Genome-Wide Identification, Expression Analysis and Functional Study of CCT Gene Family in Medicago truncatula
Abstract
1. Introduction
2. Results
2.1. Identification of CCT Genes in M. truncatula Genome
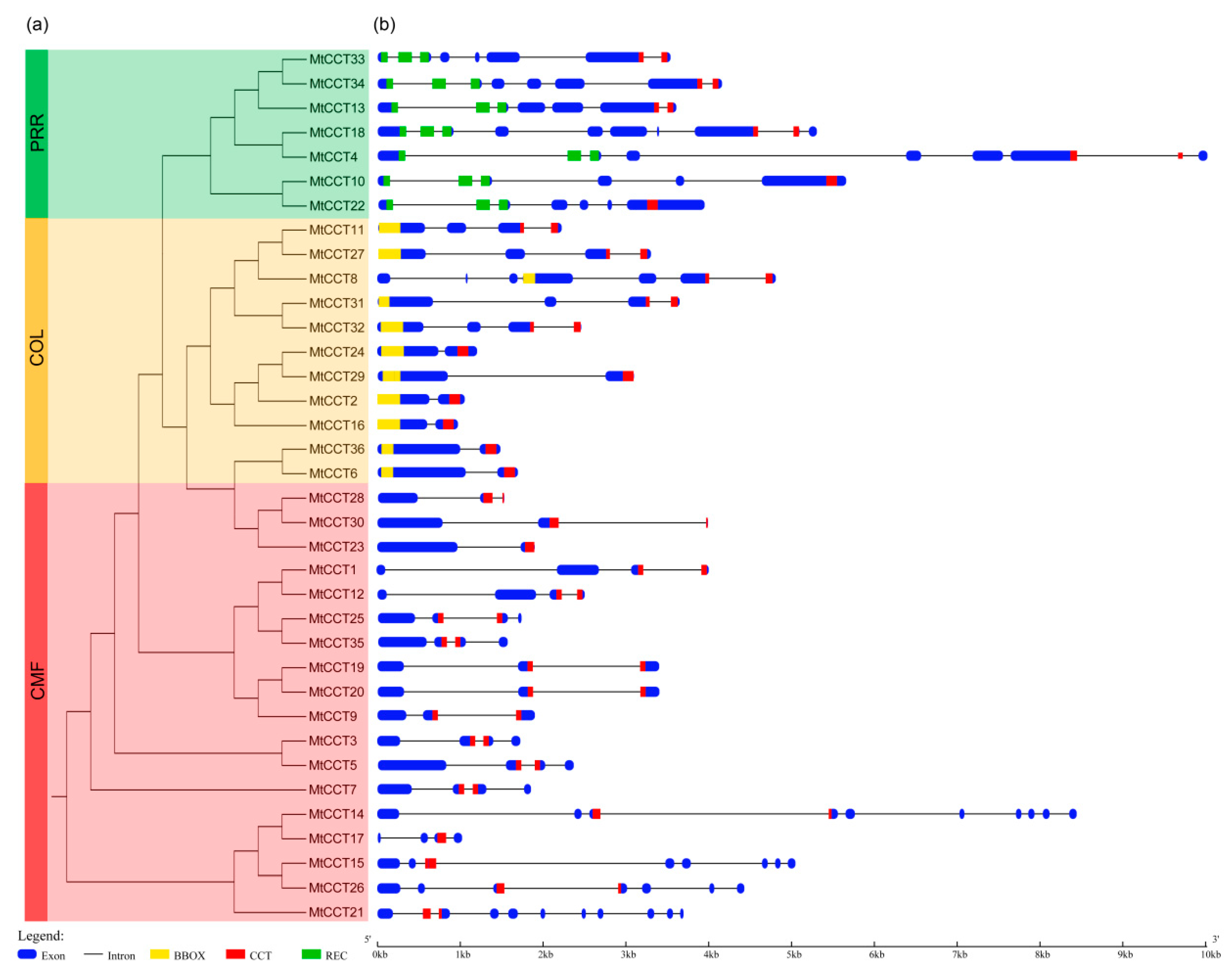
2.2. Analyses of Gene Structure, Conserved Domain and Phylogenetic Relationship of MtCCT Genes
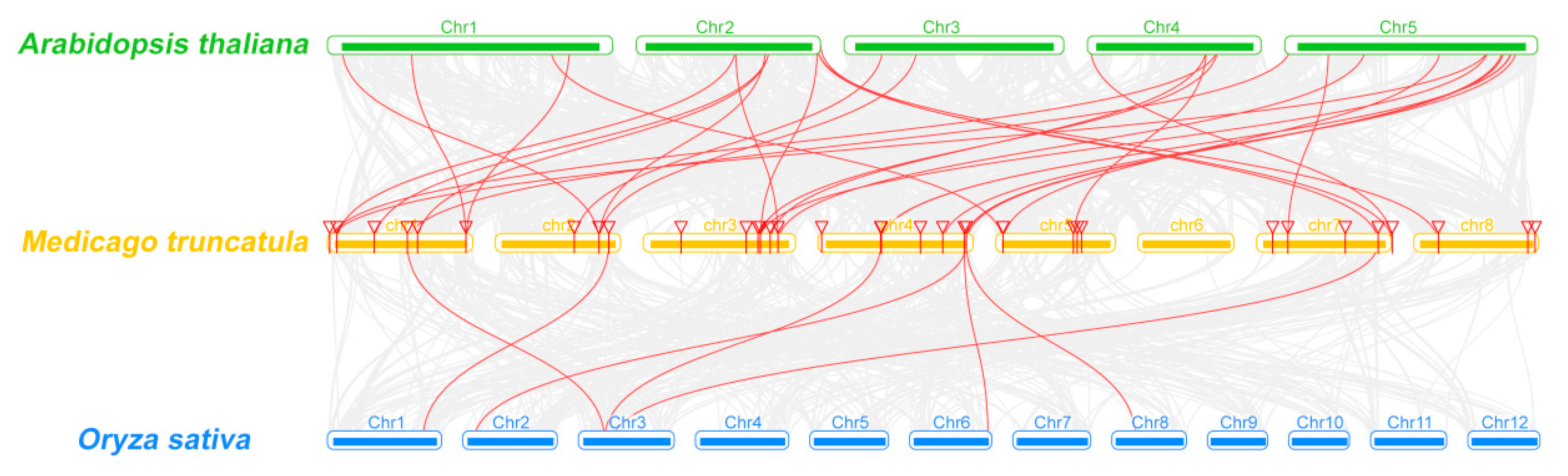
2.3. Analyses of Chromosomal Distribution and Synteny of MtCCT Genes
2.4. Phylogenetic Analysis of CCT Proteins in Plants
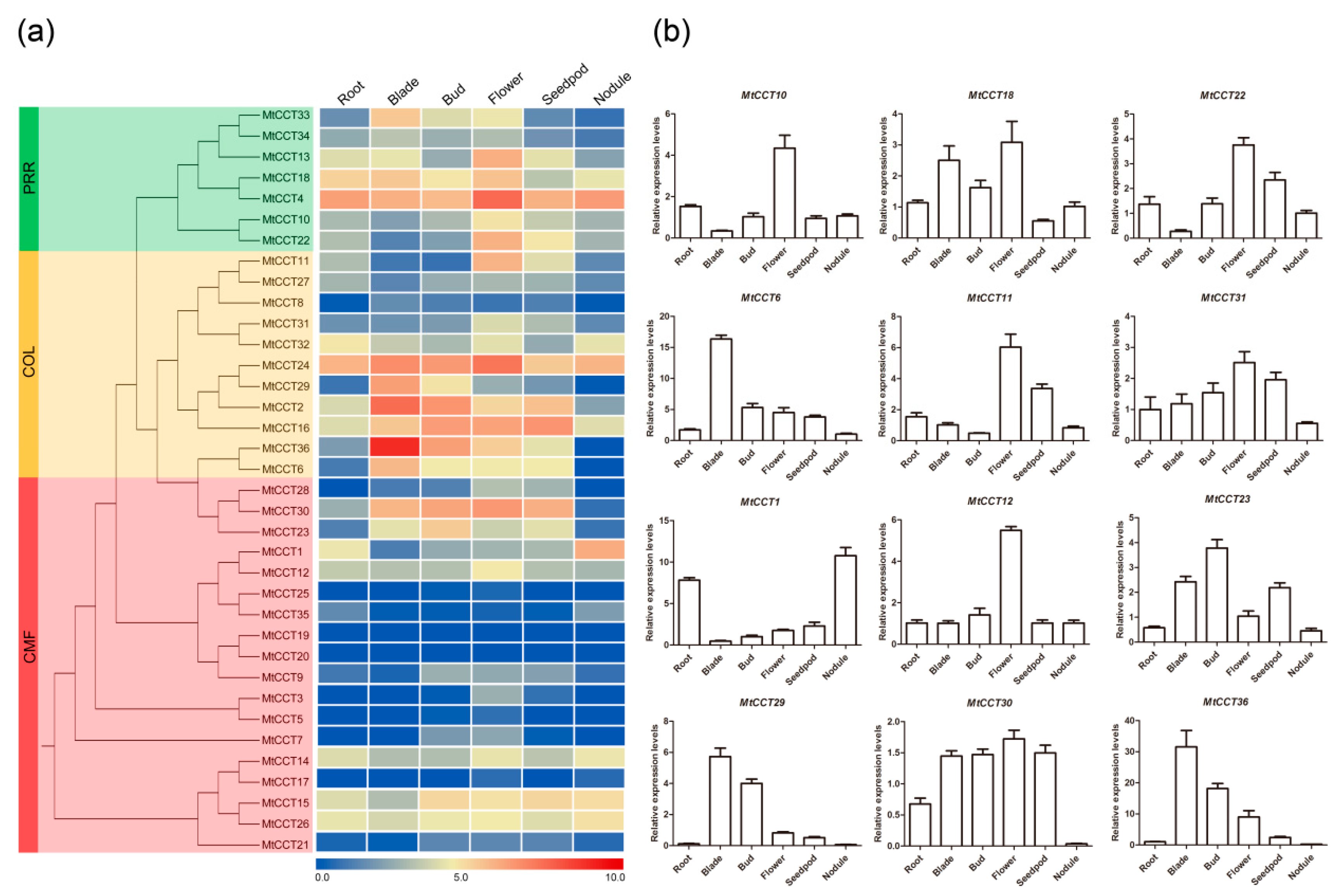
2.5. Temporal and Spatial Expression Profiling of MtCCT Genes
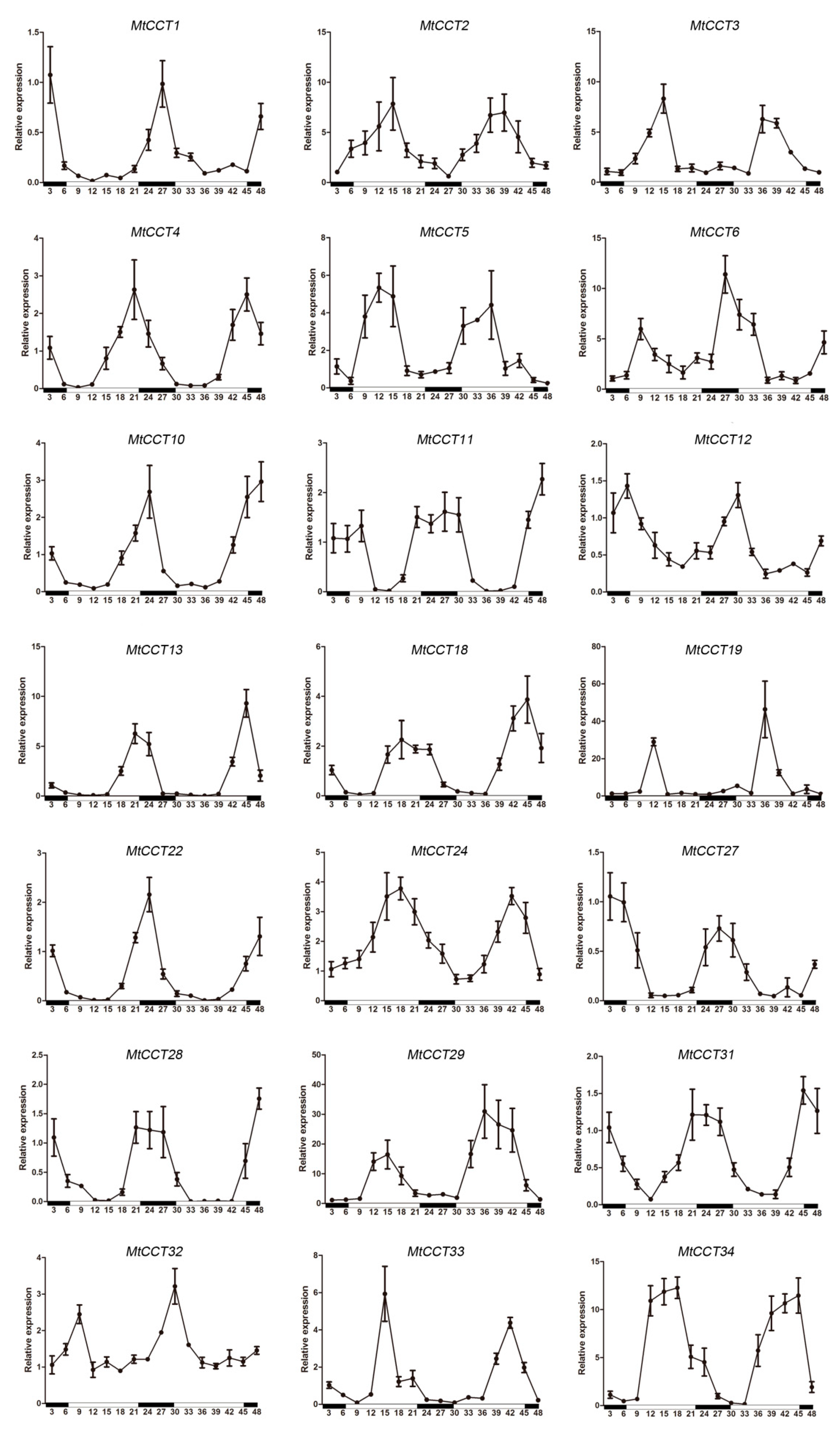
2.6. Expression Patterns of MtCCT Genes under Light Circle
2.7. Expression Patterns of MtCCT Genes in Response to Hormones
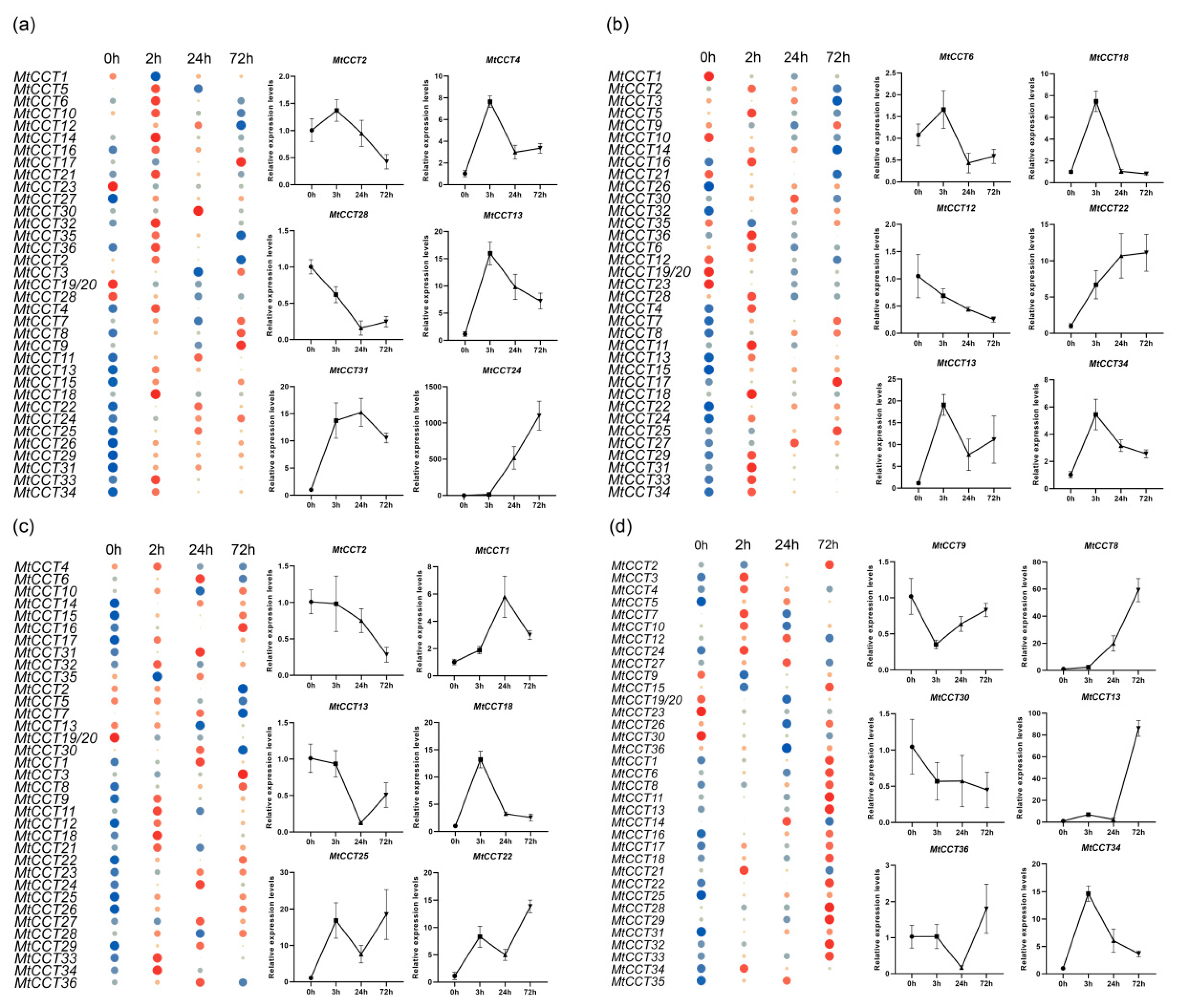
2.8. Expression of MtCCT Genes Which Responded to Abiotic Stresses
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Identification of CCT Genes in M. truncatula
4.3. Analyses of Gene Structure and Conserved Motif
4.4. Analyses of Chromosomal Distribution, Synteny and Cis-Elements of MtCCT Genes
4.5. Phylogenetic Analysis of CCT Proteins in Representative Plants
4.6. RNA Extraction and Real-Time Quantitative PCR
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Blumel, M.; Dally, N.; Jung, C. Flowering time regulation in crops—What did we learn from Arabidopsis? Curr. Opin. Biotechnol. 2015, 32, 121–129. [Google Scholar] [CrossRef]
- Putterill, J.; Laurie, R.; Macknight, R. It’s time to flower: The genetic control of flowering time. Bioessays 2004, 26, 363–373. [Google Scholar] [CrossRef]
- Srikanth, A.; Schmid, M. Regulation of flowering time: All roads lead to Rome. Cell Mol. Life Sci. 2011, 68, 2013–2037. [Google Scholar] [CrossRef]
- Yan, W.; Liu, H.; Zhou, X.; Li, Q.; Zhang, J.; Lu, L.; Liu, T.; Liu, H.; Zhang, C.; Zhang, Z.; et al. Natural variation in Ghd7.1 plays an important role in grain yield and adaptation in rice. Cell Res. 2013, 23, 969–971. [Google Scholar] [CrossRef]
- Li, Y.P.; Xu, M.L. CCT family genes in cereal crops: A current overview. Crop J. 2017, 5, 449–458. [Google Scholar] [CrossRef]
- Liu, C.; Qu, X.; Zhou, Y.; Song, G.; Abiri, N.; Xiao, Y.; Liang, F.; Jiang, D.; Hu, Z.; Yang, D. OsPRR37 confers an expanded regulation of the diurnal rhythms of the transcriptome and photoperiodic flowering pathways in rice. Plant Cell Environ. 2018, 41, 630–645. [Google Scholar] [CrossRef]
- Putterill, J.; Robson, F.; Lee, K.; Simon, R.; Coupland, G. The Constans gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc-finger transcription tactors. Cell 1995, 80, 847–857. [Google Scholar] [CrossRef]
- Robson, F.; Costa, M.M.R.; Hepworth, S.R.; Vizir, I.; Pineiro, M.; Reeves, P.H.; Putterill, J.; Coupland, G. Functional importance of conserved domains in the flowering-time gene CONSTANS demonstrated by analysis of mutant alleles and transgenic plants. Plant J. 2001, 28, 619–631. [Google Scholar] [CrossRef]
- Wenkel, S.; Turck, F.; Singer, K.; Gissot, L.; Le Gourrierec, J.; Samach, A.; Coupland, G. CONSTANS and the CCAAT box binding complex share a functionally important domain and interact to regulate flowering of Arabidopsis. Plant Cell 2006, 18, 2971–2984. [Google Scholar] [CrossRef]
- Cockram, J.; Thiel, T.; Steuernagel, B.; Stein, N.; Taudien, S.; Bailey, P.C.; O’Sullivan, D.M. Genome dynamics explain the evolution of flowering time CCT domain gene families in the Poaceae. PLoS ONE 2012, 7, e45307. [Google Scholar] [CrossRef]
- Zhang, L.; Li, Q.; Dong, H.; He, Q.; Liang, L.; Tan, C.; Han, Z.; Yao, W.; Li, G.; Zhao, H.; et al. Three CCT domain-containing genes were identified to regulate heading date by candidate gene-based association mapping and transformation in rice. Sci. Rep. 2015, 5, 7663. [Google Scholar] [CrossRef]
- Jin, M.L.; Liu, X.G.; Jia, W.; Liu, H.J.; Li, W.Q.; Peng, Y.; Du, Y.F.; Wang, Y.B.; Yin, Y.J.; Zhang, X.; et al. ZmCOL3, a CCT gene represses flowering in maize by interfering with the circadian clock and activating expression of ZmCCT. J. Integr. Plant Biol. 2018, 60, 465–480. [Google Scholar] [CrossRef]
- Yan, L.L.; Loukoianov, A.; Blechl, A.; Tranquilli, G.; Ramakrishna, W.; SanMiguel, P.; Bennetzen, J.L.; Echenique, V.; Dubcovsky, J. The wheat VRN2 gene is a flowering repressor down-regulated by vernalization. Science 2004, 303, 1640–1644. [Google Scholar] [CrossRef]
- Corbesier, L.; Vincent, C.; Jang, S.; Fornara, F.; Fan, Q.; Searle, I.; Giakountis, A.; Farrona, S.; Gissot, L.; Turnbull, C.; et al. FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis. Science 2007, 316, 1030–1033. [Google Scholar] [CrossRef]
- Yano, M.; Katayose, Y.; Ashikari, M.; Yamanouchi, U.; Monna, L.; Fuse, T.; Baba, T.; Yamamoto, K.; Umehara, Y.; Nagamura, Y.; et al. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell 2000, 12, 2473–2483. [Google Scholar] [CrossRef]
- Hayama, R.; Yokoi, S.; Tamaki, S.; Yano, M.; Shimamoto, K. Adaptation of photoperiodic control pathways produces short-day flowering in rice. Nature 2003, 422, 719–722. [Google Scholar] [CrossRef]
- Wu, W.; Zheng, X.M.; Lu, G.; Zhong, Z.; Gao, H.; Chen, L.; Wu, C.; Wang, H.J.; Wang, Q.; Zhou, K.; et al. Association of functional nucleotide polymorphisms at DTH2 with the northward expansion of rice cultivation in Asia. Proc. Natl. Acad. Sci. USA 2013, 110, 2775–2780. [Google Scholar] [CrossRef]
- Yan, L.L.; Loukoianov, A.; Tranquilli, G.; Helguera, M.; Fahima, T.; Dubcovsky, J. Positional cloning of the wheat vernalization gene VRN1. Proc. Natl. Acad. Sci. USA 2003, 100, 6263–6268. [Google Scholar] [CrossRef]
- Min, J.H.; Chung, J.S.; Lee, K.H.; Kim, C.S. The CONSTANS-like 4 transcription factor, AtCOL4, positively regulates abiotic stress tolerance through an abscisic acid-dependent manner in Arabidopsis. J. Integr. Plant Biol. 2015, 57, 313–324. [Google Scholar] [CrossRef]
- Liu, J.; Shen, J.; Xu, Y.; Li, X.; Xiao, J.; Xiong, L. Ghd2, a CONSTANS-like gene, confers drought sensitivity through regulation of senescence in rice. J. Exp. Bot. 2016, 67, 5785–5798. [Google Scholar] [CrossRef]
- Hung, H.Y.; Shannon, L.M.; Tian, F.; Bradbury, P.J.; Chen, C.; Flint-Garcia, S.A.; McMullen, M.D.; Ware, D.; Buckler, E.S.; Doebley, J.F.; et al. ZmCCT and the genetic basis of day-length adaptation underlying the post domestication spread of maize. Proc. Natl. Acad. Sci. USA 2012, 109, 1913–1921. [Google Scholar] [CrossRef]
- Wang, C.; Yang, Q.; Wang, W.; Li, Y.; Guo, Y.; Zhang, D.; Ma, X.; Song, W.; Zhao, J.; Xu, M. A transposon-directed epigenetic change in ZmCCT underlies quantitative resistance to Gibberella stalk rot in maize. New Phytol. 2017, 215, 1503–1515. [Google Scholar] [CrossRef]
- Xu, G.; Wang, X.; Huang, C.; Xu, D.; Li, D.; Tian, J.; Chen, Q.; Wang, C.; Liang, Y.; Wu, Y.; et al. Complex genetic architecture underlies maize tassel domestication. New Phytol. 2017, 214, 852–864. [Google Scholar] [CrossRef]
- Xue, W.Y.; Xing, Y.Z.; Weng, X.Y.; Zhao, Y.; Tang, W.J.; Wang, L.; Zhou, H.J.; Yu, S.B.; Xu, C.G.; Li, X.H.; et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 2008, 40, 761–767. [Google Scholar] [CrossRef]
- Weng, X.; Wang, L.; Wang, J.; Hu, Y.; Du, H.; Xu, C.; Xing, Y.; Li, X.; Xiao, J.; Zhang, Q. Grain number, plant height, and heading date7 is a central regulator of growth, development, and stress response. Plant Physiol. 2014, 164, 735–747. [Google Scholar] [CrossRef]
- Strayer, C.; Oyama, T.; Schultz, T.F.; Raman, R.; Somers, D.E.; Mas, P.; Panda, S.; Kreps, J.A.; Kay, S.A. Cloning of the Arabidopsis clock cone TOC1, an auto regulatory response regulator homolog. Science 2000, 289, 768–771. [Google Scholar] [CrossRef]
- Alabadi, D.; Oyama, T.; Yanovsky, M.J.; Harmon, F.G.; Mas, P.; Kay, S.A. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 2001, 293, 880–883. [Google Scholar] [CrossRef]
- Gao, H.; Jin, M.; Zheng, X.M.; Chen, J.; Yuan, D.; Xin, Y.; Wang, M.; Huang, D.; Zhang, Z.; Zhou, K.; et al. Days to heading 7, a major quantitative locus determining photoperiod sensitivity and regional adaptation in rice. Proc. Natl. Acad. Sci. USA 2014, 111, 16337–16342. [Google Scholar] [CrossRef]
- Nishida, H.; Yoshida, T.; Kawakami, K.; Fujita, M.; Long, B.; Akashi, Y.; Laurie, D.A.; Kato, K. Structural variation in the 5’ upstream region of photoperiod-insensitive alleles Ppd-A1a and Ppd-B1a identified in hexaploid wheat (Triticum aestivum L.), and their effect on heading time. Mol. Breed. 2013, 31, 27–37. [Google Scholar] [CrossRef]
- Sun, H.; Guo, Z.; Gao, L.; Zhao, G.; Zhang, W.; Zhou, R.; Wu, Y.; Wang, H.; An, H.; Jia, J. DNA methylation pattern of photoperiod-B1 is associated with photoperiod insensitivity in wheat (Triticum aestivum). New Phytol. 2014, 204, 682–692. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, X.M.; Zhang, H.; Ma, L.; Zhao, H.M.; Jones, C.S.; Chen, J.; Liu, G.B. A genome-wide association study approach to the identification of candidate genes underlying agronomic traits in alfalfa (Medicago sativa L.). Plant Biotechnol. J. 2019, 18, 611–613. [Google Scholar] [CrossRef] [PubMed]
- Lorenzo, C.D.; Garcia-Gagliardi, P.; Antonietti, M.S.; Sanchez-Lamas, M.; Mancini, E.; Dezar, C.A.; Vazquez, M.; Watson, G.; Yanovsky, M.J.; Cerdan, P.D. Improvement of alfalfa forage quality and management through the down-regulation of MsFTa1. Plant Biotechnol. J. 2019. [Google Scholar] [CrossRef]
- Adhikari, L.; Makaju, S.O.; Missaoui, A.M. QTL mapping of flowering time and biomass yield in tetraploid alfalfa (Medicago sativa L.). BMC Plant Biol. 2019, 19, 359. [Google Scholar] [CrossRef]
- Zhou, C.E.; Han, L.; Pislariu, C.; Nakashima, J.; Fu, C.X.; Jiang, Q.Z.; Quan, L.; Blancaflor, E.B.; Tang, Y.H.; Bouton, J.H.; et al. From model to crop: Functional analysis of a STAY-GREEN gene in the model legume Medicago truncatula and effective use of the gene for alfalfa improvement. Plant Physiol. 2011, 157, 1483–1496. [Google Scholar] [CrossRef]
- Gou, J.Q.; Debnath, S.; Sun, L.; Flanagan, A.; Tang, Y.H.; Jiang, Q.Z.; Wen, J.Q.; Wang, Z.Y. From model to crop: Functional characterization of SPL8 in M. truncatula led to genetic improvement of biomass yield and abiotic stress tolerance in alfalfa. Plant Biotechnol. J. 2018, 16, 951–962. [Google Scholar] [CrossRef]
- Bendix, C.; Marshall, C.M.; Harmon, F.G. Circadian clock genes universally control key agricultural traits. Mol. Plant 2015, 8, 1135–1152. [Google Scholar] [CrossRef]
- Liu, Y.Q.; Wu, H.; Chen, H.; Liu, Y.L.; He, J.; Kang, H.Y.; Sun, Z.G.; Pan, G.; Wang, Q.; Hu, J.L.; et al. A gene cluster encoding lectin receptor kinases confers broad-spectrum and durable insect resistance in rice. Nat. Biotechnol. 2015, 33, 301–306. [Google Scholar] [CrossRef]
- Walters, L.A.; Escobar, M.A. The AtGRXS3/4/5/7/8 glutaredoxin gene cluster on Arabidopsis thaliana chromosome 4 is coordinately regulated by nitrate and appears to control primary root growth. Plant Signal Behav. 2016, 11, e1171450. [Google Scholar] [CrossRef]
- Kryvoruchko, I.S.; Sinharoy, S.; Torres-Jerez, I.; Sosso, D.; Pislariu, C.I.; Guan, D.; Murray, J.; Benedito, V.A.; Frommer, W.B.; Udvardi, M.K. MtSWEET11, a nodule-specific sucrose transporter of Medicago truncatula. Plant Physiol. 2016, 171, 554–565. [Google Scholar] [CrossRef]
- Satge, C.; Moreau, S.; Sallet, E.; Lefort, G.; Auriac, M.C.; Rembliere, C.; Cottret, L.; Gallardo, K.; Noirot, C.; Jardinaud, M.F.; et al. Reprogramming of DNA methylation is critical for nodule development in Medicago truncatula. Nat. Plants 2016, 2, 16166. [Google Scholar] [CrossRef]
- Nakamichi, N.; Kiba, T.; Henriques, R.; Mizuno, T.; Chua, N.H.; Sakakibara, H. PSEUDO-RESPONSE REGULATORS 9, 7, and 5 are transcriptional repressors in the Arabidopsis circadian clock. Plant Cell 2010, 22, 594–605. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Li, X.; Ge, C.; Chang, J.; Shi, M.; Chen, J.; Qiao, L.; Chang, Z.; Zheng, J.; Zhang, J. Characterization of the CCT family and analysis of gene expression in Aegilops tauschii. PLoS ONE 2017, 12, e0189333. [Google Scholar] [CrossRef] [PubMed]
- Corbesier, L.; Coupland, G. The quest for florigen: A review of recent progress. J. Exp. Bot. 2006, 57, 3395–3403. [Google Scholar] [CrossRef]
- Moon, J.; Suh, S.S.; Lee, H.; Choi, K.R.; Hong, C.B.; Paek, N.C.; Kim, S.G.; Lee, I. The SOC1 MADS-box gene integrates vernalization and gibberellin signals for flowering in Arabidopsis. Plant J. 2003, 35, 613–623. [Google Scholar] [CrossRef]
- Mai, Y.X.; Wang, L.; Yang, H.Q. A gain-of-function mutation in IAA7/AXR2 confers late flowering under short-day light in Arabidopsis. J. Integr. Plant Biol. 2011, 53, 480–492. [Google Scholar] [CrossRef]
- Riboni, M.; Test, A.R.; Galbiati, M.; Tonelli, C.; Conti, L. ABA-dependent control of GIGANTEA signalling enables drought escape via up-regulation of FLOWERING LOCUS T in Arabidopsis thaliana. J. Exp. Bot. 2016, 67, 6309–6322. [Google Scholar] [CrossRef]



| Gene | Chromosome | CDS (Coding Sequence) | Protein (aa) | Molecular Weight (D) | Theoretical Pi | Subfamily |
|---|---|---|---|---|---|---|
| MtCCT1 | Chr1 | Medtr1g008220.1 | 282 | 31927 | 4.94 | CMF |
| MtCCT2 | Chr1 | Medtr1g013450.1 | 316 | 35496 | 6.31 | COL |
| MtCCT3 | Chr1 | Medtr1g044785.1 | 230 | 25360 | 5.6 | CMF |
| MtCCT4 | Chr1 | Medtr1g067110.1 | 748 | 81610 | 7.27 | PRR |
| MtCCT5 | Chr1 | Medtr1g073350.1 | 414 | 45719 | 4.47 | CMF |
| MtCCT6 | Chr1 | Medtr1g110870.1 | 436 | 48793 | 5.45 | COL |
| MtCCT7 | Chr2 | Medtr2g068730.1 | 266 | 30816 | 5.21 | CMF |
| MtCCT8 | Chr2 | Medtr2g088900.1 | 521 | 57571 | 5.85 | COL |
| MtCCT9 | Chr2 | Medtr2g096080.1 | 251 | 28657 | 6.17 | CMF |
| MtCCT10 | Chr3 | Medtr3g037390.1 | 575 | 64621 | 6.05 | PRR |
| MtCCT11 | Chr3 | Medtr3g082630.2 | 411 | 45277 | 4.89 | COL |
| MtCCT12 | Chr3 | Medtr3g091340.1 | 281 | 31916 | 5.23 | CMF |
| MtCCT13 | Chr3 | Medtr3g092780.1 | 685 | 75851 | 6.11 | PRR |
| MtCCT14 | Chr3 | Medtr3g100040.1 | 359 | 38847 | 4.47 | CMF |
| MtCCT15 | Chr3 | Medtr3g100050.1 | 309 | 34206 | 5.62 | CMF |
| MtCCT16 | Chr3 | Medtr3g105710.1 | 290 | 31869 | 7.11 | COL |
| MtCCT17 | Chr4 | Medtr4g008090.1 | 121 | 13589 | 9.86 | CMF |
| MtCCT18 | Chr4 | Medtr4g061360.1 | 796 | 87246 | 6.72 | PRR |
| MtCCT19 | Chr4 | Medtr4g061823.1 | 242 | 27580 | 5.93 | CMF |
| MtCCT20 | Chr4 | Medtr4g061910.1 | 242 | 27580 | 5.93 | CMF |
| MtCCT21 | Chr4 | Medtr4g093730.1 | 334 | 37563 | 5.04 | CMF |
| MtCCT22 | Chr4 | Medtr4g108880.1 | 585 | 68781 | 5.80 | PRR |
| MtCCT23 | Chr4 | Medtr4g127420.1 | 379 | 42006 | 9.00 | CMF |
| MtCCT24 | Chr4 | Medtr4g128930.1 | 375 | 41119 | 5.86 | COL |
| MtCCT25 | Chr5 | Medtr5g010120.1 | 251 | 29195 | 5.83 | CMF |
| MtCCT26 | Chr5 | Medtr5g066510.1 | 286 | 31424 | 5.53 | CMF |
| MtCCT27 | Chr5 | Medtr5g069480.1 | 410 | 45027 | 5.70 | COL |
| MtCCT28 | Chr5 | Medtr5g072780.1 | 217 | 25897 | 5.07 | CMF |
| MtCCT29 | Chr7 | Medtr7g018170.1 | 396 | 43757 | 5.10 | COL |
| MtCCT30 | Chr7 | Medtr7g032240.1 | 352 | 40433 | 9.53 | CMF |
| MtCCT31 | Chr7 | Medtr7g083540.1 | 390 | 44311 | 6.28 | COL |
| MtCCT32 | Chr7 | Medtr7g108150.1 | 372 | 42581 | 6.21 | COL |
| MtCCT33 | Chr7 | Medtr7g118260.1 | 596 | 66654 | 6.54 | PRR |
| MtCCT34 | Chr8 | Medtr8g024260.1 | 640 | 73001 | 6.74 | PRR |
| MtCCT35 | Chr8 | Medtr8g098725.1 | 324 | 37326 | 5.34 | CMF |
| MtCCT36 | Chr8 | Medtr8g104190.1 | 416 | 47041 | 5.10 | COL |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, L.; Yi, D.; Yang, J.; Liu, X.; Pang, Y. Genome-Wide Identification, Expression Analysis and Functional Study of CCT Gene Family in Medicago truncatula. Plants 2020, 9, 513. https://doi.org/10.3390/plants9040513
Ma L, Yi D, Yang J, Liu X, Pang Y. Genome-Wide Identification, Expression Analysis and Functional Study of CCT Gene Family in Medicago truncatula. Plants. 2020; 9(4):513. https://doi.org/10.3390/plants9040513
Chicago/Turabian StyleMa, Lin, Dengxia Yi, Junfeng Yang, Xiqiang Liu, and Yongzhen Pang. 2020. "Genome-Wide Identification, Expression Analysis and Functional Study of CCT Gene Family in Medicago truncatula" Plants 9, no. 4: 513. https://doi.org/10.3390/plants9040513
APA StyleMa, L., Yi, D., Yang, J., Liu, X., & Pang, Y. (2020). Genome-Wide Identification, Expression Analysis and Functional Study of CCT Gene Family in Medicago truncatula. Plants, 9(4), 513. https://doi.org/10.3390/plants9040513





