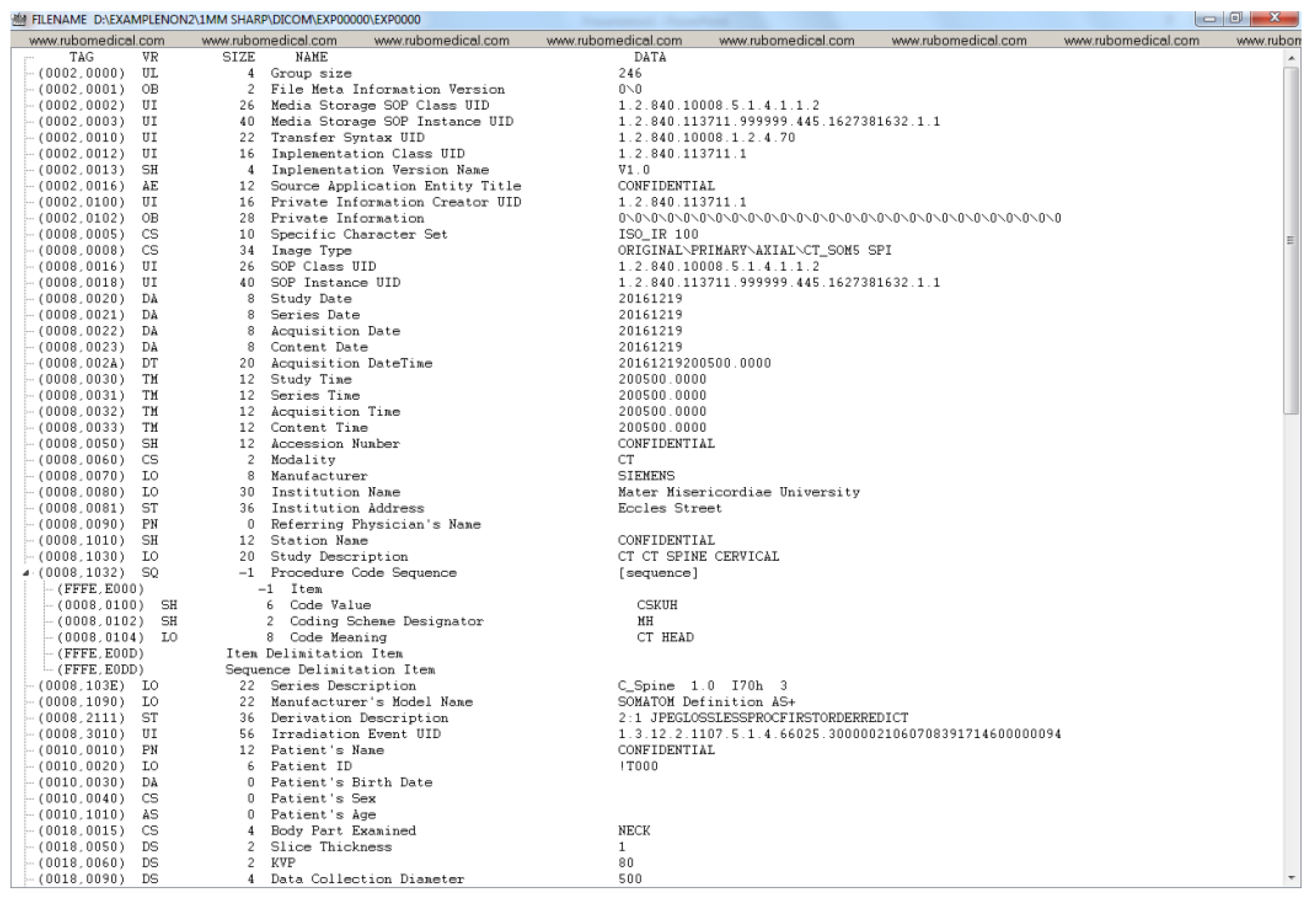

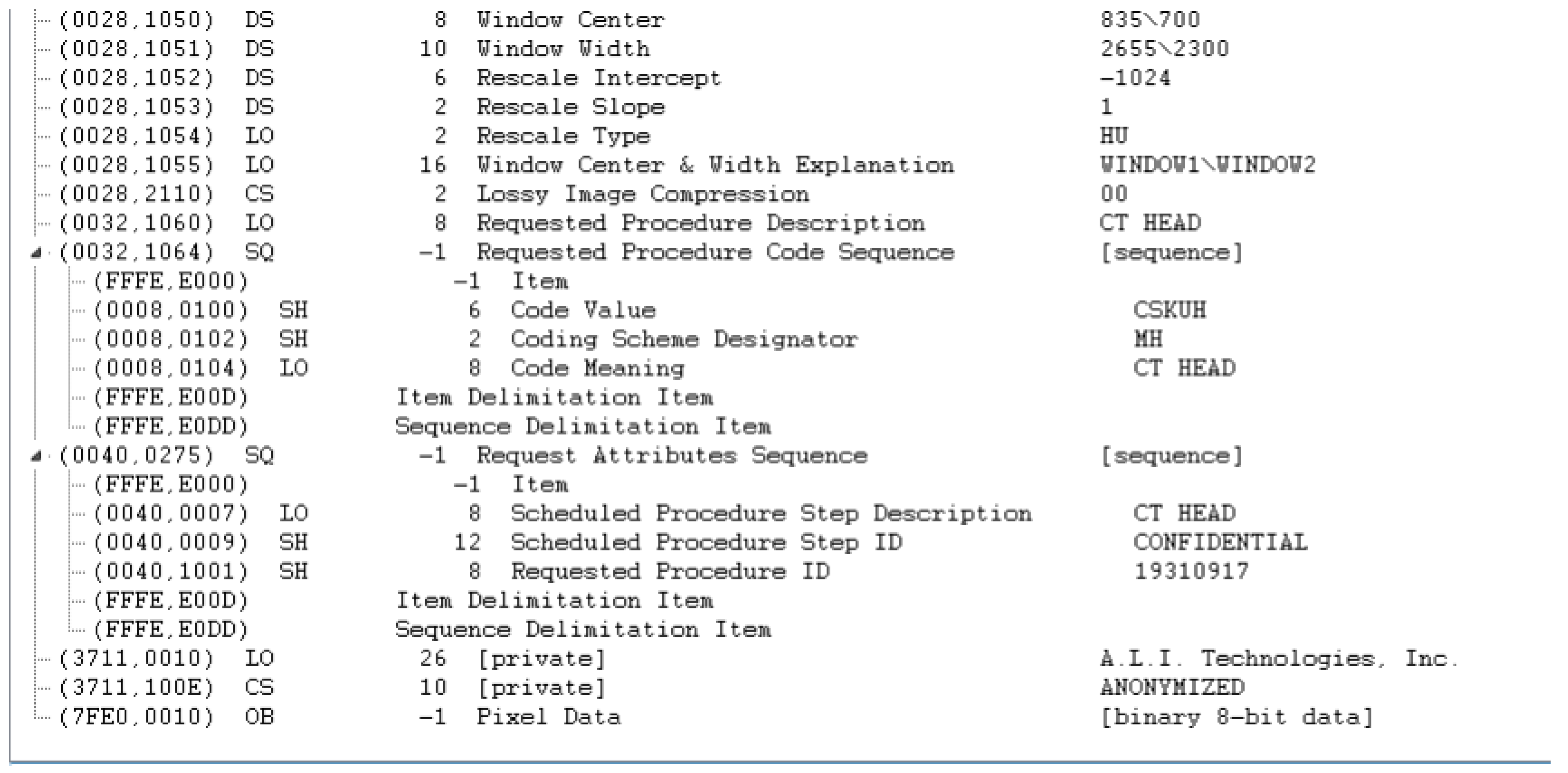
Appendix B. Detailed Examination of DICOM Headers from an Expert Radiologist Perspective
| Name | Value | De-Identification | Value Across | Value Across |
|---|---|---|---|---|
| Status | All Patients | a Given Patient | ||
| File Meta Information Group Length | 246 | SAFE | ||
| File Meta Information Version | b’\x00\x01’ | SAFE | ||
| Media Storage SOP Class UID | 1.2.840.10008.5.1.4.1.1.2 | SAFE | same | same |
| Media Storage SOP Instance UID | 1.2.840.113711.999999.255.1620909105.1.1 | SAFE | different | different |
| Transfer Syntax UID | 1.2.840.10008.1.2.4.70 | SAFE | same | same |
| Implementation Class UID | 1.2.840.113711.1 | SAFE | same | same |
| Implementation Version Name | V1.0 | SAFE | ||
| Source Application Entity Title | CONFIDENTIAL | de-identified by | ||
| Export Process | ||||
| Private Information Creator UID | 1.2.840.113711.1 | SAFE | same | same |
| Private Information | b’ALI DICOM OEM version 3.7.21’ | SAFE | ||
| Specific Character Set | ISO_IR 100 | SAFE | ||
| Image Type | ORIGINAL; PRIMARY; | SAFE | ||
| AXIAL; CT_SOM5 SPI | ||||
| SOP Class UID | 1.2.840.10008.5.1.4.1.1.2 | SAFE | same | same |
| SOP Instance UID | 1.2.840.113711.999999.255.1620909105.1.1 | SAFE | different | different |
| Series Time | 164,557 | RANDOMIZED by | ||
| Export Process | ||||
| Acquisition Time | 164,557 | RANDOMIZED by | ||
| Export Process | ||||
| Content Time | 164,557 | RANDOMIZED by | ||
| Export Process | ||||
| Accession Number | CONFIDENTIAL | de-identified by | ||
| Export Process | ||||
| Modality | CT | SAFE | ||
| Manufacturer | SIEMENS | SAFE | ||
| Institution Name | Mater Misericordiae University | NOT SAFE | ||
| Institution Address | Eccles Street Hospital Dublin IE | NOT SAFE | ||
| Referring Physician’s Name | de-identified by | |||
| Export Process | ||||
| Station Name | CONFIDENTIAL | de-identified by | ||
| Export Process | ||||
| Name | Value | de-identification | Value Across | Value Across |
| Status | All Patients | a Given Patient | ||
| Study Description | CT CT SPINE CERVICAL | SAFE | ||
| Procedure Code Sequence | Code Value: CSKUH—Coding | NOT SAFE | different | |
| Scheme Designator: MH—Code | same | |||
| Meaning: CT HEAD | ||||
| Series Description | C_Spine 1.0 I30s 3 | SAFE | ||
| Manufacturer’s Model Name | SOMATOM Definition AS+ | SAFE | ||
| Derivation Description | 2:1 JPEGLOSSLESS | SAFE | ||
| PROCFIRSTORDERREDICT |
| Irradiation Event UID | 1.3.12.2.1107.5.1.4.66025. | NOT SAFE | different | same |
| 3000002105071355105980 | ||||
| Patient’s Name | CONFIDENTIAL | de-identified by Export Process | ||
| Patient ID | !T000 | de-identified by Export Process | ||
| Patient’s Birth Date | de-identified by Export Process | |||
| Patient’s Sex | de-identified by Export Process | |||
| Patient’s Age | de-identified by Export Process | |||
| Body Part Examined | NECK | SAFE | ||
| Slice Thickness | 1 | SAFE | ||
| KVP | 100 | SAFE | ||
| Data Collection Diameter | 500 | SAFE | ||
| Device Serial Number | CONFIDENTIAL | de-identified by Export Process | ||
| Software Versions | syngo CT 2012B | SAFE | ||
| Protocol Name | SpiraL_Head | SAFE | ||
| Reconstruction Diameter | 146 | SAFE | ||
| Distance Source to Detector | 1085.6 | SAFE | ||
| Distance Source to Patient | 595 | SAFE | ||
| Gantry/Detector Tilt | 0 | SAFE | ||
| Table Height | 193.5 | SAFE | ||
| Rotation Direction | CW | SAFE | ||
| Exposure Time | 1000 | SAFE | ||
| X-Ray Tube Current | 198 | SAFE | ||
| Exposure | 247 | SAFE | ||
| Filter Type | FLAT | SAFE | ||
| Generator Power | 32 | SAFE | ||
| Focal Spot(s) | 1.2 | SAFE | ||
| Name | Value | de-identification Status | Value Across All Patients | Value Across a Given Patient |
| Single Collimation Width | 0.6 | SAFE | ||
| Total Collimation Width | 38.4 | SAFE | ||
| Table Speed | 30.7 | SAFE | ||
| Table Feed per Rotation | 30.7 | SAFE | ||
| Spiral Pitch Factor | 0.8 | SAFE | ||
| Data Collection Center (Patient) | 0.142578125; | SAFE | ||
| −193.357421875; −569.4 | ||||
| Reconstruction Target Center (Patient) | 37.142578125; | SAFE | ||
| −144.357421875; −569.4 | ||||
| Exposure Modulation Type | XYZ_EC | SAFE | ||
| Estimated Dose Saving | 43.1912 | SAFE | ||
| CTDIvol | 9.747946319 | SAFE | ||
| Study Instance UID | 1.2.840.113711.999999. | NOT SAFE | different | same |
| 255.1620909105 | ||||
| Series Instance UID | 1.2.840.113711.999999. | NOT SAFE | different | same |
| 255.1620909105.1 | ||||
| Study ID | CONFIDENTIAL | de-identified by Export Process | same | same |
| Series Number | 10 | SAFE | ||
| Acquisition Number | 4 | SAFE | ||
| Instance Number | 1 | SAFE | ||
| Image Position (Patient) | −35.857421875; | SAFE | ||
| −217.357421875; −569.4 |
| Image Orientation (Patient) | 1; 0; 0; 0; 1; 0 | SAFE | ||
| Frame of Reference UID | 1.3.12.2.1107.5.1.4.66025. | SAFE | different | same |
| 3000002105071355105980 | ||||
| Position Reference Indicator | SAFE | |||
| Slice Location | −569.4 | SAFE | ||
| Image Comments | NOT SAFE | |||
| Samples per Pixel | 1 | SAFE | ||
| Photometric Interpretation | MONOCHROME2 | SAFE | ||
| Rows | 512 | SAFE | ||
| Columns | 512 | SAFE | ||
| Pixel Spacing | 0.28515625; 0.28515625 | SAFE | ||
| Bits Allocated | 16 | SAFE | ||
| Bits Stored | 12 | SAFE | ||
| High Bit | 11 | SAFE | ||
| Pixel Representation | 0 | SAFE | ||
| Smallest Image Pixel Value | 0 | SAFE | ||
| Largest Image Pixel Value | 2921 | SAFE | ||
| Window Center | 60; 50 | SAFE | ||
| Window Width | 420; 400 | SAFE | ||
| Name | Value | de-identification Status | Value Across | Value Across |
| All Patients | a Given Patient | |||
| Rescale Intercept | −1024 | SAFE | ||
| Rescale Slope | 1 | SAFE | ||
| Rescale Type | HU | SAFE | ||
| Window Center & Width Explanation | WINDOW1; WINDOW2 | SAFE | ||
| Lossy Image Compression | 0 | SAFE | ||
| Requested Procedure Description | CT HEAD | SAFE | ||
| Requested Procedure Code Sequence | Code Value: CSKUH—Coding | NOT SAFE | different same | |
| Scheme Designator: MH—Code | ||||
| Meaning: CT HEAD | ||||
| Request Attributes Sequence | Scheduled Procedure Step Description: | NOT SAFE | same different | |
| CT HEAD—Scheduled Procedure Step | ||||
| ID: CONFIDENTIAL—Requested | ||||
| Procedure ID: 19,930,118 | ||||
| Date of Last Calibration | 20,210,507 | NOT SAFE | ||
| Time of Last Calibration | 120,556 | NOT SAFE | ||
| Convolution Kernel | I30s; 3 | SAFE | ||
| Patient Position | HFS | SAFE | ||
| Study Date | 20,170,822 | RANDOMIZED by Export Process | ||
| Series Date | 20,170,822 | RANDOMIZED by Export Process | ||
| Acquisition Date | 20,170,822 | RANDOMIZED by Export Process | ||
| Content Date | 20,170,822 | RANDOMIZED by Export Process | ||
| Acquisition DateTime | 2.02 × 10 | RANDOMIZED by Export Process | ||
| Study Time | 164,557 | RANDOMIZED by Export Process |
References
- Oren, O.; Gersh, B.J.; Bhatt, D.L. Artificial intelligence in medical imaging: Switching from radiographic pathological data to clinically meaningful endpoints. Lancet Digit. Health 2020, 2, e486–e488. [Google Scholar] [CrossRef]
- Bresnick, J. Top 5 Use Cases for Artificial Intelligence in Medical Imaging. 2018. Available online: https://healthitanalytics.com/news/top-5-use-cases-for-artificialintelligence-in-medical-imaging (accessed on 12 June 2019).
- Al-Haj, A. Providing integrity, authenticity, and confidentiality for header and pixel data of DICOM images. J. Digit. Imaging 2015, 28, 179–187. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shahbaz, S.; Mahmood, A.; Anwar, Z. SOAD: Securing oncology EMR by anonymizing DICOM images. In Proceedings of the 2013 11th International Conference on Frontiers of Information Technology, Islamabad, Pakistan, 16–18 December 2013; pp. 125–130. [Google Scholar]
- Treacy, C.; Loane, J.; McCaffery, F. A Developer Driven Framework for Security and Privacy in the Internet of Medical Things. In European Conference on Software Process Improvement; Springer: Berlin/Heidelberg, Germany, 2020; pp. 107–119. [Google Scholar]
- Magdziarczyk, M. Right to be forgotten in light of regulation (eu) 2016/679 of the european parliament and of the council of 27 april 2016 on the protection of natural persons with regard to the processing of personal data and on the free movement of such data, and repealing directive 95/46/ec. In Proceedings of the 6th International Multidisciplinary Scientific Conference on Social Sciences and Art Sgem 2019, Vienna, Austria, 11–14 April 2019; pp. 177–184. [Google Scholar]
- De Francesco, G.P. The General Data Protection Regulation’s Practical Impact on Software Architecture. Computer 2019, 52, 32–39. [Google Scholar] [CrossRef]
- Galvez, R.; Gurses, S. The odyssey: Modeling privacy threats in a brave new world. In Proceedings of the 2018 IEEE European Symposium on Security and Privacy Workshops (EuroS&PW), London, UK, 23–27 April 2018; pp. 87–94. [Google Scholar]
- Mildenberger, P.; Eichelberg, M.; Martin, E. Introduction to the DICOM standard. Eur. Radiol. 2002, 12, 920–927. [Google Scholar] [CrossRef]
- Sweeney, L. Simple demographics often identify people uniquely. Health 2000, 671, 1–34. [Google Scholar]
- Muschelli, J. Recommendations for processing head CT data. Front. Neuroinform. 2019, 13, 61. [Google Scholar] [CrossRef] [Green Version]
- Moore, S.M.; Maffitt, D.R.; Smith, K.E.; Kirby, J.S.; Clark, K.W.; Freymann, J.B.; Vendt, B.A.; Tarbox, L.R.; Prior, F.W. De-identification of medical images with retention of scientific research value. Radiographics 2015, 35, 727–735. [Google Scholar] [CrossRef] [Green Version]
- Burbridge, B. Dicom Image Anonymization and Transfer to Create a Diagnostic Radiology Teaching File. Int. J. Radiol. Imaging Technol. 2020, 6, 065. [Google Scholar]
- Newhauser, W.; Jones, T.; Swerdloff, S.; Newhauser, W.; Cilia, M.; Carver, R.; Halloran, A.; Zhang, R. Anonymization of DICOM electronic medical records for radiation therapy. Comput. Biol. Med. 2014, 53, 134–140. [Google Scholar] [CrossRef] [Green Version]
- Haak, D.; Page, C.E.; Reinartz, S.; Kruger, T.; Deserno, T.M. DICOM for clinical research: PACS-integrated electronic data capture in multi-center trials. J. Digit. Imaging 2015, 28, 558–566. [Google Scholar] [CrossRef] [Green Version]
- Aryanto, K.; Oudkerk, M.; van Ooijen, P. Free DICOM de-identification tools in clinical research: Functioning and safety of patient privacy. Eur. Radiol. 2015, 25, 3685–3695. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El Jaouhari, S.; Pasquier, G.; Cordonnier, E. On-the-fly DICOM-RTV metadata pseudonymization during a real-time streaming. In Proceedings of the 2020 International Conference on Information Networking (ICOIN), Barcelona, Spain, 7–10 January 2020; pp. 249–254. [Google Scholar]
- Beasley, D.; Eiben, B.; Doran, S.; Darcy, J.; Petts, J.; Patel, R.; White, M.; Miles, E.; McClelland, J. Secure-DICOM-Uploader: A platform for anonymising and transferring imaging data from hospital sites to remote repositories. In Proceedings of the 19th International Conference on the Use of Computers in Radiation Therapy, International Conference on the Use of Computers in Radiation Therapy, Montreal, QC, Canada, 17–20 June 2019. [Google Scholar]
- Yi, T.; Pan, I.; Collins, S.; Chen, F.; Cueto, R.; Hsieh, B.; Hsieh, C.; Smith, J.L.; Yang, L.; Liao, W.H.; et al. DICOM image analysis and archive (DIANA): An open-source system for clinical AI applications. J. Digit. Imaging 2021, 34, 1405–1413. [Google Scholar] [CrossRef] [PubMed]
- Ooijen, P.; Aryanto, K.Y.E. Pseudonymization and Anonymization of Radiology Data. In Basic Knowledge of Medical Imaging Informatics; Springer: Berlin/Heidelberg, Germany, 2021; pp. 83–97. [Google Scholar]
- Yepes-Calderon, F.; Bluml, S.; Erberich, S.; Nelson, M.D.; McComb, J.G. Improving the picture archiving and communication system: Towards one-click clinical quantifying applications. Comput. Methods Biomech. Biomed. Eng. Imaging Vis. 2019, 7, 154–161. [Google Scholar] [CrossRef]
- Natu, P.; Natu, S.; Agrawal, U. Privacy Issues in Medical Image Analysis. In Data Protection and Privacy in Healthcare; CRC Press: Boca Raton, FL, USA, 2021; pp. 51–64. [Google Scholar]
- Robinson, J.D. Beyond the DICOM header: Additional issues in deidentification. Am. J. Roentgenol. 2014, 203, W658–W664. [Google Scholar] [CrossRef] [PubMed]
- Monteiro, E.; Costa, C.; Oliveira, J.L. A de-identification pipeline for ultrasound medical images in DICOM format. J. Med. Syst. 2017, 41, 89. [Google Scholar] [CrossRef]
- Vcelak, P.; Kryl, M.; Kratochvil, M.; Kleckova, J. Identification and classification of DICOM files with burned-in text content. Int. J. Med. Inform. 2019, 126, 128–137. [Google Scholar] [CrossRef]
- Rutherford, M.; Mun, S.K.; Levine, B.; Bennett, W.; Smith, K.; Farmer, P.; Jarosz, Q.; Wagner, U.; Freyman, J.; Blake, G.; et al. A DICOM dataset for evaluation of medical image de-identification. Sci. Data 2021, 8, 1–8. [Google Scholar] [CrossRef]
- González, D.R.; Carpenter, T.; van Hemert, J.I.; Wardlaw, J. An open source toolkit for medical imaging de-identification. Eur. Radiol. 2010, 20, 1896–1904. [Google Scholar] [CrossRef]
- Clunie, D.A.; Gebow, D. Block selective redaction for minimizing loss during de-identification of burned in text in irreversibly compressed JPEG medical images. J. Med. Imaging 2015, 2, 016501. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; Wang, J.Z. DDIT-A tool for DICOM brain images de-Identification. In Proceedings of the 2011 5th International Conference on Bioinformatics and Biomedical Engineering, Wuhan, China, 10–12 May 2011; pp. 1–4. [Google Scholar]
- Kundu, S.; Chakraborty, S.; Chatterjee, S.; Das, S.; Achari, R.B.; Mukhopadhyay, J.; Das, P.P.; Mallick, I.; Arunsingh, M.; Bhattacharyyaa, T.; et al. De-identification of Radiomics data retaining longitudinal temporal information. J. Med. Syst. 2020, 44, 1–15. [Google Scholar]
- Robb, R.A.; Hanson, D.P.; Karwoski, R.; Larson, A.; Workman, E.; Stacy, M. Analyze: A comprehensive, operator-interactive software package for multidimensional medical image display and analysis. Comput. Med. Imaging Graph. 1989, 13, 433–454. [Google Scholar] [CrossRef]
- Whitcher, B.; Schmid, V.J.; Thorton, A. Working with the DICOM and NIfTI Data Standards in R. J. Stat. Softw. 2011, 44, 1–29. [Google Scholar] [CrossRef]
- Rew, R.; Davis, G. NetCDF: An interface for scientific data access. IEEE Comput. Graph. Appl. 1990, 10, 76–82. [Google Scholar] [CrossRef]
- Vincent, R.D.; Neelin, P.; Khalili-Mahani, N.; Janke, A.L.; Fonov, V.S.; Robbins, S.M.; Baghdadi, L.; Lerch, J.; Sled, J.G.; Adalat, R.; et al. MINC 2.0: A flexible format for multi-modal images. Front. Neuroinform. 2016, 10, 35. [Google Scholar] [CrossRef] [PubMed]
- Diaz, O.; Kushibar, K.; Osuala, R.; Linardos, A.; Garrucho, L.; Igual, L.; Radeva, P.; Prior, F.; Gkontra, P.; Lekadir, K. Data preparation for artificial intelligence in medical imaging: A comprehensive guide to open-access platforms and tools. Phys. Medica 2021, 83, 25–37. [Google Scholar] [CrossRef] [PubMed]
- Bazargani, M. De-Identification Tool for DICOM Image Header. Available online: https://csgitlab.ucd.ie/mldawn/dicom_de_identifier_public (accessed on 4 April 2022).
- European Society of Radiology (ESR). The new EU General Data Protection Regulation: What the radiologist should know. Insights Imaging 2017, 8, 295–299. [Google Scholar] [CrossRef] [Green Version]
- Kaissis, G.A.; Makowski, M.R.; Rückert, D.; Braren, R.F. Secure, privacy-preserving and federated machine learning in medical imaging. Nat. Mach. Intell. 2020, 2, 305–311. [Google Scholar] [CrossRef]
- Council of the European Union. General Data Protection Regulation (GDPR); Council of the European Union: Brussels, Belgium, 2016.
- Lotan, E.; Tschider, C.; Sodickson, D.K.; Caplan, A.L.; Bruno, M.; Zhang, B.; Lui, Y.W. Medical imaging and privacy in the era of artificial intelligence: Myth, fallacy, and the future. J. Am. Coll. Radiol. 2020, 17, 1159–1162. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shahid, A.; Bazargani, M.H.; Banahan, P.; Mac Namee, B.; Kechadi, T.; Treacy, C.; Regan, G.; MacMahon, P. A Two-Stage De-Identification Process for Privacy-Preserving Medical Image Analysis. Healthcare 2022, 10, 755. https://doi.org/10.3390/healthcare10050755
Shahid A, Bazargani MH, Banahan P, Mac Namee B, Kechadi T, Treacy C, Regan G, MacMahon P. A Two-Stage De-Identification Process for Privacy-Preserving Medical Image Analysis. Healthcare. 2022; 10(5):755. https://doi.org/10.3390/healthcare10050755
Chicago/Turabian StyleShahid, Arsalan, Mehran H. Bazargani, Paul Banahan, Brian Mac Namee, Tahar Kechadi, Ceara Treacy, Gilbert Regan, and Peter MacMahon. 2022. "A Two-Stage De-Identification Process for Privacy-Preserving Medical Image Analysis" Healthcare 10, no. 5: 755. https://doi.org/10.3390/healthcare10050755
APA StyleShahid, A., Bazargani, M. H., Banahan, P., Mac Namee, B., Kechadi, T., Treacy, C., Regan, G., & MacMahon, P. (2022). A Two-Stage De-Identification Process for Privacy-Preserving Medical Image Analysis. Healthcare, 10(5), 755. https://doi.org/10.3390/healthcare10050755








