Using Data-Compressors for Classification Hunting Behavioral Sequences in Rodents as “Ethological Texts”
Abstract
1. Introduction
2. The Suggested Method
3. The Procedure
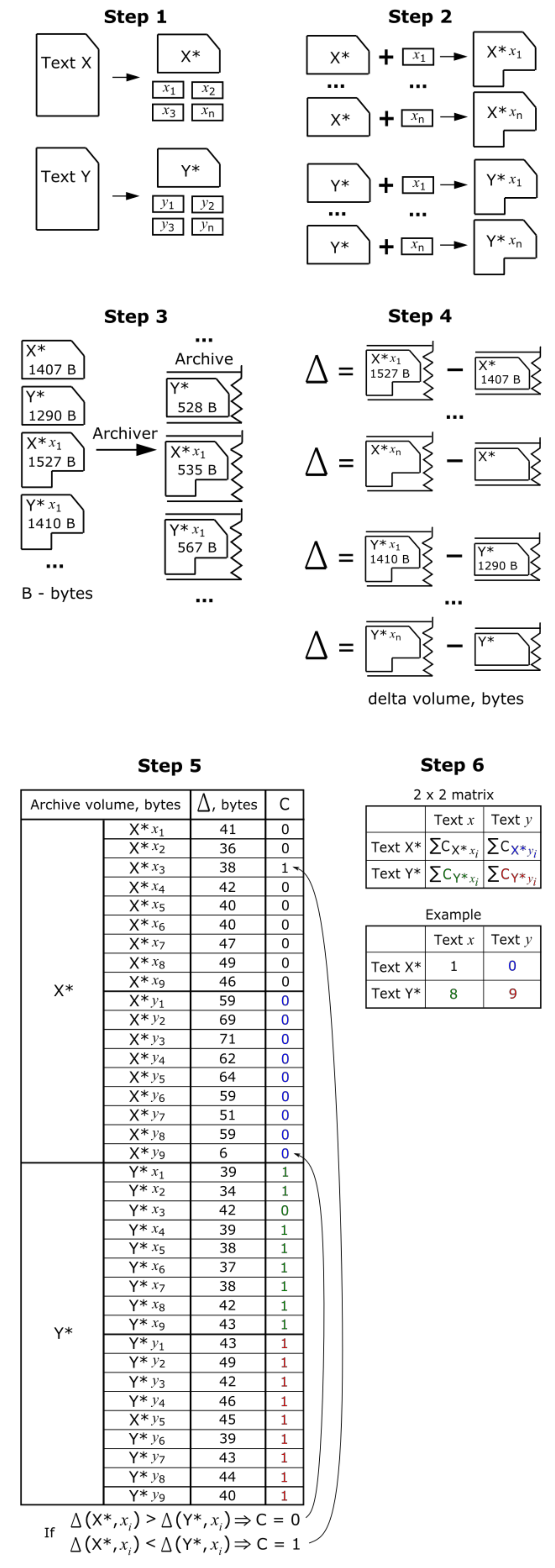
3.1. Notions and Data Encoding
3.2. Constructing Sequences for Hypothesis Testing
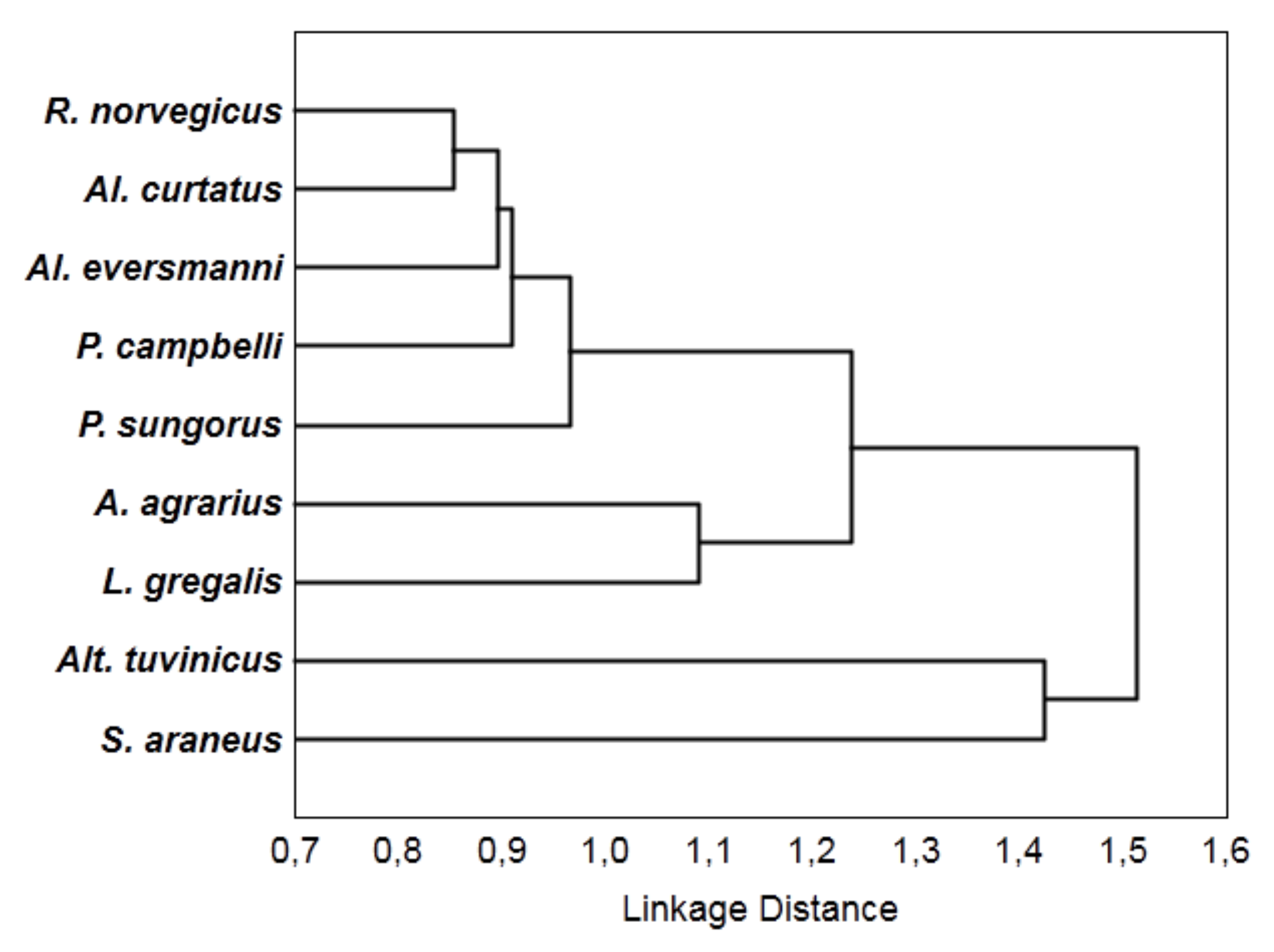
4. Results
5. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
Animals and Housing
Experimental Scheme
| Symbols | Behavioural Elements |
|---|---|
| Q | Running |
| S | Walking |
| W | Bite |
| E | Capturing the prey by forepaws (only in rodents) |
| R | Handling (only in rodents) |
| H | Nibbling insects’ legs |
| G | Carrying the prey in teeth |
| D | Sniffing |
| N | Pinning the prey down to the ground by one paw (only in shrew) |
| M | The same, by two paws (only in shrew) |
| C | Freezing |
| V | Turning a body to 90° |
| B | U-turn |
| F | Turning a head |
| Y | Rearing against the wall |
| U | Backwards movement |
| X | Self-grooming |
| J | Jump |
| I | Free-standing rearing |
References
- Li, C.; Zhang, X.; Cao, Z. Triangular and Fibonacci number patterns driven by stress on core/shell microstructures. Science 2005, 309, 909–911. [Google Scholar] [CrossRef] [PubMed]
- Lorenz, K.Z. The comparative method in studying innate behavior patterns. In Society for Experimental Biology, Physiological Mechanisms in Animal Behavior (Society’s Symposium IV); Cambridge University Press: Cambridge, UK, 1950; pp. 221–268. [Google Scholar]
- Blomberg, S.P.; Garland, T., Jr.; Ives, A.R. Testing for phylogenetic signal in comparative data: Behavioral traits are more labile. Evolution 2003, 57, 717–745. [Google Scholar] [CrossRef]
- Malange, J.; Alberts, C.C.; Oliveira, E.S.; Japyassú, H.F. The evolution of behavioural systems: A study of grooming in rodents. Behaviour 2013, 150, 1295–1324. [Google Scholar] [CrossRef]
- West-Eberhard, M.J. Developmental Plasticity and Evolution; Oxford University Press: New York, NY, USA, 2003. [Google Scholar]
- Li, M.; Chen, X.; Li, X.; Ma, B.; Vitányi, P.M. The similarity metric. IEEE Trans. Inf. Theory 2004, 50, 3250–3264. [Google Scholar] [CrossRef]
- Cilibrasi, R.; Vitányi, P.M. Clustering by compression. IEEE Trans. Inf. Theory 2005, 51, 1523–1545. [Google Scholar] [CrossRef]
- Xie, X.; Guan, J.; Zhou, S. Similarity evaluation of DNA sequences based on frequent patterns and entropy. BMC Genomics 2015, 16, S5. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Huo, H.; Chen, X.; Guo, X.; Vitter, J.S. Efficient compression and indexing for highly repetitive DNA sequence collections. IEEE/ACM Trans. Comput. Biol. Bioinform. 2020, 14, 1–14. [Google Scholar] [CrossRef]
- Forrester, G.S. A multidimensional approach to investigations of behaviour: Revealing structure in animal communication signals. Anim. Behav. 2008, 76, 1749–1760. [Google Scholar] [CrossRef]
- Asher, L.; Collins, L.M.; Ortiz-Pelaez, A.; Drewe, J.A.; Nicol, C.J.; Pfeiffer, D.U. Recent advances in the analysis of behavioural organization and interpretation as indicators of animal welfare. J. R. Soc. Interface 2009, 6, 1103–1119. [Google Scholar] [CrossRef]
- Gadbois, S.; Sievert, O.; Reeve, C.; Harrington, F.H.; Fentress, J.C. Revisiting the concept of behavior patterns in animal behavior with an example from food-caching sequences in Wolves (Canis lupus), Coyotes (Canis latrans), and Red Foxes (Vulpes vulpes). Behav. Process. 2015, 110, 3–14. [Google Scholar] [CrossRef]
- Kershenbaum, A.; Blumstein, D.T.; Roch, M.A.; Akçay, Ç.; Backus, G.; Bee, M.A.; Coen, M.; Cao, Y.; Bohn, K.; Carter, G.; et al. Acoustic sequences in non-human animals: A tutorial review and prospectus. Biol. Rev. 2016, 91, 13–52. [Google Scholar] [CrossRef] [PubMed]
- Moore, T.Y.; Cooper, K.L.; Biewener, A.A.; Vasudevan, R. Unpredictability of escape trajectory explains predator evasion ability and microhabitat preference of desert rodents. Nat. Commun. 2017, 8, 1–9. [Google Scholar] [CrossRef]
- Whishaw, I.Q.; Faraji, J.; Kuntz, J.R.; Agha, B.M.; Metz, G.A.; Mohajerani, M.H. The syntactic organization of pasta-eating and the structure of reach movements in the head-fixed mouse. Sci. Rep. 2017, 7, 10987. [Google Scholar] [CrossRef] [PubMed]
- Casarrubea, M.; Aiello, S.; Di Giovanni, G.; Santangelo, A.; Palacino, M.; Crescimanno, G. Combining quantitative and qualitative data in the study of feeding behavior in male Wistar rats. Front. Psychol. 2019, 10, 881. [Google Scholar] [CrossRef] [PubMed]
- McCowan, B.; Doyle, L.R.; Hanser, S.F. Using information theory to assess the diversity, complexity, and development of communicative repertoires. J. Comp. Psychol. 2002, 116, 166–172. [Google Scholar] [CrossRef] [PubMed]
- Kadota, M.; White, E.J.; Torisawa, S.; Komeyama, K.; Takagi, T. Employing relative entropy techniques for assessing modifications in animal behavior. PLoS ONE 2011, 6, e28241. [Google Scholar] [CrossRef] [PubMed]
- Peng, Z.; Genewein, T.; Braun, D.A. Assessing randomness and complexity in human motion trajectories through analysis of symbolic sequences. Front. Hum. Neurosci. 2014, 8, 168. [Google Scholar] [CrossRef]
- Gauvrit, N.; Singmann, H.; Soler-Toscano, F.; Zenil, H. Algorithmic complexity for psychology: A user-friendly implementation of the coding theorem method. Behav. Res. Methods 2016, 48, 314–329. [Google Scholar] [CrossRef]
- Fisher, R.A. Statistical Methods, Experimental Design, and Scientific Inference; Oliver & Boyd: Edinburgh, UK, 1956. [Google Scholar]
- Reznikova, Z.; Levenets, J.; Panteleeva, S.; Ryabko, B. Studying hunting behaviour in the striped field mouse using data compression. Acta Ethol. 2017, 20, 165–173. [Google Scholar] [CrossRef]
- Ryabko, B.; Reznikova, Z.; Druzyaka, A.; Panteleeva, S. Using ideas of Kolmogorov complexity for studying biological texts. Theory Comput. Syst. 2013, 52, 133–147. [Google Scholar] [CrossRef]
- Reznikova, Z.; Levenets, J.; Panteleeva, S.; Novikovskaya, A.; Ryabko, B.; Feoktistova, N.; Gureeva, A.; Surov, A. Using the data-compression method for studying hunting behavior in small mammals. Entropy 2019, 21, 368. [Google Scholar] [CrossRef]
- Levenets, J.V.; Panteleeva, S.N.; Reznikova, Z.I.; Gureeva, A.V.; Feoktistova, N.Y.; Surov, A.V. Experimental Comparative Analysis of Hunting Behavior in Four Species of Cricetinae Hamsters. Biol. Bull. 2019, 46, 1182–1191. [Google Scholar] [CrossRef]
- Ryabko, B.; Guskov, A.; Selivanova, I. Using data-compressors for statistical analysis of problems on homogeneity testing and classification. In Proceedings of the 2017 IEEE International Symposium on Information Theory (ISIT), Aachen, Germany, 25–30 June 2017; IEEE: Piscataway, NJ, USA, 2017; pp. 121–125. [Google Scholar] [CrossRef]
- Fisher, R.A. On the interpretation of χ2 from contingency tables, and the calculation of P. J. R. Stat. Soc. 1922, 85, 87–94. [Google Scholar] [CrossRef]
- Fisher, R.A. Statistical Methods for Research Workers; Oliver and Boyd: New York, NY, USA, 1950. [Google Scholar]
| x | y | |
|---|---|---|
| X* | 1 | 0 |
| Y* | 8 | 9 |
| x | z | |
|---|---|---|
| X* | 9 | 0 |
| Z* | 0 | 9 |
| y | z | |
|---|---|---|
| Y* | 5 | 0 |
| Z* | 4 | 9 |
| Species | Sizes of a Source Text Files (Bytes) | Numbers of Sequences in Source Text Files | Sizes of the First Parts of the Source Text Files (Bytes) | Number of the Sample Files Obtained |
|---|---|---|---|---|
| Rattus norvegicus | 2572 | 108 | 1290 | 9 |
| Apodemus agrarius | 3343 | 83 | 1672 | 9 |
| Phodopus campbelli | 1715 | 43 | 801 | 4 |
| P. sungorus | 1585 | 76 | 792 | 6 |
| Allocricetulus eversmanni | 1463 | 60 | 731 | 5 |
| Al. curtatus | 2814 | 115 | 1407 | 9 |
| Lasiopodomys gregalis | 1086 | 34 | 543 | 3 |
| Alticola tuvinicus | 1319 | 157 | 659 | 5 |
| Sorex araneus | 1637 | 61 | 818 | 5 |
| Species | A. agrarius | L. gregalis |
|---|---|---|
| A. agrarius | 6 | 0 |
| L. gregalis | 3 | 3 |
| Species | R. nor. | A. ag. | P. cam. | P. sun. | Al. ev. | Al. cur. | L. gr. | Alt. tuv. | S. ar. |
|---|---|---|---|---|---|---|---|---|---|
| R. norvegicus | 0 | 0.58 | 1 | 0.74 | 0.37 | 0.24 | 1 | 0.85 | 1 |
| A. agrarius | 0.58 | 0 | 0.28 | 0.87 | 0.85 | 1 | 0.58 | 0.86 | 0.93 |
| P. campbelli | 1 | 0.28 | 0 | 0.53 | 0 | 0.44 | 0.73 | 1 | 1 |
| P. sungorus | 0.74 | 0.87 | 0.53 | 0 | 0.83 | 0.49 | 1 | 1 | 1 |
| Al. eversmanni | 0.37 | 0.85 | 0 | 0.83 | 0 | 0.45 | 0.6 | 1 | 1 |
| Al. curtatus | 0.24 | 1 | 0.44 | 0.49 | 0.45 | 0 | 0.82 | 1 | 1 |
| L. gregalis | 1 | 0.58 | 0.73 | 1 | 0.6 | 0.82 | 0 | 1 | 1 |
| Alt. tuvinicus | 0.85 | 0.86 | 1 | 1 | 1 | 1 | 1 | 0 | 1 |
| S. araneus | 1 | 0.93 | 1 | 1 | 1 | 1 | 1 | 1 | 0 |
| Species | R. nor. | A. ag. | P. cam. | P. sun. | Al. ev. | Al. cur. | L. gr. | Alt. tuv. | S. ar. |
|---|---|---|---|---|---|---|---|---|---|
| R. norvegicus | X | 0.029* | 0.001 ** | 0.011 * | 0.360 | 1.000 | 0.005 ** | 0.005 ** | 0.001 ** |
| A. agrarius | X | 1.000 | 0.002 ** | 0.005 ** | 0.001 ** | 0.180 | 0.003 ** | 0.003 ** | |
| P. campbelli | X | 0.200 | 1.000 | 0.230 | 0.140 | 0.008 ** | 0.009 ** | ||
| P. sungorus | X | 0.015* | 0.100 | 0.020* | 0.002 ** | 0.002 ** | |||
| Al. eversmanni | X | 0.150 | 0.190 | 0.008 ** | 0.008 ** | ||||
| Al. curtatus | X | 0.020* | 0.001 ** | 0.001 ** | |||||
| L. gregalis | X | 0.020 * | 0.020 * | ||||||
| Alt. tuvinicus | X | 0.008 ** | |||||||
| S. araneus | X |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Levenets, J.; Novikovskaya, A.; Panteleeva, S.; Reznikova, Z.; Ryabko, B. Using Data-Compressors for Classification Hunting Behavioral Sequences in Rodents as “Ethological Texts”. Mathematics 2020, 8, 579. https://doi.org/10.3390/math8040579
Levenets J, Novikovskaya A, Panteleeva S, Reznikova Z, Ryabko B. Using Data-Compressors for Classification Hunting Behavioral Sequences in Rodents as “Ethological Texts”. Mathematics. 2020; 8(4):579. https://doi.org/10.3390/math8040579
Chicago/Turabian StyleLevenets, Jan, Anna Novikovskaya, Sofia Panteleeva, Zhanna Reznikova, and Boris Ryabko. 2020. "Using Data-Compressors for Classification Hunting Behavioral Sequences in Rodents as “Ethological Texts”" Mathematics 8, no. 4: 579. https://doi.org/10.3390/math8040579
APA StyleLevenets, J., Novikovskaya, A., Panteleeva, S., Reznikova, Z., & Ryabko, B. (2020). Using Data-Compressors for Classification Hunting Behavioral Sequences in Rodents as “Ethological Texts”. Mathematics, 8(4), 579. https://doi.org/10.3390/math8040579






