Objective Diagnosis for Histopathological Images Based on Machine Learning Techniques: Classical Approaches and New Trends
Abstract
1. Introduction
2. Histopathological Image Overview
2.1. Types of ML Systems in HI Analysis
2.1.1. Computer-Aided Diagnosis (CAD)
2.1.2. Content-based on Image Retrieval (CBIR)
2.1.3. Finding New Clinicopathological Associations
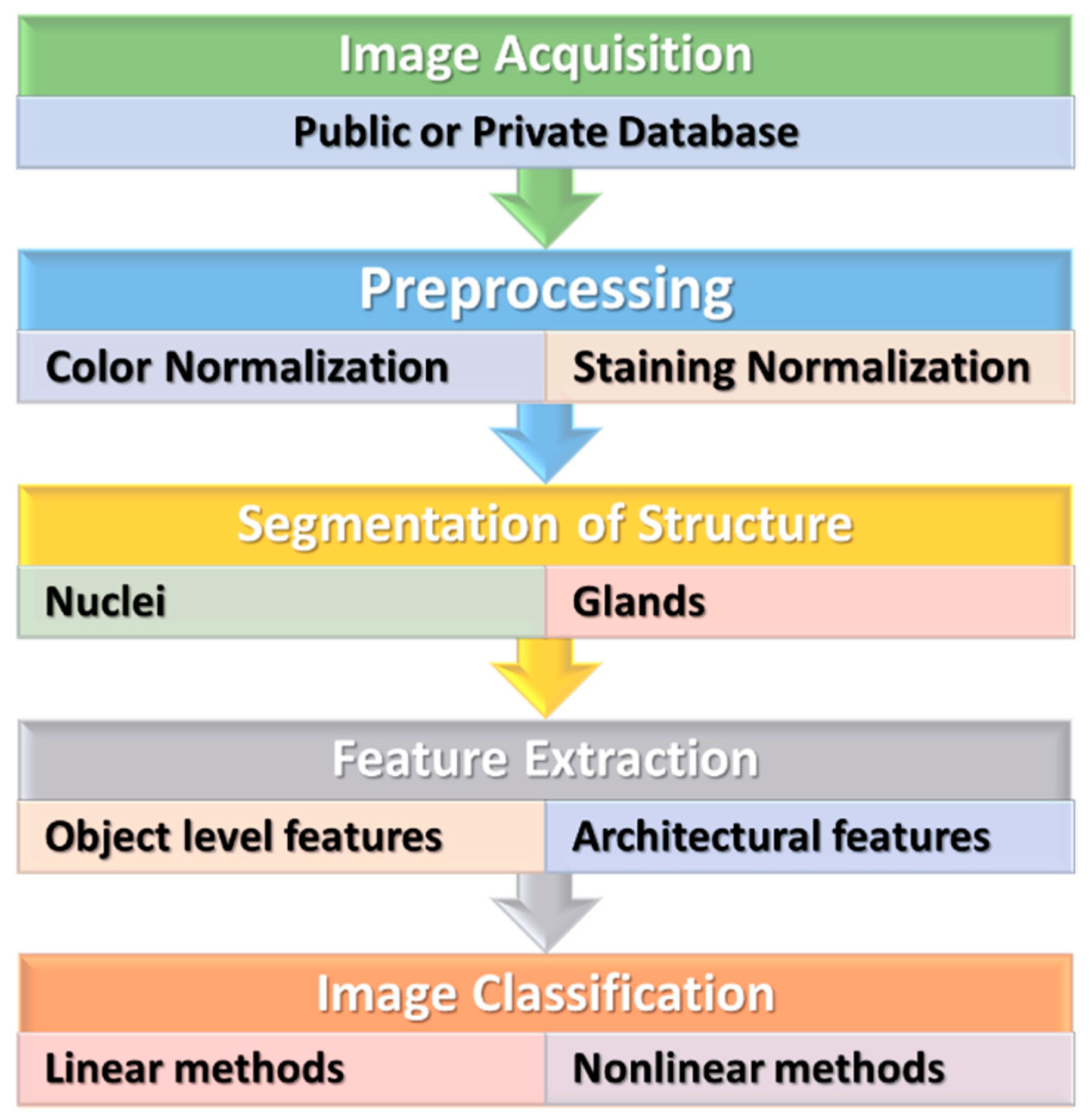
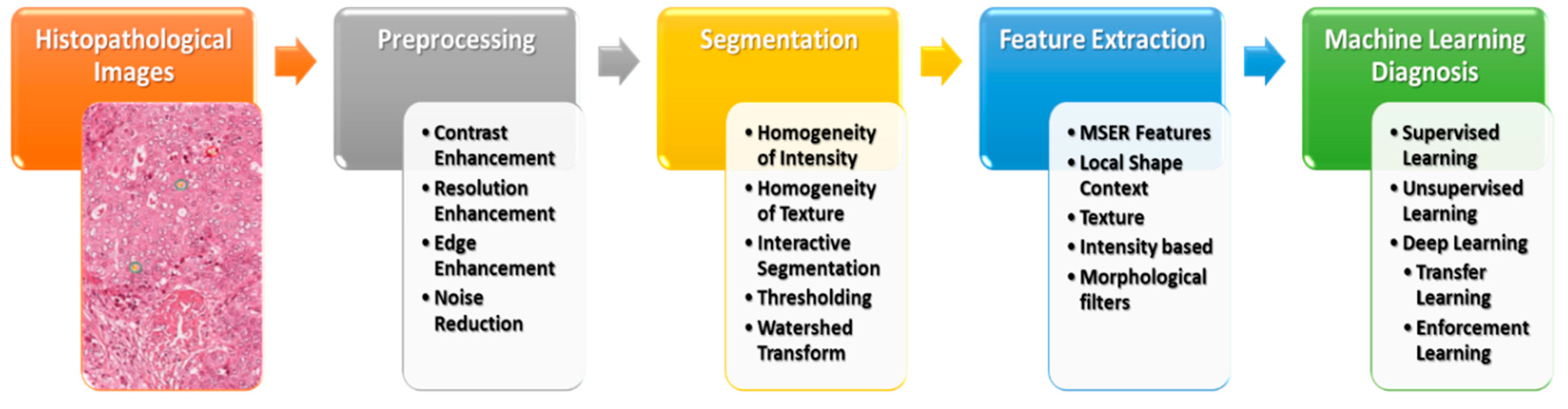
3. Conventional Machine Learning Methods
3.1. Preprocessing
3.1.1. Staining Normalization
3.1.2. Color Normalization
3.2. Recognition and Segmentation of Structures
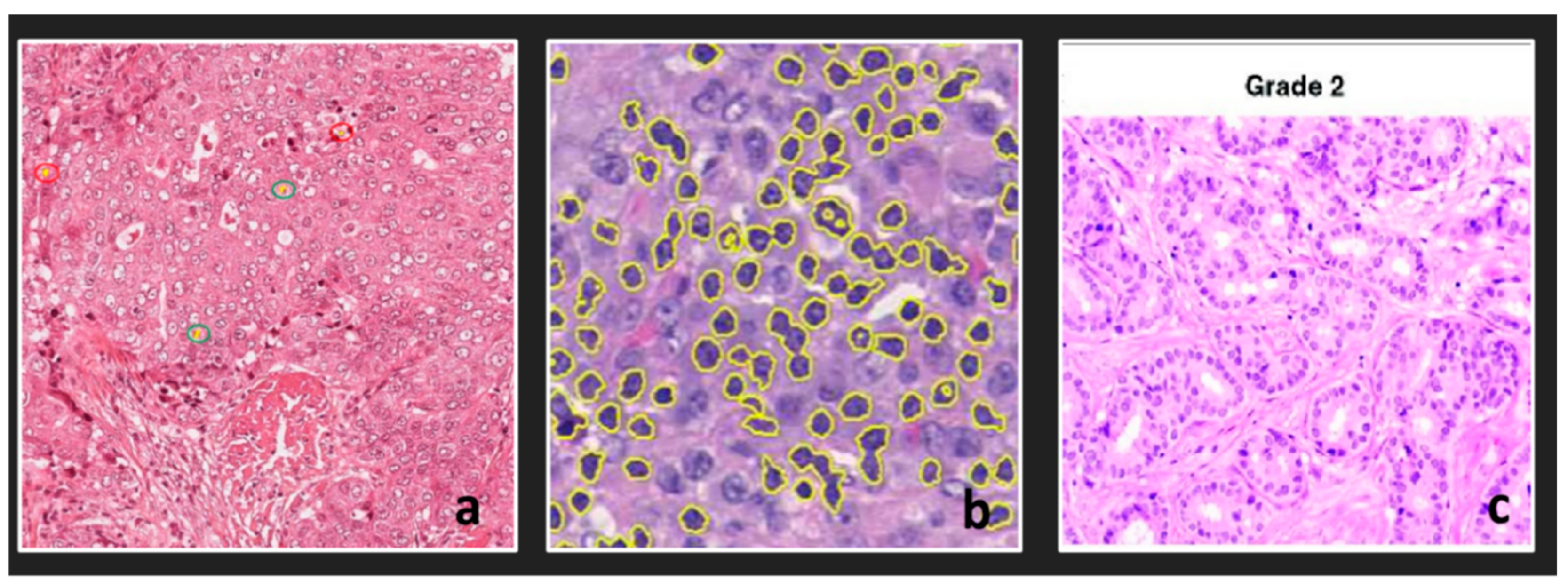
3.2.1. Nuclei and Cells
3.2.2. Glands
3.3. Feature Extraction
3.3.1. Object-Level Characteristics
3.3.2. Structural Characteristics
3.4. Classification
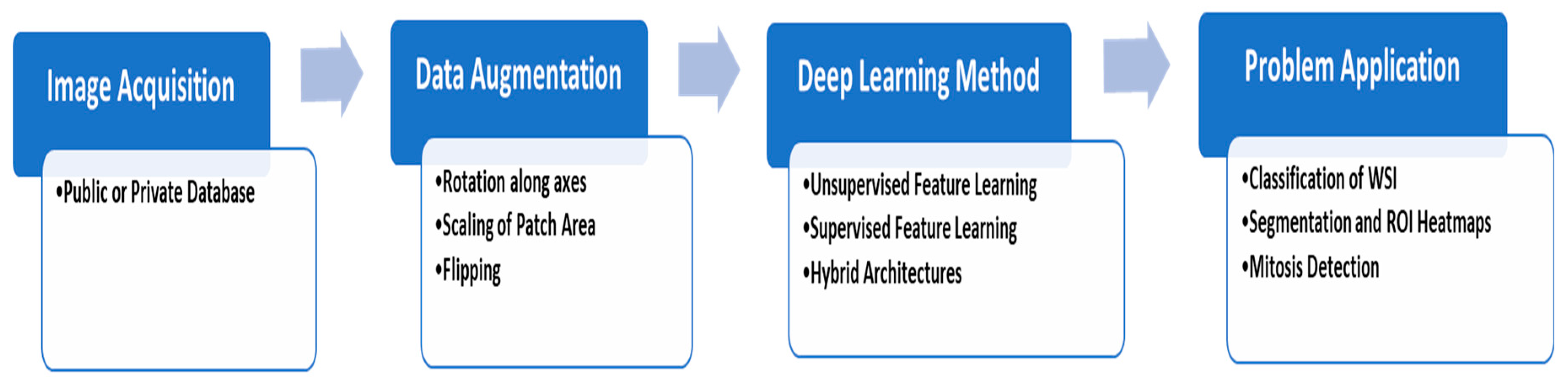
4. Deep Learning Methods
5. Datasets
| Datasets | No Slides | Staining | Diseases |
|---|---|---|---|
| TCGA [33,113] | 18,462 | H&E | Cancer |
| GTE [109] | 25,380 | H&E | Normal |
| TMAD [110,114] | 3726 | H&E/IHC | various tissue |
| TUPAC16 [115] | 821 from TCGA | H&E | Breast cancer |
| Camelyon17 [111] | 1000 | H&E | Breast cancer (lymph node metastasis) |
| Köbel et al. [116,117] | 80 | H&E | Ovarian carcinoma |
| KIMIA Path24 [118] | 24 | H&E/IHC | various tissue |
| Datasets | No of Images | Staining | Organs | Potential Usage |
|---|---|---|---|---|
| KIMIA960 [119,120] | 960 | H&E/IHC | Different tissue | Classification |
| Bio-segmentation [121,122] | 58 | H&E | Breast | Classification |
| Bioimaging challenge 2015 [123] | 269 | H&E | Breast | Classification |
| GlaS [124] | 165 | H&E | Colorectal | Gland segmentation |
| BreakHis [112] | 7909 | H&E | Breast | Classification |
| Jakob Nikolas et al. [120,125] | 100 | IHC | Colorectal | Detection of blood vessel |
| MITOS-ATYPIA-14 [126] | 4240 | H&E | Breast | Detection of mitosis, classification |
| Kumar et al. [119,127] | 30 | H&E | Different cancer | Segmentation of Nuclear |
| MITOS [20] | 100 | H&E | Breast | Detection of mitosis |
| Janowczyk et al. [128,129] | 374 | H&E | Lymphoma | classification |
| Janowczyk et al. [128,129] | 85 | H&E | Colorectal | Segmentation of gland |
| Ma et al. [130] | 81 | IHC | Breast | TIL analysis |
| Linder et al. [131,132] | 1377 | IHC | Colorectal | Segmentation of epithelium and stroma |
6. Discussion and Histopathological Tasks
Tasks for Histopathology Image
- Recognition of Mitosis
- Segmentation of structure
- Classification of images
- ▪
- For nuclear atypia rating, a points-based scheme is used.
- ▪
- The region under the curve of the receiver operating characteristic (ROC) to classify slides of lymph node comprising metastasis or not.
- ▪
- Approval with ground truth calculated with Spearman’s correlation or quadratic weighted Cohen’s kappa to grade cancer.
7. Limitations and Future Trends
- Novel Objects Discovery
- Interpretable DL Model
- Intraoperative Diagnosis
- Tumor-Infiltrating Immune Cell Analysis
- Challenges in HI analysis
- Quality of training
- Clinical translation
- Synthesis rather than marking
- Translation of morphology
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; van der Laak, J.A.; van Ginneken, B.; Sanchez, I.C. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef]
- Ker, J.; Wang, L.; Rao, J.; Lim, T. Deep Learning Applications in Medical Image Analysis. IEEE Access 2017, 6, 9375–9389. [Google Scholar] [CrossRef]
- Perez, H.; Tah, J. Improving the Accuracy of Convolutional Neural Networks by Identifying and Removing Outlier Images in Datasets Using t-SNE. Mathematics 2020, 8, 662. [Google Scholar] [CrossRef]
- Suzuki, K. Overview of deep learning in medical imaging. Radiol. Phys. Technol. 2017, 10, 257–273. [Google Scholar] [CrossRef]
- Pantanowitz, L. Digital images and the future of digital pathology. J. Pathol. Inform. 2010, 1. [Google Scholar] [CrossRef]
- Gurcan, M.N.; Boucheron, L.E.; Can, A.; Madabhushi, A.; Rajpoot, N.M.; Yener, B. Histopathological image analysis: A review. IEEE Rev. Biomed. Eng. 2009, 2, 147–171. [Google Scholar] [CrossRef]
- Greenspan, H.; Ginneken, B.; van Summers, R.M. Guest Editorial Deep Learning in Medical Imaging: Overview and Future Promise of an Exciting New Technique. IEEE Trans Med. Imaging 2016, 35, 1153–1159. [Google Scholar] [CrossRef]
- Rubin, R.; Strayer, D.S.; Rubin, E. Rubin’s Pathology: Clinicopathologic Foundations of Medicine; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2008. [Google Scholar]
- Hewitson, T.; Darby, I.; Walker, J. Histology Protocols. In Methods in Molecular Biology; Humana Press: Totowa, NJ, USA, 2010. [Google Scholar]
- Li, C.; Chen, H.; Li, X.; Xu, N.; Hu, Z.; Xue, D.; Qi, S.; Ma, H.; Zhang, L.; Sun, H. A review for cervical histopathology image analysis using machine vision approaches. Artif. Intell. Rev. 2020, 53, 4821–4862. [Google Scholar] [CrossRef]
- He, L.; Long, L.R.; Antani, S.; Thoma, G.R. Histology image analysis for carcinoma detection and grading. Comput. Methods Programs Biomed. 2012, 107, 538–556. [Google Scholar] [CrossRef]
- Ghaznavi, F.; Evans, A.; Madabhushi, A.; Feldman, M. Digital Imaging in Pathology: Whole-Slide Imaging and Beyond. Annu. Rev. Pathol. Mech. Dis. 2013, 8, 331–359. [Google Scholar] [CrossRef]
- Demir, C.; Yener, B. Automated Cancer Diagnosis Based on Histopathological Images: A Systematic Survey; Technical Report for Rensselaer Polytechnic Institute: New York, NY, USA, 2005. [Google Scholar]
- Belsare, A. Histopathological Image Analysis Using Image Processing Techniques: An Overview. Signal Image Process. Int. J. 2012, 3, 23–36. [Google Scholar] [CrossRef]
- Spanhol, F.A.; Oliveira, L.S.; Petitjean, C.; Heutte, L. Breast cancer histopathological image classification using Convolutional Neural Networks. In Proceedings of the 2016 International Joint Conference on Neural Networks (IJCNN), Vancouver, BC, Canada, 24–29 January 2016; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2016; pp. 2560–2567. [Google Scholar]
- Kieffer, B.; Babaie, M.; Kalra, S.; Tizhoosh, H.R. Convolutional neural networks for histopathology image classification: Training vs. using pre-trained networks. Proceeding of the 2017 Seventh International Conference on Image Processing Theory, Tools, and Applications (IPTA), Montreal, QC, Canada, 28 November–1 December 2017; IEEE: Piscataway, NJ, USA, 2017; pp. 1–6. [Google Scholar]
- Mungle, T.; Tewary, S.; Das, D.; Arun, I.; Basak, B.; Agarwal, S.; Ahmed, R.; Chatterjee, S.; Chakraborty, C. MRF-ANN: A machine learning approach for automated ER scoring of breast cancer immunohistochemical images. J. Microsc. 2017, 267, 117–129. [Google Scholar] [CrossRef]
- Sheikhzadeh, F.; Ward, R.K.; van Niekerk, D.; Guillaud, M. Automatic labeling of molecular biomarkers of immunohistochemistry images using fully convolutional networks. PLoS ONE 2018, 13, e0190783. [Google Scholar] [CrossRef]
- Wang, D.; Foran, D.J.; Ren, J.; Zhong, H.; Kim, I.Y.; Qi, X. Exploring automatic prostate histopathology image gleason grading via local structure modeling. In Proceedings of the 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Milan, Italy, 25–29 August 2015; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2015; Volume 2015, pp. 2649–2652. [Google Scholar]
- Roux, L.; Racoceanu, D.; Loménie, N.; Kulikova, M.; Irshad, H.; Klossa, J.; Capron, F.; Genestie, C.; Le Naour, G.; Gurcan, M.N. Mitosis detection in breast cancer histological images An ICPR 2012 contest. J. Pathol. Inform. 2013, 4, 8. [Google Scholar] [CrossRef]
- Shah, M.; Wang, D.; Rubadue, C.; Suster, D.; Beck, A. Deep learning assessment of tumor proliferation in breast cancer histological images. In Proceedings of the 2017 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Kansas City, MO, USA, 13–16 November 2017; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2017; pp. 600–603. [Google Scholar]
- Chen, H.; Qi, X.; Yu, L.; Heng, P.-A. DCAN: Deep Contour-Aware Networks for Accurate Gland Segmentation. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2016; pp. 2487–2496. [Google Scholar]
- Gertych, A.; Ing, N.; Ma, Z.; Fuchs, T.J.; Salman, S.; Mohanty, S.; Bhele, S.; Velásquez-Vacca, A.; Amin, M.B.; Knudsen, B.S. Machine learning approaches to analyze histological images of tissues from radical prostatectomies. Comput. Med. Imaging Graph. 2015, 46, 197–208. [Google Scholar] [CrossRef]
- Caicedo, J.C.; González, F.A.; Romero, E. Content-based histopathology image retrieval using a kernel-based semantic annotation framework. J. Biomed. Inform. 2011, 44, 519–528. [Google Scholar] [CrossRef]
- Caie, P.D.; Turnbull, A.K.; Farrington, S.M.; Oniscu, A.; Harrison, D.J. Quantification of tumour budding, lymphatic vessel density and invasion through image analysis in colorectal cancer. J. Transl. Med. 2014, 12, 156. [Google Scholar] [CrossRef]
- Sirinukunwattana, K.; Pluim, J.P.; Chen, H.; Qi, X.; Heng, P.-A.; Guo, Y.B.; Wang, L.Y.; Matuszewski, B.J.; Bruni, E.; Sanchez, U.; et al. Gland segmentation in colon histology images: The glas challenge contest. Med. Image Anal. 2017, 35, 489–502. [Google Scholar] [CrossRef]
- Qi, X.; Wang, D.; Rodero, I.; Diaz-Montes, J.; Gensure, R.H.; Xing, F.; Zhong, H.; Goodell, L.; Parashar, M.; Foran, D.J.; et al. Content-based histopathology image retrieval using CometCloud. BMC Bioinform. 2014, 15, 1–17. [Google Scholar] [CrossRef]
- Sparks, R.; Madabhushi, A. Out-of-Sample Extrapolation utilizing Semi-Supervised Manifold Learning (OSE-SSL): Content Based Image Retrieval for Histopathology Images. Sci. Rep. 2016, 6, 27306. [Google Scholar] [CrossRef]
- Sridhar, A.; Doyle, S.; Madabhushi, A. Content-based image retrieval of digitized histopathology in boosted spectrally embedded spaces. J. Pathol. Inform. 2015, 6. [Google Scholar] [CrossRef]
- Vanegas, J.A.; Arevalo, J.; González, F.A. Unsupervised feature learning for content-based histopathology image retrieval. In 2014 12th International Workshop on Content-Based Multimedia Indexing (CBMI), Klagenfurt, Austria, 18–20 June 2014; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2014; pp. 1–6. [Google Scholar]
- Zhang, X.; Liu, W.; Dundar, M.; Badve, S.; Zhang, S. Towards Large-Scale Histopathological Image Analysis: Hashing-Based Image Retrieval. IEEE Trans. Med. Imaging 2014, 34, 496–506. [Google Scholar] [CrossRef]
- Flahou, B.; Haesebrouck, F.; Smet, A. Non-Helicobacter pylori Helicobacter Infections in Humans and Animals. In Helicobacter Pylori Research; Springer: Tokyo, Japan, 2016; pp. 233–269. [Google Scholar] [CrossRef]
- Weinstein, J.N.; Collisson, E.A.; Mills, G.B.; Shaw, K.R.M.; Ozenberger, B.A.; Ellrott, K.; Shmulevich, I.; Sander, C.; Stuart, J.M. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 2013, 45, 1113–1120. [Google Scholar] [CrossRef]
- Molin, M.D.; Matthaei, H.; Wu, J.; Blackford, A.; Debeljak, M.; Rezaee, N.; Wolfgang, C.L.; Butturini, G.; Salvia, R.; Bassi, C.; et al. Clinicopathological Correlates of Activating GNAS Mutations in Intraductal Papillary Mucinous Neoplasm (IPMN) of the Pancreas. Ann. Surg. Oncol. 2013, 20, 3802–3808. [Google Scholar] [CrossRef]
- Yoshida, A.; Tsuta, K.; Nakamura, H.; Kohno, T.; Takahashi, F.; Asamura, H.; Sekine, I.; Fukayama, M.; Shibata, T.; Furuta, K.; et al. Comprehensive Histologic Analysis of ALK-Rearranged Lung Carcinomas. Am. J. Surg. Pathol. 2011, 35, 1226–1234. [Google Scholar] [CrossRef]
- Arevalo, J.; Cruz-Roa, A. Histopathology image representation for automatic analysis: A state-of-the-art review. Rev. Med. 2014, 22, 79–91. [Google Scholar] [CrossRef]
- Lyon, H.O.; de Leenheer, A.P.; Horobin, R.W.; Lambert, W.E.; Schulte, E.K.W.; van Liedekerke, B.; Wittekind, D.H. Standardization of reagents and methods used in cytological and histological practice with emphasis on dyes, stains and chromogenic reagents. J. Mol. Histol. 1994, 26, 533–544. [Google Scholar] [CrossRef]
- Khan, A.M.; Rajpoot, N.; Treanor, D.; Magee, D. A Nonlinear Mapping Approach to Stain Normalization in Digital Histopathology Images Using Image-Specific Color Deconvolution. IEEE Trans. Biomed. Eng. 2014, 61, 1729–1738. [Google Scholar] [CrossRef]
- Anghel, A.; Stanisavljevic, M.; Andani, S.; Papandreou, N.; Rüschoff, J.H.; Wild, P.; Gabrani, M.; Pozidis, H. A High-Performance System for Robust Stain Normalization of Whole-Slide Images in Histopathology. Front. Med. 2019, 6. [Google Scholar] [CrossRef]
- Can, A.; Bello, M.; Cline, H.E.; Tao, X.; Ginty, F.; Sood, A.; Gerdes, M.; Montalto, M. Multi-modal imaging of histological tissue sections. In Proceedings of the 2008 5th IEEE International Symposium on Biomedical Imaging: From Nano to Macro, Paris, France, 14–18 May 2008; IEEE: Piscataway, NJ, USA, 2008; pp. 288–291. [Google Scholar] [CrossRef]
- Casiraghi, E.; Cossa, M.; Huber, V.; Tozzi, M.; Rivoltini, L.; Villa, A.; Vergani, B. MIAQuant, a novel system for automatic segmentation, measurement, and localization comparison of different biomarkers from serialized histological slices. Eur. J. Histochem. 2017, 61, 61. [Google Scholar] [CrossRef]
- Casiraghi, E.; Huber, V.; Frasca, M.; Cossa, M.; Tozzi, M.; Rivoltini, L.; Leone, B.E.; Villa, A.; Vergani, B. A novel computational method for automatic segmentation, quantification and comparative analysis of immunohistochemically labeled tissue sections. BMC Bioinform. 2018, 19, 357–397. [Google Scholar] [CrossRef]
- Roy, S.; Jain, A.K.; Lal, S.; Kini, J. A study about color normalization methods for histopathology images. Micron 2018, 114, 42–61. [Google Scholar] [CrossRef]
- Mărginean, R.; Andreica, A.; Dioşan, L.; Bálint, Z. Feasibility of Automatic Seed Generation Applied to Cardiac MRI Image Analysis. Mathematics 2020, 8, 1511. [Google Scholar] [CrossRef]
- Gleason, D.F. Histologic grading of prostate cancer: A perspective. Hum. Pathol. 1992, 23, 273–279. [Google Scholar] [CrossRef]
- Kong, J.; Sertel, O.; Shimada, H.; Boyer, K.L.; Saltz, J.H.; Gurcan, M.N. Computer-aided evaluation of neuroblastoma on whole-slide histology images: Classifying grade of neuroblastic differentiation. Pattern Recognit. 2009, 42, 1080–1092. [Google Scholar] [CrossRef]
- Washington, K.; Berlin, J.; Branton, P.; Burgart, L.J.; Carter, D.K.; Fitzgibbons, P.L.; Halling, K.; Frankel, W.; Jessup, J.; Kakar, S.; et al. Protocol for the examination of specimens from patients with primary carcinoma of the colon and rectum. Arch. Pathol. Lab. Med. 2009, 133, 1539–1551. [Google Scholar] [CrossRef]
- Irshad, H.; Veillard, A.; Roux, L.; Racoceanu, D. Methods for Nuclei Detection, Segmentation, and Classification in Digital Histopathology: A Review—Current Status and Future Potential. IEEE Rev. Biomed. Eng. 2013, 7, 97–114. [Google Scholar] [CrossRef]
- Dalle, J.-R.; Li, H.; Huang, C.-H.; Leow, W.K.; Racoceanu, D.; Putti, T. Nuclear pleomorphism scoring by selective cell nuclei detection. In Proceedings of the Workshop on Applications of Computer Vision, Snowbird, UT, USA, 7–8 December 2009. [Google Scholar]
- Wahlby, C.; Sintorn, I.-M.; Erlandsson, F.; Borgefors, G.; Bengtsson, E. Combining intensity, edge and shape information for 2D and 3D segmentation of cell nuclei in tissue sections. J. Microsc. 2004, 215, 67–76. [Google Scholar] [CrossRef]
- Jung, C.; Kim, C. Segmenting Clustered Nuclei Using H-minima Transform-Based Marker Extraction and Contour Parameterization. IEEE Trans. Biomed. Eng. 2010, 57, 2600–2604. [Google Scholar] [CrossRef]
- Cosatto, E.; Miller, M.; Graf, H.P.; Meyer, J.S. Grading Nuclear Pleomorphism on Histological Micrographs. In Proceedings of the 2008 19th International Conference on Pattern Recognition, Tampa, FL, USA, 8–11 December 2008; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2008; pp. 1–4. [Google Scholar]
- Al-Kofahi, Y.; Lassoued, W.; Lee, W.; Roysam, B. Improved Automatic Detection and Segmentation of Cell Nuclei in Histopathology Images. IEEE Trans. Biomed. Eng. 2009, 57, 841–852. [Google Scholar] [CrossRef]
- Veta, M.; Huisman, A.; Viergever, M.; van Diest, P.J.; Pluim, J. Marker-controlled watershed segmentation of nuclei in H&E stained breast cancer biopsy images. In Proceedings of the IEEE International Symposium on Biomedical Imaging: From Nano to Macro, Chicago, IL, USA, 30 March–2 April 2011; IEEE: Piscataway, NJ, USA, 2011; pp. 618–621. [Google Scholar] [CrossRef]
- Aptoula, E.; Courty, N.; Lefèvre, S. Mitosis detection in breast cancer histological images with mathematical morphology. In Proceedings of the 2013 21st Signal Processing and Communications Applications Conference (SIU), Haspolat, Turkey, 24–26 April 2013; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2013; pp. 1–4. [Google Scholar]
- Ciresan, D.C.; Giusti, A.; Gambardella, L.M.; Schmidhuber, J. Mitosis Detection in Breast Cancer Histology Images with Deep Neural Networks. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Nagoya, Japan, 22–26 September 2013; Springer: Berlin/Heidelberg, Germany, 2013; Volume 2, pp. 411–418. [Google Scholar]
- Petushi, S.; Garcia, F.U.; Haber, M.M.; Katsinis, C.; Tozeren, A. Large-scale computations on histology images reveal grade-differentiating parameters for breast cancer. BMC Med Imaging 2006, 6, 14. [Google Scholar] [CrossRef]
- Rittscher, J.; Machiraju, R.; Wong, S. Microscopic Image Analysis for Life Science Applications; Artech House: Norwood, MA, USA, 2008. [Google Scholar]
- Boucheron, L.E. Object- and Spatial-Level Quantitative Analysis of Multispectral Histopathology Images for Detection and Characterization of Cancer; University of California at Santa Barbara: Santa Barbara, CA, USA, 2008. [Google Scholar]
- Kuse, M.; Sharma, T.; Gupta, S. A Classification Scheme for Lymphocyte Segmentation in H&E Stained Histology Images. In Static Analysis; Springer Science and Business Media LLC: Berlin/Heidelberg, Germany, 2010; pp. 235–243. [Google Scholar]
- Chekkoury, A.; Khurd, P.; Ni, J.; Bahlmann, C.; Kamen, A.; Patel, A.; Grady, L.; Singh, M.; Groher, M.; Navab, N.; et al. Automated malignancy detection in breast histopathological images. In Medical Imaging 2012: Computer-Aided Diagnosis; SPIE: Bellingham, WA, USA, 2012; pp. 332–344. [Google Scholar]
- Doyle, S.; Hwang, M.; Shah, K.; Madabhushi, A.; Feldman, M.D.; Tomaszeweski, J.E. Automated Grading of Prostate Cancer Using Architectural and Textural Image Features. In Proceedings of the 2007 4th IEEE International Symposium on Biomedical Imaging: From Nano to Macro, Arlington, VA, USA, 12–15 April 2007; IEEE: Piscataway, NJ, USA, 2007; pp. 1284–1287. [Google Scholar]
- Di Franco, M.D.; O’Hurley, G.; Kay, E.W.; Watson, R.W.G.; Cunningham, P. Ensemble-based system for whole-slide prostate cancer probability mapping using color texture features. Comput. Med. Imaging Graphics 2011, 35, 629–645. [Google Scholar] [CrossRef]
- Huang, P.-W.; Lai, Y.H. Effective segmentation and classification for HCC biopsy images. Pattern Recognit. 2010, 43, 1550–1563. [Google Scholar] [CrossRef]
- Alexandratou, E.; Atlamazoglou, V.; Thireou, T.; Agrogiannis, G.; Togas, D.; Kavantzas, N.; Patsouris, E.; Yova, D. Evaluation of machine learning techniques for prostate cancer diagnosis and Gleason grading. Int. J. Comput. Intell. Bioinform. Syst. Biol. 2010, 1, 297. [Google Scholar] [CrossRef]
- Basavanhally, A.; Yu, E.; Xu, J.; Ganesan, S.; Feldman, M.; Tomaszeweski, J.E.; Madabhushi, A. Incorporating domain knowledge for tubule detection in breast histopathology using O’Callaghan neighborhoods. Inter. Soc. Optics Photonics 2011, 7963, 796310. [Google Scholar]
- Al-Kadi, O.S. Texture measures combination for improved meningioma classification of histopathological images. Pattern Recognit. 2010, 43, 2043–2053. [Google Scholar] [CrossRef]
- Demir, C.; Kandemir, M.; Tosun, A.B.; Sokmensuer, C. Automatic segmentation of colon glands using object-graphs. Med. Image Anal. 2010, 14, 1–12. [Google Scholar] [CrossRef]
- Tosun, A.B.; Demir, C. Graph Run-Length Matrices for Histopathological Image Segmentation. IEEE Transact. Med. Imaging 2011, 30, 721–732. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet classification with deep convolutional neural networks. In Proceedings of the Advances in Neural Information Processing Systems (NIPS), Lake Tahoe, NV, USA, 3–6 December 2012; pp. 1097–1105. [Google Scholar]
- Le Cun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef]
- Lecun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef]
- Arevalo, J.; Cruz-Roa, A.; González, F.A. Hybrid image representation learning model with invariant features for basal cell carcinoma detection. In Proceedings of the IX International Seminar on Medical Information Processing and Analysis, Mexico City, Mexico, 11–14 November2013; SPIE: Bellingham, WA, USA, 2013; Volume 8922, p. 89220M. [Google Scholar]
- Nayak, N.; Chang, H.; Borowsky, A.; Spellman, P.T.; Parvin, B. Classification of tumor histopathology via sparse feature learning. In Proceedings of the 2013 IEEE 10th International Symposium on Biomedical Imaging, San Francisco, CA, USA, 7–11 April 2013; IEEE: Piscataway, NJ, USA, 2013; pp. 410–413. [Google Scholar]
- Malon, C.D.; Cosatto, E. Classification of mitotic figures with convolutional neural networks and seeded blob features. J. Pathol. Informat. 2013, 4, 9. [Google Scholar] [CrossRef]
- Xu, Y.; Mo, T.; Feng, Q.; Zhong, P.; Lai, M.; Chang, E.I.-C. Deep learning of feature representation with multiple instance learning for medical image analysis. In Proceedings of the 2014 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), Florence, Italy, 4–9 May 2014; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2014; pp. 1626–1630. [Google Scholar]
- Hou, L.; Samaras, D.; Kurç, T.M.; Gao, Y.; Davis, J.E.; Saltz, J.H. Efficient Multiple Instance Convolutional Neural Networks for Gigapixel Resolution Image Classification. arXiv 2015, arXiv:abs/1504.07947. [Google Scholar]
- Arevalo, J.; Cruz-Roa, A.; Arias, V.; Romero, E.; González, F.A. An unsupervised feature learning framework for basal cell carcinoma image analysis. Artif. Intell. Med. 2015, 64, 131–145. [Google Scholar] [CrossRef]
- Chang, H.; Zhou, Y.; Borowsky, A.; Barner, K.; Spellman, P.; Parvin, B. Stacked Predictive Sparse Decomposition for Classification of Histology Sections. Int. J. Comput. Vis. 2014, 113, 3–18. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Han, J.; Fontenay, G.V.; Wang, Y.; Mao, J.-H.; Chang, H. Phenotypic characterization of breast invasive carcinoma via transferable tissue morphometric patterns learned from glioblastoma multiforme. Proceeding of the 2016 IEEE 13th International Symposium on Biomedical Imaging (ISBI), Prague, Czech Republic, 13–16 April 2016; IEEE: Piscataway, NJ, USA, 2016; pp. 1025–1028. [Google Scholar]
- Noël, H.; Roux, L.; Lu, S.; Boudier, T. Detection of high-grade atypia nuclei in breast cancer imaging. In Medical Imaging; SPIE: Bellingham, WA, USA, 2015; p. 94200R. [Google Scholar]
- Romo-Bucheli, D.; Janowczyk, A.; Gilmore, H.; Romero, E.; Madabhushi, A. Automated Tubule Nuclei Quantification and Correlation with Oncotype DX risk categories in ER+ Breast Cancer Whole Slide Images. Sci. Rep. 2016, 6, 32706. [Google Scholar] [CrossRef] [PubMed]
- Chen, T. Deep Learning Based Automatic Immune Cell Detection for Immunohistochemistry Images. In International Workshop on Machine Learning in Medical Imaging; Springer: Cham, Switzerlnad, 2014; pp. 17–24. [Google Scholar]
- Srinidhi, C.L.; Ciga, O.; Martel, A.L. Deep neural network models for computational histopathology: A survey. Med Image Anal. 2020, 101813. [Google Scholar] [CrossRef] [PubMed]
- Sumi, P.S.; Delhibabu, R. Glioblastoma Multiforme Classification On High Resolution Histology Image Using Deep Spatial Fusion Network. In Proceedings of theCEUR Workshop, Como, Italy, 9–11 September 2019. [Google Scholar]
- Zhang, L.; Wu, Y.; Zheng, B.; Su, L.; Chen, Y.; Ma, S.; Hu, Q.; Zou, X.; Yao, L.; Yang, Y.; et al. Rapid histology of laryngeal squamous cell carcinoma with deep-learning based stimulated Raman scattering microscopy. Theranostics 2019, 9, 2541–2554. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, A. GPU Based Digital Histopathology and Diagnostic Support System for Breast Cancer Detection: A Comparison of CNN Models and Machine Learning Models. Nature Rev. Drug Discov. 2019, 18, 463–477. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015; pp. 234–241. [Google Scholar]
- Tschuchnig, M.E.; Oostingh, G.J.; Gadermayr, M. Generative Adversarial Networks in Digital Pathology: A Survey on Trends and Future Potential. arXiv 2020, arXiv:abs/2004.14936. [Google Scholar] [CrossRef]
- Litjens, G.J.S.; Sánchez, C.I.; Timofeeva, N.; Hermsen, M.; Nagtegaal, I.D.; Kovacs, I.; Kaa, C.H.; van de Bult, P.; Ginneken, B.; van Laak, J. Deep learning as a tool for increased accuracy and efficiency of histopathological diagnosis. Sci. Reports 2016, 6. [Google Scholar] [CrossRef]
- Nagpal, K.; Foote, D.; Liu, Y.; Chen, P.-H.C.; Wulczyn, E.; Tan, F.; Olson, N.; Smith, J.L.; Mohtashamian, A.; Wren, J.H.; et al. Publisher Correction: Development and validation of a deep learning algorithm for improving Gleason scoring of prostate cancer. npj Digit. Med. 2019, 2, 2. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Lin, H.; Chen, H.; Heng, P.-A. PFA-ScanNet: Pyramidal Feature Aggregation with Synergistic Learning for Breast Cancer Metastasis Analysis. Lecture Notes Comput. Sci. 2019, 586–594. [Google Scholar]
- Xing, F.; Xie, Y.; Yang, L. An Automatic Learning-Based Framework for Robust Nucleus Segmentation. IEEE Trans. Med Imaging 2015, 35, 550–566. [Google Scholar] [CrossRef]
- Gu, F.; Burlutskiy, N.; Andersson, M.; Wilén, L.K. Multi-resolution Networks for Semantic Segmentation in Whole Slide Images. In Lecture Notes in Computer Science; Springer Science and Business Media LLC: Berlin/Heidelberg, Germany, 2018; pp. 11–18. [Google Scholar]
- Tellez, D.; Balkenhol, M.; Otte-Holler, I.; van de Loo, R.; Vogels, R.; Bult, P.; Wauters, C.; Vreuls, W.; Mol, S.; Karssemeijer, N.; et al. Whole-Slide Mitosis Detection in H&E Breast Histology Using PHH3 as a Reference to Train Distilled Stain-Invariant Convolutional Networks. IEEE Trans. Med Imaging 2018, 37, 2126–2136. [Google Scholar] [CrossRef]
- Wei, J.W.; Tafe, L.J.; Linnik, Y.A.; Vaickus, L.J.; Tomita, N.; Hassanpour, S. Pathologist-level classification of histologic patterns on resected lung adenocarcinoma slides with deep neural networks. Sci. Rep. 2019, 9, 1–8. [Google Scholar] [CrossRef]
- Song, Y.; Tan, E.-L.; Jiang, X.; Cheng, J.-Z.; Ni, D.; Chen, S.; Lei, B.Y.; Wang, T. Accurate Cervical Cell Segmentation from Overlapping Clumps in Pap Smear Images. IEEE Transact. Med. Imaging 2017, 36, 288–300. [Google Scholar] [CrossRef]
- Agarwalla, A.; Shaban, M.; Rajpoot, N.M. Representation-Aggregation Networks for Segmentation of Multi-Gigapixel Histology Images. arXiv 2017, arXiv:1707-08814. [Google Scholar]
- Ding, H.; Pan, Z.; Cen, Q.; Li, Y.; Chen, S. Multi-scale fully convolutional network for gland segmentation using three-class classification. Neurocomputing 2020, 380, 150–161. [Google Scholar] [CrossRef]
- Bejnordi, B.E.; Zuidhof, G.C.A.; Balkenhol, M.; Hermsen, M.; Bult, P.; Ginneken, B.; van Karssemeijer, N.; Litjens, G.J.S.; Laak, J. Context-aware stacked convolutional neural networks for classification of breast carcinomas in whole-slide histopathology images. J. Med. Imaging 2017, 4, 044504. [Google Scholar] [CrossRef]
- Seth, N.; Akbar, S.; Nofech-Mozes, S.; Salama, S.; Martel, A.L. Automated Segmentation of DCIS in Whole Slide Images. In Case-Based Reasoning Research and Development; Springer Science and Business Media LLC: Berlin/Heidelberg, Germany, 2019; pp. 67–74. [Google Scholar]
- Xu, J.; Xiang, L.; Liu, Q.; Gilmore, H.; Wu, J.; Tang, J.; Madabhushi, A. Stacked Sparse Autoencoder (SSAE) for Nuclei Detection on Breast Cancer Histopathology Images. IEEE Trans. Med Imaging 2015, 35, 119–130. [Google Scholar] [CrossRef]
- Bulten, W.; Litjens, G. Unsupervised Prostate Cancer Detection on H&E using Convolutional Adversarial Autoencoders. arXiv 2018, arXiv:1804.07098. [Google Scholar]
- Hou, L.; Agarwal, A.; Samaras, D.; Kurc, T.M.; Gupta, R.R.; Saltz, J.H. Robust Histopathology Image Analysis: To Label or to Synthesize? In Proceedings of the 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Long Beach, CA, USA, 16–20 June 2019; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2019; pp. 8525–8534. [Google Scholar]
- Sari, C.T.; Gunduz-Demir, C. Unsupervised Feature Extraction via Deep Learning for Histopathological Classification of Colon Tissue Images. IEEE Transact. Med. Imaging 2019, 38, 1139–1149. [Google Scholar] [CrossRef] [PubMed]
- Gadermayr, M.; Gupta, L.; Appel, V.; Boor, P.; Klinkhammer, B.M.; Merhof, D. Generative Adversarial Networks for Facilitating Stain-Independent Supervised and Unsupervised Segmentation: A Study on Kidney Histology. IEEE Trans. Med Imaging 2019, 38, 2293–2302. [Google Scholar] [CrossRef] [PubMed]
- Gadermayr, M.; Gupta, L.; Klinkhammer, B.M.; Boor, P.; Merhof, D. Unsupervisedly Training GANs for Segmenting Digital Pathology with Automatically Generated Annotations. arXiv 2018, arXiv:1805.10059. [Google Scholar]
- Komura, D.; Ishikawa, S. Machine Learning Methods for Histopathological Image Analysis. Comput. Struct. Biotechnol. J. 2018, 16, 34–42. [Google Scholar] [CrossRef]
- Search Home—Biospecimen Research Database. Available online: https://brd.nci.nih.gov/brd/image-search/searchhome (accessed on 1 October 2020).
- TMAD Main Menu. Available online: https://tma.im/cgi-bin/home.pl (accessed on 1 October 2020).
- Home—CAMELYON17—Grand Challenge. Available online: https://camelyon17.grand-challenge.org/ (accessed on 1 October 2020).
- Breast Cancer Histopathological Database (BreakHis)—Laboratório Visão Robótica e Imagem. Available online: https://web.inf.ufpr.br/vri/databases/breast-cancer-histopathological-database-breakhis/ (accessed on 1 October 2020).
- Search GDC. Available online: https://portal.gdc.cancer.gov/legacy-archive/search/f (accessed on 1 October 2020).
- Marinelli, R.J.; Montgomery, K.; Liu, C.L.; Shah, N.H.; Prapong, W.; Nitzberg, M.; Zachariah, Z.K.; Sherlock, G.; Natkunam, Y.; West, R.B.; et al. The Stanford Tissue Microarray Database. Nucleic Acids Res. 2007, 36, D871–D877. [Google Scholar] [CrossRef]
- Dataset Tumor Proliferation Assessment Challenge 2016. Available online: http://tupac.tue-image.nl/node/3 (accessed on 1 October 2020).
- Bentaieb, A.; Li-Chang, H.; Huntsman, D.; Hamarneh, G. A structured latent model for ovarian carcinoma subtyping from histopathology slides. Med Image Anal. 2017, 39, 194–205. [Google Scholar] [CrossRef]
- Ovarian Carcinomas Histopathology Dataset. Available online: http://ensc-mica-www02.ensc.sfu.ca/download/ (accessed on 1 October 2020).
- Babaie, M.; Kalra, S.; Sriram, A.; Mitcheltree, C.; Zhu, S.; Khatami, A.; Rahnamayan, S.; Tizhoosh, H.R. Classification and Retrieval of Digital Pathology Scans: A New Dataset. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), Honolulu, HI, USA, 21–16 July 2017; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2017; pp. 760–768. [Google Scholar]
- Kumar, N.; Verma, R.; Sharma, S.; Bhargava, S.; Vahadane, A.; Sethi, A. A Dataset and a Technique for Generalized Nuclear Segmentation for Computational Pathology. IEEE Trans. Med Imaging 2017, 36, 1550–1560. [Google Scholar] [CrossRef]
- Pathology Images: KIMIA Path960—Kimia Lab. Available online: https://kimialab.uwaterloo.ca/kimia/index.php/pathology-images-kimia-path960/ (accessed on 1 October 2020).
- Gelasca, E.D.; Byun, J.; Obara, B.; Manjunath, B.S. Evaluation and benchmark for biological image segmentation. In Proceedings of the 2008 15th IEEE International Conference on Image Processing, San Diego, CA, USA, 12–15 October 2008; Institute of Electrical and Electronics Engineers (IEEE): Pisctaway, NJ, USA, 2008; pp. 1816–1819. [Google Scholar]
- Bio-Segmentation Center for Bio-Image Informatics UC Santa Barbara. Available online: https://bioimage.ucsb.edu/research/bio-segmentation (accessed on 1 October 2020).
- Bioimaging Challenge 2015 Breast Histology Dataset—Datasets CKAN. Available online: https://rdm.inesctec.pt/dataset/nis-2017-003 (accessed on 1 October 2020).
- BIALab@Warwick: GlaS Challenge Contest. Available online: https://warwick.ac.uk/fac/sci/dcs/research/tia/glascontest/ (accessed on 1 October 2020).
- Kather, J.N.; Marx, A.; Reyes-Aldasoro, C.C.; Schad, L.R.; Zoellner, F.G.; Weis, C.-A. Continuous representation of tumor microvessel density and detection of angiogenic hotspots in histological whole-slide images. Oncotarget 2015, 6, 19163–19176. [Google Scholar] [CrossRef]
- Dataset—MITOS-ATYPIA-14—Grand Challenge. Available online: https://mitos-atypia-14.grand-challenge.org/dataset/ (accessed on 1 October 2020).
- Nucleisegmentation. Available online: https://nucleisegmentationbenchmark.weebly.com/ (accessed on 1 October 2020).
- Janowczyk, A.; Madabhushi, A. Deep learning for digital pathology image analysis: A comprehensive tutorial with selected use cases. J. Pathol. Informat. 2016, 7. [Google Scholar] [CrossRef]
- Andrew Janowczyk—Tidbits from Along the Way. Available online: http://www.andrewjanowczyk.com/ (accessed on 1 October 2020).
- Ma, Z.; Shiao, S.L.; Yoshida, E.J.; Swartwood, S.; Huang, F.; Doche, M.E.; Chung, A.P.; Knudsen, B.S.; Gertych, A. Data integration from pathology slides for quantitative imaging of multiple cell types within the tumor immune cell infiltrate. Diagn. Pathol. 2017, 12. [Google Scholar] [CrossRef] [PubMed]
- Linder, N.; Konsti, J.; Turkki, R.; Rahtu, E.; Lundin, M.; Nordling, S.; Haglund, C.; Ahonen, T.; Pietikäinen, M.; Lundin, J. Identification of tumor epithelium and stroma in tissue microarrays using texture analysis. Diagn. Pathol. 2012, 7, 22. [Google Scholar] [CrossRef] [PubMed]
- Egfr Colon Stroma Classification. Available online: http://fimm.webmicroscope.net/supplements/epistroma (accessed on 1 October 2020).
- Jimenez-del-Toro, O.; Otálora, S.; Andersson, M.; Eurén, K.; Hedlund, M.; Rousson, M.; Müller, H.; Atzori, M. Chapter 10—Analysis of Histopathology Images: From Traditional Machine Learning to Deep Learning; Academic Press: Cambridge, MA, USA, 2017; pp. 281–314. [Google Scholar]
- Zhang, Y.; Zhang, B.; Coenen, F.; Xiao, J.; Lu, W. One-class kernel subspace ensemble for medical image classification. EURASIP J. Adv. Signal Process. 2014, 2014, 17. [Google Scholar] [CrossRef]
- Xia, Y.; Cao, X.; Wen, F.; Hua, G.; Sun, J. Learning Discriminative Reconstructions for Unsupervised Outlier Removal. In Proceedings of the 2015 IEEE International Conference on Computer Vision (ICCV), Las Condes, Chile, 11–15 December 2015; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2015; pp. 1511–1519. [Google Scholar]
- Samek, W.; Binder, A.; Montavon, G.; Lapuschkin, S.; Muller, K.-R. Evaluating the Visualization of What a Deep Neural Network Has Learned. IEEE Trans. Neural Networks Learn. Syst. 2016, 28, 2660–2673. [Google Scholar] [CrossRef] [PubMed]
- Zintgraf, L.M.; Cohen, T.S.; Adel, T.; Welling, M. Visualizing Deep Neural Network Decisions: Prediction Difference Analysis. arXiv 2017, arXiv:1702.04595. [Google Scholar]
- Koh, P.W.; Liang, P. Understanding Black-box Predictions via Influence Functions. arXiv 2017, arXiv:abs/1703.04730. [Google Scholar]
- Abas, F.S.; Gokozan, H.; Goksel, B.; Otero, J.J. Intraoperative Neuropathology of Glioma Recurrence: Cell Detection and Classification; SPIE: Bellingham, WA, USA, 2016; p. 979109. [Google Scholar] [CrossRef]
- Chen, J.; Srinivas, C. Automatic Lymphocyte Detection in H&E Images with Deep Neural Networks. arXiv 2016, arXiv:abs/1612.03217. [Google Scholar]
- Turkki, R.; Linder, N.; Kovanen, P.E.; Pellinen, T.; Lundin, J. Antibody-supervised deep learning for quantification of tumor-infiltrating immune cells in hematoxylin and eosin stained breast cancer samples. J. Pathol. Informat. 2016, 7, 38. [Google Scholar] [CrossRef]
- Feng, Z.; Bethmann, D.; Kappler, M.; Ballesteros-Merino, C.; Eckert, A.; Bell, R.B.; Cheng, A.; Bui, T.; Leidner, R.; Urba, W.J.; et al. Multiparametric immune profiling in HPV—Oral squamous cell cancer. JCI Insight 2017, 2, e93652. [Google Scholar] [CrossRef]
- Basavanhally, A.N.; Ganesan, S.; Agner, S.; Monaco, J.P.; Feldman, M.D.; Tomaszewski, J.E.; Bhanot, G.; Madabhushi, A. Computerized Image-Based Detection and Grading of Lymphocytic Infiltration in HER2+ Breast Cancer Histopathology. IEEE Trans. Biomed. Eng. 2009, 57, 642–653. [Google Scholar] [CrossRef]
- Li, J.; Li, W.; Gertych, A.; Knudsen, B.S.; Speier, W.; Arnold, C. An attention-based multi-resolution model for prostate whole slide imageclassification and localization. arXiv 2019, arXiv:1905.13208. [Google Scholar]
- Bera, K.; Schalper, K.A.; Rimm, D.L.; Velcheti, V.; Madabhushi, A. Artificial intelligence in digital pathology new tools for diagnosis and precision oncology. Nat. Rev. Clin. Oncol. 2019, 16, 703–715. [Google Scholar] [CrossRef] [PubMed]
- Niazi, M.K.K.; Parwani, A.V.; Gurcan, M.N. Digital pathology and artificial intelligence. Lancet Oncol. 2019, 20, 253–261. [Google Scholar] [CrossRef]
- Tellez, D.; Litjens, G.J.S.; Bándi, P.; Bulten, W.; Bokhorst, J.-M.; Ciompi, F.; Laak, J. Quantifying the effects of data augmentation and stain color normalization in convolutional neural networks for computational pathology. Med. Image Anal. 2019, 58, 101544. [Google Scholar] [CrossRef] [PubMed]
- Steiner, D.F.; Macdonald, R.; Liu, Y.; Truszkowski, P.; Hipp, J.D.; Gammage, C.; Thng, F.; Peng, L.; Stumpe, M.C. Impact of Deep Learning Assistance on the Histopathologic Review of Lymph Nodes for Metastatic Breast Cancer. Am. J. Surg. Pathol. 2018, 42, 1636–1646. [Google Scholar] [CrossRef]
- Kamnitsas, K.; Ferrante, E.; Parisot, S.; Ledig, C.; Nori, A.V.; Criminisi, A.; Rueckert, D.; Glocker, B. DeepMedic for Brain Tumor Segmentation; Springer: Berlin/Heidelberg, Germany, 2016. [Google Scholar]
- Voets, M.; Møllersen, K.; Bongo, L.A. Replication study: Development and validation of deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. arXiv 2018, arXiv:abs/1803.04337. [Google Scholar]


| Study | Organ | Method | Results | Problem with Method |
|---|---|---|---|---|
| Basavanhallya et al. [66] | Breast | Hierarchical Normalized cut | 89% accuracy of segmentation | Discovers false-positive errors because of lumen presence. |
| Khadi [67] | Meningioma | Classification of texture applying fractal features | 92.5% accuracy applying individual texture measurement to meningioma tissue | Misclassification results because of the non-uniformity cell construct. |
| Demir et al. [68] | Colon | Object graph approach for segmentation | 87.59% accuracy of compatible images | Requires variable optimization that reduced results for segmentation |
| Tosum et al. [69] | Breast | Diagram run-length models to segmentation of the image | The novel descriptor of texture for unsupervised classification had 99.0% accuracy for gland segmentation | Complexity relies on the number of primitives in picture |
| Chekkoury et al. [61] | Breast | A novel hybrid between features based on texton and morphologic system | 86% classification accuracy combining textural features and morphometric | The effects of image compression on classification accuracy |
| Doyle et al. [62] | Prostate | mix of characteristics | 90% accuracy in characterizing cancer and benign tissue | Limited Feature Set |
| DiFranco et al. [63] | Prostate | classified into seven classes (benign hyperplasia, inflammation, Gleason rank 3,4, and intraepithelial neoplasia) | 83% median accuracy based on some characteristics like sum average, contrast | The computation time required for reading and writing data to and from disk, particularly in feature extraction |
| Study | Organ | Staining | Potential Usage | Method |
|---|---|---|---|---|
| Supervised Learning | ||||
| Litjens et al. [90] | different tissue | H&E | Prostate and breast carcinoma detection | Convolutional Neural Network based on pixel classifier |
| Nagpal et al. [91] | Prostate | H&E | Anticipating Gleason indicator | CNN based on sectional Gleason model classifier + k-nearst neighbors (KNN) based on Gleason grade anticipation |
| Zhao et al. [92] | Breast | H&E | Metastasis Detection + classification | Characteristic pyramid collecting based on the fully convolutional network (FCN) system with the synergistic training technique |
| Xing et al. [93] | different tissue | H&E, Immunohistochemistry (IHC) | Segmentation of nuclei | CNN + selection based on sparse form Pattern |
| Gu et al. [94] | Breast | H&E | Tumor detection | U-Net based on multiple resolution model with multiple encoders and a singular decoder system |
| Tellez et al. [95] | Breast | H&E | Detection of Mitosis | Train of Convolutional Network applying H&E registered to PHH3 slides as a reference |
| Wei et al. [96] | Lung | H&E | Histological subtypes of lung gland classifier | ResNet-18 on the basis of patch classification |
| Song et al. [97] | Cervix | Papanicolaou (Pap), H&E | Cells Segmentation | Multiple level CNN system |
| Agarwalla et al. [98] | Breast | H&E | Segmentation of tumor | CNN and 2D- Long short-term memory (LSTM) to representing training and context collecting |
| Ding et al. [99] | Colon | H&E | Glands segmentation | Multiple level FCN network with a high-resolution section to avoid the lost in highest pooling layers |
| Bejnordi et al. [100] | Breast | H&E | Invasive Carcinoma detection | Multiple level CNN which first determines tumor-associated stromal modifications and more categorize into normal/benign versus invasive carcinoma |
| Seth et al. [101] | Breast | H&E | Ductal carcinoma in-situ (DCIS) segmentation | Compared UNets learned in many resolutions |
| Unsupervised Learning | ||||
| Xu et al. [102] | Breast | H&E | Segmentation of nuclei | Stacked sparse autoencoders |
| Bulten and Litjens [103] | Prostate | H&E | Tumor classification | Convolutional adversarial Autoencoders |
| Hou et al. [104] | Breast | H&E | Segmentation and detection of nuclei | Sparse autoencoder |
| Sari and Gunduz-Demir [105] | Colon | H&E | Feature extraction and classification | Restricted Boltzmann + clustering |
| Gadermayr et al. [106] | Kidney | Stain agnostic | Object of interest segmentation in WSIs | CycleGAN + UNet segmentation |
| Gadermayr et al. [107] | Kidney | Periodic acid–Schiff (PAS), H&E | Glomeruli segmentation | CycleGAN |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Elazab, N.; Soliman, H.; El-Sappagh, S.; Islam, S.M.R.; Elmogy, M. Objective Diagnosis for Histopathological Images Based on Machine Learning Techniques: Classical Approaches and New Trends. Mathematics 2020, 8, 1863. https://doi.org/10.3390/math8111863
Elazab N, Soliman H, El-Sappagh S, Islam SMR, Elmogy M. Objective Diagnosis for Histopathological Images Based on Machine Learning Techniques: Classical Approaches and New Trends. Mathematics. 2020; 8(11):1863. https://doi.org/10.3390/math8111863
Chicago/Turabian StyleElazab, Naira, Hassan Soliman, Shaker El-Sappagh, S. M. Riazul Islam, and Mohammed Elmogy. 2020. "Objective Diagnosis for Histopathological Images Based on Machine Learning Techniques: Classical Approaches and New Trends" Mathematics 8, no. 11: 1863. https://doi.org/10.3390/math8111863
APA StyleElazab, N., Soliman, H., El-Sappagh, S., Islam, S. M. R., & Elmogy, M. (2020). Objective Diagnosis for Histopathological Images Based on Machine Learning Techniques: Classical Approaches and New Trends. Mathematics, 8(11), 1863. https://doi.org/10.3390/math8111863







