Generating Mathematical Expressions for Estimation of Atomic Coordinates of Carbon Nanotubes Using Genetic Programming Symbolic Regression
Abstract
:1. Introduction
- Is it possible to apply the GPSR algorithm on the publicly available dataset to obtain MEs that could estimate the calculated coordinates of CNTs with high estimation accuracy?
- Is it possible to develop and implement the RHVS method to find the optimal combination of GPSR hyperparameter values using which the MEs are obtained with high estimation accuracy?
- Is it possible to obtain a robust set of MEs and prevent overfitting by training of GPSR algorithm using the 5FCV training process?
2. Materials and Methods
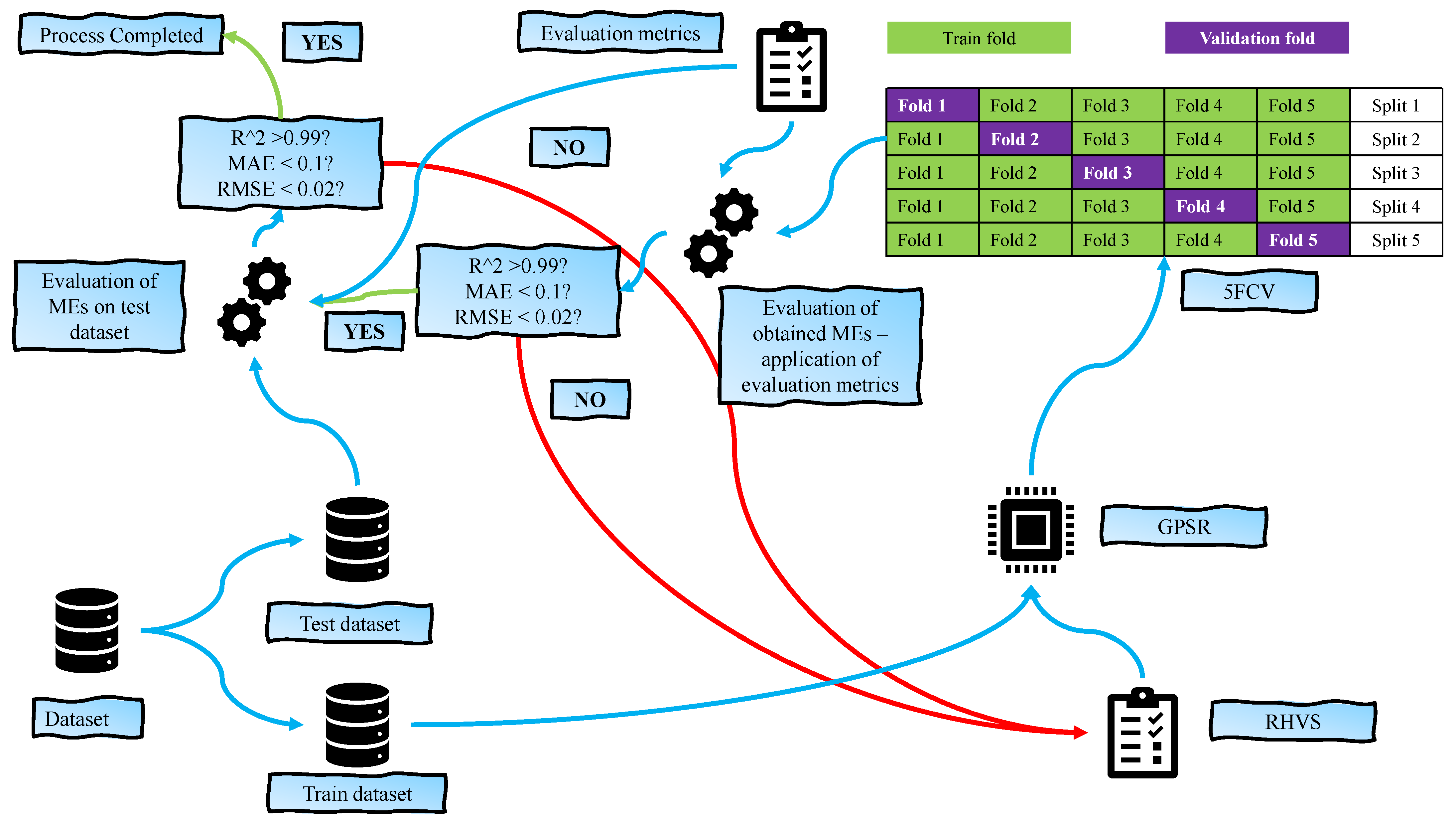
2.1. Research Methodology
- Dataset—Perform initial dataset analysis (correlation analysis, outliers analysis) to see if application of addition preprocessing techniques is required. After analysis, divide the dataset into train and test parts.
- Genetic Programming Symbolic Regressor—Application of Genetic Programming Symbolic Regressor method with random hyperparameter selection, and trained using 5-fold cross-validation process.
- Results—Application of evaluation metric methods on obtained MEs to determine which one of them has the highest classification performance and plotting the results.
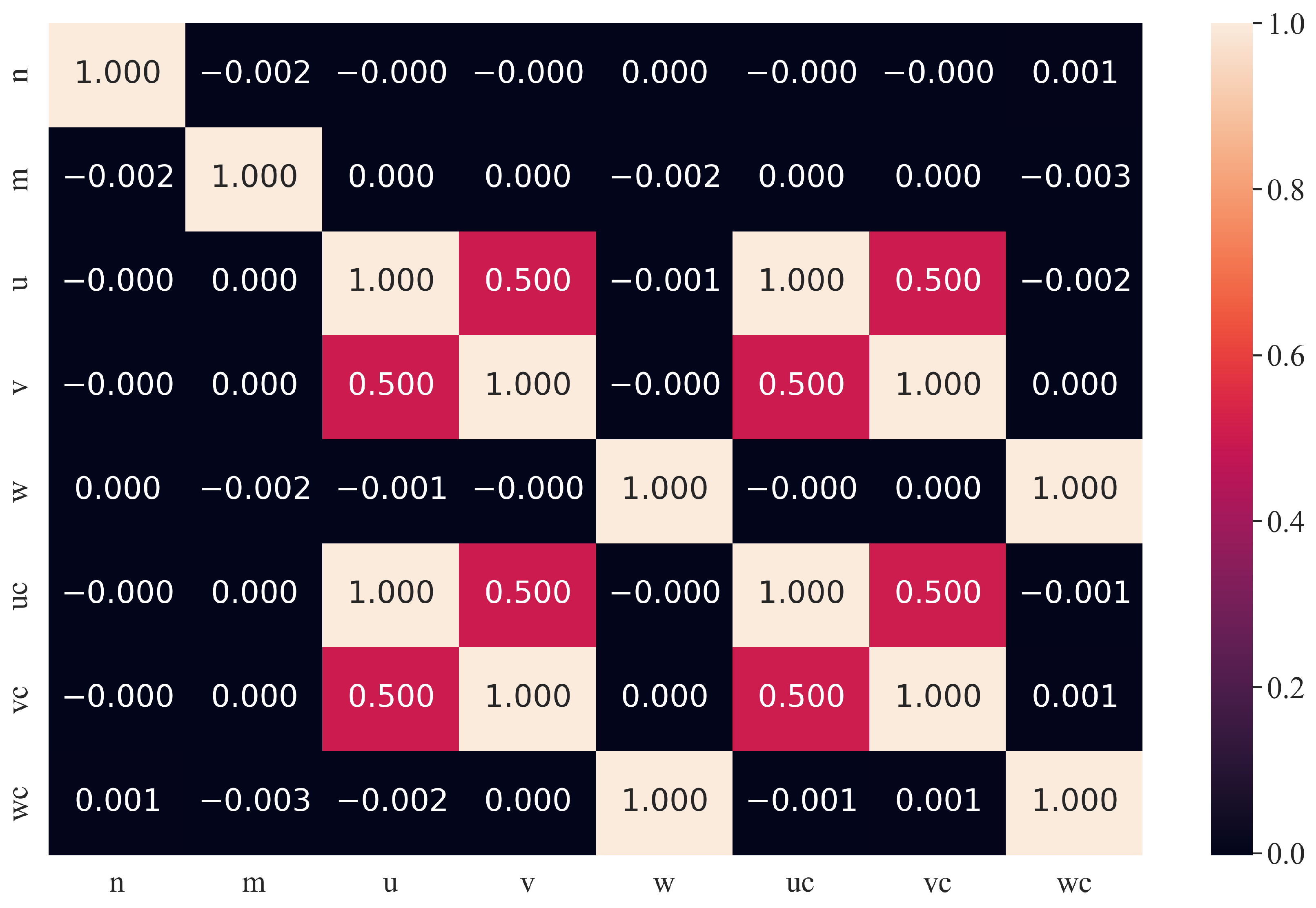
2.2. Dataset Description and Statistical Analysis
- chiral indices n—the parameter of the selected chiral vector;
- chiral indices m—the parameter of the selected chiral vector;
- initial atomic coordinate u—randomly generated parameter of the initial atomic coordinates of all carbon atoms;
- initial atomic coordinate v—randomly generated parameter of the initial atomic coordinate of all carbon atoms;
- initial atomic coordinate w—randomly generated parameters of the initial atomic coordinate of all carbon atoms;
- calculated atomic coordinates ;
- calculated atomic coordinates ;
- calculated atomic coordinates .
2.3. Genetic Programming Symbolic Regression
2.4. Evaluation Metrics
2.5. Training and Testing Process
3. Results
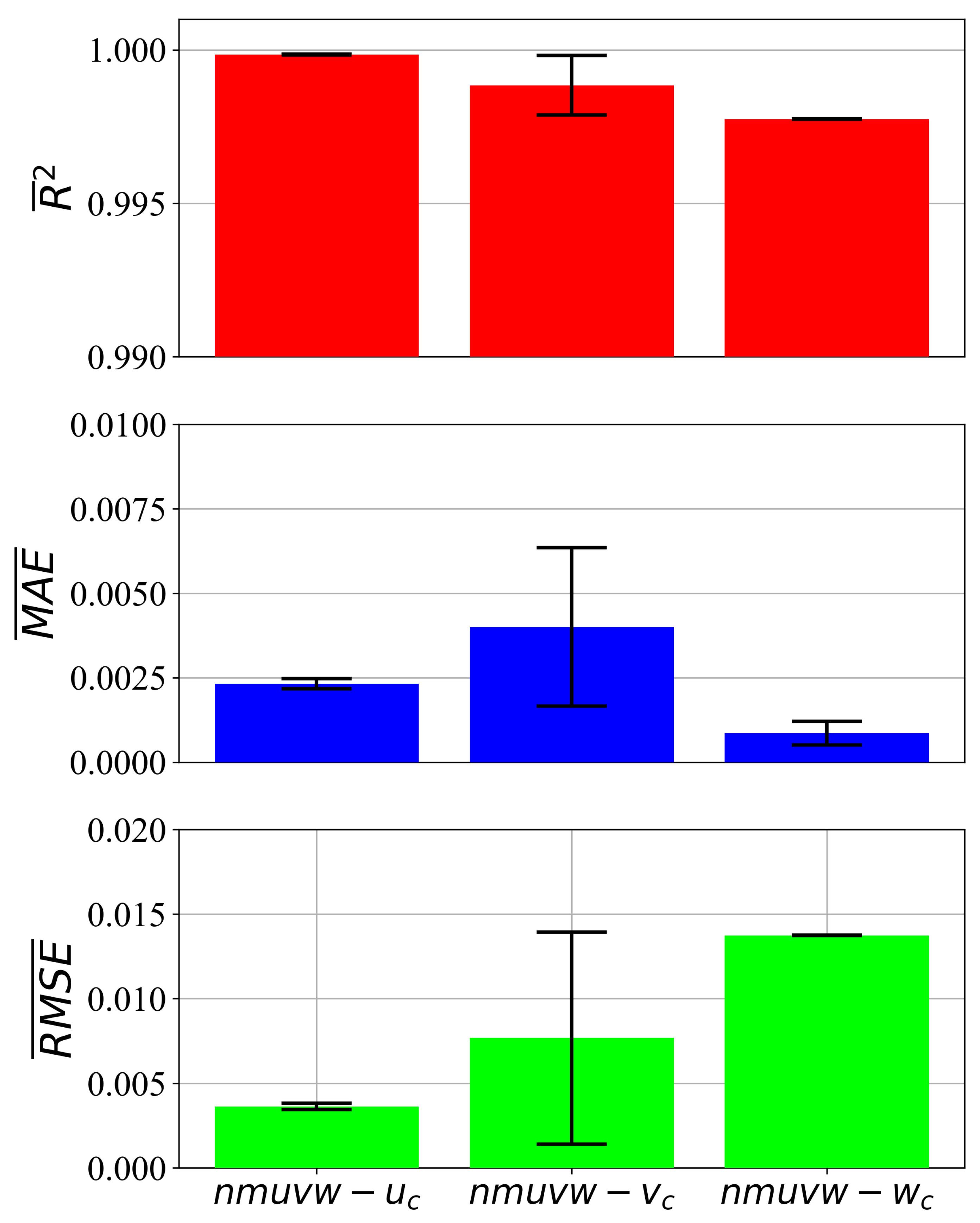
3.1. All Input Variables
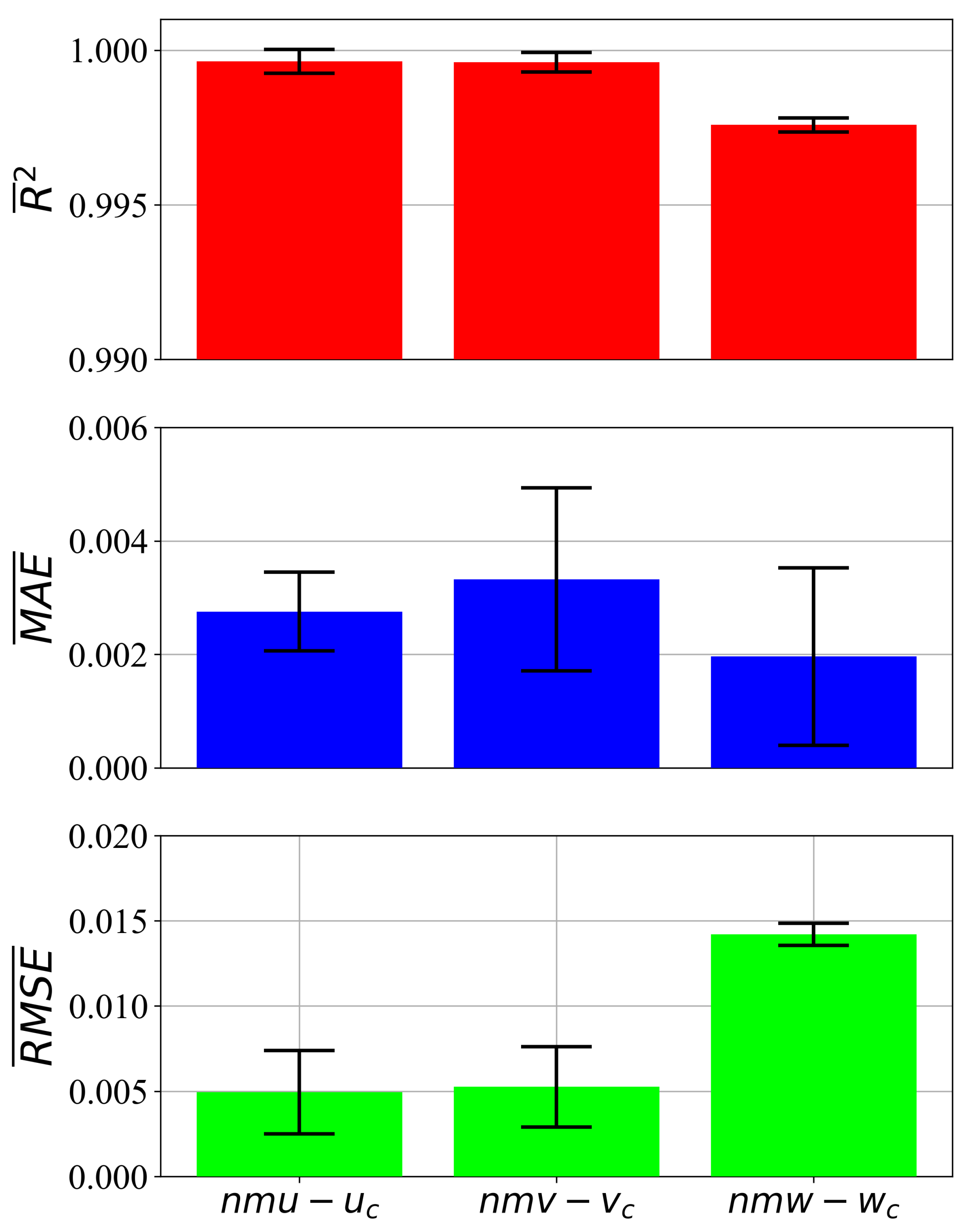
3.2. Selected Input Variables
4. Discussion
5. Conclusions
- The GPSR can be applied to obtain MEs that can estimate the calculated atomic coordinates of CNTs with high estimation accuracy.
- Using GPSR with RHVS, the optional GPSR hyperparameter values were randomly selected, using which a robust set of MEs were obtained that can estimate the calculated coordinates of CNTs with high accuracy.
- By training GPSR with 5FCV, the robust sets of MEs were obtained in each case with high mean values of evaluation metrics and small standard deviation values, which proves that overfitting did not occur.
- The main benefit of GPSR is that ME is obtained after training that connects input variables with an output variable that can be easily used and requires low computational resources to predict the output.
- The application of the RHVS method in the GPSR algorithm is in most cases a quicker way of finding the optimal combination of hyperparameter values when compared to the classic grid search method, since it does not execute all possible combinations of defined hyperparameters.
- The training of GPSR using 5FCV when compared to the classic training procedure is a good way of obtaining a robust system/set of MEs, since, on each split of 5FCV, one ME is obtained, and then on 5FCV, a total of five MEs are obtained. A larger number of obtained MEs can generally prevent potential ovefitting, which might occur in classic AI algorithm training, and the predicted result with one ME can be solidified multiple times.
- Generally, the GPSR combination of hyperparameters can greatly prolong its execution, so finely tuning its hyperparameters is mandatory.
- Sometimes, the application of GPSR and RHVS can take a couple of days to find the GPSR optimal hyperparameter values. The prerequisite for finding optimal hyperparameters is a good correlation between input and output dataset variables and a dataset without outliers, if possible.
- Defining and testing the boundaries of the GPSR algorithm is the prerequisite process for the creation of the RHVS method. However, this process can take some time, since it is required that each boundary is tested and adjusted if necessary.
- The combination of GPSR with the RHVS method trained using 5FCV is a time-consuming process, especially of boundaries of Size_pop and max_gen values used in this research. The very large population and large number of generations can generally lead to long GPSR execution.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
Appendix A.1. The Modification of Mathematical Functions Used in GPSR
Appendix A.2. How to Obtain and Use Generated MEs in This Research
References
- Dresselhaus, M.S.; Dresselhaus, G.; Eklund, P.; Rao, A. Carbon Nanotubes; Springer: Berlin/Heidelberg, Germany, 2000. [Google Scholar]
- Popov, V.N. Carbon nanotubes: Properties and application. Mater. Sci. Eng. R Rep. 2004, 43, 61–102. [Google Scholar] [CrossRef]
- He, M.; Zhang, S.; Wu, Q.; Xue, H.; Xin, B.; Wang, D.; Zhang, J. Designing catalysts for chirality-selective synthesis of single-walled carbon nanotubes: Past success and future opportunity. Adv. Mater. 2019, 31, 1800805. [Google Scholar] [CrossRef] [PubMed]
- Takakura, A.; Beppu, K.; Nishihara, T.; Fukui, A.; Kozeki, T.; Namazu, T.; Miyauchi, Y.; Itami, K. Strength of carbon nanotubes depends on their chemical structures. Nat. Commun. 2019, 10, 3040. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Z.; Wang, G.; Wolan, B.F.; Wu, N.; Wang, C.; Zhao, S.; Yue, S.; Li, B.; He, W.; Liu, J.; et al. Printable aligned single-walled carbon nanotube film with outstanding thermal conductivity and electromagnetic interference shielding performance. Nano-Micro Lett. 2022, 14, 179. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, K.S. Carbon nanotubes? Properties and applications: A review. Carbon Lett. 2013, 14, 131–144. [Google Scholar] [CrossRef]
- Jafari, S. Engineering applications of carbon nanotubes. In Carbon Nanotube-Reinforced Polymers; Elsevier: Amsterdam, The Netherlands, 2018; pp. 25–40. [Google Scholar]
- Nurazzi, N.; Sabaruddin, F.; Harussani, M.; Kamarudin, S.; Rayung, M.; Asyraf, M.; Aisyah, H.; Norrrahim, M.; Ilyas, R.; Abdullah, N.; et al. Mechanical performance and applications of cnts reinforced polymer composites—A review. Nanomaterials 2021, 11, 2186. [Google Scholar] [CrossRef]
- Anantram, M.; Leonard, F. Physics of carbon nanotube electronic devices. Rep. Prog. Phys. 2006, 69, 507. [Google Scholar] [CrossRef]
- Bianco, A.; Kostarelos, K.; Prato, M. Applications of carbon nanotubes in drug delivery. Curr. Opin. Chem. Biol. 2005, 9, 674–679. [Google Scholar] [CrossRef]
- Talla, J.A.; Salman, S.A. Electronic structure tuning and band gap engineering of carbon nanotubes: Density functional theory. Nanosci. Nanotechnol. Lett. 2015, 7, 381–386. [Google Scholar] [CrossRef]
- Sun, G.; Kürti, J.; Rajczy, P.; Kertesz, M.; Hafner, J.; Kresse, G. Performance of the Vienna ab initio simulation package (VASP) in chemical applications. J. Mol. Struct. Theochem 2003, 624, 37–45. [Google Scholar] [CrossRef]
- Giannozzi, P.; Baroni, S.; Bonini, N.; Calandra, M.; Car, R.; Cavazzoni, C.; Ceresoli, D.; Chiarotti, G.L.; Cococcioni, M.; Dabo, I.; et al. Quantum espresso: A modular and open-source software project for quantum simulations of materials. J. Phys. Condens. Matter 2009, 21, 395502. [Google Scholar] [CrossRef]
- Acı, M.; Avcı, M. Artificial neural network approach for atomic coordinate prediction of carbon nanotubes. Appl. Phys. A 2016, 122, 1–14. [Google Scholar] [CrossRef]
- Aci, M.; Avci, M.; Aci, Ç. Destek Vektör regresyonu yöntemiyle karbon nanotüp benzetim süresinin kisaltilmasi. Gazi Üniversitesi Mühendislik Mimarlık Fakültesi Dergisi 2017, 32. [Google Scholar] [CrossRef]
- Obilor, E.I.; Amadi, E.C. Test for significance of Pearson’s correlation coefficient. Int. J. Innov. Math. Stat. Energy Policies 2018, 6, 11–23. [Google Scholar]
- Vinutha, H.; Poornima, B.; Sagar, B. Detection of outliers using interquartile range technique from intrusion dataset. In Information and Decision Sciences, Proceedings of the 6th International Conference on FICTA, Bhubaneswar, Odisha, 14 October 2017; Springer: Singapore, 2018; pp. 511–518. [Google Scholar]
- Poli, R.; Langdon, W.B.; McPhee, N.F. A Field Guide to Genetic Programming; Lulu Press: Morrisville, NC, USA, 2008; 250p, ISBN 978-1-4092-0073-4. [Google Scholar]
- Luke, S.; Panait, L. A survey and comparison of tree generation algorithms. In Proceedings of the 3rd Annual Conference on Genetic and Evolutionary Computation, San Francisco, CA, USA, 7–11 July 2001; pp. 81–88. [Google Scholar]
- Chai, T.; Draxler, R.R. Root mean square error (RMSE) or mean absolute error (MAE). Geosci. Model Dev. Discuss. 2014, 7, 1525–1534. [Google Scholar]
- Di Bucchianico, A. Coefficient of determination (R2). In Encyclopedia of Statistics in Quality and Reliability; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2008. [Google Scholar]
- Anđelić, N.; Baressi Šegota, S. Development of Symbolic Expressions Ensemble for Breast Cancer Type Classification Using Genetic Programming Symbolic Classifier and Decision Tree Classifier. Cancers 2023, 15, 3411. [Google Scholar] [CrossRef]
- Awan, S.E.; Shamim, R.; Awais, M.; Irum, S.; Shoaib, M.; Raja, M.A.Z. Convective flow dynamics with suspended carbon nanotubes in the presence of magnetic dipole: Intelligent solution predicted Bayesian regularization networks. Tribol. Int. 2023, 187, 108685. [Google Scholar] [CrossRef]
- Awan, S.E.; Ali, F.; Awais, M.; Shoaib, M.; Raja, M.A.Z. Intelligent Bayesian regularization-based solution predictive procedure for hybrid nanoparticles of AA7072-AA7075 oxide movement across a porous medium. ZAMM-J. Appl. Math. Mech. Angew. Math. Mech. 2023, 103, e202300043. [Google Scholar] [CrossRef]
- Raja, M.A.Z.; Sabati, M.; Parveen, N.; Awais, M.; Awan, S.E.; Chaudhary, N.I.; Shoaib, M.; Alquhayz, H. Integrated intelligent computing application for effectiveness of Au nanoparticles coated over MWCNTs with velocity slip in curved channel peristaltic flow. Sci. Rep. 2021, 11, 22550. [Google Scholar] [CrossRef]
- Awan, S.E.; Awais, M.; Shamim, R.; Raja, M.A.Z. Novel design of intelligent Bayesian networks to study the impact of magnetic field and Joule heating in hybrid nanomaterial flow with applications in medications for blood circulation. Tribol. Int. 2023, 189, 108914. [Google Scholar] [CrossRef]
- Awan, S.E.; Awais, M.; Raja, M.A.Z.; Rehman, S.U.; Shu, C.M. Bayesian regularization knack-based intelligent networks for thermo-physical analysis of 3D MHD nanofluidic flow model over an exponential stretching surface. Eur. Phys. J. Plus 2023, 138, 2. [Google Scholar] [CrossRef]
- Awan, S.E.; Raja, M.A.Z.; Awais, M.; Shu, C.M. Intelligent Bayesian regularization networks for bio-convective nanofluid flow model involving gyro-tactic organisms with viscous dissipation, stratification and heat immersion. Eng. Appl. Comput. Fluid Mech. 2021, 15, 1508–1530. [Google Scholar] [CrossRef]
- Wahid, M.A.; Bukhari, S.H.R.; Maqsood, M.; Aadil, F.; Khan, M.I.; Awan, S.E. Parametric estimation scheme for aircraft fuel consumption using machine learning. Neural Comput. Appl. 2023, 35, 24925–24946. [Google Scholar] [CrossRef]
- Anđelić, N.; Lorencin, I.; Glučina, M.; Car, Z. Mean Phase Voltages and Duty Cycles Estimation of a Three-Phase Inverter in a Drive System Using Machine Learning Algorithms. Electronics 2022, 11, 2623. [Google Scholar] [CrossRef]
- Baressi Šegota, S.; Mrzljak, V.; Anđelić, N.; Poljak, I.; Car, Z. Use of Synthetic Data in Maritime Applications for the Problem of Steam Turbine Exergy Analysis. J. Mar. Sci. Eng. 2023, 11, 1595. [Google Scholar] [CrossRef]


| Reference | AI-Algorithms | Estimation Perofrmance |
|---|---|---|
| [14] | FFNN | |
| FITNET | ||
| CFNN | ||
| GRNN |
| Count | Mean | Std | Min | Max | |
|---|---|---|---|---|---|
| n | 10,721 | 8.225725 | 2.138919 | 2 | 12 |
| m | 3.337189 | 1.683881 | 1 | 6 | |
| u | 0.500064 | 0.286524 | 0.045149 | 0.954851 | |
| v | 0.500072 | 0.286495 | 0.045149 | 0.954851 | |
| w | 0.499637 | 0.288503 | 6.10 | 0.999411 | |
| 0.500064 | 0.290935 | 0.038504 | 0.961496 | ||
| 0.500072 | 0.291012 | 0.03893 | 0.96107 | ||
| 0.499834 | 0.289095 | 0 | 1 |
| Case | Input Variables | Output Variable | ||||
|---|---|---|---|---|---|---|
| - | n | m | u | v | w | |
| Variables represented in GPSR | ||||||
| - | n | m | u | v | w | |
| Variables represented in GPSR | ||||||
| - | n | m | u | v | w | |
| Variables represented in GPSR | ||||||
| - | n | m | u | / | / | |
| Variables represented in GPSR | / | / | ||||
| - | n | m | v | / | / | |
| Variables represented in GPSR | / | / | ||||
| - | n | m | w | / | / | |
| Variables represented in GPSR | / | / | ||||
| Hyperparameter Name | Lower Boundary | Upper Boundary |
|---|---|---|
| Size_Pop | 1000 | 2000 |
| cnst_vals | −10,000 | 10,000 |
| init_depth | 3 | 15 |
| t_size | 100 | 500 |
| crossover | 0.001 | 1 |
| point mutation | 0.001 | 1 |
| hoist mutation | 0.001 | 1 |
| subtree mutation | 0.001 | 1 |
| crit_stop | 0 | |
| max_gen | 300 | 500 |
| max_samp | 0.99 | 1 |
| CO-Pars | 0 |
| Case | GPSR Hyperparameter Values (Size_Pop, Max_Gen, t_Size, Init_Depth Crossover, Subtree Mutation, Hoist Mutation, Point Mutation, Crit_Stop, Max_Samp, CO-Pars) |
|---|---|
| - | 1166, 380, 366, (3, 9), 0.021, 0.953, 0.02, 0.0039, , 0.99, (−2929.06, 8732.5), 0.00086 |
| - | 1580, 465, 419, (7, 8), 0.032, 0.96, 0.0042, 0.0021, , 0.99, (−2239.26, 5430.02), |
| - | 1658, 481, 249, (3, 15), 0.027, 0.95, 0.0054, 0.012, , 0.99, (−4451.28, 2031.56), 0.00043 |
| Dataset Type | Depth | Length | Mean Depth | Mean Length |
|---|---|---|---|---|
| - | 16/28/17/19/21 | 64/142/58/36/72 | 20.2 | 74.5 |
| - | 38/41/34/34/31 | 285/517/224/250/186 | 35.6 | 292.4 |
| - | 19/7/13/10/17 | 212/18/30/16/51 | 13.2 | 65.4 |
| Case | GPSR Hyperparameter Values (Size_Pop, Max_Gen, t_Size, Init_Depth Crossover, Subtree Mutation, Hoist Mutation, Point Mutation, Crit_Stop, Max_Samp, CO-Pars) |
|---|---|
| - | 1091, 492, 483, (7, 13), 0.01, 0.97, 0.0022, 0.013, , 0.99, (−6987.98, 9366.61), 0.00052 |
| - | 1762, 455, 234, (6, 9), 0.0022, 0.966, 0.01168, 0.018, , 0.995, (−7321.94, 8132.74), 0.00038 |
| - | 1455, 383, 124, (3, 13), 0.0017, 0.95, 0.024, 0.02, , 0.996, (−7726.074, 3767.204), 0.00031 |
| Dataset Type | Depth | Length | Mean Depth | Mean Length |
|---|---|---|---|---|
| - | 20/28/28/20/22 | 137/335/68/113/168 | 23.6 | 164.2 |
| - | 38/30/20/22/19 | 333/458/160/157/133 | 25.8 | 248.2 |
| - | 3/36/19/24/21 | 11/123/62/181/61 | 20.5 | 87.6 |
| Reference | AI-Algorithms | Estimation Perofrmance | ||
|---|---|---|---|---|
| [14] | FFNN | |||
| FITNET | ||||
| CFNN | ||||
| GRNN | ||||
| This Paper | GPSR + RHVS | Estimation of using n, m, and u as input: | Estimation of using n, m, and v as input: | Estimation of using n, m, and w as input: |
| Estimation of using all input variables: | Estimation of using all input variables: | Estimation of using all input variables: | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Anđelić, N.; Baressi Šegota, S. Generating Mathematical Expressions for Estimation of Atomic Coordinates of Carbon Nanotubes Using Genetic Programming Symbolic Regression. Technologies 2023, 11, 185. https://doi.org/10.3390/technologies11060185
Anđelić N, Baressi Šegota S. Generating Mathematical Expressions for Estimation of Atomic Coordinates of Carbon Nanotubes Using Genetic Programming Symbolic Regression. Technologies. 2023; 11(6):185. https://doi.org/10.3390/technologies11060185
Chicago/Turabian StyleAnđelić, Nikola, and Sandi Baressi Šegota. 2023. "Generating Mathematical Expressions for Estimation of Atomic Coordinates of Carbon Nanotubes Using Genetic Programming Symbolic Regression" Technologies 11, no. 6: 185. https://doi.org/10.3390/technologies11060185
APA StyleAnđelić, N., & Baressi Šegota, S. (2023). Generating Mathematical Expressions for Estimation of Atomic Coordinates of Carbon Nanotubes Using Genetic Programming Symbolic Regression. Technologies, 11(6), 185. https://doi.org/10.3390/technologies11060185







