PCDM and PCDM4MP: New Pairwise Correlation-Based Data Mining Tools for Parallel Processing of Large Tabular Datasets
Abstract
:1. Introduction
2. Materials and Methods
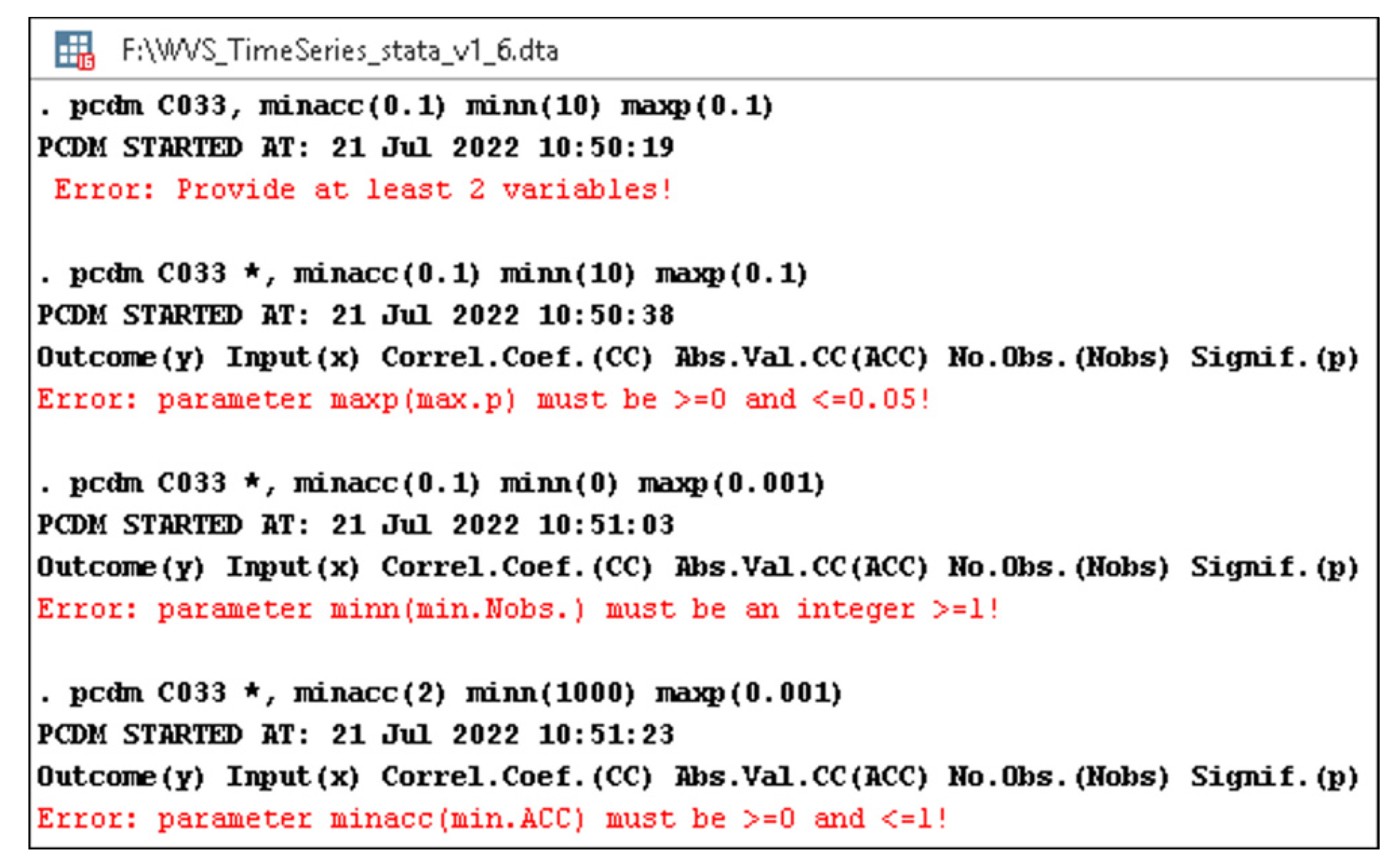
- minacc—the minimum accepted absolute value (lines 21–29 and 59, Listing A1) of the correlation coefficient (its default value is 0—line 18, Listing A1);
- minn—the minimum accepted number of observations (lines 30–38 and 59, Listing A1) for each response-predictor pair (its default value is 1—line 19, Listing A1);
- maxp—the maximum tolerated p-value (lines 39–47 and 59, Listing A1) for a significance threshold, usually 0.05 or less (therefore, its default value is 0.05—line 20, Listing A1).
- Intel Xeon Gold 6240 CascadeLake CPU (Central Processing Unit) with 36 virtual processors/logical cores/threads and 18 physical ones, Socket 3647 LGA, 14 nm technology, 2.6 GHz and 32 GB of RAM (Random Access Memory), SCSI Disk, on a Windows Server Datacenter 2019 Virtual Machine (VM—CPU’s bus/core ratio/clock multiplier locked inside the VM, and maximum 32 virtual processors (https://drive.google.com/file/d/1LbbB9Jz3C9SYJHsRUCkwmSREKoI-_ejJ/view, accessed on 1 June 2022, configured for use) in a private cloud (https://cloud.raas.uaic.ro, accessed on 1 June 2022) managed using OpenStack on Ubuntu.
- Intel Core i7–4710HQ CPU (8 logical cores, 4 physical ones), Socket 1364 BGA, 22 nm technology, up to 3.5 GHz and 32 GB of RAM, SSD, on a Physical Machine (PM—CPU’s bus/core ratio not locked) using Windows 8.1 Professional x64.
- Intel Atom N550 dual-core CPU (4 logical cores), Socket 559 FCBGA8, 45 nm technology, 1.5 GHz and 2 GB of RAM, SATA HDD, on a PM using the same type of Windows 8.1 above.
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
- Listing A1. The source script of PCDM with numbered lines—numbers displayed separately, as when opened with the Stata editor.
- Listing A2. The source script of PCDM4MP with numbered lines.
| Variable | Question | Coding |
|---|---|---|
| C033 | Job satisfaction—DEPENDENT VARIABLE | 1-Dissatisfied … 10-Satisfied |
| C033_bin | Job satisfaction (binary format)—DEPENDENT VARIABLE | 1 if C033!=. & C033>=6 0 if C033!=. & C033<6 & C033>0 |
| A170 | Satisfaction with your life | 1-Dissatisfied … 10-Satisfied |
| A170_bin | Satisfaction with your life (binary format) | 1 if A170!=. & A170>=6 0 if A170!=. & A170<6 & A170>0 |
| C006 | Satisfaction with the financial situation of household | 1-Dissatisfied … 10-Satisfied |
| C006_bin | Satisfaction with the financial situation of household (binary format) | 1 if C006!=. & C006>=6 0 if C006!=. & C006<6 & C006>0 |
| C031 | Degree of pride in your work | 1-A great deal … 4-None |
| C031_bin | Degree of pride in your work (binary format) | 1 if C031!=. & C031<=2 & C031>0 0 if C031!=. & C031>2 |
| C034 | Freedom of decision taking in the job | 1-Not at all … 10-A great deal |
| C034_bin | Freedom of decision taking in the job (binary format) | 1 if C034!=. & C034>=6 0 if C034!=. & C034<6 & C034>0 |
| D002 | Satisfaction with home life | 1-Dissatisfied … 10-Satisfied |
| D002_bin | Satisfaction with home life (binary format) | 1 if D002!=. & D002>=6 0 if D002!=. & D002<6 & D002>0 |
| Variable | N | Mean | Std.Dev. | Min | Median | Max |
|---|---|---|---|---|---|---|
| C033 | 15,968 | 7.27 | 2.31 | 1 | 8 | 10 |
| C033_bin | 15,968 | 0.77 | 0.42 | 0 | 1 | 1 |
| A170 | 420,669 | 6.7 | 2.42 | 1 | 7 | 10 |
| A170_bin | 420,669 | 0.69 | 0.46 | 0 | 1 | 1 |
| C006 | 411,461 | 5.75 | 2.58 | 1 | 6 | 10 |
| C006_bin | 411,461 | 0.54 | 0.5 | 0 | 1 | 1 |
| C031 | 14,988 | 1.73 | 0.87 | 1 | 2 | 4 |
| C031_bin | 14,988 | 0.51 | 0.5 | 0 | 1 | 1 |
| C034 | 17,900 | 6.54 | 2.79 | 1 | 7 | 10 |
| C034_bin | 17,900 | 0.65 | 0.48 | 0 | 1 | 1 |
| D002 | 25,653 | 7.72 | 2.24 | 1 | 8 | 10 |
| D002_bin | 25,653 | 0.83 | 0.38 | 0 | 1 | 1 |
| Model | (1) | (2) | (3) | (4) | (5) | (6) | (7) | (8) | (9) | (10) |
|---|---|---|---|---|---|---|---|---|---|---|
| Predictors/Response var. | C033_bin | A170_bin | C033_bin | C006_bin | C033_bin | C031_bin | C033_bin | C034_bin | C033_bin | D002_bin |
| A170 | 0.3973 *** | |||||||||
| (0.0097) | ||||||||||
| C006 | 0.3300 *** | |||||||||
| (0.0084) | ||||||||||
| C031 | −1.2461 *** | |||||||||
| (0.0263) | ||||||||||
| C034 | 0.3233 *** | |||||||||
| (0.0077) | ||||||||||
| D002 | 0.3264 *** | |||||||||
| (0.0092) | ||||||||||
| C033 | 0.3480 *** | 0.3049 *** | 0.5306 *** | 0.3800 *** | 0.3360 *** | |||||
| (0.0089) | (0.0081) | (0.0118) | (0.0088) | (0.0096) | ||||||
| _cons | −1.3840 *** | −1.2868 *** | −0.5840 *** | −1.8448 *** | 3.5825 *** | −1.8024 *** | −0.7322 *** | −1.9542 *** | −1.1913 *** | −0.6141 *** |
| (0.0646) | (0.0618) | (0.0473) | (0.0604) | (0.0576) | (0.0729) | (0.0475) | (0.0638) | (0.0690) | (0.0644) | |
| N | 15,848 | 15,848 | 15,811 | 15,811 | 14,900 | 14,900 | 15,811 | 15,811 | 15,752 | 15,752 |
| chi2 | 1681.6969 | 1511.4602 | 1558.7477 | 1406.8425 | 2237.2495 | 2034.7577 | 1771.5264 | 1851.9322 | 1253.1181 | 1212.2401 |
| p | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 |
| pseudo R2 | 0.1258 | 0.1060 | 0.1046 | 0.0800 | 0.1833 | 0.2168 | 0.1244 | 0.1204 | 0.0880 | 0.0993 |
| AUCROC | 0.7443 | 0.7129 | 0.7272 | 0.6797 | 0.7667 | 0.8095 | 0.7377 | 0.7280 | 0.6912 | 0.7193 |
| AIC | 14,832.3486 | 15,908.2089 | 15,176.6194 | 19,733.4346 | 13,249.4465 | 10,656.0603 | 14,786.6602 | 17,607.5324 | 15,391.2067 | 12,641.8199 |
| BIC | 14,847.6902 | 15,923.5505 | 15,191.9563 | 19,748.7715 | 13,264.6647 | 10,671.2786 | 14,801.9971 | 17,622.8693 | 15,406.5362 | 12,657.1493 |
| Model | (1) | (2) | (3) | (4) | (5) | (6) | (7) | (8) | (9) | (10) | (11) | (12) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Regression Type | logit | OLS | logit | OLS | logit | logit | OLS | OLS | logit | logit | OLS | OLS |
| Filter Condition | N/A | N/A | N/A | N/A | if C006!=. | if A170!=. | if C006!=. | if A170!=. | N/A | N/A | N/A | N/A |
| Predictors/Response var. | C033_bin | |||||||||||
| A170 | 0.1867 *** | 0.0258 *** | 0.2667 *** | 0.0416 *** | 0.3433 *** | 0.0535 *** | 0.3441 *** | 0.0536 *** | ||||
| (0.0132) | (0.0018) | (0.0111) | (0.0017) | (0.0103) | (0.0016) | (0.0102) | (0.0015) | |||||
| C006 | 0.1423 *** | 0.0169 *** | 0.1851 *** | 0.0254 *** | 0.2780 *** | 0.0409 *** | 0.2776 *** | 0.0409 *** | ||||
| (0.0115) | (0.0015) | (0.0102) | (0.0014) | (0.0092) | (0.0013) | (0.0091) | (0.0013) | |||||
| C031 | −0.9285 *** | −0.1497 *** | ||||||||||
| (0.0294) | (0.0044) | |||||||||||
| C034 | 0.1925 *** | 0.0260 *** | 0.2579 *** | 0.0394 *** | 0.2784 *** | 0.2765 *** | 0.0432 *** | 0.0449 *** | 0.2791 *** | 0.2768 *** | 0.0433 *** | 0.0451 *** |
| (0.0093) | (0.0013) | (0.0084) | (0.0013) | (0.0083) | (0.0081) | (0.0013) | (0.0013) | (0.0083) | (0.0081) | (0.0013) | (0.0013) | |
| D002 | 0.0907* ** | 0.0137 *** | ||||||||||
| (0.0127) | (0.0019) | |||||||||||
| _cons | −0.8378 *** | 0.4666 *** | −3.1023 *** | 0.0649 *** | −2.7134 *** | −1.9714 *** | 0.1076 *** | 0.2276 *** | −2.7222 *** | −1.9723 *** | 0.1061 *** | 0.2267 *** |
| (0.1301) | (0.0208) | (0.0866) | (0.0127) | (0.0813) | (0.0672) | (0.0126) | (0.0112) | (0.0810) | (0.0669) | (0.0125) | (0.0111) | |
| N | 14,375 | 14,375 | 15,576 | 15,576 | 15,576 | 15,576 | 15,576 | 15,576 | 15,705 | 15,671 | 15,705 | 15,671 |
| chi2 | 2803.3215 | 2541.0448 | 2376.3159 | 2285.7133 | 2400.0313 | 2306.7919 | ||||||
| p | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 | 0.0000 |
| R2 | 0.3060 | 0.2279 | 0.2101 | 0.1900 | 0.2111 | 0.1906 | ||||||
| pseudo R2 | 0.2966 | 0.2231 | 0.2021 | 0.1842 | 0.2030 | 0.1846 | ||||||
| RMSE | 0.3519 | 0.3668 | 0.3710 | 0.3757 | 0.3710 | 0.3759 | ||||||
| maxAbsVPMCC | 0.5211 | 0.5211 | 0.4696 | 0.4696 | 0.2763 | 0.2754 | 0.2763 | 0.2754 | 0.2765 | 0.2759 | 0.2765 | 0.2759 |
| OLSmaxAcceptVIF | 1.4410 | 1.2951 | 1.2659 | 1.2346 | 1.2676 | 1.2355 | ||||||
| OLSmaxComputVIF | 1.5793 | 1.3203 | 1.0872 | 1.0878 | 1.0873 | 1.0882 | ||||||
| AUCROC | 0.8532 | 0.8166 | 0.8022 | 0.7919 | 0.8028 | 0.7922 | ||||||
| AIC | 10,973.3805 | 10,770.2185 | 12,902.2457 | 12,963.9656 | 13,249.4442 | 13,546.3796 | 13,317.1563 | 13,707.6119 | 13,353.6160 | 13,640.0974 | 13,423.0338 | 13,806.9150 |
| BIC | 11,018.8200 | 10,815.6580 | 12,932.8596 | 12,994.5796 | 13,272.4047 | 135,69.3401 | 13,340.1168 | 13,730.5724 | 13,376.6012 | 13,663.0761 | 13,446.0190 | 13,829.8937 |
References
- Baker, M. Why scientists must share their research code. Nature 2016. [Google Scholar] [CrossRef]
- Matarese, V. Kinds of replicability: Different terms and different functions. Axiomathes 2022, 1–24. [Google Scholar] [CrossRef]
- Homocianu, D.; Plopeanu, A.-P.; Ianole-Calin, R. A Robust Approach for Identifying the Major Components of the Bribery Tolerance Index. Mathematics 2021, 9, 1570. [Google Scholar] [CrossRef]
- Rajiah, K.; Sivarasa, S.; Maharajan, M.K. Impact of Pharmacists’ Interventions and Patients’ Decision on Health Outcomes in Terms of Medication Adherence and Quality Use of Medicines among Patients Attending Community Pharmacies: A Systematic Review. Int. J. Environ. Res. Public Health 2021, 18, 4392. [Google Scholar] [CrossRef] [PubMed]
- Sadeghi, A.R.; Bahadori, Y. Urban Sustainability and Climate Issues: The Effect of Physical Parameters of Streetscape on the Thermal Comfort in Urban Public Spaces; Case Study: Karimkhan-e-Zand Street, Shiraz, Iran. Sustainability 2021, 13, 10886. [Google Scholar] [CrossRef]
- Thanh, M.T.G.; Van Toan, N.; Toan, D.T.T.; Thang, N.P.; Dong, N.Q.; Dung, N.T.; Hang, P.T.T.; Anh, L.Q.; Tra, N.T.; Ngoc, V.T.N. Diagnostic Value of Fluorescence Methods, Visual Inspection and Photographic Visual Examination in Initial Caries Lesion: A Systematic Review and Meta-Analysis. Dent. J. 2021, 9, 30. [Google Scholar] [CrossRef]
- Wang, L.; Ling, C.-H.; Lai, P.-C.; Huang, Y.-T. Can The ‘Speed Bump Sign’ Be a Diagnostic Tool for Acute Appendicitis? Evidence-Based Appraisal by Meta-Analysis and GRADE. Life 2022, 12, 138. [Google Scholar] [CrossRef] [PubMed]
- Damasceno, E.; Azevedo, A.; Pérez-Cota, M. Data mining, business intelligence, grid and utility computing: A bibliometric review of the literature from 2015 to 2020. In Proceedings of the 23rd International Conference on Enterprise Information Systems, Prague, Czech Republic, 26–28 April 2021; Volume 1, pp. 367–373. [Google Scholar] [CrossRef]
- Kopf, O.; Homocianu, D. The Business Intelligence Based Business Process Management Challenge. Inform. Econ. J. 2016, 20, 7–19. [Google Scholar] [CrossRef]
- Studer, S.; Bui, T.B.; Drescher, C.; Hanuschkin, A.; Winkler, L.; Peters, S.; Müller, K.-R. Towards CRISP-ML(Q): A Machine Learning Process Model with Quality Assurance Methodology. Mach. Learn. Knowl. Extr. 2021, 3, 392–413. [Google Scholar] [CrossRef]
- Bendel, R.B.; Afifi, A.A. Comparison of stopping rules in forward “stepwise” regression. J. Am. Stat. Assoc. 1977, 72, 46. [Google Scholar] [CrossRef]
- Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. Ser. B Methodol. 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Sanchez, J.D.; Rêgo, L.C.; Ospina, R. Prediction by Empirical Similarity via Categorical Regressors. Mach. Learn. Knowl. Extr. 2019, 1, 641–652. [Google Scholar] [CrossRef] [Green Version]
- Ahrens, A.; Hansen, C.B.; Schaffer, M.E. Lassopack: Model selection and prediction with regularized regression in Stata. Stata J. Promot. Commun. Stat. Stata 2020, 20, 176–235. [Google Scholar] [CrossRef] [Green Version]
- Bilger, M. Overfit: Stata module to calculate shrinkage statistics to measure overfitting as well as out- and in-sample predictive bias. Stat Soft. Comp. 2015, S457950. Available online: https://EconPapers.repec.org/RePEc:boc:bocode:s457950 (accessed on 1 June 2022).
- Gao, Y.; Cowling, M. Introduction to Panel Data, Multiple Regression Method, and Principal Components Analysis Using Stata: Study on the Determinants of Executive Compensation—A Behavioral Approach Using Evidence from Chinese Listed Firms; SAGE Publications Ltd.: Thousand Oaks, CA, USA, 2019. [Google Scholar] [CrossRef]
- De Luca, G.; Magnus, J.R. Bayesian model averaging and weighted-average least squares: Equivariance, stability, and numerical issues. Stata J. Promot. Commun. Stat. Stata 2011, 11, 518–544. [Google Scholar] [CrossRef]
- Karabulut, E.M.; Ibrikci, T. Analysis of cardiotocogram data for fetal distress determination by decision tree based adaptive boosting approach. J. Comput. Commun. 2014, 2, 32–37. [Google Scholar] [CrossRef] [Green Version]
- Schonlau, M. Boosted regression (boosting): An introductory tutorial and a Stata plugin. Stata J. Promot. Commun. Stat. Stata 2005, 5, 330–354. [Google Scholar] [CrossRef]
- Zlotnik, A.; Abraira, V. A general-purpose nomogram generator for predictive logistic regression models. Stata J. Promot. Commun. Stat. Stata 2015, 15, 537–546. [Google Scholar] [CrossRef] [Green Version]
- Zdravevski, E.; Lameski, P.; Kulakov, A.; Filiposka, S.; Trajanov, D.; Jakimovski, B. Parallel computation of information gain using Hadoop and mapreduce. Ann. Comput. Sci. Inf. Syst. 2015. [Google Scholar] [CrossRef] [Green Version]
- Oancea, B.; Dragoescu, R.M. Integrating R and Hadoop for Big Data Analysis, Romanian Statistical Review. arXiv 2014, arXiv:1407.4908. [Google Scholar] [CrossRef]
- Meng, X.; Bradley, J.; Yavuz, B.; Sparks, E.; Venkataraman, S.; Liu, D.; Freeman, J.; Tsai, D.B.; Amde, M.; Owen, S.; et al. MLlib: Machine Learning in Apache Spark. arXiv 2015, arXiv:1505.06807. [Google Scholar] [CrossRef]
- Fotache, M.; Cluci, M.-I. Big Data Performance in private clouds. In Some initial findings on Apache Spark Clusters deployed in OpenStack. In Proceedings of the 2021 20th RoEduNet Conference: Networking in Education and Research (RoEduNet), Iasi, Romania, 4–6 November 2021. [Google Scholar] [CrossRef]
- Li, J.; Zhang, C.; Zhang, J.; Qin, X.; Hu, L. MICS-P:parallel mutual-information computation of big categorical data on Spark. J. Parallel Distrib. Comput. 2022, 161, 118–129. [Google Scholar] [CrossRef]
- Khoshaba, F.; Kareem, S.; Awla, H.; Mohammed, C. Machine learning algorithms in Bigdata analysis and its applications: A Review. In Proceedings of the 2022 International Congress on Human-Computer Interaction, Optimization and Robotic Applications (HORA), Ankara, Turkey, 9–11 June 2022; pp. 1–8. [Google Scholar] [CrossRef]
- Murty, C.S.; Saradhi Varma, G.P.; Satyanarayana, C. Content-based collaborative filtering with hierarchical agglomerative clustering using user/item based ratings. J. Interconnect. Netw. 2022. [Google Scholar] [CrossRef]
- Aldabbas, H.; Albashish, D.; Khatatneh, K.; Amin, R. An architecture of IOT-aware healthcare smart system by leveraging machine learning. Int. Arab. J. Inf. Technol. 2022, 19, 160–172. [Google Scholar] [CrossRef]
- Alhussan, A.A.; AlEisa, H.N.; Atteia, G.; Solouma, N.H.; Seoud, R.A.; Ayoub, O.S.; Ghoneim, V.F.; Samee, N.A. ForkJoinPcc algorithm for computing the PCC matrix in gene co-expression networks. Electronics 2022, 11, 1174. [Google Scholar] [CrossRef]
- Huckvale, E.D.; Hodgman, M.W.; Greenwood, B.B.; Stucki, D.O.; Ward, K.M.; Ebbert, M.T.; Kauwe, J.S.; Miller, J.B. Pairwise Correlation Analysis of the Alzheimer’s disease neuroimaging initiative (ADNI) dataset reveals significant feature correlation. Genes 2021, 12, 1661. [Google Scholar] [CrossRef] [PubMed]
- Ye, R.; Fang, B.; Du, W.; Luo, K.; Lu, Y. Bootstrap Tests for the Location Parameter under the Skew-Normal Population with Unknown Scale Parameter and Skewness Parameter. Mathematics 2022, 10, 921. [Google Scholar] [CrossRef]
- Airinei, D.; Homocianu, D. The Importance of Video Tutorials for Higher Education—The Example of Business Information Systems. In Proceedings of the 6th International Seminar on the Quality Management in Higher Education, Tulcea, Romani, 8–9 July 2010; Available online: https://ssrn.com/abstract=2381817 (accessed on 1 June 2022).
- Michelucci, U.; Venturini, F. Estimating Neural Network’s Performance with Bootstrap: A Tutorial. Mach. Learn. Knowl. Extr. 2021, 3, 357–373. [Google Scholar] [CrossRef]
- Airinei, D.; Homocianu, D. The Geographical Dimension of DSS Applications. Sci. Ann. Alexandru Ioan Cuza Univ. Iasi 2009, 56, 637–642. Available online: https://econpapers.repec.org/RePEc:aic:journl:y:2009:v:56:p:637-642 (accessed on 1 June 2022).
- Hayashi, K.; Llorca, L.P.; Bugayong, I.D.; Agustiani, N.; Capistrano, A.O.V. Evaluating the Predictive Accuracy of the Weather-Rice-Nutrient Integrated Decision Support System (WeRise) to Improve Rainfed Rice Productivity in Southeast Asia. Agriculture 2021, 11, 346. [Google Scholar] [CrossRef]
- Peña, M.; Biscarri, F.; Personal, E.; León, C. Decision Support System to Classify and Optimize the Energy Efficiency in Smart Buildings: A Data Analytics Approach. Sensors 2022, 22, 1380. [Google Scholar] [CrossRef]
- Goodwin, J.L.; Williams, A.L.; Snell Herzog, P. Cross-Cultural Values: A Meta-Analysis of Major Quantitative Studies in the Last Decade (2010–2020). Religions 2020, 11, 396. [Google Scholar] [CrossRef]
- Ortega-Gil, M.; Mata García, A.; ElHichou-Ahmed, C. The Effect of Ageing, Gender and Environmental Problems in Subjective Well-Being. Land 2021, 10, 1314. [Google Scholar] [CrossRef]
- Miniesy, R.S.; AbdelKarim, M. Generalized Trust and Economic Growth: The Nexus in MENA Countries. Economies 2021, 9, 39. [Google Scholar] [CrossRef]
- Lim, S.B.; Malek, J.A.; Yigitcanlar, T. Post-Materialist Values of Smart City Societies: International Comparison of Public Values for Good Enough Governance. Future Internet 2021, 13, 201. [Google Scholar] [CrossRef]
- Vo, T.T.D.; Tuliao, K.V.; Chen, C.-W. Work Motivation: The Roles of Individual Needs and Social Conditions. Behav. Sci. 2022, 12, 49. [Google Scholar] [CrossRef]
- Sánchez-García, J.; Gil-Lacruz, A.I.; Gil-Lacruz, M. The influence of gender equality on volunteering among European senior citizens. Volunt. Int. J. Volunt. Nonprofit Organ. 2022. [Google Scholar] [CrossRef]
- Fakih, A.; Makdissi, P.; Marrouch, W.; Tabri, R.V.; Yazbeck, M. A stochastic dominance test under survey nonresponse with an application to comparing trust levels in Lebanese public institutions. J. Econom. 2022, 228, 342–358. [Google Scholar] [CrossRef]
- Freund, R.J.; Wilson, W.J. Regression Analysis: Statistical Modeling of a Response Variable, 2nd ed.; Academic Press: Cambridge, UK, 2006. [Google Scholar]
- Vatcheva, P.K.; Lee, M.; McCormick, J.B.; Rahbar, M.H. Multicollinearity in regression analyses conducted in epidemiologic studies. Epidemiol. Sunnyvale Open Access 2016, 6, 227. [Google Scholar] [CrossRef] [Green Version]
- Arabameri, A.; Asadi Nalivan, O.; Chandra Pal, S.; Chakrabortty, R.; Saha, A.; Lee, S.; Pradhan, B.; Tien Bui, D. Novel Machine Learning Approaches for Modelling the Gully Erosion Susceptibility. Remote Sens. 2020, 12, 2833. [Google Scholar] [CrossRef]
- Pepe, M.S.; Cai, T.; Longton, G. Combining predictors for classification using the area under the receiver operating characteristic curve. Biometrics 2005, 62, 221–229. [Google Scholar] [CrossRef]
- Carreras, J.; Hamoudi, R. Artificial Neural Network Analysis of Gene Expression Data Predicted Non-Hodgkin Lymphoma Subtypes with High Accuracy. Mach. Learn. Knowl. Extr. 2021, 3, 720–739. [Google Scholar] [CrossRef]
- Espinheira, P.L.; da Silva, L.C.M.; Silva, A.d.O.; Ospina, R. Model Selection Criteria on Beta Regression for Machine Learning. Mach. Learn. Knowl. Extr. 2019, 1, 427–449. [Google Scholar] [CrossRef] [Green Version]
- Dziak, J.J.; Coffman, D.L.; Lanza, S.T.; Li, R.; Jermiin, L.S. Sensitivity and specificity of information criteria. Brief. Bioinform. 2019, 21, 553–565. [Google Scholar] [CrossRef]
- Jimenez, J.; Navarro, L.; Quintero, M.C.G.; Pardo, M. Multivariate Statistical Analysis for Training Process Optimization in Neural Networks-Based Forecasting Models. Appl. Sci. 2021, 11, 3552. [Google Scholar] [CrossRef]
- Sayers, A. QSUB: Stata Module to Emulate a Cluster Environment Using Your Desktop PC. EconPapers. 2017. Available online: https://EconPapers.repec.org/RePEc:boc:bocode:s458366 (accessed on 1 June 2022).
- Pearson, K. Mathematical contributions to the theory of evolution—III. Regression, heredity, and panmixia. Philos. Trans. R. Soc. Lond. Ser. A 1896, 187, 253–318. [Google Scholar]
- Pearson, K.; Filon, L.N.G. Mathematical contributions to the theory of evolution. IV. On the probable errors of frequency constants and on the influence of random selection on variation and correlation. Philos. Trans. R. Soc. Lond. Ser. A 1898, 191, 229–311. [Google Scholar]
- Rauchwerger, L.; Padua, D. Parallelizing while loops for multiprocessor systems. In Proceedings of the 9th International Parallel Processing Symposium, Santa Barbara, CA, USA, 25–28 April 1995; pp. 347–356. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.-K.; Li, W.; Tong, X. Parallelization of AdaBoost algorithm on multi-core processors. In Proceedings of the 2008 IEEE Workshop on Signal Processing Systems 2008, Washington, DC, USA, 8–10 October 2008; pp. 275–280. [Google Scholar] [CrossRef]
- Williams, G. Data Mining with Rattle and R: The Art of Excavating Data for Knowledge Discovery; Springer: Berlin/Heidelberg, Germany, 2011; pp. 269–291. [Google Scholar]
- Munafò, M.R.; Smith, G.D. Robust research needs many lines of evidence. Nature 2018, 553, 399–401. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schober, P.; Boer, C.; Schwarte, L.A. Correlation coefficients. Anesth. Analg. 2018, 126, 1763–1768. [Google Scholar] [CrossRef]
- Mukaka, M.M. Statistics corner: A guide to appropriate use of correlation coefficient in medical research. Malawi Med. J. 2012, 24, 69–71. [Google Scholar]
- Corlett, M.T.; Pethick, D.W.; Kelman, K.R.; Jacob, R.H.; Gardner, G.E. Consumer Perceptions of Meat Redness Were Strongly Influenced by Storage and Display Times. Foods 2021, 10, 540. [Google Scholar] [CrossRef]
- Lace, J.W.; Handal, P.J. Psychometric Properties of the Daily Spiritual Experiences Scale: Support for a Two-Factor Solution, Concurrent Validity, and Its Relationship with Clinical Psychological Distress in University Students. Religions 2017, 8, 123. [Google Scholar] [CrossRef] [Green Version]
- Berthold, D.P.; Morikawa, D.; Muench, L.N.; Baldino, J.B.; Cote, M.P.; Creighton, R.A.; Denard, P.J.; Gobezie, R.; Lederman, E.; Romeo, A.A.; et al. Negligible Correlation between Radiographic Measurements and Clinical Outcomes in Patients Following Primary Reverse Total Shoulder Arthroplasty. J. Clin. Med. 2021, 10, 809. [Google Scholar] [CrossRef] [PubMed]
- Roberts, D.R.; Bahn, V.; Ciuti, S.; Boyce, M.S.; Elith, J.; Guillera-Arroita, G.; Hauenstein, S.; Lahoz-Monfort, J.J.; Schröder, B.; Thuiller, W.; et al. Cross-validation strategies for data with temporal, spatial, hierarchical, or phylogenetic structure. Ecography 2017, 40, 913–929. [Google Scholar] [CrossRef]
- Link, W.A.; Sauer, J.R. Bayesian Cross-Validation for Model Evaluation and Selection, with Application to the North American Breeding Survey. Ecology 2015, 97, 1746–1758. [Google Scholar] [CrossRef] [PubMed]
- Bayerl, P.S.; Akhgar, B. Surveillance and falsification implications for open source intelligence investigations. Commun. ACM 2015, 58, 62–69. [Google Scholar] [CrossRef]
- Giacomello, G.; Martinelli, D. Crystal Clear: Investigating Databases for Research, the Case of Drone Strikes. Data 2021, 6, 124. [Google Scholar] [CrossRef]
- Sierras-Davo, M.C.; Lillo-Crespo, M.; Verdu, P.; Karapostoli, A. Transforming the Future Healthcare Workforce across Europe through Improvement Science Training: A Qualitative Approach. Int. J. Environ. Res. Public Health 2021, 18, 1298. [Google Scholar] [CrossRef]


| Platform/ No.of Allocated Logical Cores (nalc) | Intel Xeon Gold 6240 CascadeLake, 2.6 GHz (VM), SCSI Disk | Intel Xeon Gold 6240 CascadeLake, 2.6 GHz (VM), ImDisk RAMdisk | Intel Core i7 4710HQ, 3.5 GHz (PM),SSD | Intel Core i7 4710HQ, 3.5 GHz (PM), ImDisk RAMdisk | Atom N550, 1.5 GHz (PM), SATA HDD |
|---|---|---|---|---|---|
| 1 (PCDM) | 124 (between 00:02:32 as hh:mm:ss and 00:04:36 in the 3rd recorded simulation, namely 3.pcdm-RaaS-IS(1x).mp4 *) | 115 ** | 85 | 85 | 800 |
| 2 | 51 | 50 *** | 38 | 36 | 421 |
| 4 | 36 | 32 | 29 | 27 | 380 |
| 6 | 36 | 33 (between 00:04:23 and 00:04:56 in the 4th recorded simulation, namely 4.pcdm4mp-RaaS-IS(6x)RAMdisks.mp4 ****) | 30 | 28 | N/A |
| 8 | 66 | 47 | 31 | 29 | N/A |
| 10 | 69 | 64 | N/A | N/A | N/A |
| 12 | 85 | 74 | N/A | N/A | N/A |
| 14 | 94 | 86 | N/A | N/A | N/A |
| 16 (15 really used) | 112 (between 00:02:08 and 00:04:00 in the 5th recorded simulation, namely 5.pcdm4mp-RaaS-IS(16x).mp4 *****) | 92 | N/A | N/A | N/A |
| Task No. | Var.Chunk (Starting Letter in var. Names) | No.of.Vars. in the Chunk | Processing Time (Xeon CPU, 1st Config.) | Processing Time (Core i7 CPU, 2nd Config.) | Processing Time (Atom CPU, 3rd Config.) |
|---|---|---|---|---|---|
| 1 | A | 204 | 25 | 20 | 173 |
| 2 | B | 25 | 2 | 2 | 14 |
| 3 | C | 43 | 6 | 5 | 45 |
| 4 | D | 56 | 6 | 6 | 49 |
| 5 | E | 305 | 28 | 21 | 184 |
| 6 | F | 129 | 22 | 13 | 117 |
| 7 | G | 124 | 13 | 9 | 88 |
| 8 | H | 30 | 1 | 1 | 10 |
| 9 | I | 2 | 0 | 0 | 1 |
| 10 | S | 20 | 4 | 3 | 27 |
| 11 | T | 1 | 0 | 0 | 1 |
| 12 | V | 7 | 0 | 0 | 2 |
| 13 | W | 11 | 1 | 1 | 3 |
| 14 | X | 51 | 7 | 6 | 50 |
| 15 | Y | 37 | 8 | 6 | 58 |
| Total | - | 1045 | 123 | 93 | 822 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Homocianu, D.; Airinei, D. PCDM and PCDM4MP: New Pairwise Correlation-Based Data Mining Tools for Parallel Processing of Large Tabular Datasets. Mathematics 2022, 10, 2671. https://doi.org/10.3390/math10152671
Homocianu D, Airinei D. PCDM and PCDM4MP: New Pairwise Correlation-Based Data Mining Tools for Parallel Processing of Large Tabular Datasets. Mathematics. 2022; 10(15):2671. https://doi.org/10.3390/math10152671
Chicago/Turabian StyleHomocianu, Daniel, and Dinu Airinei. 2022. "PCDM and PCDM4MP: New Pairwise Correlation-Based Data Mining Tools for Parallel Processing of Large Tabular Datasets" Mathematics 10, no. 15: 2671. https://doi.org/10.3390/math10152671
APA StyleHomocianu, D., & Airinei, D. (2022). PCDM and PCDM4MP: New Pairwise Correlation-Based Data Mining Tools for Parallel Processing of Large Tabular Datasets. Mathematics, 10(15), 2671. https://doi.org/10.3390/math10152671







