1. Introduction
Consider a finite graph consisting of vertices , and edges E. Throughout this paper, a graph is undirected and connected. The neighbour of the vertex is denoted by , where means that i and j are adjacent. The degree of the vertex i is denoted by , which is the cardinality of the set . An independent set of is a subset of V such that no two are adjacent. In other words, a set of vertices is independent if and only if it is a clique in the graph complement of .
If two vertices of a graph have precisely the same neighbour, throughout this paper, we call the graph obtained by identifying these two vertices with keeping the adjacency a reduced graph of .
Let
be the totality of probability measures on the simplex
equipped with the topology of weak convergence. Itoh et al. [
1] discussed a diffusion-taking value in probability measures on graph
, whose state is identified with probability measure
and the masses on each vertex
which starts from
and satisfies the stochastic differential equation of the form
where
are independent standard Brownian motions with skew symmetry
. The generator of the diffusion
L operates on a function
as
where
,
,
, and
otherwise, and
is the totality of functions with a continuous derivative up to the second order and compact support in
.
We say a face of the simplex corresponds to a set of vertices if the face is the interior of the convex hall of denoted by , where is the set of standard basis of the vector space . If U consists of a single vertex, say , should be read as . An observation on the diffusion is as follows.
Proposition 1. Every point in a face of the simplex corresponding to an independent set of a graph is a fixed point of the stochastic differential Equation (1). Namely,is a fixed point. Proof. For each
,
is a martingale with respect to the natural filtration generated by the diffusion
. The condition (
3) implies
for each
,
, and thus
,
almost surely. Then, for each
, (
1) reduces to
, and we have
,
. Therefore, (
3) is a fixed point of (
1). □
Itoh et al. [
1] discussed the diffusion
as an approximation of the following discrete stochastic model. Consider a system of
N particles, where each particle is assigned to one vertex in
V, and there is a continuous-time Markov chain
Here,
is the number of particles assigned to the vertex
at time
s. At each instant of time, two of the
N particles are chosen uniformly randomly. If particles at
vertices are chosen, one of the particles is chosen with equal probabilities and assigned to another vertex. This causes a transition from
to
or
with equal probabilities. This stochastic model seems to have various applications. Tainaka et al. [
2] discussed this stochastic model as a model of a speciation process caused by geography. Ben-Haim et al. [
3] considered a slightly modified version, in which a transition from
to
,
occurs on the one-dimensional graph, where
,
. They called it a “compromise process” because if we regard the vertices as political positions, then a transition is a compromise. The process
converges weakly to the diffusion
: if
, then
as
in the space of right continuous functions with left limits (see Theorem 10.3.5 of [
4]).
Apart from an approximation of the discrete stochastic model discussed above, the diffusion
seems to appear in various contexts. The diffusion on a complete graph appears as an approximation of a quite different discrete stochastic model, called the Wright–Fisher model in population genetics, which evolves by repetition of multinomial sampling of a fixed number of particles (see, e.g., Chapter 10 of [
4], for details). The class of measure-valued diffusions is called Fleming–Viot processes. Such diffusions appear as prior distributions in Bayesian statistics.
As we will see below, the support of an extremal stationary state of the semigroup associated with the diffusion
is a face of
which corresponds to an independent set of
. In this sense, the diffusion can be regarded as independent set-finding in graph theory. Some problems related with independent set-finding, such as the maximum independent set problem, are known to be NP-hard, so it is believed that there is no efficient algorithm to solve them. Therefore, algorithms to find maximal independent sets are useful to obtain practical solutions. For example, Luby [
5] discussed a parallel algorithm for finding maximal independent sets that was derived from a stochastic algorithm to find a maximal independent set. His algorithm is based on a step to find an independent set based on random permutation executed by
processors in time
for large
.
Let us assume there exists a strongly continuous Markov semigroup
associated with the diffusion
governed by the generator (
2) such that
where
is the totality of continuous functions on
, and
The existence of such a semigroup for complete graphs was proven by Ethier [
6]. For the solution of the stochastic differential Equation (
1), we have
Let us denote by
the adjoint semigroup on
induced by
. Consider the totality of all fixed points of
:
We call each element of
a
stationary state of
. A stationary state
satisfies
The set
is non-empty and convex. The totality of the extremal elements of stationary states is denoted by
. Namely, a stationary state
is uniquely represented as
for some
,
. In Theorem 1, we see that support of an extremal stationary state of the diffusion
is a face of
corresponding to an independent set of
.
In this paper, we use the term support in a sloppy sense. Namely, positivity of a stationary state is not assumed unless otherwise stated. In fact, Proposition 1 implies that if the diffusion starts from any point x in , the stationary state is , or an atom at x. In this situation, we say the support is . In other words, if a stationary state does not have probability mass anywhere in an open set, then we say the set is not the support of the stationary state. Some examples are as follows.
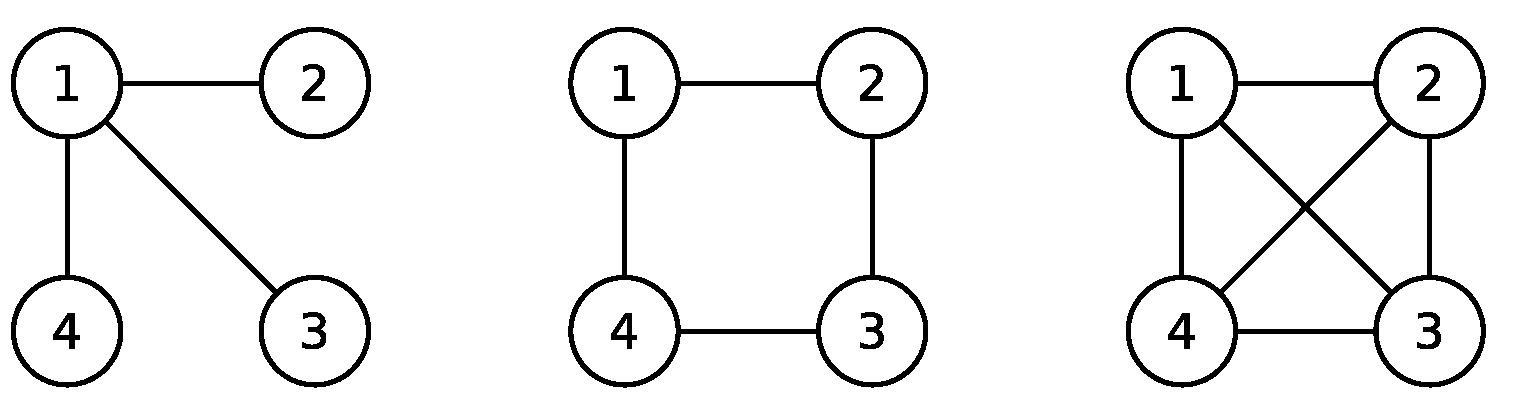
Example 1. Let , which is a complete graph consisting of r vertices. Since each vertex is maximally independent, a stationary state is represented as , where . For the solution of the stochastic differential Equation (1), is the absorption probability for the vertex . Since is a martingale, we know . Example 2. Let for an even positive integer r, which is a cycle graph consisting of r vertices, i.e., . The maximal independent sets are the set of all even integer vertices and that of all odd integer vertices. When , we have independent sets , , , , , and . The supports of extremal stationary states are the faces , , , and . The totality of the extremal stationary states is , where and are densities (not necessarily strictly positive) on and , respectively. Therefore, a stationary state is represented as .
Example 3. Let for positive integers r and s, which is a complete bipartite graph consisting of two disjointed and maximally independent sets of r and s vertices. For a graph whose maximal independent sets are and , a stationary state may be represented as .
Obtaining an explicit expression for the stationary states is a challenging problem. Itoh et al. [
1] successfully obtained an explicit expression for the stationary states for a star graph
, where a star graph
,
is a complete bipartite graph
, and the vertices of
are numbered such that
,
. A stationary state may be represented as
. If we identify vertices
, the star graph
is reduced to a complete graph
. Using the arguments for a complete graph in Example 1, we know
and
. Itoh et al. [
1] obtained an explicit expression for the diffusion starting from
:
by using martingales introduced in
Section 2. This result is for a specific graph but can be applied to other graphs reducible to
. For example, the four-cycle graph
discussed in Example 2 can be reduced to
. Explicit expressions for
,
are immediately obtained.
This paper is organized as follows. In
Section 2, the martingales used by Itoh et al. [
1] are revisited in a slightly generalized form. An interpretation of the martingales is presented in
Section 3. A duality relation between the Markov semigroup associated with the diffusion and a Markov chain on the set of ordered integer partitions is established. The dual Markov chain is studied and used to show that the support of an extremal stationary state of the adjoint semigroup is an independent set of the graph. In
Section 4, we investigate the diffusion with a linear drift, which gives a killing of the dual Markov chain on a finite integer lattice. The Markov chain is studied and used to study the unique stationary state of the diffusion, which generalizes the Dirichlet distribution. In
Section 5, two applications of the diffusions are discussed: analysis of an algorithm to find an independent set of a graph and Bayesian graph selection based on computation of the probability of a sample by using coupling from the past.
Section 6 is devoted to discussion of open problems.
3. Dual Process on Integer Partitions
To study diffusions
governed by the generator (
2), we employ a tool called duality, which is a familiar tool in the study of interacting particle systems (see, e.g., Chapter 2 of [
7]).
Consider a graph
and let
,
be a continuous-time Markov chain on the set of ordered non-negative integer partitions of
n with
, which is denoted by
, by the rate matrix
:
where
The backward equation for the transition probability
is
We have the following duality relation between the Markov semigroup and the Markov chain .
Lemma 1. The Markov semigroup associated with the generator (2) and the Markov chain with the rate matrix (10) satisfyfor each , where the killing rate is Proof. Noting that
and the operation (
4), we see that
satisfies the differential equation
for each
. This is uniquely solved by means of the Feynman–Kac formula, and the assertion follows. □
Since the total number of particles is kept, i.e.,
,
, the killing rate (
13) is bounded. The killing rate is not positive definite; however, a key observation is that if the support of a monomial
a denoted by
is an independent set of
, then the killing rate is non-positive:
. The converse is not always true.
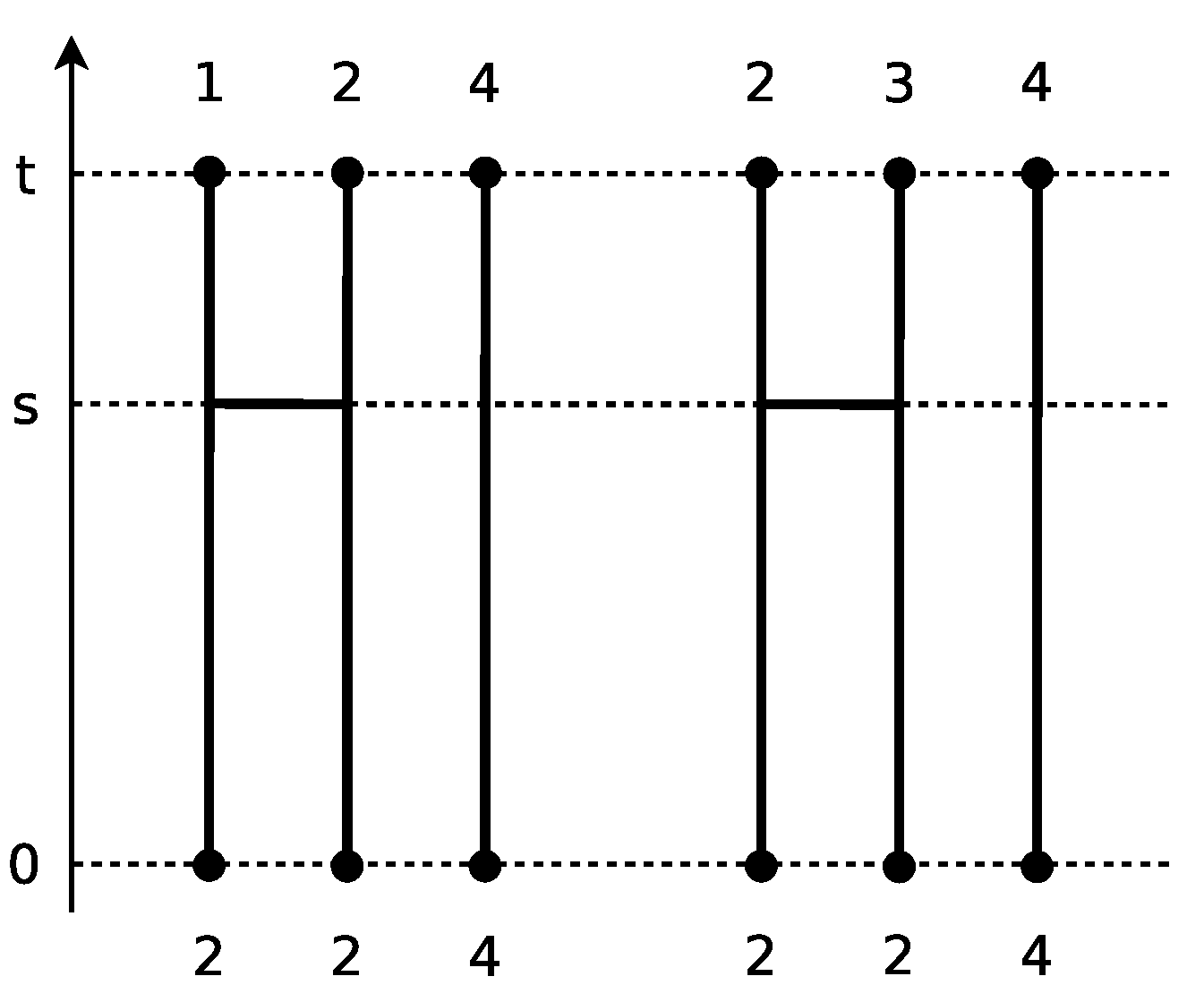
To illustrate the Markov chain
, let us ask specific questions. What is the moment of
for the cycle graph
discussed in Example 2? For a chain that starts from
, there are two possible transitions: the one is absorbed into the state
and the other is absorbed into the state
, where the rates are unities (see
Figure 1).
The waiting time for the occurrence of either of these two transitions follows the exponential distribution of rate two. Since
and
, the right side of the duality relation (
12) can be computed as
where
s is the time that one of the two possible transitions occurs. The first term corresponds to the case that no transition occurs until time
t. Let us call a transition event a
collision.
Remark 1. An analogous dual Markov chain of the diffusion approximation of a kind of Wright–Fisher model was effectively used by Shiga [8,9], where a transition occurs with the same rate as in (10). Such a transition event is called “coalescent”. In contrast to a collision, the total number of particles decreases by a coalescent. Here, we have a simple observation about the invariants among moments discussed in
Section 2. If a chain
starts from a state
a such that
is a maximal independent set
of the graph
, then the killing rate is non-positive, and the duality relation (
12) is reduced to
Corollary 1 implies cancellation of the second term among moments. Moreover, considering a case that the diffusion
starts from a point
x in the face corresponding to
, by Proposition 1, we know that after a collision,
must contain a vertex that is not contained in
.
Let us ask another question. What is the moment of
for the star graph
discussed in
Section 1? Some consideration reveals that a chain
starting from
a will never be absorbed, and transitions occur among the three states:
a,
, and
. Since
and
, the duality relation (
12) gives
where
is 1 if the argument
is true and zero otherwise. Solving the backward Equation (
11), we immediately obtain
However, computation of the right side of Equation (
14) does not seem easy because the expectation depends on a sample path of the chain
. Nevertheless, the moments can be obtained easily by solving the system of differential equations (
9). In fact, we have
where
The observations above lead to the following proposition on the fate of the Markov chain .
Proposition 3. Consider the Markov chain on the set of the ordered non-negative integer positive partitions .
If a chain starts from a state a satisfying , then it is absorbed into an element of ;
If a chain starts from a state a satisfying , then the transition probability converges to the uniform distribution on the set of ordered positive integer partitions
Proof.
- (i)
A state
a is absorbing if and only if the row vector of the rate matrix (
10) is zero, which implies
. Consider the set of vertices
. Then,
,
is a death process and is absorbed into the state
at Markov time
with respect to the Markov chain
, where
for
. Let us show that the process eventually decreases if
. If
, at least one vertex, say
, satisfies
. If the vertex
j is connected with a vertex in
, say
k, the transition
occurs with positive probability, and it makes
. Otherwise, the vertex
j should be connected with at least one vertex in
, say
l. The transition
makes
. If the vertex
l is connected with a vertex in
, the assertion is shown. Otherwise, the vertex
l should be connected with at least one vertex in
. The proof is completed by repeating this procedure until we reach a vertex in
that is connected with a vertex in
.
- (ii)
If a chain starts from a state in the set
, the argument for
, i.e.,
is a death process, shows that in a finite time the chain reaches a state, say
a, in the set
and never exits from
. For such a case, consider restarting the chain from the state
a. For simplicity, we consider the case of
. Then, a state
can be identified with the unique vertex
satisfying
. Since we are considering a connected graph, there exists a path
for any
with some
, where
,
. Since all of the transitions occur with rate unity, the sample path has a positive probability. Hence, the Markov chain is irreducible and ergodic, and there exists the unique stationary distribution on the set
. Since the uniform distribution on
satisfies the backward Equation (
11), it is the stationary distribution. Cases of
can be shown in a similar manner by showing that up to
particles can be moved from one vertex to another vertex.
□
Proposition 3 shows that if the Markov chain
starts from a state
a satisfying
, convergence of the chain can be divided into two phases: (1) exit from the set
and (2) convergence to the uniform distribution on the set
. The largest eigenvalue of the rate matrix is zero. To consider the mixing, we have to know the second-largest eigenvalue, say
. By spectral decomposition of the transition probability, the mixing time of Phase 2, or the infimum of
t satisfying
for
is less than
for some constant
, where
is the total variation distance between probability measures
and
.
Example 5. Consider the case that (see the proof of Proposition 3 ()). It can be shown that the rate matrix reduces to the negative of the Laplacian matrix of , whose second- largest eigenvalue is called the algebraic connectivity of . For a connected graph, it is known that the largest eigenvalue is zero and the second-largest eigenvalue is bounded below by [10], where is the diameter of , or the maximum of the shortest path lengths between any pair of vertices in . For the star graph and the fourth-order monomials discussed above, the rate matrix is the negative of the Laplacian matrix:and the eigenvalues are 0, , and , where , . The two eigenvalues appear in the transition probabilities (15). For a generic graph with large r, the mixing time is . Assessment of Phase 1 seems harder. The death process
,
with
is not a Markov process. The death rate is bounded below:
To obtain a rough estimate of the right side, let us suppose the state
follows the uniform distribution on the set of ordered positive integer partitions
, where
Then, the expectation of the lower bound is
, where
. When
n is large, the dominant contribution to the expectation of the waiting time for the exit comes from the period
, and it is
. Hence, the expectation of the waiting time for the exit would be
.
Shiga [
8,
9] and Shiga and Uchiyama [
11] studied structures of extremal stationary states of the diffusion approximation of a kind of Wright–Fisher model in
for a countable set
S by using its dual Markov chain. Extremal states of the adjoint Markov semigroup
on
induced by
associated with the diffusion
can be studied by using the dual Markov chain
. Note that positivity of a stationary state is not assumed, as explained in
Section 1.
Theorem 1. The support of an extremal stationary state of the adjoint Markov semigroup is a face of the simplex corresponding to an independent set of the graph , namely, .
Proof. Consider a Markov chain
with rate matrix (
10) starting from a state
. According to Proposition 3
, such an
a is an absorbing state and the chain stays there. Lemma 1 gives
A stationary state
satisfies
for any
. If
, this condition reduces to
Therefore, if
is not an independent set, then
. Since we are considering a connected graph
,
V is not an independent set of
. The condition reduces to
. Since
the condition
excludes
from the support of
. Suppose there exists a vertex
such that
is still not an independent set. Set
a is such that
if
and
otherwise for each
. Since
the condition
excludes
from the support of
. Repeating this procedure yields an independent set
for some
, and the face
is not excluded from the support of
. □
The steps in the above proof appear in the following example.
Example 6. Let , which is a cycle graph consisting of four vertices. The support of an extremal stationary state appearing in Example 2 is confirmed as follows. Remove the vertex from the vertex set V. Since is not an independent set, the face is excluded from the support of extremal stationary states. Then, remove from . Since is an independent set, the face is the support of an extremal stationary state.
A direct consequence of Theorem 1 on the moments is as follows.
Corollary 2. For each , if the limit of an n-th order moment of the diffusion on a graph is positive, namely,then is an independent set of . 4. Diffusion with Linear Drift
In this section, we consider the diffusion
taking value in probability measures on a graph
,
satisfying the following stochastic differential equation with linear drift:
for
. The drift term,
, gives a killing of the dual process with a linear rate. As shown below, behaviours of the diffusion and the dual Markov chain are significantly different from those without drift discussed in previous sections.
In Itoh et al.’s discrete stochastic model described in
Section 1, this drift corresponds to adding the following dynamics: at each instant of time, one of
N particles is chosen uniformly randomly and assigned to another vertex chosen uniformly randomly with rate
. In the Wright–Fisher model, this drift corresponds to a mutation mechanism [
4].
Let
be a continuous-time Markov chain on a finite integer lattice, or the set of non-negative integers
by the rate matrix
:
where
The backward equation for the transition probability
is
The following duality relation between the Markov semigroup associated with the diffusion denoted by and the Markov chain can be shown in the same manner as Lemma 1.
Lemma 2. The Markov semigroup and the Markov chain with the rate matrix (17) satisfyfor each , where the killing rate is In contrast to the rate matrix
in (
10), particles are erased, and it makes the total number of particles
decrease. It is clear from the rate matrix (
17) that 0 is the unique absorbing state.
Proposition 4. Let , which is a Markov time with respect to the Markov chain with the rate matrix (17). Then, Proof. Since
if and only if
, we consider the Markov chain of the cardinality
. According to the rate matrix (
17), it is a linear death process with rate
. Noting that
is the convolution of exponential random variables of rates
,
, we have the assertion. □
To illustrate the Markov chain
, let us ask a specific question. What is the moment of
from the cycle graph
? For a chain that starts from
a, there are four possible sample paths (
Figure 2):
- (i)
No particles are erased;
- (ii)
Particle 1 is erased, but Particle 2 survives;
- (iii)
Particle 2 is erased, but Particle 1 survives;
- (iv)
Both particles are erased.
The waiting time for either of the two particles to be erased follows the exponential distribution of rate
. Since
and
, the right side of the duality relation (
18) can be computed as
where
and
are the times that a particle is erased.
The stationary state of the adjoint Markov semigroup on induced by consists of the unique probability measure.
Theorem 2. For the adjoint Markov semigroup , there exists the unique stationary state satisfyingfor every . Proof. Since the Markov chain
with the rate matrix (
17) is absorbed into 0, Lemma 2 and Proposition 4 give
for all
. Since
for each
, there exists a unique probability measure
satisfying
for all
. □
The stationary state converges weakly to a common limit as irrespective of the graph.
Corollary 3. The stationary state of the adjoint Markov semigroup satisfies Proof. Since the killing rate (
19) is bounded and
for large
, the leading contribution to the expression (
20) can be evaluated by the death process
considered in Proposition 4, whose waiting time follows the exponential distribution of rate
. Let
. We have
where
is a constant satisfying
for all
b satisfying
. In the same way,
is bounded below. The assertion follows by taking the limit
of these bounds. □
Moreover, the stationary state has a continuous and strictly positive density.
Theorem 3. For the adjoint Markov semigroup , the unique stationary state is absolutely continuous with respect to the Lebesgue measure on and admits a probability density that is strictly positive in and is of -class.
Proof. We first show that
has a density of
-class. By Theorem 2, we have
for each
. Therefore,
has a
-density represented as
We next show that the density
is strictly positive in
. Consider an approximation of
by polynomials:
satisfying
. Suppose there exist a point
satisfying
. Since the polynomials
are strictly positive, for any small positive constants
and
, there exists an
N such that
and
is covered by open balls:
for all
. Since
is smooth, for every point
, we can find a ball containing
x and
for some constant
c. This implies
,
, but it contradicts the fact that
,
followed by the expression (
20) because the killing rate
is bounded and the Markov time satisfies
by Proposition 4. □
An immediate consequence is the following corollary, which is an analogous result to Corollary 2.
Corollary 4. The moments of the stationary state of the adjoint Markov semigroup are positive, namely, The moments of the stationary state can be obtained by the condition for the stationary state (
5). It gives a system of recurrence relations:
for each
with the boundary condition
. In contrast to the system of ordinary differential equations (
9), this system is not closed among the moments of the same order. Prior to solving the system for the moment of a given monomial
a, we have to solve the systems for the moments of lower orders than
a. Therefore, it seems a formidable task to solve System (
22). Diffusion on a complete graph is an exception.
Example 7. Let , which is the complete graph consisting of r vertices discussed in Example 1. The unique solution of the system of recurrence relations (22) isMoreover, since this expression is the moments of the symmetric Dirichlet distribution of parameter α, the stationary state is the Dirichlet distribution: Remark 2. The limit of the moments (23) is known as the Dirichlet-multinomial distribution up to multiplication of the multinomial coefficient. Renumbering the set by and taking the limit and with , the expression (23) reduces to the formwhich is known as the Ewens sampling formula, or the exchangeable partition probability function of the Dirichlet prior process in Bayesian statistics (see, e.g., [12] for an introduction). Karlin and McGregor [13] derived this formula by using a system of recurrence relations based on coalescents mentioned in Remark 1. In this sense, we have found an alternative system of recurrence relations (22) the Formula (24) satisfies based on collisions.