Impact of Spontaneous Haploid Genome Doubling in Maize Breeding
Abstract
1. Overview of Doubled Haploid Technology in Maize
1.1. History and Uses of DH Technology
1.2. Methods for Producing DHs in Maize
2. Biology and Genetic Architecture of Spontaneous Haploid Genome Doubling
2.1. Biology of Spontaneous Haploid Genome Doubling
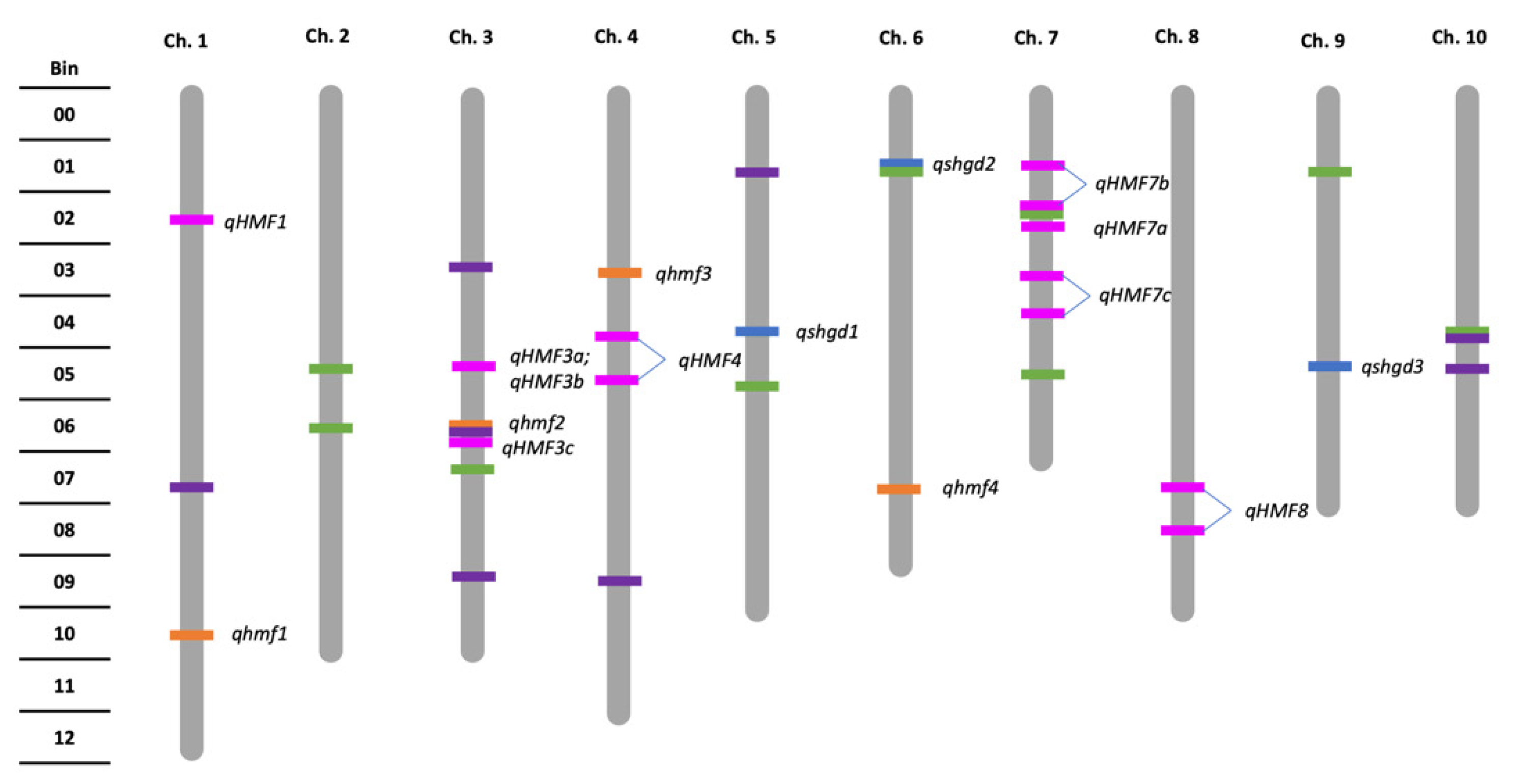
2.2. Genetic Architecture and Candidate Genes of Spontaneous Haploid Genome Doubling
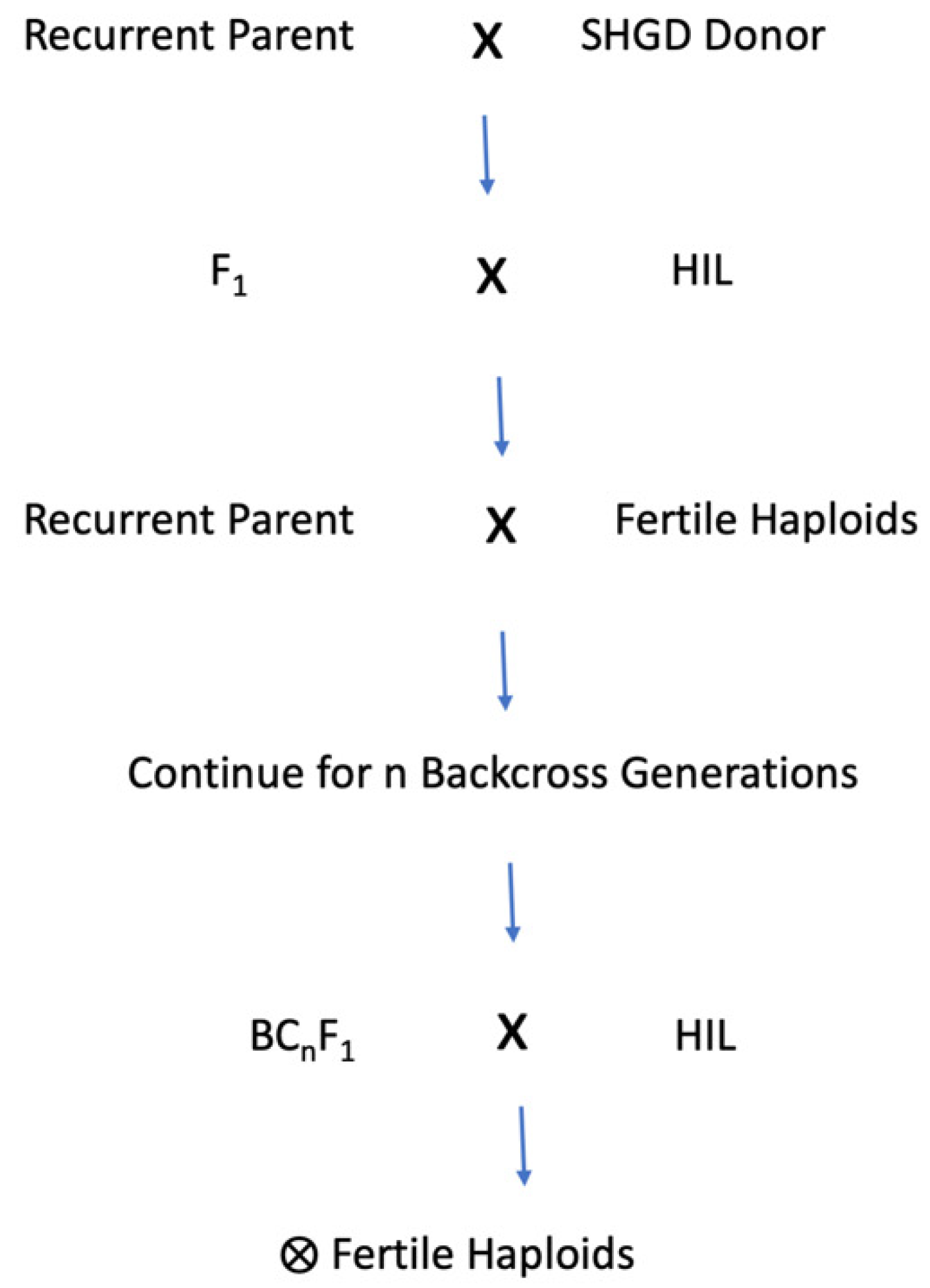
3. Introgression of Spontaneous Haploid Genome Doubling
4. Implications of Using SHGD in Maize Breeding
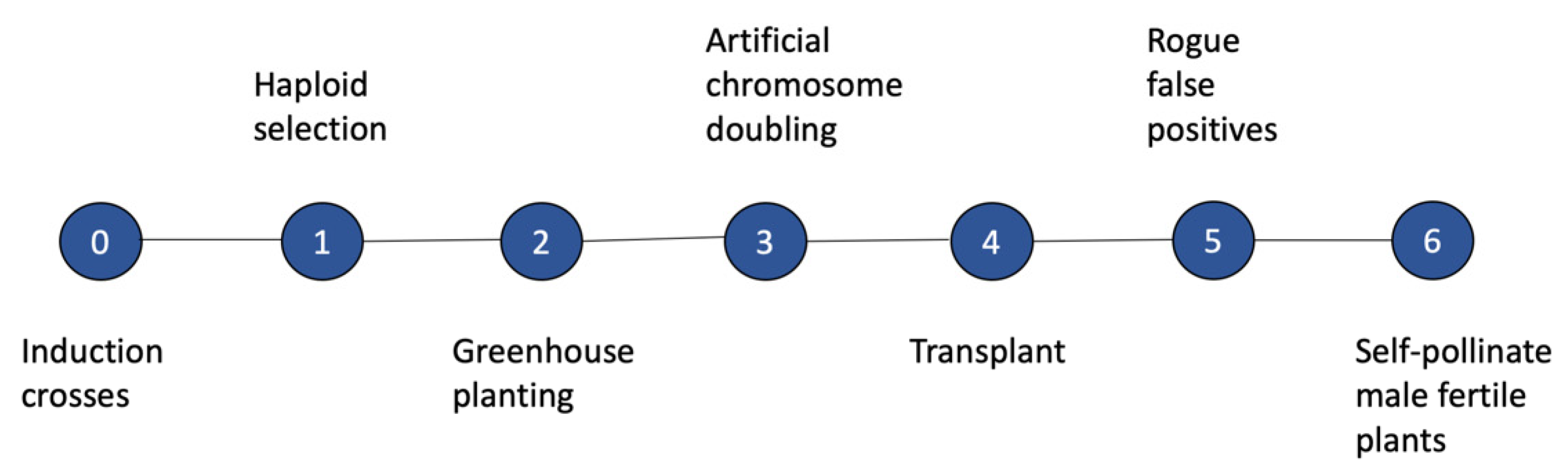
4.1. Utilization of SHGD within a Maize Breeding Program
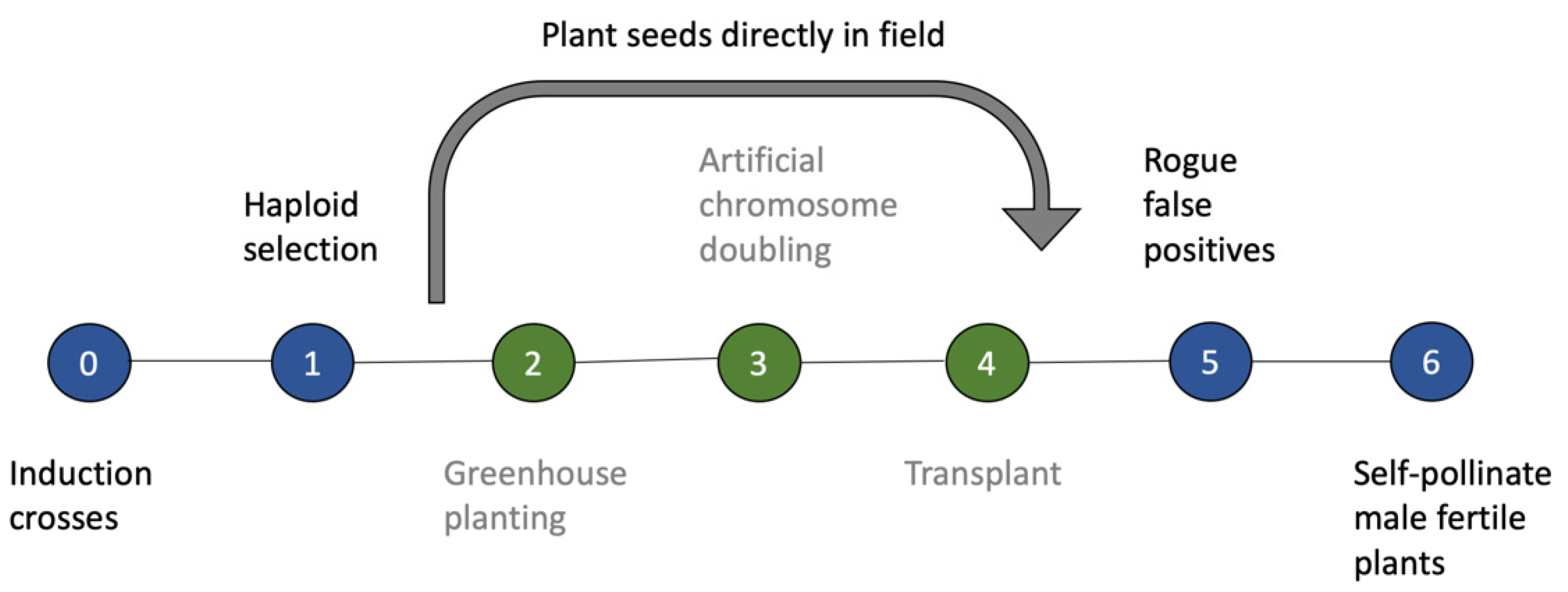
4.2. Advanced Applications of SHGD in Maize Breeding
Author Contributions
Funding
Conflicts of Interest
References
- Randolph, L.F. Some effects of high temperature on polyploidy and other variations in maize. Proc. Natl. Acad. Sci. USA 1932, 18, 22–229. [Google Scholar] [CrossRef] [PubMed]
- Blakeslee, A.F.; Avery, A.G. Methods of inducing doubling of chromosomes in plants by treatment with colchicine. J. Hered. 1937, 12, 393–411. [Google Scholar] [CrossRef]
- Chase, S.S. Production of homozygous diploids of maize from monoploids. Agronomia 1952, 44, 263–267. [Google Scholar] [CrossRef]
- Geiger, H.H.; Gordillo, G.A. Doubled haploids in hybrid maize breeding. Maydica 2009, 54, 485–499. [Google Scholar]
- Seitz, G. The use of doubled haploids in corn breeding. In Proceedings of the Forty First Annual ILLINOIS Corn Breeders’ School, Urbana-Champaign, Champaign, IL, USA, 7–8 March 2005; pp. 1–7. [Google Scholar]
- Strigens, A.; Schipprack, W.; Reif, J.C.; Melchinger, A. Unlocking the genetic diversity of maize landraces with doubled haploids opens new avenues for breeding. PLoS ONE 2013, 8, e57234. [Google Scholar] [CrossRef]
- Griffing, B. Efficiency changes due to use of doubles haploids in recurrent selection methods. Theor. Appl. Genet. 1975, 46, 367–386. [Google Scholar] [CrossRef]
- Choo, T.M. Doubled haploids for estimating mean and variance of recombination values. Genetics 1981, 97, 165–172. [Google Scholar]
- Snape, J.W.; Simpson, E. The utilization of doubled haploid lines in quantitative genetics. Bull. Soc. Bot. Fr. 1986, 4, 59–66. [Google Scholar]
- Hallauer, A.R.; Russell, W.A.; Lamkey, K.R. Corn breeding. In Corn and Corn Improvement, 3rd ed.; ASA-CSSA-SSSA: Madison, WI, USA, 1988; pp. 463–564. [Google Scholar]
- Gallais, A.; Bordes, J. The use of doubled haploids in recurrent selection and hybrid development in maize. Crop Sci. 2007, 47, 190–201. [Google Scholar] [CrossRef]
- Forster, B.P.; Thomas, W.T.B. Doubled haploids in genetics and plant breeding. In Plant Breeding Reviews; Janick, J., Ed.; John, C. Wiley and Sons, Inc.: Hoboken, NJ, USA, 2005; Volume 25, pp. 57–88. [Google Scholar]
- Lübberstedt, T.; Frei, U.K. Application of doubled haploids for target gene fixation in backcross programmes of maize. Plant Breed. 2012, 131, 449–452. [Google Scholar] [CrossRef]
- Andorf, C.; Beavis, W.D.; Hufford, M.; Smith, S.; Suza, W.P.; Wang, K.; Woodhouse, M.; Yu, J.; Lübberstedt, T. Technological advances in maize breeding: Past, present and future. Theor. Appl. Genet. 2019, 132, 817–849. [Google Scholar] [CrossRef] [PubMed]
- Bernardo, R.; Yu, J. Prospects for genomewide selection for quantitative traits in maize. Crop Sci. 2007, 47, 1082–1090. [Google Scholar] [CrossRef]
- Mayor, P.J.; Bernardo, R. Genomewide selection and marker-asisted selection in doubled haploid versus F2 Populations. Crop Sci. 2009, 49, 1719–1725. [Google Scholar] [CrossRef]
- Wu, Y.; Frei, U.K.; Liu, H.; De La Fuente, G.; Huang, K.; Wei, Y.; Lübberstedt, T. Combining genomic selection and doubled haploid technology increases efficiency of maize breeding. In Biotechnology Volume 2: Plant Biotechnology, 1st ed.; Kumar, P.A., Ed.; Studium Press LLC: New Delhi, India, 2014; Volume 2. [Google Scholar]
- Melchinger, A.E.; Molenaar, W.S.; Mirdita, V.; Schipprack, W. Colchicine alternatives for chromosome doubling in maize haploids for doubled-haploid production. Crop Sci. 2016, 56, 559–569. [Google Scholar] [CrossRef]
- Molenaar, W.S.; Schipprack, W.; Brauner, P.C.; Melchinger, A.E. Haploid male fertility and spontaneous chromosome doubling evaluated in a diallel and recurrent selection experiment in maize. Theor. Appl. Genet. 2019, 132, 2273–2284. [Google Scholar] [CrossRef] [PubMed]
- De La Fuente, G.N.; Frei, U.K.; Trampe, B.; Ren, J.; Bohn, M.; Yana, N.; Verzegnazzi, A.; Murray, S.C.; Lübberstedt, T. A diallel analysis of a maize donor population response to in vivo maternal haploid induction II: Haploid male fertility. Crop Sci. 2019, in press. [Google Scholar] [CrossRef]
- Ren, J.; Boerman, N.A.; Liu, R.; Vanous, K.; Trampe, B.; Frei, U.K.; Chen, S.; Lübberstedt, T. Mapping of QTL and identification of candidate genes conferring spontaneous haploid genome doubling in maize [Zea mays L.]. Plant Sci. 2019, 293. [Google Scholar] [CrossRef]
- Trampe, B.; Goncalvez, I.; Frei, U.K.; Ren, J.; Chen, S.; Lübberstedt, T. QTL mapping of Spontaneous Haploid Genome Doubling using Genotype by Sequencing Approach in maize. Theor. Appl. Genet. submitted.
- Ren, J.; Wu, P.; Tian, X.; Lübberstedt, T.; Chen, S. QTL mapping for haploid male fertility by a segregation distortion method and fine mapping of a key QTL qhmf4 in maize. Theor. Appl. Genet. 2017, 130, 1349–1359. [Google Scholar] [CrossRef]
- Chalyk, S.T. Properties of maternal haploid maize plants and potential application to maize breeding. Euphytica 1994, 79, 13–18. [Google Scholar] [CrossRef]
- Geiger, H.H.; Braun, M.D.; Gordillo, G.A.; Koch, S.; Jesse, J.; Krutzefeldt, B.A.E. Variation for female fertility among haploid maize lines. Maize Genet. Newsl. 2006, 80, 28–29. [Google Scholar]
- Röber, F.K.; Gordillo, G.A.; Geiger, H.H. In vivo haploid induction in maize-performance of new inducers and significance of doubled haploid lines in hybrid breeding. Maydica 2005, 50, 275–283. [Google Scholar]
- Eder, J.; Chalyk, S. In vivo haploid induction in maize. Theor. Appl. Genet. 2002, 104, 703–708. [Google Scholar] [CrossRef] [PubMed]
- Melchinger, A.E.; Schipprack, W.; Würschum, T.; Chen, S.; Technow, F. Rapid and accurate identification of in vivo-induced haploid seeds based on oil content in maize. Sci. Rep. 2013, 3, e2129. [Google Scholar] [CrossRef] [PubMed]
- Jones, R.W.; Reinot, T.; Frei, U.K.; Tseng, Y.; Lübberstedt, T.; McClelland, J.F. Selection of haploid maize kernels from hybrid kernels for plant breeding using near-infrared spectroscopy and SIMCA analysis. Appl. Spectrosc. 2012, 66, 444–450. [Google Scholar] [CrossRef]
- Bartels, P.G.; Hilton, J.L. Comparison of trifluralin, oryzalin, pronamide, propham, and colchicine treatments on microtubules. Pestic. Biochem. Phys. 1973, 3, 462–472. [Google Scholar] [CrossRef]
- Morejohn, L.C.; Fosket, D.E. Inhibition of plant microtubule polymerization in vitro by the phosphoric amide herbicide amiprophos-methyl. Science 1984, 224, 874–876. [Google Scholar] [CrossRef]
- Kato, A. Chromosome Doubling Method. U.S. Patent 7,135,615, 14 November 2006. [Google Scholar]
- Ma, H.; Li, G.; Würschum, T.; Zhang, Y.; Zheng, D.; Yang, X.; Li, J.; Liu, W.; Yan, J.; Chen, S. Genome-wide association study of haploid male fertility in maize (Zea mays L.). Front. Plant Sci. 2018, 9, 974. [Google Scholar] [CrossRef]
- Chaikam, V.; Gowda, M.; Nair, S.K.; Melchinger, A.E.; Boddupalli, P.M. Genome-wide association study to identify genomic regions influencing spontaneous fertility in maize haploids. Euphytica 2019, 215, 138. [Google Scholar] [CrossRef]
- Kleiber, D.; Prigge, V.; Melchinger, A.E.; Burkard, F.; San Vicente, F.; Palomino, G.; Gordillo, G.A. Haploid fertility in temperate and tropical maize germplasm. Crop Sci. 2012, 52, 623–630. [Google Scholar] [CrossRef]
- Wu, P.; Ren, J.; Li, L.; Chen, S. Early spontaneous diploidization of maternal maize haploids generated by in vivo haploid induction. Euphytica 2014, 200, 127–138. [Google Scholar] [CrossRef]
- Sugihara, N.; Higashigawa, T.; Aramoto, D.; Kato, A. Haploid plants carrying a sodium azide-induced mutation (fdr1) produce fertile pollen grains due to first division restitution (FDR) in maize (Zea mays L.). Theor. Appl. Genet. 2013, 126, 2931–2941. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Qu, Y.; Chen, Q.; Tang, J.; Lübberstedt, T.; Li, H.; Liu, Z. Genetic dissection of haploid male fertility in maize (Zea mays L.). Plan. Breed. 2019, 138, 259–265. [Google Scholar] [CrossRef]
- Li, S.; Sun, T.; Ren, H. The functions of the cytoskeleton and associated proteins during mitosis and cytokinesis in plants cells. Front. Plant Sci. 2015, 6, 282. [Google Scholar] [CrossRef] [PubMed]
- Frisch, M.; Bohn, M.; Melchinger, A.E. Comparison of strategies for marker-assisted backcrossing of a gene. Crop Sci. 1999, 39, 1295–1301. [Google Scholar] [CrossRef]
- Frisch, M.; Melchinger, A.E. Marker-assisted backcrossing for simultaneous introgression of two genes. Crop Sci. 2001, 41, 1716–1725. [Google Scholar] [CrossRef]
- Cameron, J.; Han, Y.; Wang, L.; Beavis, W.D. Systematic design for trait introgression projects. Theor. Appl. Genet. 2017, 130, 1993–2004. [Google Scholar] [CrossRef]
- Kelliher, T.; Starr, D.; Su, X.; Tang, G.; Chen, Z.; Carter, J.; Wittich, P.E.; Dong, S.; Green, J.; Burch, E.; et al. One-step genome editing of elite crop germplasm during haploid induction. Nat. Biotech. 2019, 37, 287–292. [Google Scholar] [CrossRef]
- Jenko, J.; Gorjanc, G.; Cleveland, M.A.; Varshney, R.K.; Whitelaw, C.B.A.; Woolliams, J.A.; Hickey, J.M. Potential of promotion of alleles by genome editing to improve quantitative traits in livestock breeding programs. Genet. Sel. Evol. 2015, 47, 55. [Google Scholar] [CrossRef]
- Bortesi, L.; Fischer, R. The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol. Adv. 2015, 33, 41–52. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Boerman, N.A.; Frei, U.K.; Lübberstedt, T. Impact of Spontaneous Haploid Genome Doubling in Maize Breeding. Plants 2020, 9, 369. https://doi.org/10.3390/plants9030369
Boerman NA, Frei UK, Lübberstedt T. Impact of Spontaneous Haploid Genome Doubling in Maize Breeding. Plants. 2020; 9(3):369. https://doi.org/10.3390/plants9030369
Chicago/Turabian StyleBoerman, Nicholas A., Ursula K. Frei, and Thomas Lübberstedt. 2020. "Impact of Spontaneous Haploid Genome Doubling in Maize Breeding" Plants 9, no. 3: 369. https://doi.org/10.3390/plants9030369
APA StyleBoerman, N. A., Frei, U. K., & Lübberstedt, T. (2020). Impact of Spontaneous Haploid Genome Doubling in Maize Breeding. Plants, 9(3), 369. https://doi.org/10.3390/plants9030369




