Flower Colour Polymorphism, Pollination Modes, Breeding System and Gene Flow in Anemone coronaria
Abstract
1. Introduction
2. Results
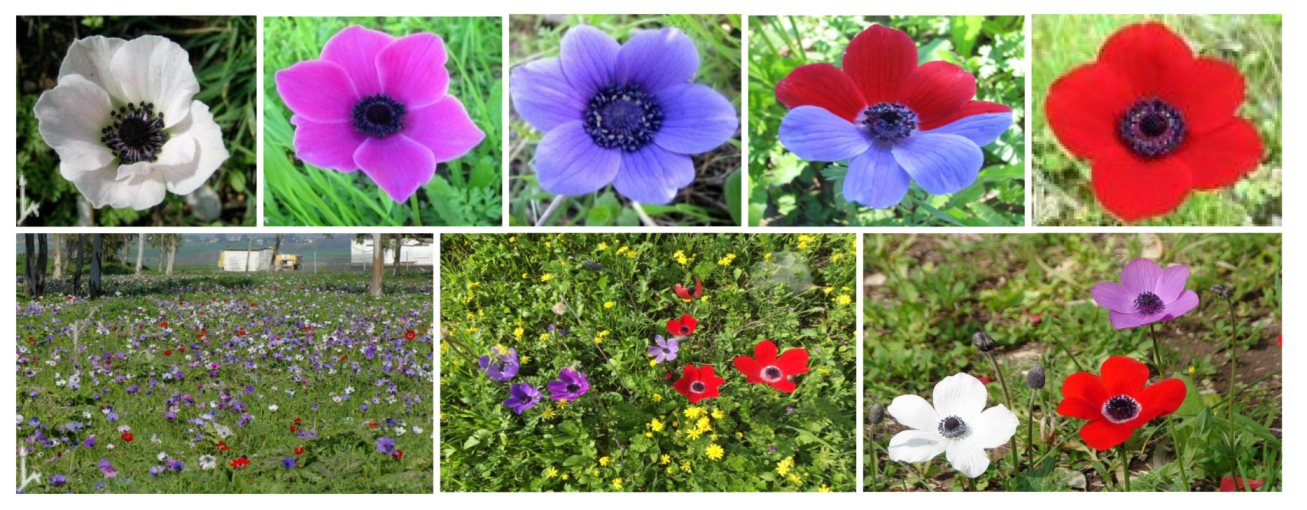
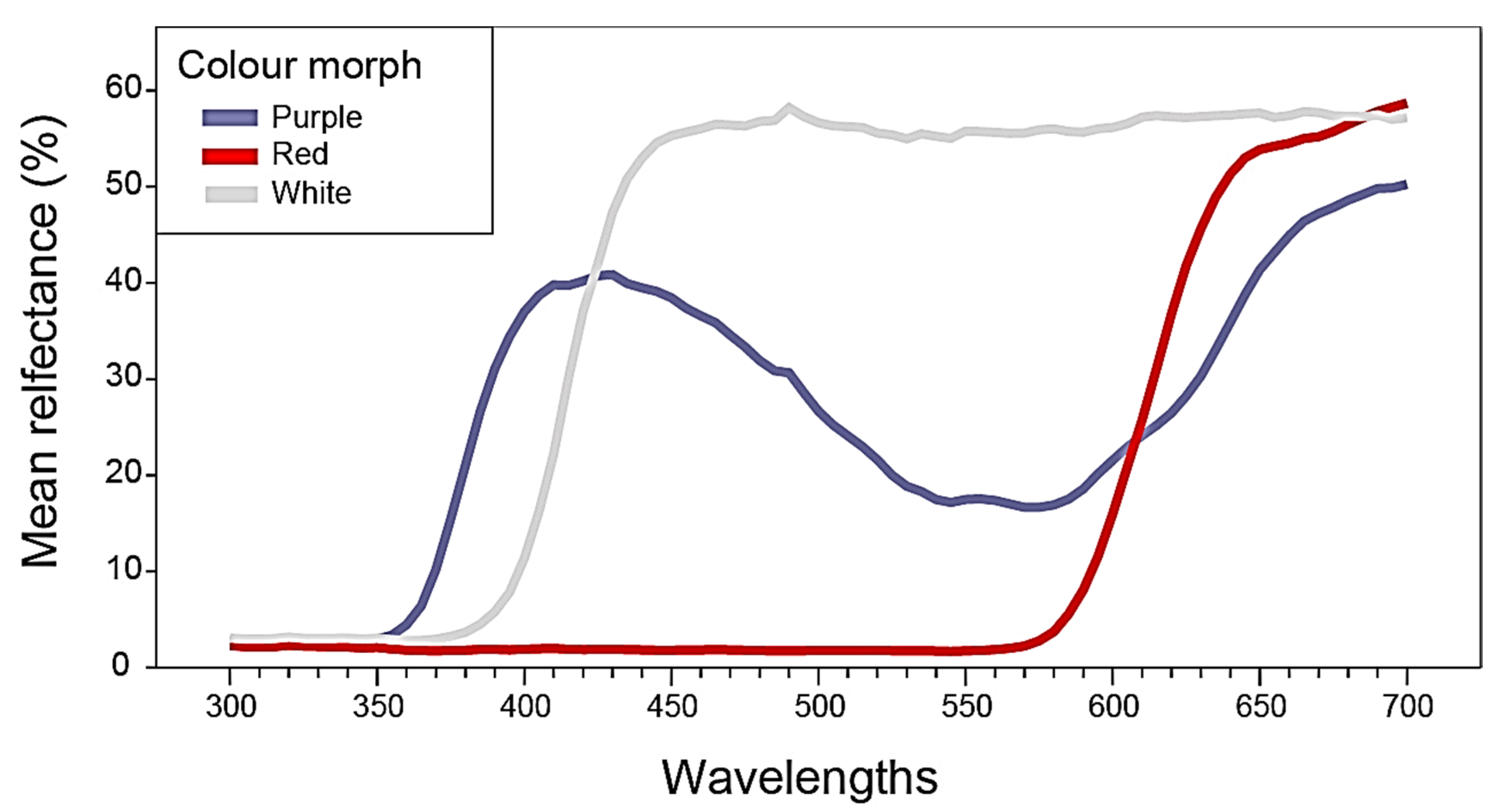
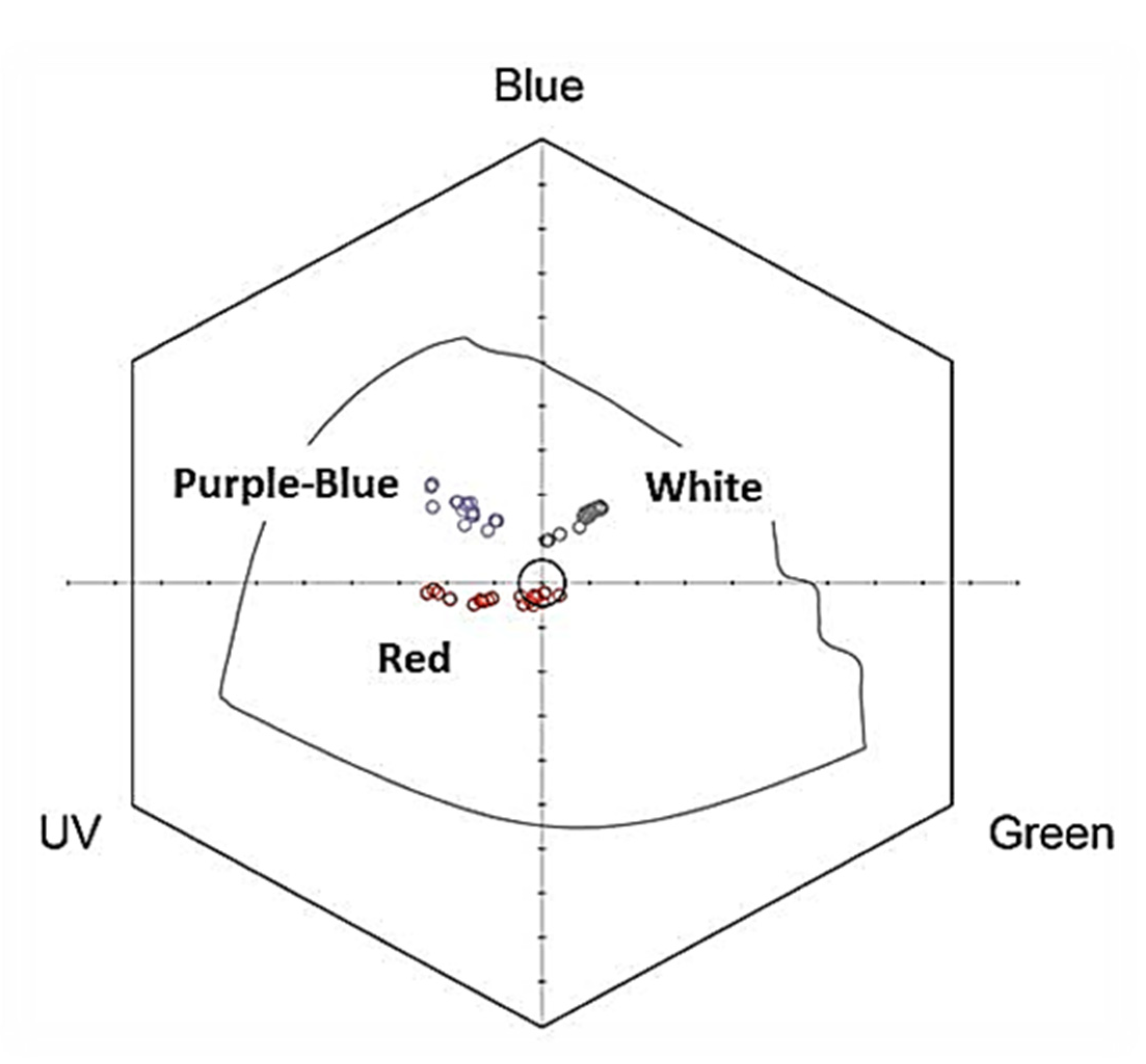
2.1. Flower Reflectance and Perception
2.2. Number of Carpels
2.3. Pollination Modes
2.3.1. Spontaneous Self-Pollination
2.3.2. Insect and Wind Pollination
2.4. Flower Visitors
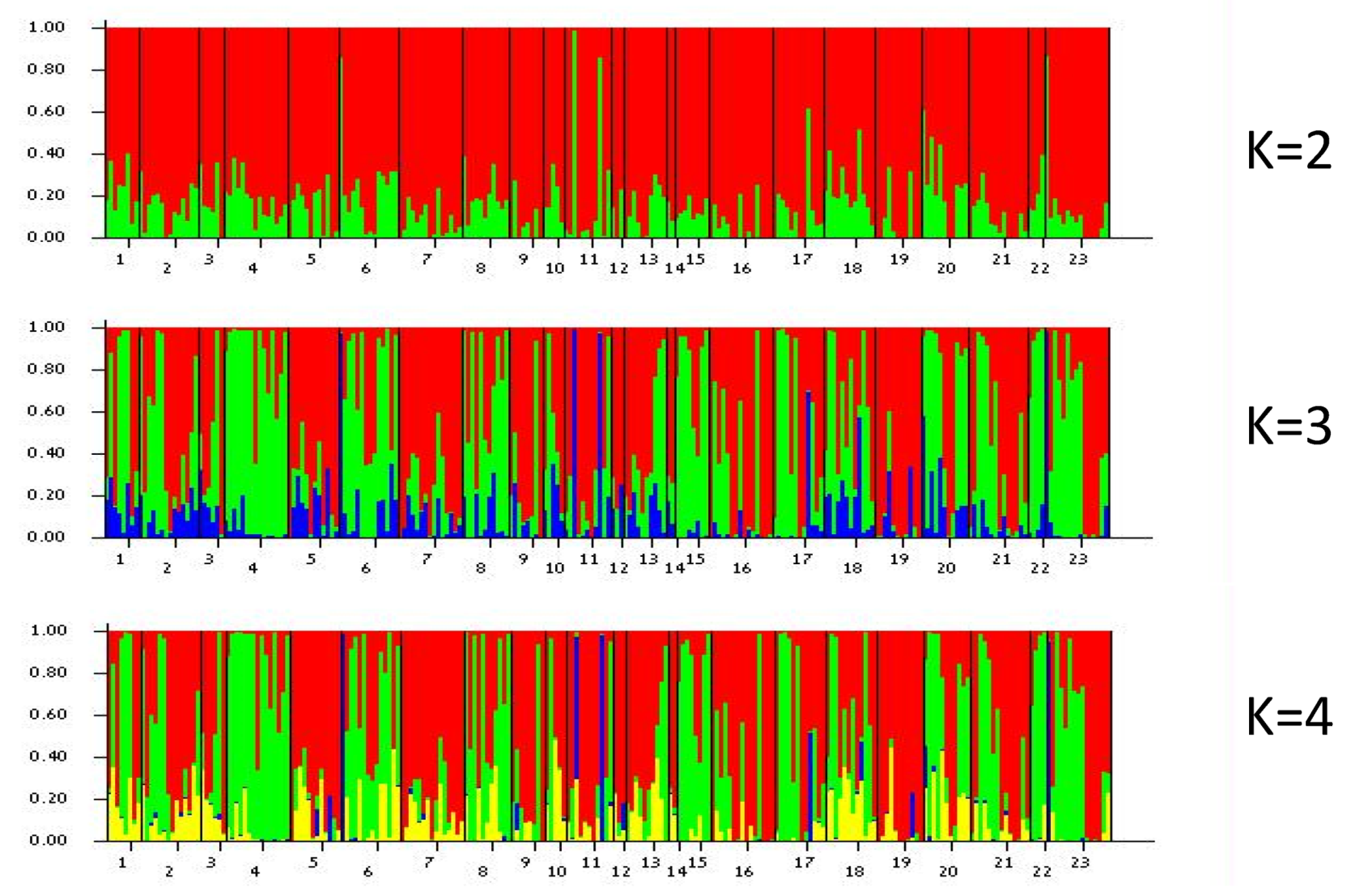
2.5. Genetic Structure of Populations and Colour Morphs
3. Discussion
3.1. Colour Discrimination by Visiting Insects
3.2. Spontaneous Self-Pollination and Wind Pollination
3.3. Flowers Visitors
3.4. Population Genetic Analysis
3.5. The Evolution and Maintenance of Flower Colour Polymorphism
4. Materials and Methods
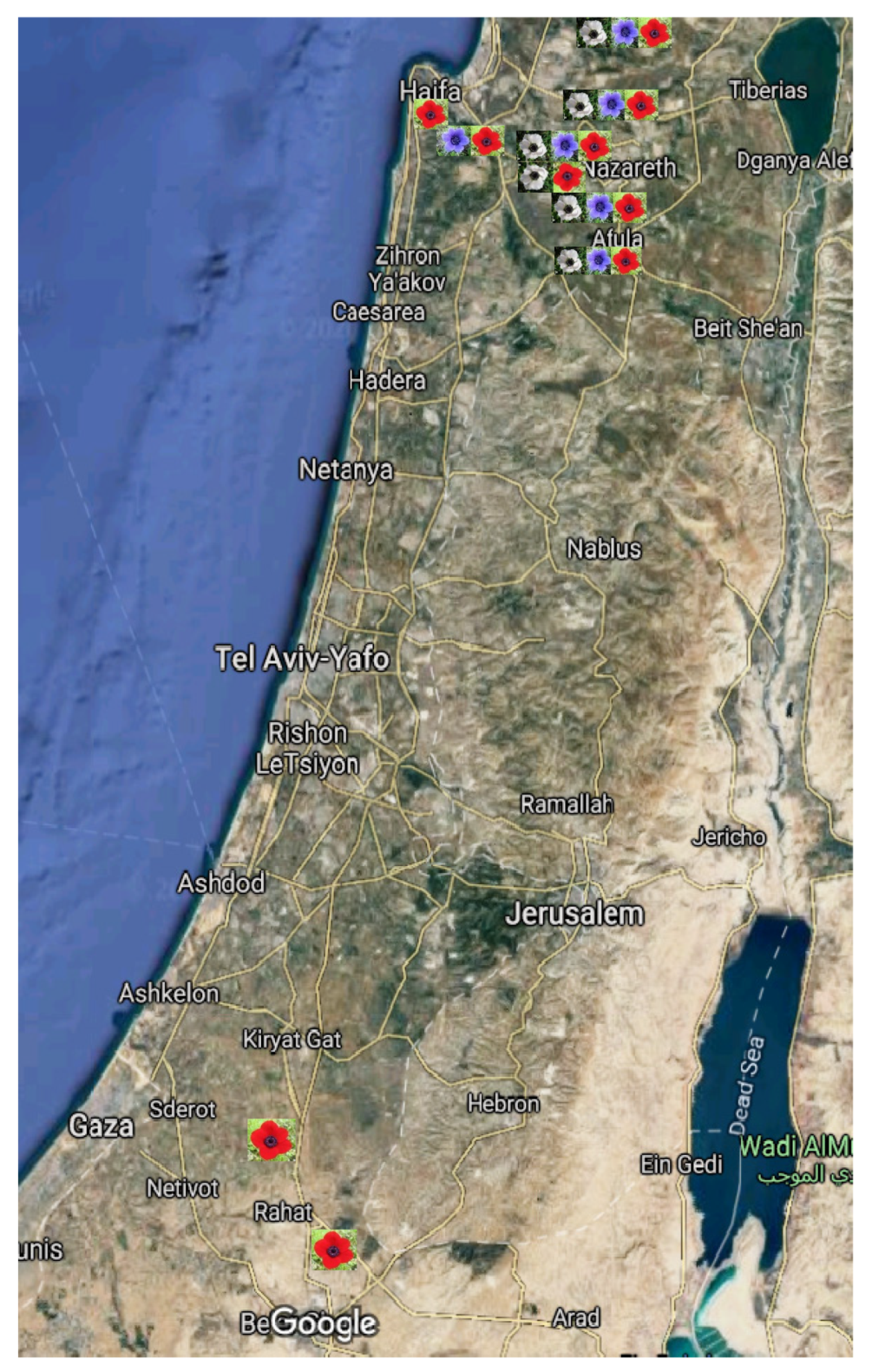
4.1. Study Sites
4.2. Flower Reflectance and Its Perception
4.3. Number of Carpels
4.4. Pollination Modes
4.4.1. Spontaneous Self-Pollination
4.4.2. Wind Pollination
4.5. Flower Visitors
4.6. Genetic Structure of Populations and Colour Morphs
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Joseph, N.; Siril, E.A. Floral color polymorphism and reproductive success in annatto (Bixa orellana L.). Trop. Plant Biol. 2013, 6, 217–227. [Google Scholar] [CrossRef]
- Wang, H.; Conchou, L.; Bessière, J.M.; Cazals, G.; Schatz, B.; Imbert, E. Flower color polymorphism in Iris lutescens (Iridaceae), biochemical analyses in light of plant–insect interactions. Phytochemistry 2013, 94, 123–134. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Talavera, M.; Min, Y.; Flaven, E.; Imbert, E. Neutral processes contribute to patterns of spatial variation for flower colour in the Mediterranean Iris lutescens (Iridaceae). Ann. Bot. 2016, 117, 995–1007. [Google Scholar] [CrossRef] [PubMed]
- Schemske, D.W.; Bierzychudek, P. Evolution of flower color in the desert annual Linanthus parryae, Wright revisited. Evolution 2001, 55, 1269–1282. [Google Scholar] [CrossRef] [PubMed]
- Schemske, D.W.; Bierzychudek, P. Spatial differentiation for flower color in the desert annual Linanthus parryae, was right? Evolution 2007, 61, 2528–2543. [Google Scholar] [CrossRef] [PubMed]
- Narbona, E.; Wang, H.; Ortiz, P.L.; Arista, M.; Imbert, E. Flower colour polymorphism in the Mediterranean Basin, occurrence, maintenance and implications for speciation. Plant Biol. 2017, 20, 8–20. [Google Scholar] [CrossRef]
- Malerba, R.; Nattero, J. Pollinator response to flower color polymorphism and floral display in a plant with a single-locus floral color polymorphism, consequences for plant reproduction. Ecol. Res. 2012, 27, 377–385. [Google Scholar] [CrossRef]
- Streisfeld, M.A.; Kohn, J. Environment and pollinator-mediated selection on parapatric floral races of Mimulus aurantiacus. J. Evol. Biol. 2007, 20, 122–132. [Google Scholar] [CrossRef]
- Hopkins, R.; Rausher, M.D. The cost of reinforcement, selection on flower color in allopatric populations of Phlox drummondii. Am. Nat. 2014, 183, 693–710. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chittka, L.; Thomson, J.D.; Waser, N.M. Flower constancy, insect psychology, and plant evolution. Naturwissenschaften 1999, 86, 361–377. [Google Scholar] [CrossRef]
- Jones, K.N.; Reithel, J.S. Pollinator-mediated selection on a flower color polymorphism in experimental populations of Antirrhinum (Scrophulariaceae). Am. J. Bot. 2001, 88, 447–454. [Google Scholar] [CrossRef]
- Irwin, R.E.; Strauss, S.Y. Flower color microevolution in wild radish, evolutionary response to pollinator-mediated selection. Am. Nat. 2005, 165, 225–237. [Google Scholar] [CrossRef] [PubMed]
- Caruso, C.M.; Scott, S.L.; Wray, J.C.; Walsh, C.A. Pollinators, herbivores, and the maintenance of flower color variation, a case study with Lobelia siphilitica. Int. J. Plant Sci. 2010, 171, 1020–1028. [Google Scholar] [CrossRef]
- Strauss, S.Y.; Irwin, R.E.; Lambrix, V.M. Optimal defence theory and flower petal colour predict variation in the secondary chemistry of wild radish. J. Ecol. 2004, 92, 132–141. [Google Scholar] [CrossRef]
- Arista, M.; Talavera, M.; Berjano, R.; Ortiz, P.L. Abiotic factors may explain the geographical distribution of flower colour morphs and the maintenance of colour polymorphism in the scarlet pimpernel. J. Ecol. 2013, 101, 1613–1622. [Google Scholar] [CrossRef]
- Levin, D.A.; Watkins, L. Assortative mating in Phlox. Heredity 1984, 53, 595–602. [Google Scholar] [CrossRef]
- Cresswell, J.E.; Galen, C. Frequency-dependent selection and adaptive surfaces for floral character combinations, the pollination of Polemonium viscosum. Am. Nat. 1991, 138, 1342–1353. [Google Scholar] [CrossRef]
- Smithson, A.; MacNair, M.R. Negative frequency-dependent selection by pollinators on artificial flowers without rewards. Evolution 1997, 51, 715–723. [Google Scholar] [CrossRef]
- Gegear, R.J.; Laverty, T.M. The effects of variation among floral traits on the flower constancy of pollinators. In Cognitive Ecology of Pollination; Chittka, L., Thomson, J.D., Eds.; Cambridge University Press: Cambridge, UK, 2001; pp. 297–317. [Google Scholar]
- Gigord, L.D.B.; MacNair, M.R.; Smithson, A. Negative frequency-dependent selection maintains a dramatic flower color polymorphism in the rewardless orchid Dactylorhiza sambucina (L.) Soo. Proc. Natl. Acad. Sci. USA 2001, 98, 6253–6255. [Google Scholar] [CrossRef]
- Eckhart, V.M.; Rushing, N.S.; Hart, G.M.; Hansen, J.D. Frequency-dependent pollinator foraging in polymorphic Clarkia xantiana ssp. Xantiana populations, implications for flower colour evolution and pollinator interactions. Oikos 2006, 112, 412–421. [Google Scholar] [CrossRef]
- Rausher, M.D.; Fry, J.D. Effects of a locus affecting floral pigmentation in Ipomoea purpurea on female fitness components. Genetics 1993, 134, 1237–1247. [Google Scholar]
- Subramaniam, B.; Rausher, M.D. Balancing selection on a floral polymorphism. Evolution 2000, 54, 691–695. [Google Scholar] [CrossRef] [PubMed]
- Levin, D.A.; Brack, E.T. Natural selection against white petals in Phlox. Evolution 1995, 49, 1017–1022. [Google Scholar] [CrossRef]
- Carlson, J.E.; Holsinger, K.E. Natural selection on inflorescence color polymorphisms in wild Protea populations, the role of pollinators, seed predators and inter-trait correlations. Am. J. Bot. 2010, 97, 934–944. [Google Scholar] [CrossRef] [PubMed]
- Carlson, J.E.; Holsinger, K.E. Developmental plasticity in Protea as an evolutionary response to environmental clines in the Cape Floristic Region. PLoS ONE 2012, 7, e52035. [Google Scholar] [CrossRef] [PubMed]
- Bernhardt, P.; Edens-Meier, R.; Jocson, D.; Zweck, J.; Ren, Z.X.; Camilo, G.R.; Arduser, M. Comparative floral ecology of bicolor and concolor morphs of Viola pedata L. (Violaceae) following controlled burns. J. Pollinat. Ecol. 2016, 19, 57–70. [Google Scholar] [CrossRef]
- Warren, J.; Mackenzie, S. Why are all colour combinations not equally represented as flower-colour polymorphisms? New Phytol. 2001, 151, 237–241. [Google Scholar] [CrossRef]
- Steyn, W.J.; Wand, S.J.E.; Holcroft, D.M.; Jacobs, G. Anthocyanins in vegetative tissues, a proposed unified function in photoprotection. New Phytol. 2002, 155, 349–361. [Google Scholar] [CrossRef]
- Brown, B.A.; Clegg, M.T. The influence of flower color polymorphisms on genetic transmission in a natural population of the common morning glory, Ipomoea purpurea. Evolution 1984, 38, 796–803. [Google Scholar] [CrossRef]
- Frey, F.M. Opposing natural selection from herbivores and pathogens may maintain floral-color variation in Claytonia virginica (Portulacaceae). Evolution 2004, 58, 2426–2437. [Google Scholar] [CrossRef]
- Raguso, R.A. Wake up and smell the roses, the ecology and evolution of floral scent. Annu. Rev. Ecol. Evol. Syst. 2008, 39, 549–569. [Google Scholar] [CrossRef]
- Johnson, E.T.; Berhow, M.A.; Dowd, P.F. Colored and white sectors from star-patterned petunia flowers display differential resistance to corn earworm and cabbage looper larvae. J. Chem. Ecol. 2008, 34, 757–765. [Google Scholar] [CrossRef] [PubMed]
- Irwin, R.E.; Strauss, S.Y.; Storz, S.; Emerson, A.; Guibert, G. The role of herbivores in the maintenance of a flower color polymorphism in wild radish. Ecology 2003, 84, 1733–1743. [Google Scholar] [CrossRef]
- Whitney, K.D.; Stanton, M.L. Insect seed predators as novel agents of selection on fruit color. Ecology 2004, 84, 2153–2160. [Google Scholar] [CrossRef]
- Brody, A.K.; Mitchell, R.J. Effects of experimental manipulation of inflorescence size on pollination and pre-dispersal seed predation in the hummingbird-pollinated plant Ipomopsis aggregata. Oecologia 1997, 110, 86–93. [Google Scholar] [CrossRef]
- Galen, C.; Cuba, J. Down the tube, pollinators, predators, and the evolution of flower shape in the alpine skypilot, Polemonium viscosum. Evolution 2001, 55, 1963–1971. [Google Scholar] [CrossRef]
- Adler, L.S.; Bronstein, J.L. Attracting antagonists, does floral nectar increase leaf herbivory? Ecology 2004, 85, 1519–1526. [Google Scholar] [CrossRef]
- Strauss, S.Y.; Whittall, J.B. Non-pollinating agents of selection on floral traits. In Ecology and Evolution of Flowers; Harder, L.D., Barrett, S.C.H., Eds.; Oxford University Press: Oxford, UK, 2006; pp. 120–138. [Google Scholar]
- Fineblum, W.L.; Rausher, M.D. Do floral pigmentation genes also influence resistance to enemies? The W locus in Ipomoea purpurea. Ecology 1997, 78, 1646–1654. [Google Scholar] [CrossRef]
- Simms, E.L.; Bucher, M.A. Pleiotropic effects of flower-color intensity on herbivore performance on Ipomoea purpurea. Evolution 1996, 50, 957–963. [Google Scholar] [CrossRef]
- Armbruster, W.S. Can indirect selection and genetic context contribute to trait diversification? A transition-probability study of blossom-colour evolution in two genera. J. Evol. Biol. 2002, 15, 468–486. [Google Scholar] [CrossRef]
- Strauss, S.Y.; Irwin, R.E. Ecological and evolutionary consequences of multispecies plant–animal interactions. Annu. Rev. Ecol. Evol. Syst. 2004, 35, 435–466. [Google Scholar] [CrossRef]
- Bowman, R.N. Cryptic self-incompatibility and the breeding system of Clarkia unguiculata (Onagraceae). Am. J. Bot. 1987, 74, 471–476. [Google Scholar] [CrossRef]
- Grace, S.C.; Logan, B.A. Energy dissipation and radical scavenging by the plant phenylpropanoid pathway. Philos. Trans. R. Soc. B Biol. Sci. 2000, 355, 1402–1499. [Google Scholar] [CrossRef]
- Koes, R.E.; Quattrocchio, F.; Mol, J.N.M. The flavonoid biosynthetic pathway in plants, function and evolution. Bioessays 1994, 16, 123–132. [Google Scholar] [CrossRef]
- Coberly, L.C.; Rausher, M.D. Analysis of a chalcone synthase mutant in Ipomoea purpurea reveals a novel function for flavonoids, amelioration of heat stress. Mol. Ecol. 2003, 12, 1113–1124. [Google Scholar] [CrossRef]
- Horovitz, A.; Galil, J.; Zohary, D. Biological flora of Israel, 6. Anemone coronaria L. Isr. J. Bot. 1975, 24, 26–41. [Google Scholar]
- Harvey, D.M. Phenotypic variation in flower colour within the Anemone coronaria cultivars. Ann. Bot. 1971, 35, 1–8. [Google Scholar] [CrossRef]
- Horovitz, A. The local gene pool of Anemone coronria. Ornam. Plant Improv. Class. Mol. 1995, 420, 144–146. [Google Scholar] [CrossRef]
- Horovitz, A.; Zohary, D. Spontaneous variegation for perianth colour in wild Anemone coronaria. Heredity 1966, 21, 513–515. [Google Scholar] [CrossRef]
- Horovitz, A. The pollination syndrome of Anemone coronaria L.; an insect-biased mutualism. VI Int. Symp. Pollinat. 1990, 288, L283–L287. [Google Scholar] [CrossRef]
- Dafni, A.; Bernhardt, P.; Shmida, A.; Ivri, Y.; Greenbaum, S.; O’Toole, C.; Losito, L. Red bowl-shaped flowers, convergence for beetle pollination in the Mediterranean region. Isr. J. Bot. 1990, 39, 81–92. [Google Scholar]
- Keasar, T.; Shmida, A.; Zylbertal, A. Pollination Ecology of the Red Anemone coronaria (Ranunclaceae), Honeybees May Select for Early Flowering; Center for the Study of Rationality: Jerusalem, Israel, 2008; Available online: https://www.researchgate.net/profile/Avi_Shmida/publication/23531070 (accessed on 6 February 2020).
- Keasar, T.; Harari, A.R.; Sabatinelli, G.; Keith, D.; Dafni, A.; Shavit, O.; Shmida, A. Red anemone guild flowers as focal places for mating and feeding by Levant glaphyrid beetles. Biol. J. Linn. Soc. 2010, 99, 808–817. [Google Scholar] [CrossRef]
- Keasar, T.; Kishinevsky, M.; Shmida, A.; Gerchman, Y.; Chinkov, N.; Koplovich, A.; Katzir, G. Plant-derived visual signals may protect beetle herbivores from bird predators. Behav. Ecol. Sociobiol. 2013, 67, 1613–1622. [Google Scholar] [CrossRef][Green Version]
- Perevolotsky, A.; Schwartz-Tzachor, R.; Yonathan, R.; Ne’eman, G. Geophytes–herbivore interactions, reproduction and population dynamics of Anemone coronaria L. Plant Ecol. 2011, 212, 563–571. [Google Scholar] [CrossRef]
- Wallach, A.D.; Inbar, M.; Shanas, U. Roe deer and decapitated Anemone flowers. Isr. J. Plant Sci. 2009, 57, 103–106. [Google Scholar] [CrossRef]
- Chittka, L. The colour hexagon, a chromaticity diagram based on photoreceptor excitations as a generalized representation of colour opponency. J. Comp. Physiol. A Neuroethol. Sens. Neural Behav. Physiol. 1992, 170, 533–543. [Google Scholar] [CrossRef]
- Peakall, R.; Smouse, P.E. GENALEX 6, genetic analysis in excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 6, 288–295. [Google Scholar] [CrossRef]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar]
- Falush, D.; Stephens, M.; Pritchard, J.K. Inference of population structure using multilocus genotype data, dominant markers and null alleles. Mol. Ecol. Notes 2007, 7, 574–578. [Google Scholar] [CrossRef]
- Dyer, A.G.; Chittka, L. Biological significance of distinguishing between similar colours in spectrally variable illumination, bumblebees (Bombus terrestris) as a case study. J. Comp. Physiol. 2004, 190, 105–114. [Google Scholar] [CrossRef]
- Dyer, A.G.; Chittka, L. Fine color discrimination requires differential conditioning in bumblebees. Naturwissenschaften 2004, 91, 224–227. [Google Scholar] [CrossRef] [PubMed]
- Chittka, L.; Gumbert, A.; Kunze, J. Foraging dynamics of bumble bees, correlates of movements within and between plant species. Behav. Ecol. 1997, 8, 239–249. [Google Scholar] [CrossRef]
- Lunau, K. Visual ecology of flies with particular reference to colour vision and colour preferences. J. Comp. Physiol. A 2014, 200, 497–512. [Google Scholar] [CrossRef]
- Martinez-Harms, J.; Vorobyev, M.; Schorn, J.; Shmida, A.; Keasar, T.; Homberg, U.; Schmeling, F.; Menzel, R. Evidence of red sensitive photoreceptors in Pygopleurus israelitus (Glaphyridae, Coleoptera) and its implications for beetle pollination in the southeast Mediterranean. J. Comp. Physiol. A 2012, 198, 451–463. [Google Scholar] [CrossRef]
- Regal, P.J. Pollination by wind and animals, ecology of geographic patterns. Annu. Rev. Ecol. Syst. 1982, 13, 497–524. [Google Scholar] [CrossRef]
- Duan, Y.W.; Zhang, T.F.; He, Y.P.; Liu, J.Q. Insect and wind pollination of an alpine biennial Aconitum gymnandrum (Ranunculaceae). Plant Biol. 2009, 11, 796–802. [Google Scholar] [CrossRef]
- De la Bandera, M.C.; Traveset, A. Breeding system and spatial variation in the pollination biology of the heterocarpic Thymelaea velutina (Thymelaeaceae). Plant Syst. Evol. 2006, 257, 9–23. [Google Scholar] [CrossRef]
- Dafni, A.; Marom-Levy, T.; Jürgens, A.; Dötterl, S.; Dorchin, A.; Shimrat, Y.; Kirkpatrick, E.B. Ambophily and “super generalism” in Ceratonia siliqua pollination. In Evolution of Plant–Pollinator Relationships; Patiny, S., Ed.; Cambridge University Press: Cambridge, UK, 2012; pp. 344–373. [Google Scholar]
- Cursach, J.; Rita, J. Reproductive biology of Ranunculus weyleri (Ranunculaceae), a narrowly endemic plant from the Balearic Islands with disjunct populations. Flora Morphol. Distrib. Funct. Ecol. Plants 2012, 207, 726–735. [Google Scholar] [CrossRef]
- Kaplan, S.M.; Mulcahy, D.L. Mode of pollination and floral sexuality in Thalictrum. Evolution 1971, 25, 659–668. [Google Scholar] [CrossRef]
- Dafni, A. Autumnal and winter pollination adaptations under Mediterranean conditions. Bocconea 1996, 5, 171–181. [Google Scholar]
- Hedrick, P.W. Genetic polymorphism in heterogeneous environments, a decade later. Annu. Rev. Ecol. Syst. 1986, 17, 535–566. [Google Scholar] [CrossRef]
- Hedrick, P.W.; Ginevan, M.E.; Ewing, E.P. Genetic polymorphism in heterogeneous environments. Annu. Rev. Ecol. Syst. 1976, 7, 1–32. [Google Scholar] [CrossRef]
- Ellner, S.; Sasaki, A. Patterns of genetic polymorphism maintained by fluctuating selection with overlapping generations. Theor. Popul. Biol. 1996, 50, 31–65. [Google Scholar] [CrossRef] [PubMed]
- Rausher, M.D. Evolutionary transitions in floral color. Int. J. Plant Sci. 2008, 169, 7–21. [Google Scholar] [CrossRef]
- Dormont, L.; Delle-Vedove, R.; Bessière, J.M.; Key, H.M.; Schatz, B. Rare white flowered morphs increase the reproductive success of common purple morphs in a food deceptive orchid. New Phytol. 2010, 185, 300–310. [Google Scholar] [CrossRef]
- Suchet, C.; Dormont, L.; Schatz, B.; Giurfa, M.; Simon, V.; Raynaud, C.; Chave, J. Floral scent variation in two Antirrhinum majus subspecies influences the choice of naive bumblebees. Behav. Ecol. Sociobiol. 2011, 65, 1015–1027. [Google Scholar] [CrossRef]
- Raine, N.E.; Chittka, L. The adaptive significance of sensory bias in a foraging context, floral colour preferences in the bumblebee Bombus terrestris. PLoS ONE 2007, 2, e556. [Google Scholar] [CrossRef]
- Kay, Q.O.N. Preferential pollination of yellow-flowered morphs of Raphanus raphanistrum by Pieris and Eristalis spp. Nature 1976, 261, 230–232. [Google Scholar] [CrossRef]
- Stanton, M.L. Reproductive biology of petal color variants of wild populations of Raphanus sativus L., I. pollinator response to color morphs. Am. J. Bot. 1987, 74, 176–185. [Google Scholar] [CrossRef]
- Horovitz, A. Edaphic factors and flower colour distribution in the Anemone coronaria (Ranunculaceae). Plant Syst. Evol. 1976, 126, 239–242. [Google Scholar] [CrossRef]
- Peitsch, D.; Fietz, A.; Hertel, H.; Souza, J.; Ventura, D.F.; Menzel, R. The spectral input systems of hymenopteran insects and their receptor-based colour vision. J. Comp. Physiol. A Neuroethol. Sens. Neural Behav. Physiol. 1992, 170, 23–40. [Google Scholar] [CrossRef] [PubMed]
- Chittka, L.; Kevan, P.G. Flower colour as advertisement. In Practical Pollination Biology; Dafni, A., Kevan, P.G., Husband, B.C., Eds.; Enviroquest Ltd.: Cambridge, ON, Canada, 2005; pp. 157–196. [Google Scholar]
- Dyer, A.G.; Spaethe, J.; Prack, S. Comparative psychophysics of bumblebee and honeybee colour discrimination and object detection. J. Comp. Physiol. A 2008, 194, 617–627. [Google Scholar] [CrossRef] [PubMed]
- Vos, P.; Hogers, R.; Bleeker, M.; Reijans, M.; Lee, T.V.D.; Hornes, M.; Frijters, A.; Pot, J.; Peleman, J.; Kuiper, M. AFLP, a new technique for DNA fingerprinting. Nucl. Acids Res. 1995, 3, 4407–4414. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.P. Tools for Population Genetic Analyses (TFPGA), version 1.3; Department of Biological Sciences, Northern Arizona University: Flagstaff, AZ, USA, 1997. [Google Scholar]
- Nei, M. Estimation of Average Heterozygosity and genetic distance from a small number of individuals. Genetics 1978, 89, 583–590. [Google Scholar] [PubMed]
- Raymond, M.; Rousset, F. An exact test for population differentiation. Evolution 1995, 49, 1280–1283. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE, a simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef]
- Earl, D.A. Structural harvester, a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 2012, 4, 359–361. [Google Scholar] [CrossRef]

| Visitor | White Flowers | Purple-Blue Flowers | Red Flowers | Total | P |
|---|---|---|---|---|---|
| Honeybee | 520 | 252 | 0 | 772 | <0.005 |
| Andrena sp. | 75 | 16 | 0 | 91 | <0.005 |
| Syrphid flies | 24 | 7 | 0 | 31 | 0.056 |
| Beetles | 0 | 0 | 1 | 1 | |
| Total visits | 619 | 275 | 1 | 895 | |
| Number of flowers | 694 | 377 | 115 | 1015 |
| All | Red | White | Purple-Blue | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Population | N | He | SE | P(%) | N | He | SE | N | He | SE | N | He | SE |
| Mt. Meron | 28 | 0.156 | 0.007 | 87.7 | 14 | 0.139 | 0.007 | 6 | 0.133 | 0.008 | 8 | 0.191 | 0.008 |
| Zippori | 41 | 0.141 | 0.007 | 89.2 | 12 | 0.109 | 0.006 | 14 | 0.153 | 0.007 | 15 | 0.114 | 0.007 |
| Haifa University | 15 | 0.115 | 0.006 | 66.9 | 15 | 0.115 | 0.006 | -- | -- | -- | -- | -- | -- |
| Nof Carmel | 19 | 0.136 | 0.007 | 77.9 | 8 | 0.096 | 0.007 | -- | -- | -- | 11 | 0.153 | 0.007 |
| Alonim | 19 | 0.120 | 0.006 | 67.8 | 11 | 0.096 | 0.007 | 3 | 0.109 | 0.009 | 5 | 0.154 | 0.008 |
| Beit Shearim | 12 | 0.14 | 0.007 | 63.6 | 10 | 0.14 | 0.007 | 2 | 0.097 | 0.01 | -- | -- | -- |
| Balfouria | 35 | 0.094 | 0.006 | 60.6 | 15 | 0.074 | 0.005 | 12 | 0.094 | 0.007 | 8 | 0.101 | 0.008 |
| Megiddo | 34 | 0.150 | 0.006 | 93.7 | 11 | 0.077 | 0.006 | 11 | 0.177 | 0.008 | 12 | 0.178 | 0.007 |
| Purra | 18 | 0.108 | 0.007 | 60.0 | 14 | 0.097 | 0.007 | -- | -- | -- | -- | -- | -- |
| Lahav | 15 | 0.084 | 0.007 | 39.4 | 14 | 0.085 | 0.007 | -- | -- | -- | -- | -- | -- |
| Total | 236 | 0.124 | 0.002 | 70.7 | 124 | 0.102 | 0.006 | 48 | 0.136 | 0.007 | 59 | 0.146 | 0.007 |
| Site Name | GPS N E | Colour Morphs | Visitor Activity 2011-12 | Visitor Activity 2014 | DNA Analyses | Colour Analysis |
|---|---|---|---|---|---|---|
| 1. Alonei Abba | 32.73 | RBW | ✔ | |||
| 35.17 | ||||||
| 2. Alonim | 32.72 | RBW | ✔ | ✔ | ||
| 35.15 | ||||||
| 3. Balfuria | 32.63 | RBW | ✔ | |||
| 35.29 | ||||||
| 4. Beit Lehem Haglilit* | 32.74 | RBW | ✔ | |||
| 35.19 | ||||||
| 5. Beit Shearim | 32.70 | RW | ✔ | |||
| 35.13 | ||||||
| 6. Haifa University | 32.75 | R | ✔ | |||
| 35.03 | ||||||
| 7. Lehavim | 32.37 | R | ✔ | |||
| 34.85 | ||||||
| 8. Megiddo | 32.59 | RBW | ✔ | ✔ | ✔ | ✔ |
| 35.23 | ||||||
| 9. Mt. Meiron | 33.01 | RBW | ✔ | |||
| 35.40 | ||||||
| 10. Nahal Sanin | 32.64 | RBW | ✔ | |||
| 35.10 | ||||||
| 11. Nof Carmel | 32.75 | RBW | ✔ | |||
| 35.05 | ||||||
| 12. Pura | 31.50 | R | ✔ | |||
| 34.78 | ||||||
| 13. Tsippori | 32.76 | RBW | ✔ | ✔ | ||
| 35.24 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dafni, A.; Tzohari, H.; Ben-Shlomo, R.; Vereecken, N.J.; Ne’eman, G. Flower Colour Polymorphism, Pollination Modes, Breeding System and Gene Flow in Anemone coronaria. Plants 2020, 9, 397. https://doi.org/10.3390/plants9030397
Dafni A, Tzohari H, Ben-Shlomo R, Vereecken NJ, Ne’eman G. Flower Colour Polymorphism, Pollination Modes, Breeding System and Gene Flow in Anemone coronaria. Plants. 2020; 9(3):397. https://doi.org/10.3390/plants9030397
Chicago/Turabian StyleDafni, Amots, Hagai Tzohari, Rachel Ben-Shlomo, Nicolas J. Vereecken, and Gidi Ne’eman. 2020. "Flower Colour Polymorphism, Pollination Modes, Breeding System and Gene Flow in Anemone coronaria" Plants 9, no. 3: 397. https://doi.org/10.3390/plants9030397
APA StyleDafni, A., Tzohari, H., Ben-Shlomo, R., Vereecken, N. J., & Ne’eman, G. (2020). Flower Colour Polymorphism, Pollination Modes, Breeding System and Gene Flow in Anemone coronaria. Plants, 9(3), 397. https://doi.org/10.3390/plants9030397





