1. Introduction
Mathematical models of infectious disease are key tools for understanding the epidemiological dynamics and persistence of infections in humans and animals [
1,
2]. Model systems have been used to determine the expected persistence time of infection [
3,
4,
5], and thereby provide insight into ways of reducing the impact and spread of pathogens and inform policy for infectious disease management [
6,
7].
Following an outbreak, an infection may fade-out due to random fluctuations in the number of infected individuals. This can lead to epidemic fade-out where, after a major outbreak, the population is depleted to a level where the number of infected individuals is low and pathogen extinction (zero sources of infection) occurs due to stochasticity or endemic fade-out where pathogen extinction occurs due to stochasticity from a relatively stable endemic state [
1,
4]. Analysis of endemic fade-out focuses on the expected time to pathogen extinction,
, beginning from the quasi-stationary distribution of the number of susceptible and infectious individuals. The expected pathogen extinction time increases as the population size increases while also depending on the relative timescales of infection and demographic processes [
4,
8]. The analysis of an epidemic fade-out is more complicated and arises when population numbers drop to low levels following an epidemic outbreak which can temporarily limit transmission. Persistence through the outbreak depends on the replenishment of susceptible individuals and the tail of infecteds following the outbreak [
4,
8].
The mean time to pathogen extinction has been calculated for
(susceptible-infected-susceptible) and
(susceptible-infected-recovered) model frameworks that assume density-dependent transmission and a constant population size (and therefore omit the impact of disease-induced mortality on regulating host density). For an
framework, where the basic reproductive number of the infection (defined as the number of secondary cases when a single infected individual is introduced into a disease-free population),
, is greater than unity, the mean time to pathogen extinction is given by
, where the coefficients
A and
C depend on model parameters but are independent of the total population,
N, [
9,
10]. The expression also holds for
model frameworks but with
A computed numerically and indicates that recovery from infection to immunity reduces the persistence time of infection compared to
dynamics where recovery replenishes the susceptible pool [
11,
12]. These model studies emphasize the importance of the infection processes in determining the persistence of a pathogen.
The infectious disease dynamics for many real-world systems are more complex than the SIS and SIR frameworks. For example, immunity to infection may be temporary [
12], the infection may have a significant latent period in which an individual has been exposed to the pathogen but is asymptomatic and non-infectious [
13] (such as with rabies [
14]), asymptomatic individuals may be infectious (as with Epstein Barr virus infections [
15] and COVID-19 coronavirus disease [
16]) and following infection there may be a prolonged chronic stage of disease in which individuals can still transmit the infection or can revert to the infectious state (as has been suggested for African swine fever [
17,
18]). Of particular relevance is the inclusion of disease-induced mortality as it may lead to a large drop in population size following an infectious outbreak and this has an impact on pathogen persistence [
19]. Since disease-induced mortality leads to a non-constant population size it is also important to consider different forms of transmission, where, for example, density-dependent transmission can promote an initial infectious outbreak and frequency-dependent transmission can aid persistence of infection at low population levels [
20].
In this study, we develop and assess a range of infectious disease model frameworks to explore and understand how the inclusion of more complex epidemiological processes will affect the persistence of infection. In all model frameworks we include disease-induced mortality and a combination of density-dependent and frequency-dependent transmission. We extend classical infectious disease frameworks without immunity (susceptible-infected-susceptible,
) and with immunity (susceptible-infected-recovered/immune,
) to include an exposed/latent class,
E and a chronically infected class,
C. For this mathematical study we define
E as the stage prior to full infection,
I. In this exposed class,
E, individuals do not incur disease-induced mortality (which individuals in the infected class do incur) but may transmit the infection. The exposed class therefore approximates asymptomatic or presymptomatic individuals that may or may not be infectious. We examine the impact of the exposed class on pathogen persistence by comparing systems without immunity,
and
, and those with immunity
and
. In a similar manner we define the chronically infected class,
C, as the stage that follows full infection,
I. Individuals in this class may suffer disease-induced mortality and may be able to transmit the infection but at a reduced rate compared to the infected class. We examine the impact of the chronically infected class on pathogen persistence by comparing systems without immunity,
and
, and those with immunity,
and
. We estimate the persistence time of the infection numerically using a suite of stochastic continuous-time Markov chain models [
21,
22] and our aim is to understand the importance of exposed/latent and chronic infection on pathogen persistence. Although the definitions of exposed/latent and chronic infection we employ are common in the theoretical literature [
1,
2,
7,
17], we acknowledge that their definitions may not match the precise characteristics of a specific infectious disease. Notwithstanding, our model framework allows the effect of exposed/latent and chronic infection on the mean time to extinction of an infection to be isolated and allows us to infer and discuss the impact of these processes on the persistence and management of specific infectious diseases.
2. Methods
We will first outline the model frameworks in the absence of immunity. For the
framework we describe the deterministic model framework and then the corresponding stochastic (continuous-time Markov chain) model [
22,
23]. The deterministic
model is based on classical, compartmental infectious disease modelling frameworks [
2,
24] and is represented by the following system of equations:
Here,
S represents the susceptible population density,
I the infected density and
the total population density. The maximum birth rate is given by
b, which is modified due to intra-specific competition through the parameter,
, where
K denotes the carrying capacity of the population (the stable population level in the absence of infection). The natural death rate is given by
d. For the infection dynamics we denote
to be the frequency-dependent transmission rate,
the density-dependent transmission coefficient,
the recovery rate and
the disease-induced mortality incurred by infected individuals. The equivalent stochastic model is a continuous-time Markov chain [
22,
23], with transition rates given in
Table 1.
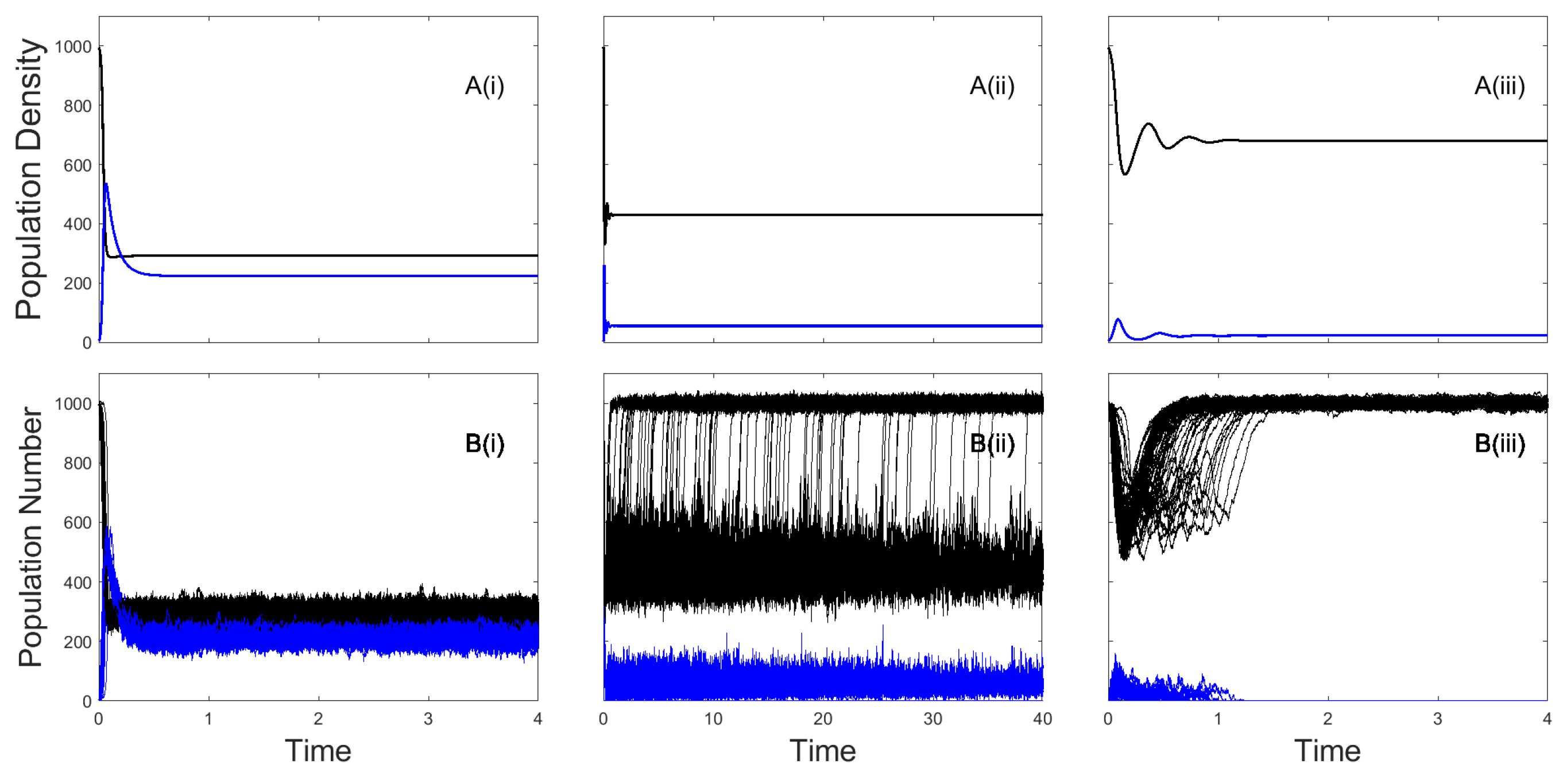
In the stochastic model of the
framework the susceptible class,
S, and infected class,
I, are integer values that represent the population number. Individual simulations can be undertaken using a Gillespie algorithm [
21,
25] that describes the dynamics for a single realization. We run 100 realizations of the stochastic model and determine the mean time to extinction of the infection,
, (pathogen extinction occurs when no sources of infection remain in the population) and the
confidence interval for the mean for each distinct model framework and set of parameters. We only consider simulations in which an initial infectious outbreak occurs, which we define as a drop in the susceptible population number below
of the pre-infection population size prior to pathogen extinction. For all simulations we choose a carrying capacity of
and use the initial conditions:
.
We choose a baseline set of parameters as defined in
Table 2. These are chosen such that the basic reproductive number of the infection,
, is greater than unity and so pathogen persistence will depend on epidemic or endemic fade-out. The
for the
framework and the other model frameworks considered in this study can be found in
supplementary information (Equations (S4)–(S6)). We compare results for a low, medium and high level of disease-induced mortality (see
Table 2) and undertake a sensitivity analysis on other key parameters to understand their impact on pathogen persistence. The
model is then extended to an
or
framework with the inclusion of an exposed/latent class,
E, (referred to as the exposed class for the remainder of the methods and results section) or a chronic class,
C, respectively. The deterministic versions of these model frameworks are shown in Equations (
2) and (
3).
In the
model (Equation (
2)) susceptible individuals progress to the exposed class,
E, following transmission and subsequently progress from the exposed class to the infected class at rate
. An exposed individual does not incur disease-induced mortality but could be infectious and transmit the pathogen at a proportion
of that of an infected,
I, individual. Our default (
Table 2) is to assume
and we consider
as part of a parameter sensitivity analysis. Following infection an individual progresses back to the susceptible class at rate
. All other parameters are as described for the
model. The deterministic
model is as follows:
In the
model (Equation (
3)) susceptible individuals progress to the infected class,
I, following transmission and then progress to the chronically infected class,
C, at rate
. A chronically infected individual can progress back to the susceptible class at rate
and may incur disease-induced mortality at a proportion,
c, of that of an infected individual. In a similar manner to the
model above we assume that a chronically infected individual may also be infectious (at a proportion
of that of an infected individual). We consider two baseline cases, one where chronically infected individuals incur disease-induced mortality but cannot transmit the infection and one where they do not incur disease-induced mortality but can transmit the infection. All other parameters are described for the
model. The deterministic
model is as follows:
For both the and model the exposed and chronically infected population are not infectious when . Note, in the limit of both models converge back to the original model. The proportional level of pathogen transmission, , of exposed and chronic individuals, the rate of progression, , from the exposed and chronic class and the proportional level of disease-induced mortality incurred by the chronic class, c, will be varied to understand their effect on the persistence of infection.
We adapt the methods described above to include immunity by considering
,
and
model frameworks. In these frameworks individuals enter a recovered and immune class
R following recovery from the infected class,
I, or chronically infected class,
C, as appropriate. These model frameworks are detailed in full in the
Supplementary Information (Equations (S1)–(S3)).
4. Discussion
Both deterministic and stochastic modelling approaches are key tools for understanding the epidemiological dynamics and persistence of infectious disease. Deterministic models have advantages of analytical tractability, where for example, determining the endemic level of infection becomes more straightforward. However, real systems are subject to random fluctuations and stochastic models developed to capture this randomness indicate that an infection can become extinct even when
[
27]. In this study we examine classical infectious disease model frameworks to consider the role of exposed/latent and chronic infection in systems with and without recovery to immunity and the role of epidemiological parameters on the mean time to extinction of an infection.
Our study confirms previous findings that compare
and
frameworks and indicate that the inclusion of immunity reduces the time to extinction of the infection and therefore the likelihood of pathogen persistence [
11,
12]. We also show that an increase in disease-induced mortality reduces the mean time to pathogen extinction for models that do not include immunity. Our work therefore extends previous findings for focused stochastic models that examine the impact of disease-induced mortality and infection induced reduction in fecundity in SI frameworks [
19]. For models that include immunity the relationship between disease-induced mortality and mean time to pathogen extinction is less clear. For a wide range of parameters the persistence time of the infection is short and has low sensitivity to changes in disease-induced mortality. However, at lower rates of recovery from infection there is evidence of a reduction in the mean time to pathogen extinction as disease-induced mortality increases. Here, the increase in the force of infection that arises from a longer duration in the infected class is offset as the level of disease-induced mortality increases. Reduced pathogen persistence for acute disease has been shown in spatial
models [
28] where the rapid turnover of infected individuals can lead to the loss of infection before it can be transmitted to neighboring areas.
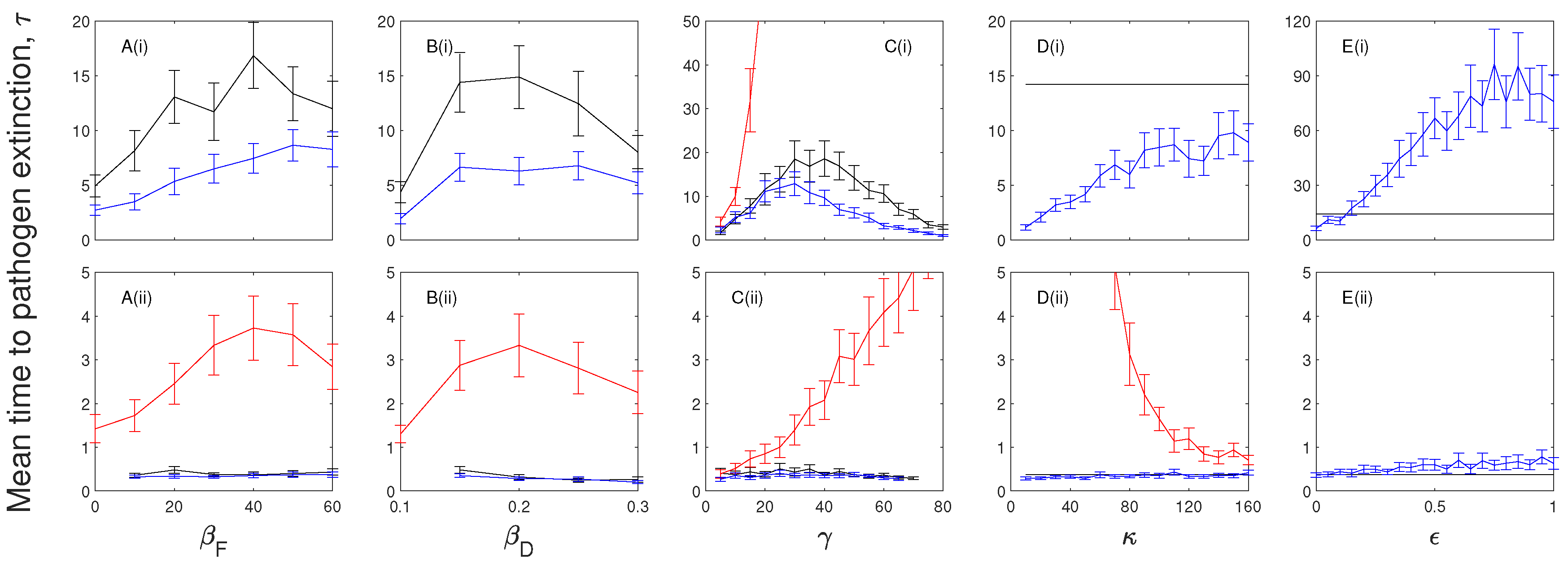
A key model assessment that we have undertaken in this study determines the impact of latent/exposed and chronic infection on the persistence of infection. Our model findings indicate that the mean time to pathogen extinction is increased when there is an exposed/latent period prior to infection. The extinction time of the infection is further increased if individuals in the exposed class are capable of transmission. Wildlife diseases have been represented by models that include an exposed class [
7], including those for infections that incur high levels of disease-induced mortality, such as rabies [
14] and Tasmanian devil facial tumor disease [
29]. Furthermore, animal tuberculosis (TB) in wild boar [
30,
31,
32] has an infection process where initial infection does not cause high levels of mortality and infected individuals have low rates of transmission (similar to the exposed class in our model where exposed individuals can transmit the infection at a reduced rate). Infected wild boar progress to generalized infection in which they incur increased mortality and have high rates of transmission (similar to the infected class in our model - with a low recovery rate,
, since individuals do not recover from TB). Under these conditions our model framework indicates that the ‘exposed’ class plays a significant role in pathogen persistence as it provides a future source of infected individuals. Theoretical assessments of the COVID-19 epidemic use
model frameworks to account for the incubation period of the virus [
33,
34], and a further key aspect of COVID-19 epidemiology is that asymptomatic individuals can transmit the infection. Although this latter process does not precisely fit one of our defined model frameworks, since for COVID-19 asymptomatic individuals that transmit the infection may progress directly to the immune stage, our model results do allow us to infer that infection from exposed individuals may be a key process in promoting persistence of this infection.
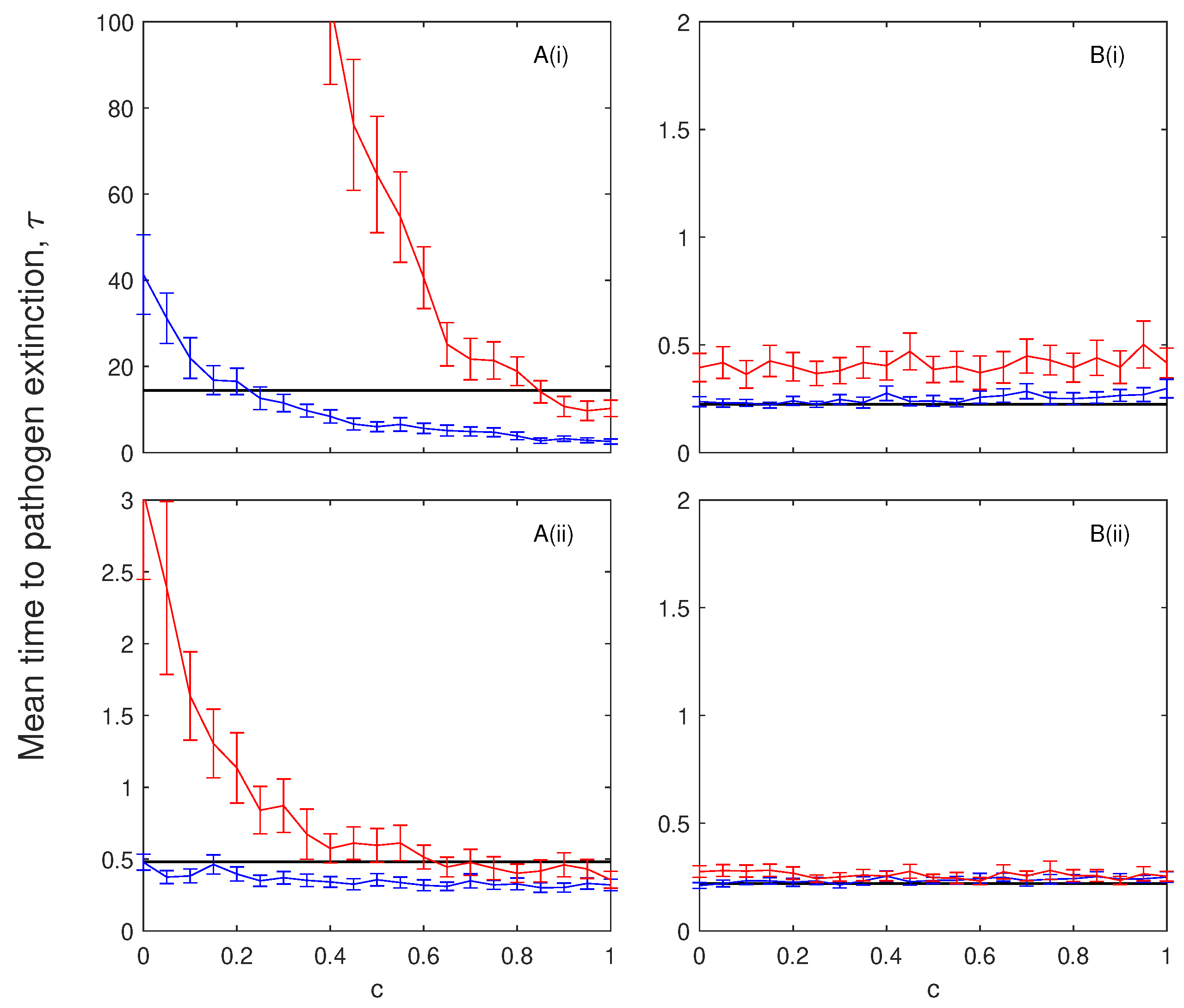
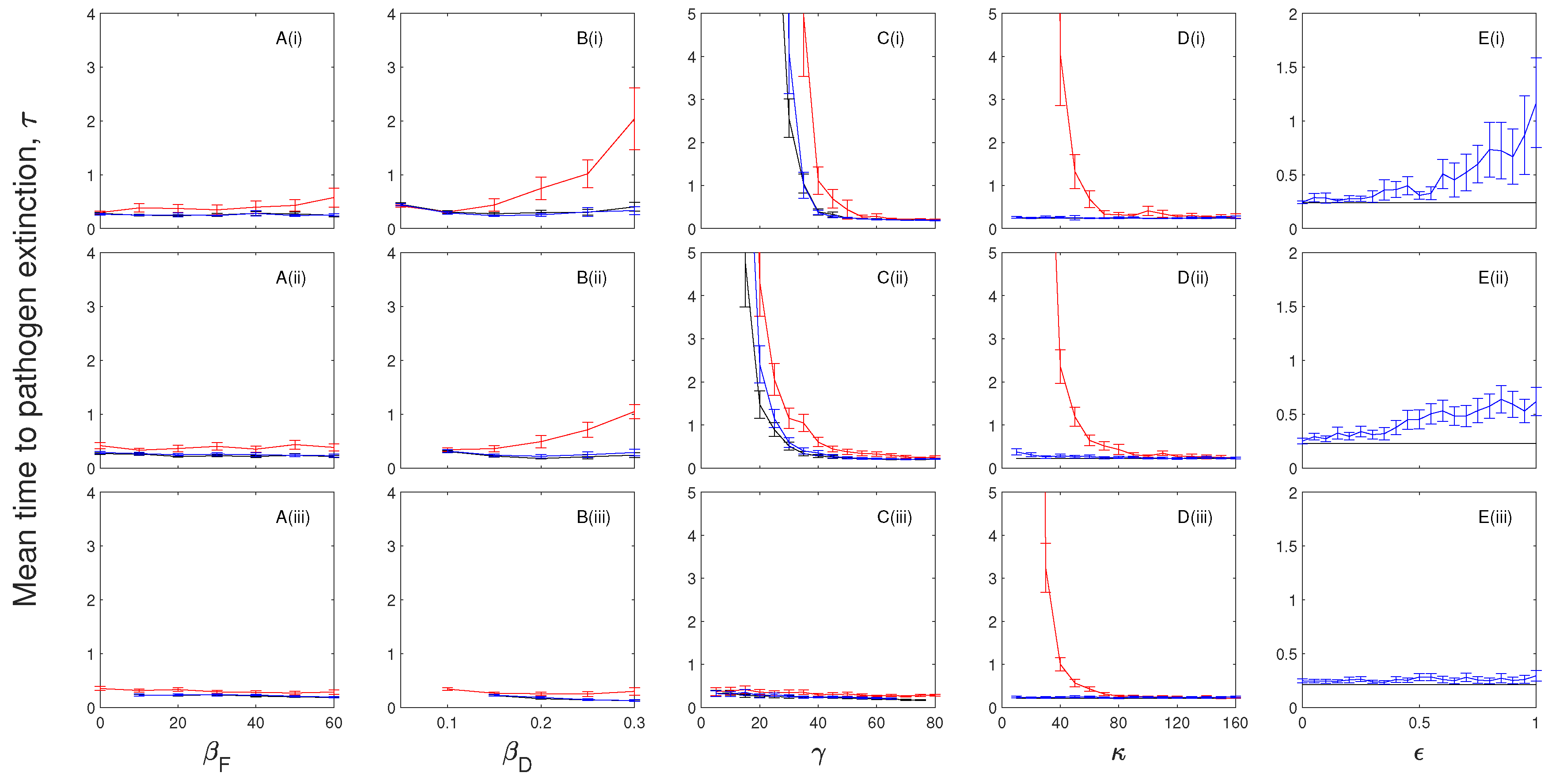
The impact of chronic infection on pathogen persistence depends on the characteristics of chronic infection. If chronically infected individuals incur disease-induced mortality but cannot transmit the infection then chronic infection can reduce the mean extinction time of the infection in models without immunity. Here, the chronic stage acts to delay the recovery while incurring the costs of disease-induced mortality. If chronically infected individuals can transmit the infection but incur low levels of disease-induced mortality then the mean persistence time of the infection is increased. The role of a chronic stage (named survivors in Stahl, et al. [
18]) has been implicated in the persistence of African swine fever where the infection persists in the long-term even though the disease is highly virulent and leads to significant losses at the population level [
17,
18]. In particular, two types of survivor are discussed [
18]. Type I survivors continue to shed the virus following infection, but they are also likely to incur additional levels of mortality compared to healthy individuals. Here, our model results indicate that the balance between transmission of infection and the level of increased mortality is a key determinant of whether pathogen persistence is enhanced. Type II survivors can revert to the infectious stage from the chronic stage and could therefore be represented by a SICI model framework. A comparison between an SI and SICI framework (see
Figure S2) indicates that the chronic stage here can lead to an increase in the mean time to extinction of the infection. Therefore, our study suggests that a chronic stage of infection could aid pathogen persistence, make the infection more difficult to eradicate and could be a mechanism that promotes the observed persistence of African swine fever following an outbreak [
17].
Our strategic model study has considered the importance of different epidemiological processes on the persistence of infectious disease. We have confirmed and extended the findings that indicate that pathogen persistence is reduced for infections that lead to immunity. We have also shown that a latent or chronic stage of infection can increase the persistence time of an infection which may make infectious diseases with these characteristics more difficult to manage and control.