Nanofibrillation is an Effective Method to Produce Chitin Derivatives for Induction of Plant Responses in Soybean
Abstract
1. Introduction
2. Results
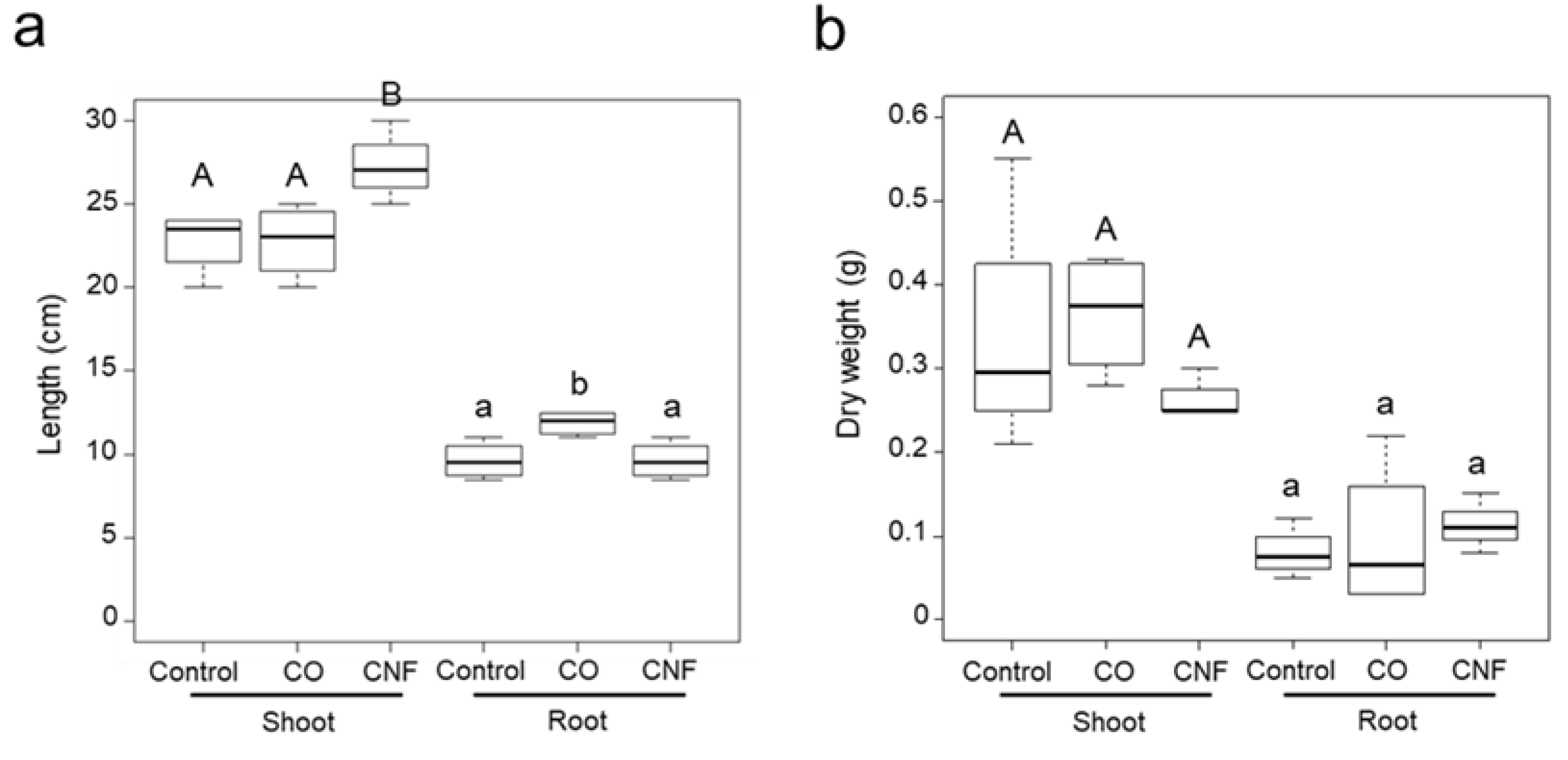
2.1. Effects of Chitins on the Plant Growth in Soybeans
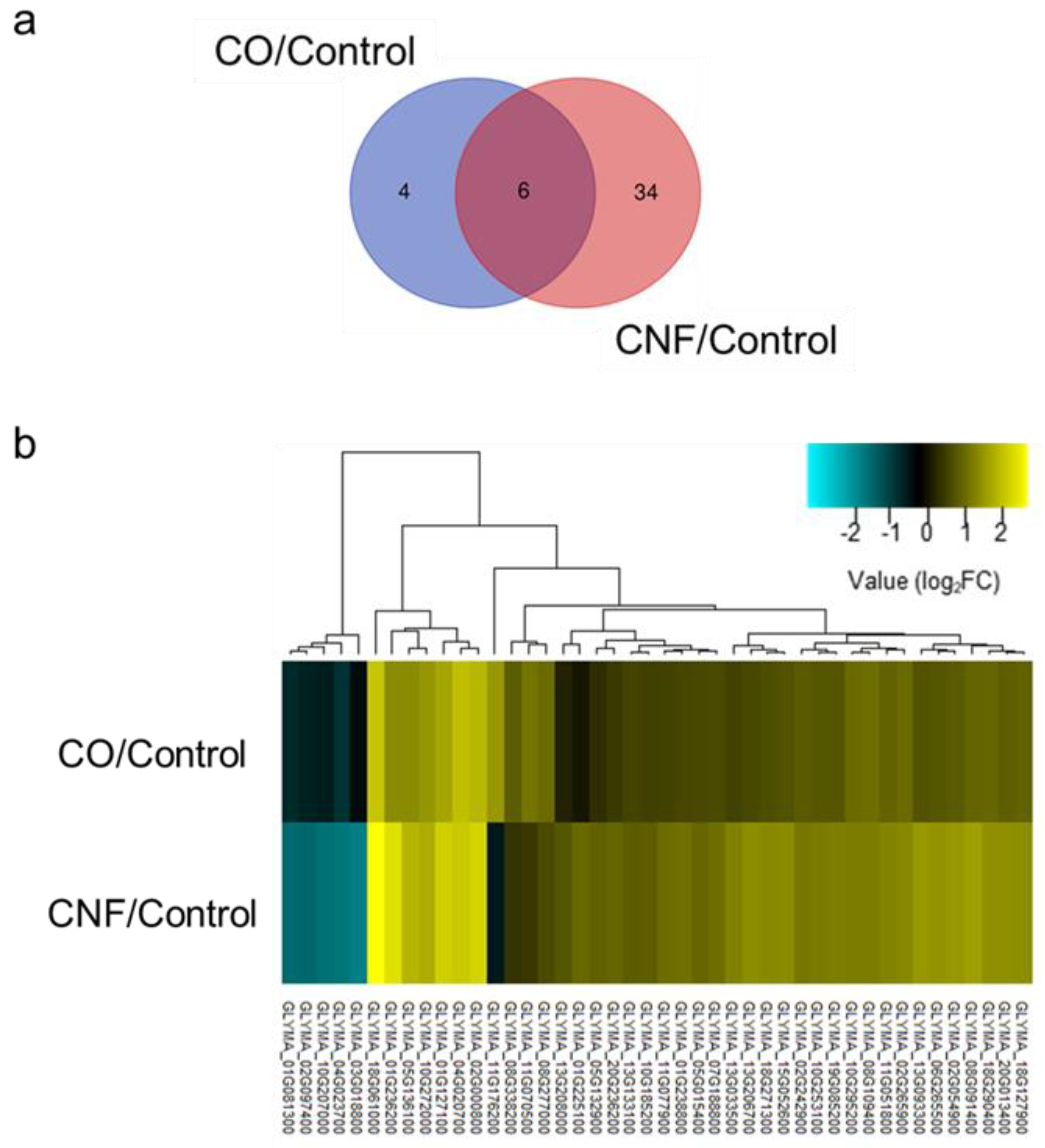
2.2. Comparison and Analysis of Differentially Expressed Genes (DEGs) in Chitin-treated Soybeans
3. Discussion
4. Materials and Methods
4.1. Preparation of Chitin Nanofiber and Chitin Oligosaccharide Water Dispersions
4.2. Plant Materials, Growth Conditions, and Measurement of Plant Growth
4.3. RNA Sequencing and Data Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sharp, R. A Review of the Applications of Chitin and Its Derivatives in Agriculture to Modify Plant-Microbial Interactions and Improve Crop Yields. Agronomy 2013, 3, 757–793. [Google Scholar] [CrossRef]
- Ifuku, S.; Saimoto, H. Chitin nanofibers: Preparations, modifications, and applications. Nanoscale 2012, 4, 3308–3318. [Google Scholar] [CrossRef]
- Pillai, C.K.S.; Paul, W.; Sharma, C.P. Chitin and chitosan polymers: Chemistry, solubility and fiber formation. Prog. Polym. Sci. 2009, 34, 641–678. [Google Scholar] [CrossRef]
- Desaki, Y.; Miyata, K.; Suzuki, M.; Shibuya, N.; Kaku, H. Plant immunity and symbiosis signaling mediated by LysM receptors. Innate Immun. 2018, 24, 92–100. [Google Scholar] [CrossRef] [PubMed]
- Egusa, M.; Matsui, H.; Urakami, T.; Okuda, S.; Ifuku, S.; Nakagami, H.; Kaminaka, H. Chitin nanofiber elucidates the elicitor activity of polymeric chitin in plants. Front. Plant Sci. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Parada, R.Y.; Egusa, M.; Aklog, Y.F.; Miura, C.; Ifuku, S.; Kaminaka, H. Optimization of nanofibrillation degree of chitin for induction of plant disease resistance: Elicitor activity and systemic resistance induced by chitin nanofiber in cabbage and strawberry. Int. J. Biol. Macromol. 2018, 118, 2185–2192. [Google Scholar] [CrossRef] [PubMed]
- Egusa, M.; Parada, R.; Aklog, Y.F.; Ifuku, S.; Kaminaka, H. Nanofibrillation enhances the protective effect of crab shells against Fusarium wilt disease in tomato. Int. J. Biol. Macromol. 2019, 128, 22–27. [Google Scholar] [CrossRef] [PubMed]
- Aklog, Y.F.; Egusa, M.; Kaminaka, H.; Izawa, H.; Morimoto, M.; Saimoto, H.; Ifuku, S. Protein/CaCO(3)/Chitin Nanofiber Complex Prepared from Crab Shells by Simple Mechanical Treatment and Its Effect on Plant Growth. Int. J. Mol. Sci. 2016, 17, 1600. [Google Scholar] [CrossRef]
- Karasov, T.L.; Chae, E.; Herman, J.J.; Bergelson, J. Mechanisms to Mitigate the Trade-Off between Growth and Defense. Plant Cell 2017, 29, 666–680. [Google Scholar] [CrossRef]
- Egusa, M.; Matsukawa, S.; Miura, C.; Nakatani, S.; Yamada, J.; Endo, T.; Ifuku, S.; Kaminaka, H. Improving nitrogen uptake efficiency by chitin nanofiber promotes growth in tomato. Int. J. Biol. Macromol. 2020, 151, 1322–1331. [Google Scholar] [CrossRef]
- Group, T.L.P.W.; Bruneau, A.; Doyle, J.J.; Herendeen, P.; Hughes, C.; Kenicer, G.; Lewis, G.; Mackinder, B.; Pennington, R.T.; Sanderson, M.J.; et al. Legume phylogeny and classification in the 21st century: Progress, prospects and lessons for other species–rich clades. TAXON 2013, 62, 217–248. [Google Scholar] [CrossRef]
- Ge, S.X.; Jung, D.; Yao, R. ShinyGO: A graphical gene-set enrichment tool for animals and plants. Bioinformatics 2019, 36, 2628–2629. [Google Scholar] [CrossRef] [PubMed]
- Miya, A.; Albert, P.; Shinya, T.; Desaki, Y.; Ichimura, K.; Shirasu, K.; Narusaka, Y.; Kawakami, N.; Kaku, H.; Shibuya, N. CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc. Natl. Acad. Sci. USA 2007, 104, 19613–19618. [Google Scholar] [CrossRef] [PubMed]
- Kaku, H.; Nishizawa, Y.; Ishii-Minami, N.; Akimoto-Tomiyama, C.; Dohmae, N.; Takio, K.; Minami, E.; Shibuya, N. Plant cells recognize chitin fragments for defense signaling through a plasma membrane receptor. Proc. Natl. Acad. Sci. USA 2006, 103, 11086–11091. [Google Scholar] [CrossRef]
- Petutschnig, E.K.; Jones, A.M.E.; Serazetdinova, L.; Lipka, U.; Lipka, V. The Lysin Motif Receptor-like Kinase (LysM-RLK) CERK1 Is a Major Chitin-binding Protein in Arabidopsis thaliana and Subject to Chitin-induced Phosphorylation. J. Biol. Chem. 2010, 285, 28902–28911. [Google Scholar] [CrossRef]
- Wan, J.; Tanaka, K.; Zhang, X.-C.; Son, G.H.; Brechenmacher, L.; Nguyen, T.H.N.; Stacey, G. LYK4, a Lysin Motif Receptor-Like Kinase, Is Important for Chitin Signaling and Plant Innate Immunity in Arabidopsis. Plant Physiol. 2012, 160, 396–406. [Google Scholar] [CrossRef]
- Bouché, N.; Fromm, H. GABA in plants: Just a metabolite? Trends Plant Sci. 2004, 9, 110–115. [Google Scholar] [CrossRef]
- Ramesh, S.A.; Tyerman, S.D.; Xu, B.; Bose, J.; Kaur, S.; Conn, V.; Domingos, P.; Ullah, S.; Wege, S.; Shabala, S.; et al. GABA signalling modulates plant growth by directly regulating the activity of plant-specific anion transporters. Nat. Commun. 2015, 6, 7879. [Google Scholar] [CrossRef]
- Boller, T.; Felix, G. A Renaissance of Elicitors: Perception of Microbe-Associated Molecular Patterns and Danger Signals by Pattern-Recognition Receptors. Annu. Rev. Plant Biol. 2009, 60, 379–406. [Google Scholar] [CrossRef]
- Torres, M.A.; Jones, J.D.G.; Dangl, J.L. Reactive oxygen species signaling in response to pathogens. Plant Physiol. 2006, 141, 373–378. [Google Scholar] [CrossRef]
- Morales, J.; Kadota, Y.; Zipfel, C.; Molina, A.; Torres, M.-A. The Arabidopsis NADPH oxidases RbohD and RbohF display differential expression patterns and contributions during plant immunity. J. Exp. Bot. 2016, 67, 1663–1676. [Google Scholar] [CrossRef] [PubMed]
- López, M.; Miranda, E.; Ramos, C.; García, H.; Neira-Carrillo, A. Activation of Early Defense Signals in Seedlings of Nicotiana benthamiana Treated with Chitin Nanoparticles. Plants 2020, 9, 607. [Google Scholar] [CrossRef] [PubMed]
- Colman, S.L.; Salcedo, M.F.; Mansilla, A.Y.; Iglesias, M.J.; Fiol, D.F.; Martín-Saldaña, S.; Alvarez, V.A.; Chevalier, A.A.; Casalongué, C.A. Chitosan microparticles improve tomato seedling biomass and modulate hormonal, redox and defense pathways. Plant Physiol. Biochem. 2019, 143, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Divya, K.; Jisha, M.S. Chitosan nanoparticles preparation and applications. Environ. Chem. Lett. 2018, 16, 101–112. [Google Scholar] [CrossRef]
- Ifuku, S.; Nogi, M.; Abe, K.; Yoshioka, M.; Morimoto, M.; Saimoto, H.; Yano, H. Preparation of Chitin Nanofibers with a Uniform Width as α-Chitin from Crab Shells. Biomacromolecules 2009, 10, 1584–1588. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2012, 29, 15–21. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2013, 30, 923–930. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2009, 26, 139–140. [Google Scholar] [CrossRef]
| Gene ID | Description | CO/Control | CNF/Control | ||
|---|---|---|---|---|---|
| log2FC | FDR a | log2FC | FDR a | ||
| CO only | |||||
| GLYMA_11G176200 | Uncharacterized protein | 1.76 | 2.74 | −0.31 | 1.00 |
| GLYMA_11G070500 | Isoflavone reductase, NmrA domain-containing protein | 1.36 | 1.98 | 0.67 | 9.43 |
| GLYMA_08G277000 | TRANSKETOLASE_1 domain-containing protein | 1.26 | 1.17 | 0.88 | 3.36 |
| GLYMA_08G338200 | Glycosyltransferase (EC 2.4.1.-) | 1.09 | 3.04 | 0.62 | 1.00 |
| CO and CNF | |||||
| GLYMA_04G020700 | Syringolide-induced protein 14-1-1 | 2.22 | 1.47 | 2.34 | 7.60 |
| GLYMA_10G272000 | Uncharacterized protein | 1.74 | 1.99 | 2.00 | 4.01 |
| GLYMA_05G136100 | Glutamate decarboxylase (EC 4.1.1.15) | 1.63 | 2.30 | 2.11 | 6.65 |
| GLYMA_08G109400 | Chalcone synthase 1 (EC 2.3.1.74) | 1.30 | 7.11 | 1.45 | 1.79 |
| GLYMA_10G295200 | C2H2-type domain-containing protein | 1.25 | 1.50 | 1.47 | 3.87 |
| GLYMA_02G265900 | Uncharacterized protein | 1.24 | 5.89 | 1.57 | 2.97 |
| CNF only | |||||
| GLYMA_18G061000 | PPM-type phosphatase domain-containing protein | 2.28 | 2.42 | 2.98 | 3.85 |
| GLYMA_01G236200 | Uncharacterized protein | 1.61 | 1.00 | 2.59 | 3.20 |
| GLYMA_02G000800 | BHLH domain-containing protein | 2.13 | 5.36 | 2.46 | 1.51 |
| GLYMA_01G127100 | Dirigent protein | 1.91 | 6.91 | 2.42 | 5.28 |
| GLYMA_08G091400 | Glutamate decarboxylase (EC 4.1.1.15) | 1.18 | 2.42 | 1.86 | 7.46 |
| GLYMA_13G093300 | PBPe domain-containing protein | 0.96 | 5.93 | 1.76 | 1.79 |
| GLYMA_02G054900 | Uncharacterized protein | 1.06 | 1.00 | 1.76 | 1.42 |
| GLYMA_06G265500 | GRAS domain-containing protein | 1.01 | 1.47 | 1.71 | 1.37 |
| GLYMA_20G013400 | Uncharacterized protein | 1.11 | 9.36 | 1.67 | 1.82 |
| GLYMA_18G290400 | Uncharacterized protein | 1.19 | 1.00 | 1.66 | 2.68 |
| GLYMA_18G127900 | TIR-NBS-LRR type disease resistance protein | 1.14 | 5.75 | 1.65 | 1.79 |
| GLYMA_13G206700 | TPT domain-containing protein | 0.85 | 8.68 | 1.65 | 5.62 |
| GLYMA_15G052600 | Peroxidase (EC 1.11.1.7) | 0.95 | 1.00 | 1.63 | 8.57 |
| GLYMA_18G271300 | Protein kinase domain-containing protein | 0.93 | 2.41 | 1.59 | 2.18 |
| GLYMA_11G051800 | Uncharacterized protein | 1.16 | 5.82 | 1.53 | 1.49 |
| GLYMA_13G033500 | Protein kinase domain-containing protein | 0.78 | 1.00 | 1.50 | 2.55 |
| GLYMA_19G085200 | Uncharacterized protein | 1.01 | 9.23 | 1.47 | 8.31 |
| GLYMA_10G253100 | Uncharacterized protein | 1.01 | 3.80 | 1.43 | 3.85 |
| GLYMA_02G242900 | RING-type E3 ubiquitin transferase (EC 2.3.2.27) | 1.06 | 5.80 | 1.39 | 3.87 |
| GLYMA_01G238800 | MFS domain-containing protein | 0.81 | 7.33 | 1.30 | 7.50 |
| GLYMA_07G188800 | Uncharacterized protein | 0.88 | 2.42 | 1.28 | 5.87 |
| GLYMA_11G077900 | Rhodanese domain-containing protein | 0.77 | 7.06 | 1.27 | 1.42 |
| GLYMA_01G225100 | PPM-type phosphatase domain-containing protein | 0.28 | 1.00 | 1.23 | 3.46 |
| GLYMA_05G015400 | Phospholipid-transporting ATPase (EC 7.6.2.1) | 0.85 | 2.42 | 1.20 | 3.20 |
| GLYMA_20G236200 | Uncharacterized protein | 0.68 | 1.00 | 1.19 | 4.31 |
| GLYMA_05G132900 | Uncharacterized protein | 0.55 | 1.00 | 1.12 | 3.87 |
| GLYMA_10G185200 | Zeta_toxin domain-containing protein | 0.77 | 5.92 | 1.12 | 3.87 |
| GLYMA_13G133100 | Uncharacterized protein | 0.79 | 5.47 | 1.08 | 4.80 |
| GLYMA_13G208000 | Peptidase A1 domain-containing protein | 0.41 | 1.00 | 1.00 | 1.42 |
| GLYMA_02G097400 | Protein kinase domain-containing protein | −0.42 | 1.00 | −1.28 | 2.68 |
| GLYMA_01G081300 | Uncharacterized protein | −0.50 | 1.00 | −1.28 | 1.88 |
| GLYMA_10G207000 | Uncharacterized protein | −0.35 | 1.00 | −1.38 | 1.79 |
| GLYMA_04G023700 | Uncharacterized protein | −0.61 | 1.00 | −1.43 | 1.49 |
| GLYMA_03G018800 | TCP domain-containing protein | −0.15 | 1.00 | −1.56 | 3.85 |
| GO Term | Category a | N | Genes |
|---|---|---|---|
| Drug binding | MF | 10 | GLYMA_02G097400 GLYMA_05G015400 GLYMA_05G132900 GLYMA_05G136100 GLYMA_07G188800 GLYMA_08G091400 GLYMA_10G185200 GLYMA_10G253100 GLYMA_13G033500 GLYMA_18G271300 |
| Cell periphery | CC | 9 | GLYMA_01G238800 GLYMA_02G097400 GLYMA_05G015400 GLYMA_07G188800 GLYMA_10G253100 GLYMA_13G093300 GLYMA_15G052600 GLYMA_20G013400 GLYMA_20G236200 |
| Plasma membrane | CC | 8 | GLYMA_01G238800 GLYMA_02G097400 GLYMA_05G015400 GLYMA_07G188800 GLYMA_10G253100 GLYMA_13G093300 GLYMA_20G013400 GLYMA_20G236200 |
| Transporter activity | MF | 6 | GLYMA_01G238800 GLYMA_05G015400 GLYMA_05G132900 GLYMA_13G093300 GLYMA_13G206700 GLYMA_19G085200 |
| Response to stimulus | BP | 5 | GLYMA_13G093300 GLYMA_13G133100 GLYMA_15G052600 GLYMA_18G127900 GLYMA_20G236200 |
| Localization | BP | 5 | GLYMA_01G238800 GLYMA_05G015400 GLYMA_05G132900 GLYMA_13G206700 GLYMA_19G085200 |
| Establishment of localization | BP | 5 | GLYMA_01G238800 GLYMA_05G015400 GLYMA_05G132900 GLYMA_13G206700 GLYMA_19G085200 |
| Transmembrane transporter activity | MF | 5 | GLYMA_01G238800 GLYMA_05G132900 GLYMA_13G093300 GLYMA_13G206700 GLYMA_19G085200 |
| Catabolic process | BP | 4 | GLYMA_05G136100 GLYMA_08G091400 GLYMA_13G208000 GLYMA_15G052600 |
| Cellular response to stimulus | BP | 4 | GLYMA_13G093300 GLYMA_15G052600 GLYMA_18G127900 GLYMA_20G236200 |
| Cofactor binding | MF | 4 | GLYMA_05G136100 GLYMA_08G091400 GLYMA_11G051800 GLYMA_15G052600 |
| Regulation of metabolic process | BP | 3 | GLYMA_03G018800 GLYMA_06G265500 GLYMA_10G295200 |
| Response to chemical | BP | 3 | GLYMA_13G133100 GLYMA_15G052600 GLYMA_20G236200 |
| DNA-binding transcription factor activity | MF | 3 | GLYMA_03G018800 GLYMA_06G265500 GLYMA_10G295200 |
| Oxidoreductase activity | MF | 3 | GLYMA_11G051800 GLYMA_15G052600 GLYMA_20G236200 |
| Signaling | BP | 2 | GLYMA_13G093300 GLYMA_18G127900 |
| Detoxification | BP | 2 | GLYMA_15G052600 GLYMA_20G236200 |
| Cellular detoxification | BP | 2 | GLYMA_15G052600 GLYMA_20G236200 |
| Extracellular region | CC | 2 | GLYMA_01G127100 GLYMA_15G052600 |
| Cell-cell junction | CC | 2 | GLYMA_02G097400 GLYMA_15G052600 |
| Endomembrane system | CC | 2 | GLYMA_05G015400 GLYMA_13G206700 |
| Cell junction | CC | 2 | GLYMA_02G097400 GLYMA_15G052600 |
| Symplast | CC | 2 | GLYMA_02G097400 GLYMA_15G052600 |
| Peroxidase activity | MF | 2 | GLYMA_15G052600 GLYMA_20G236200 |
| Antioxidant activity | MF | 2 | GLYMA_15G052600 GLYMA_20G236200 |
| Lyase activity | MF | 2 | GLYMA_05G136100 GLYMA_08G091400 |
| Carbohydrate binding | MF | 2 | GLYMA_01G081300 GLYMA_13G033500 |
| Functional Category (GO Category a or KEGG Pathway) | Genes in List | Total Genes | Enrichment FDR b |
|---|---|---|---|
| Glutamate catabolic process (BP) | 2 | 10 | 1.68 |
| Glutamate decarboxylase activity (MF) | 2 | 10 | 1.68 |
| Dicarboxylic acid catabolic process (BP) | 2 | 13 | 1.94 |
| Taurine and hypotaurine metabolism (KEGG) | 2 | 29 | 6.43 |
| Glutamine family amino acid catabolic process (BP) | 2 | 30 | 6.43 |
| Butanoate metabolism (KEGG) | 2 | 33 | 6.50 |
| Glutamate metabolic process (BP) | 2 | 37 | 7.01 |
| Plasma membrane (CC) | 8 | 2728 | 9.11 |
| Cell periphery (CC) | 9 | 3371 | 9.11 |
| MAPK signaling pathway (KEGG) | 3 | 230 | 9.11 |
| Alanine, aspartate and glutamate metabolism (KEGG) | 2 | 73 | 1.73 |
| Beta-Alanine metabolism (KEGG) | 2 | 81 | 1.95 |
| Drug binding (MF) | 10 | 4868 | 2.12 |
| Alpha-amino acid catabolic process (BP) | 2 | 110 | 3.05 |
| Carboxy-lyase activity (MF) | 2 | 119 | 3.17 |
| Cellular amino acid catabolic process (BP) | 2 | 120 | 3.17 |
| Organophosphate ester transport (BP) | 2 | 130 | 3.23 |
| Carbohydrate transmembrane transport (BP) | 2 | 139 | 3.23 |
| Dicarboxylic acid metabolic process (BP) | 2 | 142 | 3.23 |
| Magnesium-dependent protein serine/threonine phosphatase activity (MF) | 2 | 134 | 3.23 |
| Calcium ion binding (MF) | 3 | 474 | 3.23 |
| Glutamine family amino acid metabolic process (BP) | 2 | 137 | 3.23 |
| Metal ion binding (MF) | 9 | 4694 | 3.35 |
| Carbohydrate transmembrane transporter activity (MF) | 2 | 156 | 3.38 |
| Cation binding (MF) | 9 | 4733 | 3.38 |
| Carbon-carbon lyase activity (MF) | 2 | 173 | 3.82 |
| Carbohydrate transport (BP) | 2 | 170 | 3.82 |
| Organic acid catabolic process (MF) | 2 | 191 | 4.18 |
| Carboxylic acid catabolic process (MF) | 2 | 191 | 4.18 |
| Transporter activity (MF) | 6 | 2526 | 4.18 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaminaka, H.; Miura, C.; Isowa, Y.; Tominaga, T.; Gonnami, M.; Egusa, M.; Ifuku, S. Nanofibrillation is an Effective Method to Produce Chitin Derivatives for Induction of Plant Responses in Soybean. Plants 2020, 9, 810. https://doi.org/10.3390/plants9070810
Kaminaka H, Miura C, Isowa Y, Tominaga T, Gonnami M, Egusa M, Ifuku S. Nanofibrillation is an Effective Method to Produce Chitin Derivatives for Induction of Plant Responses in Soybean. Plants. 2020; 9(7):810. https://doi.org/10.3390/plants9070810
Chicago/Turabian StyleKaminaka, Hironori, Chihiro Miura, Yukiko Isowa, Takaya Tominaga, Mamu Gonnami, Mayumi Egusa, and Shinsuke Ifuku. 2020. "Nanofibrillation is an Effective Method to Produce Chitin Derivatives for Induction of Plant Responses in Soybean" Plants 9, no. 7: 810. https://doi.org/10.3390/plants9070810
APA StyleKaminaka, H., Miura, C., Isowa, Y., Tominaga, T., Gonnami, M., Egusa, M., & Ifuku, S. (2020). Nanofibrillation is an Effective Method to Produce Chitin Derivatives for Induction of Plant Responses in Soybean. Plants, 9(7), 810. https://doi.org/10.3390/plants9070810






