miRNA–mRNA Integrated Analysis Reveals Roles for miRNAs in a Typical Halophyte, Reaumuria soongorica, during Seed Germination under Salt Stress
Abstract
1. Introduction
2. Results
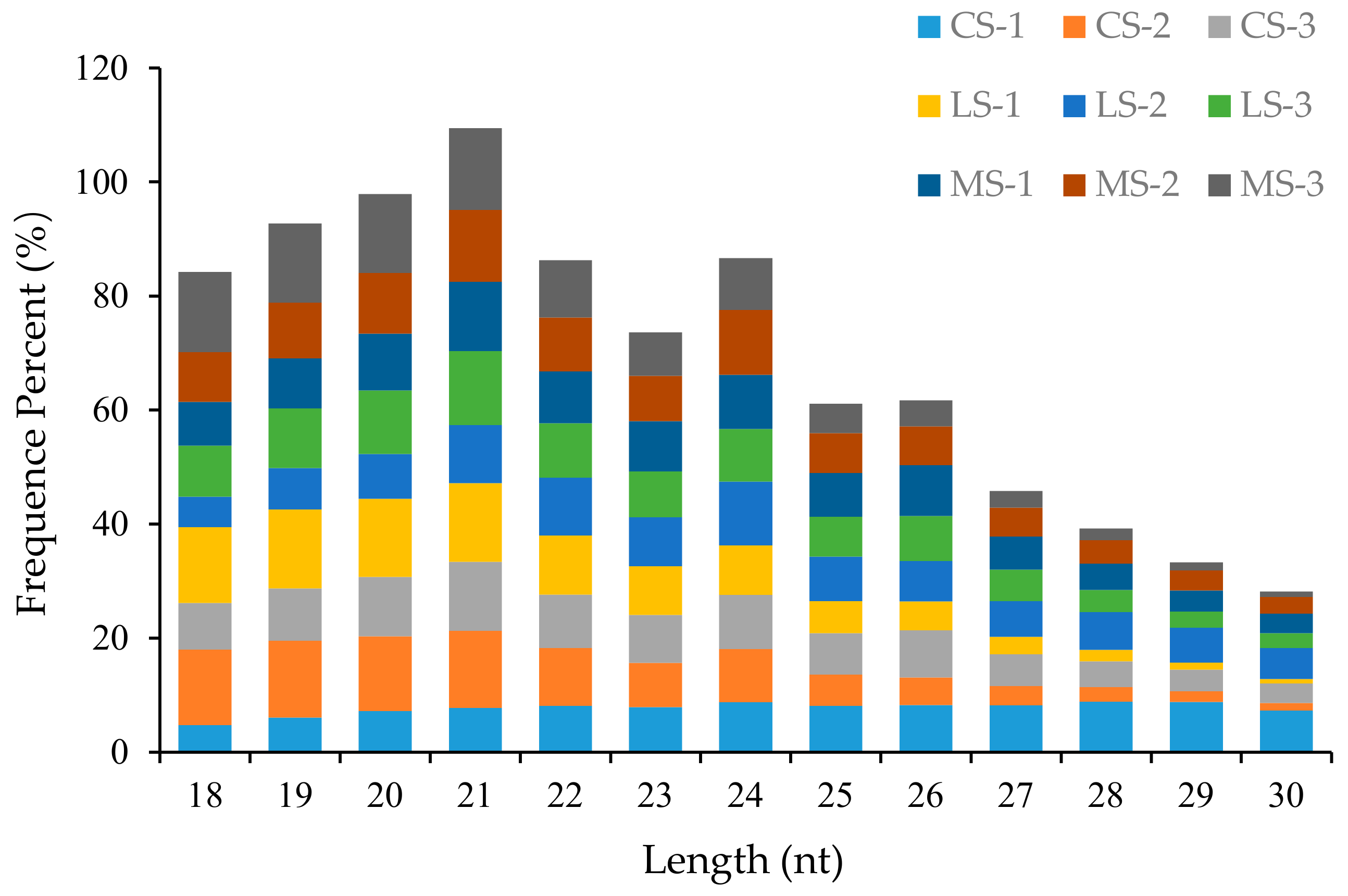
2.1. Summary of the Small-RNA Library Dataset Obtained from Deep Sequencing of R. soongorica
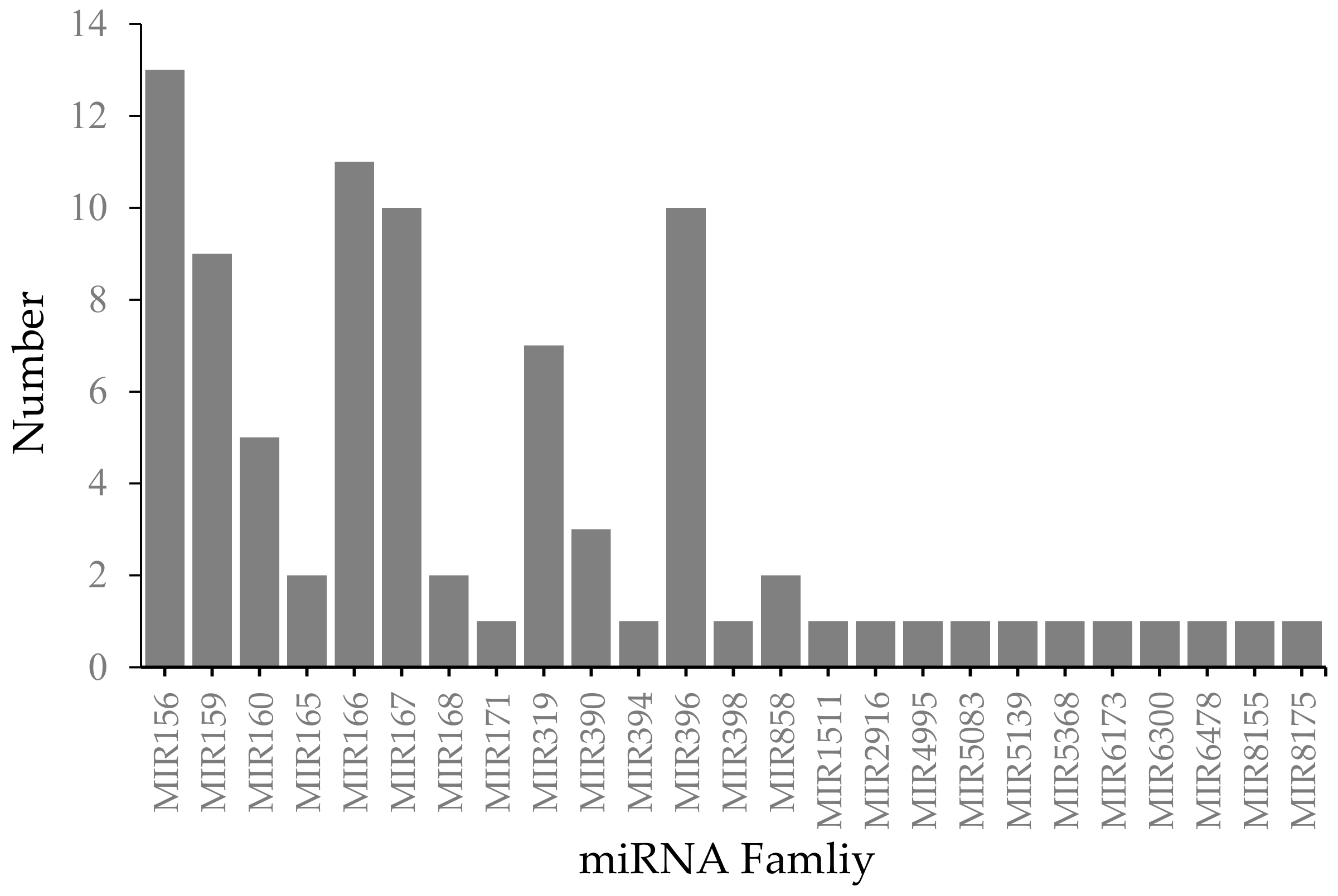
2.2. Identification of Conserved miRNAs from Known Families
2.3. Discovery of Novel miRNAs in R. soongorica Seeds
2.4. Differential Expression of miRNAs during Germination of R. soongorica Seeds Exposed to Various Salt Treatments
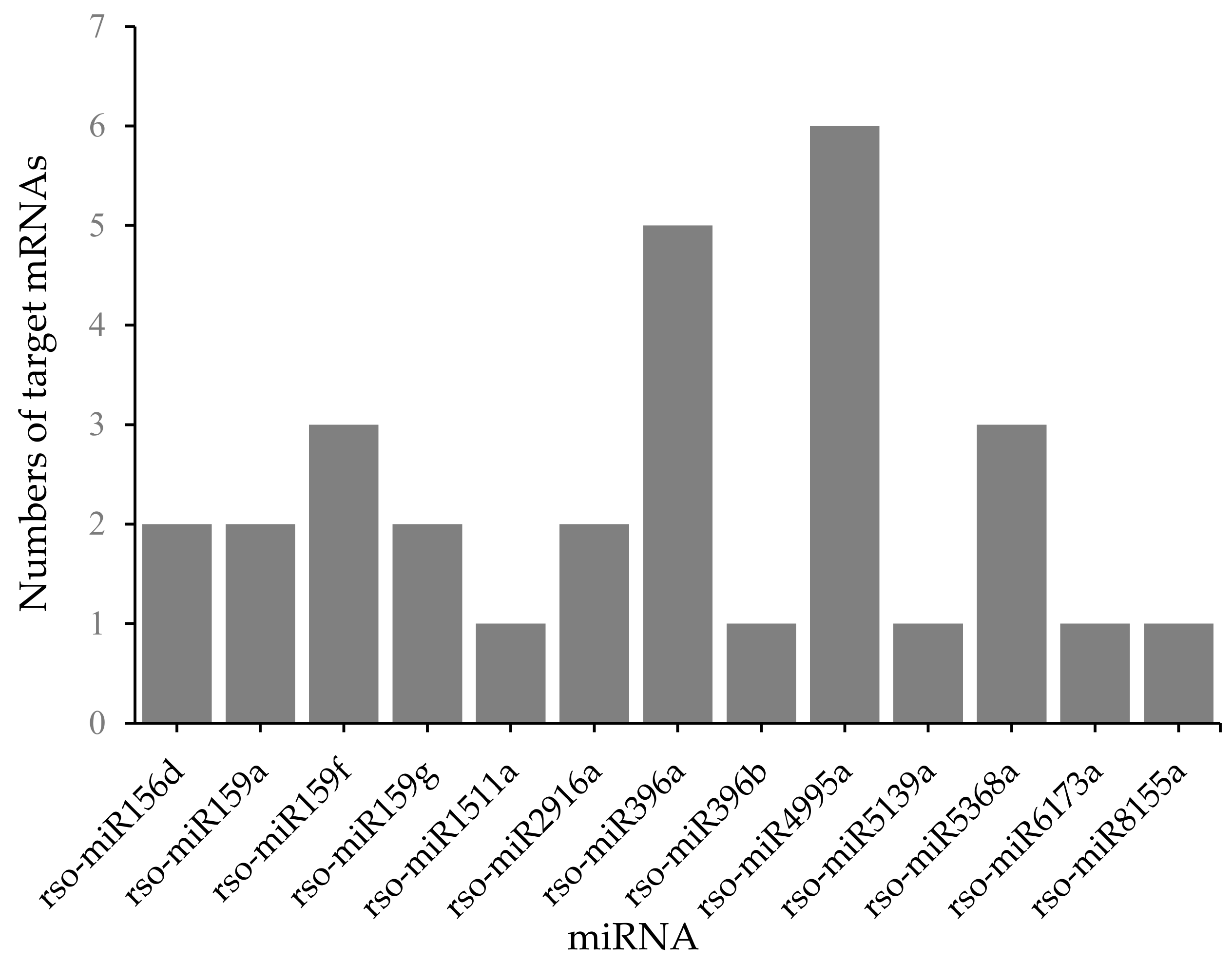
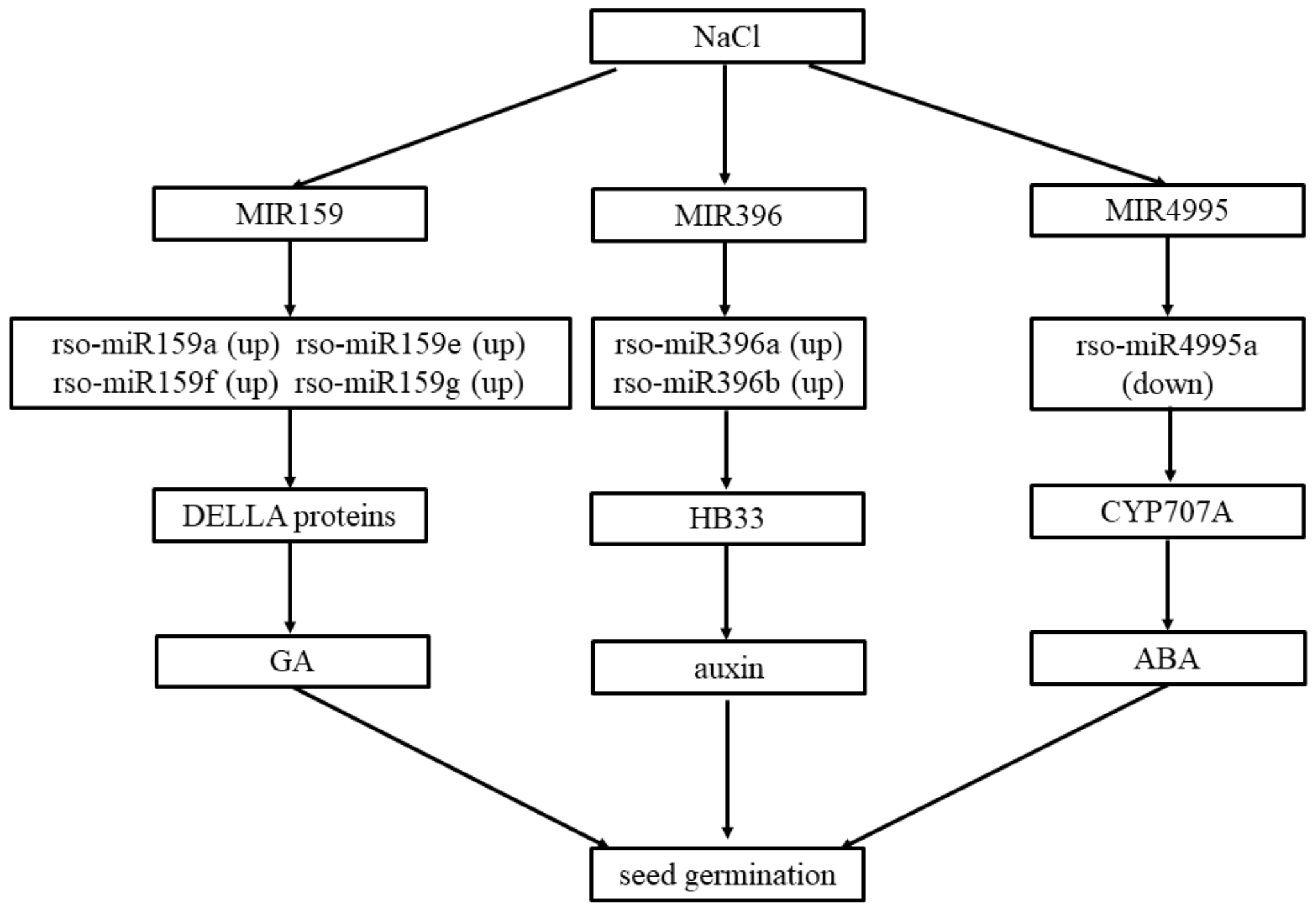
2.5. miRNA and mRNA Correlation Analysis
3. Discussion
4. Materials and Methods
4.1. Plant Material and Salt-Stress Treatment
4.2. Small-RNA Library Construction and Sequencing
4.3. Identification of Conserved and Novel miRNAs
4.4. Relative Expression between miRNA Libraries
4.5. Prediction of miRNA Target Genes
4.6. miRNA and mRNA Correlation Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Finch Savage, W.E.; Leubner Metzger, G. Seed dormancy and the control of germination. New Phytol. 2006, 171, 501–523. [Google Scholar] [CrossRef] [PubMed]
- Bai, B.; Peviani, A.; van der Horst, S.; Gamm, M.; Snel, B.; Bentsink, L.; Hanson, J. Extensive translational regulation during seed germination revealed by polysomal profiling. New Phytol. 2017, 214, 233–244. [Google Scholar] [CrossRef] [PubMed]
- Weitbrecht, K.; Muller, K.; Leubner-Metzger, G. First off the mark: Early seed germination. J. Exp. Bot. 2011, 62, 3289–3309. [Google Scholar] [CrossRef] [PubMed]
- Gu, D.; Chen, C.Y.; Zhao, M.; Zhao, L.; Duan, X.; Duan, J.; Wu, K.; Liu, X. Identification of HDA15-PIF1 as a key repression module directing the transcriptional network of seed germination in the dark. Nucleic Acids Res. 2017, 45, 7137–7150. [Google Scholar] [CrossRef]
- Cristiano, G.; Camposeo, S.; Fracchiolla, M.; Vivaldi, G.; De Lucia, B.; Cazzato, E. Salinity Differentially Affects Growth and Ecophysiology of Two Mastic Tree (Pistacia lentiscus L.) Accessions. Forests 2016, 7, 156. [Google Scholar] [CrossRef]
- Yang, X.; Rost, K.T.; Lehmkuhl, F.; Zhu, Z.; Dodson, J. The evolution of dry lands in northern China and in the Republic of Mongolia since the Last Glacial Maximum. Quat. Int. 2004, 118–119, 69–85. [Google Scholar] [CrossRef]
- Zhu, Z.D.; Wu, Z.; Liu, S.; Di, X. An Outline of Chinese Deserts; Science Press: Beijing, China, 1980; pp. 56–90. [Google Scholar]
- Yang, X.P.; Scuderi, L.; Paillou, P.; Liu, Z.T.; Li, H.W.; Ren, X.Z. Quaternary environmental changes in the drylands of China-A critical review. Quat. Sci. Rev. 2011, 30, 3219–3233. [Google Scholar] [CrossRef]
- Guo, Z.T.; Ruddiman, W.F.; Hao, Q.Z.; Wu, H.B.; Qiao, Y.S.; Zhu, R.X.; Peng, S.Z.; Wei, J.J.; Yuan, B.Y.; Liu, T.S. Onset of Asian desertification by 22 Myr ago inferred from loess deposits in China. Nature 2002, 416, 159–163. [Google Scholar] [CrossRef]
- Delectis Florae Reipublicae Popularis Sinicae Agendae Academiae Sinicae Edita. In Flora Reipublicae Popularis Sinicae; Science Press: Beijing, China, 1990; Volume 50, p. 143.
- Liu, Y.B.; Wang, G.; Liu, J.; Zhao, X.; Tan, H.J.; Li, X.R. Anatomical, morphological and metabolic acclimation in the resurrection plant Reaumuria soongorica during dehydration and rehydration. J. Arid Environ. 2007, 70, 183–194. [Google Scholar] [CrossRef]
- Liu, G.; Ren, Z. Discussion on feeding value of several main plants of Alashan desert region. Grassl. China 1990, 5, 31–34. [Google Scholar]
- Ma, M.H.; Kong, L.S. The bio-ecological characteristics of Reaumuria soongorica on the border of oasis at Hutubi, Xinjiang. Acta Phytoecol. Sin. 1998, 3, 237–244. [Google Scholar]
- Liu, J.Q.; Qiu, M.X.; Pu, J.C.; Lu, Z.M. The typical extreme xerophyte Reaumuria soongorica in the desert of China. Acta Bot. Sin. 1982, 24, 485–488. [Google Scholar]
- Li, X.L.; Chen, J.G.W. Spatial autocorrelation analysis of ISSR genetic variation of Reaumuria soongorica population in Northwest of China. J. Desert Res. 2008, 28, 468–472. [Google Scholar]
- Wu, Z.Y. Vegetation of China; Science Press: Beijing, China, 1980; pp. 583–584, 597–598. [Google Scholar]
- Zeng, Y.J.; Wang, Y.R.; Zhang, B.L.; Zhuang, G.H. Reproductive characteristics of Reaumuria soongorica populations. Acta Pratacu Lturae Sin. 2002, 11, 66–71. [Google Scholar]
- Liu, X.W.; Yang, X.Y.; Wu, H.W.; Liu, X.Y.; Zhu, J.F.; Zhang, H.X. Transcriptome analysis of differentially expressed genes in Reaumuria soongorica seeds germination under NaCl stress. J. Nanjing For. Univ. (Nat. Sci. Ed.) 2019, 43, 28–36. [Google Scholar]
- Liu, X.W.; Yang, X.Y.; Wu, H.W.; Zhi, X.R.; Zhu, J.F.; Zhang, H.X. Effects of NaCl stress on the germination of Reaumuria soongorica and evaluation of salt tolerance at germination stage. Biotechnol. Bull. 2019, 35, 27–34. [Google Scholar]
- Zeng, Y.J.; Wang, Y.R.; Zhuang, G.H.; Yang, Z.S. Seed germination responses of Reaumuria soongorica and Zygophyllum xanthoxylum to drought stress and sowing depth. Acta Ecol. Sin. 2004, 24, 1629–1634. [Google Scholar]
- Bai, J.; Xu, D.H.; Kang, H.M.; Chen, K.; Wang, G. Photoprotective function of photorespiration in Reaumuria soongorica during different levels of drought stress in natural high irradiance. Photosynthetica 2008, 46, 232–237. [Google Scholar] [CrossRef]
- Bai, J.; Gong, C.M.; Wang, G.; Kang, H.M. Antioxidative characteristics of Reaumuria soongorica under drought stress. Acta Bot. Boreali-Occident. Sin. 2010, 30, 2444–2450. [Google Scholar]
- Lv, M.T.; Yang, J.Y.; Yang, M.; Zhang, Z.R.; Ma, X. Effect of different drought stress conditions on germination of Reaumuria soongorica seeds. Chin. J. Grassl. 2010, 32, 58–63. [Google Scholar]
- Chong, P.F.; Su, S.P.; Li, Y. Comprehensive evaluation of drought resistance of Reaumuria soongorica from four geographical populations. Acta Prataculturae Sin. 2011, 20, 26–33. [Google Scholar]
- Shi, Y.; Yan, X.; Zhao, P.; Yin, H.; Zhao, X.; Xiao, H.; Li, X.; Chen, G.; Ma, X.F. Transcriptomic analysis of a tertiary relict plant, extreme xerophyte Reaumuria soongorica to identify genes related to drought adaptation. PLoS ONE 2013, 8, e63993. [Google Scholar] [CrossRef]
- Cuperus, J.T.; Fahlgren, N.; Carrington, J.C. Evolution and functional diversification of miRNA genes. Plant Cell 2011, 23, 431–442. [Google Scholar] [CrossRef] [PubMed]
- Chen, X. MicroRNA biogenesis and function in plants. FEBS Lett. 2005, 579, 5923–5931. [Google Scholar] [CrossRef]
- Mallory, A.C.; Vaucheret, H. Functions of microRNAs and related small RNAs in plants. Nat. Genet. 2006, 38, S31–S36. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, W.; Chen, J.; Liu, J.; Xia, M.; Shen, F. Identification of miRNAs and their targets in cotton inoculated with verticillium dahliae by high-throughput sequencing and degradome analysis. Int. J. Mol. Sci. 2015, 16, 14749–14768. [Google Scholar] [CrossRef]
- Zhang, B. MicroRNA: A new target for improving plant tolerance to abiotic stress. J. Exp. Bot. 2015, 66, 1749–1761. [Google Scholar] [CrossRef]
- Lv, S.; Nie, X.; Wang, L.; Du, X.; Biradar, S.S.; Jia, X.; Weining, S. Identification and characterization of microRNAs from barley (Hordeum vulgare L.) by high-throughput sequencing. Int. J. Mol. Sci. 2012, 13, 2973–2984. [Google Scholar] [CrossRef]
- Zhou, Z.S.; Zeng, H.Q.; Liu, Z.P.; Yang, Z.M. Genome-wide identification of Medicago truncatula microRNAs and their targets reveals their differential regulation by heavy metal. Plant Cell Environ. 2012, 35, 86–99. [Google Scholar] [CrossRef]
- Lertpanyasampatha, M.; Gao, L.; Kongsawadworakul, P.; Viboonjun, U.; Chrestin, H.; Liu, R.; Chen, X.; Narangajavana, J. Genome-wide analysis of microRNAs in rubber tree (Hevea brasiliensis L.) using high-throughput sequencing. Planta 2012, 236, 436–445. [Google Scholar] [CrossRef]
- Kulcheski, F.R.; de Oliveira, L.F.; Molina, L.G.; Almerao, M.P.; Rodrigues, F.A.; Marcolino, J.; Barbosa, J.F.; Stolf-Moreira, R.; Nepomuceno, A.L.; Marcelino-Guimaraes, F.C.; et al. Identification of novel soybean microRNAs involved in abiotic and biotic stresses. BMC Genom. 2011, 12, 307. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Wang, T.; Zhao, M.; Tian, Q.; Zhang, W.H. Identification of aluminum-responsive microRNAs in Medicago truncatula by genome-wide high-throughput sequencing. Planta 2012, 235, 375–386. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Zhang, Y.; Ren, Y.; Xu, J.; Zhang, Z.; Wang, Y. Genome-wide identification of cold-responsive and new microRNAs in Populus tomentosa by high-throughput sequencing. Biochem. Biophys. Res. Commun. 2012, 417, 892–896. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wan, L.; Bi, S.; Wan, X.; Li, Z.; Cao, J.; Tong, Z.; Xu, H.; He, F.; Li, X. Identification of drought-responsive micrornas from roots and leaves of alfalfa by high-throughput sequencing. Genes (Basel) 2017, 8, 119. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Zhang, Y.; Huang, N.; Liu, F.; Su, W.; Xu, L.; Ahmad, W.; Wu, Q.; Guo, J.; Que, Y. Small RNA sequencing reveals a role for sugarcane miRNAs and their targets in response to Sporisorium scitamineum infection. BMC Genom. 2017, 18, 325. [Google Scholar] [CrossRef]
- Aukerman, M.J.; Sakai, H. Regulation of flowering time and floral organ identity by a MicroRNA and its APETALA2-like target genes. Plant Cell 2003, 15, 2730–2741. [Google Scholar] [CrossRef]
- Zhou, M.; Li, D.; Li, Z.; Hu, Q.; Yang, C.; Zhu, L.; Luo, H. Constitutive expression of a miR319 gene alters plant development and enhances salt and drought tolerance in transgenic creeping bentgrass. Plant Physiol. 2013, 161, 1375–1391. [Google Scholar] [CrossRef]
- Zhao, B.; Ge, L.; Liang, R.; Li, W.; Ruan, K.; Lin, H.; Jin, Y. Members of miR169 family are induced by high salinity and transiently inhibit the NF-YA transcription factor. BMC Mol. Biol. 2009, 10, 29. [Google Scholar] [CrossRef]
- Zhou, L.; Chen, J.; Li, Z.; Li, X.; Hu, X.; Huang, Y.; Zhao, X.; Liang, C.; Wang, Y.; Sun, L.; et al. Integrated profiling of microRNAs and mRNAs: microRNAs located on Xq27.3 associate with clear cell renal cell carcinoma. PLoS ONE 2010, 5, e15224. [Google Scholar] [CrossRef]
- Zhou, L.; Liu, Y.; Liu, Z.; Kong, D.; Duan, M.; Luo, L. Genome-wide identification and analysis of drought-responsive microRNAs in Oryza sativa. J. Exp. Bot. 2010, 61, 4157–4168. [Google Scholar] [CrossRef]
- Chen, X.M. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 2004, 303, 2022–2025. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.W.; Wang, L.J.; Mao, Y.B.; Cai, W.J.; Xue, H.W.; Chen, X.Y. Control of root cap formation by MicroRNA-targeted auxin response factors in Arabidopsis. Plant Cell 2005, 17, 2204–2216. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, Z.; Deng, M.; Liu, R.; Niu, S.; Fan, G. Identification and functional analysis of microRNAs and their targets in Platanus acerifolia under lead (Pb) stress. Int. J. Mol. Sci. 2015, 16, 7098–7111. [Google Scholar] [CrossRef] [PubMed]
- Saminathan, T.; Bodunrin, A.; Singh, N.V.; Devarajan, R.; Nimmakayala, P.; Jeff, M.; Aradhya, M.; Reddy, U.K. Genome-wide identification of microRNAs in pomegranate (Punica granatum L.) by high-throughput sequencing. BMC Plant Biol. 2016, 16, 122. [Google Scholar] [CrossRef]
- Ramya, R.; Hervé, V.; Jerry, T.; Bartel, D.P. A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Evelopment 2006, 20, 3407–3425. [Google Scholar]
- Zhang, C.; Zhang, B.; Ma, R.; Yu, M.; Guo, S.; Guo, L.; Korir, N.K. Identification of known and novel microRNAs and their targets in peach (Prunus persica) fruit by high-throughput sequencing. PLoS ONE 2016, 11, e0159253. [Google Scholar] [CrossRef]
- Gao, J.; Ge, W.; Zhang, Y.; Cheng, Z.C.; Li, L.; Hou, D.; Hou, C.L. Identification and characterization of microRNAs at different flowering developmental stages in moso bamboo (Phyllostachys edulis) by high-throughput sequencing. Mol. Genet. Genom. 2015, 290, 2335–2353. [Google Scholar] [CrossRef]
- Chi, X.Y.; Yang, Q.L.; Chen, X.P.; Wang, J.Y.; Pan, L.J.; Chen, M.N.; Yang, Z.; He, Y.N.; Liang, X.Q.; Yu, S.L. Identification and characterization of microRNAs from peanut (Arachis hypogaea L.) by high-throughput sequencing. PLoS ONE 2011, 6, e27530. [Google Scholar] [CrossRef]
- Baksa, I.; Nagy, T.; Barta, E.; Havelda, Z.; Várallyay, É.; Silhavy, D.; Burgyán, J.; Szittya, G. Identification of Nicotiana benthamiana microRNAs and their targets using high throughput sequencing and degradome analysis. BMC Genom. 2015, 16, 1025. [Google Scholar] [CrossRef]
- Zhu, J.; Yang, X.; Liu, Z.; Zhang, H. Identification and target prediction of MicroRNAs in Ulmus pumila L. seedling roots under salt stress by high-throughput sequencing. Forests 2016, 7, 318. [Google Scholar] [CrossRef]
- Ma, L.; Huang, Y.; Zhu, W.; Zhou, S.; Zhou, J.; Zeng, F.; Liu, X.; Zhang, Y.; Yu, J. An integrated analysis of miRNA and mRNA expressions in non-small cell lung cancers. PLoS ONE 2011, 6, e26502. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.P.; Montgomery, T.A.; Fahlgren, N.; Kasschau, K.D.; Nonogaki, H.; Carrington, J.C. Repression of AUXIN RESPONSE FACTOR10 by microRNA160 is critical for seed germination and post-germination stages. Plant J. 2007, 52, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.; Wilen, R.W.; Robertson, A.J.; Gusta, L.V. Isolation, chromosomal localization, and differential expression of mitochondrial manganese superoxide dismutase and chloroplastic copper/zinc superoxide dismutase genes in wheat. Plant Physiol. 1999, 120, 513–520. [Google Scholar] [CrossRef] [PubMed]
- Jagadeeswaran, G.; Saini, A.; Sunkar, R. Biotic and abiotic stress down-regulate miR398 expression in Arabidopsis. Planta 2009, 229, 1009–1014. [Google Scholar] [CrossRef]
- Zhu, J.; Li, W.; Yang, W.; Qi, L.; Han, S. Identification of microRNAs in Caragana intermedia by high-throughput sequencing and expression analysis of 12 microRNAs and their targets under salt stress. Plant Cell Rep. 2013, 32, 1339–1349. [Google Scholar] [CrossRef]
- Zhu, G.; Liu, Y.; Ye, N.; Liu, R.; Zhang, J. Involvement of the abscisic acid catabolic gene CYP707A2 in the glucose-induced delay in seed germination and post-germination growth of Arabidopsis. Physiol. Plant. 2011, 143, 375–384. [Google Scholar] [CrossRef]
- Okamoto, M.; Kuwahara, A.; Seo, M.; Kushiro, T.; Asami, T.; Hirai, N.; Kamiya, Y.; Koshiba, T.; Nambara, E. CYP707A1 and CYP707A2, which encode abscisic acid 8′-hydroxylases, are indispensable for proper control of seed dormancy and germination in Arabidopsis. Plant Physiol. 2006, 141, 97–107. [Google Scholar] [CrossRef]
- Matakiadis, T.; Alboresi, A.; Jikumaru, Y.; Tatematsu, K.; Pichon, O.; Renou, J.P.; Kamiya, Y.; Nambara, E.; Truong, H.N. The Arabidopsis abscisic acid catabolic gene CYP707A2 plays a key role in nitrate control of seed dormancy. Plant Physiol. 2009, 149, 949–960. [Google Scholar] [CrossRef]
- Zhu, G.; Ye, N.; Zhang, J. Glucose-induced delay of seed germination in rice is mediated by the suppression of ABA catabolism rather than an enhancement of ABA biosynthesis. Plant Cell Physiol. 2009, 50, 644–651. [Google Scholar] [CrossRef]
- Patrick, A.; Alan, H.; Baulcombe, D.C.; Harberd, N.P. Modulation of floral development by a gibberellin-regulated microRNA. Development 2004, 131, 3357–3365. [Google Scholar]
- Millar, A.A.; Gubler, F. The Arabidopsis GAMYB-Like genes, MYB33 and MYB65, are microRNA-regulated genes that redundantly facilitate anther development. Plant Cell 2005, 17, 705–721. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhang, T.; Wen, Z.; Xiao, H.; Yang, Z.; Chen, G.; Zhao, X. The chromosome number, karyotype and genome size of the desert plant diploid Reaumuria soongorica (Pall.) Maxim. Plant Cell Rep. 2011, 30, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.Q.; Zhu, S.Q.; Yu, R.P.; Li, L.Q.; Shan, G.Z.; You, W.R.; Zeng, X.X.; Zhang, C.W.; Zhang, L.J.; Song, R.H. China Saline Soil; Science Press: Beijing, China, 1994; Volume 250. [Google Scholar]
- Zhang, D.F.; Wang, S.J. Mechanism of freeze-thaw action in land salinization process—as an sample in west Jilin province. Bull. Soil Water Conserv. 2000, 20, 15–17. [Google Scholar]
- Xiao, H.Y.; Wen, G.C.; Liu, Q.M.; Li, G.T.; Zhao, X.R. Effects of winter and spring plant residue mulching on saline soil salt in Hetao Inner Mongolia. Agric. Res. Arid Areas 2018, 36, 23–26. [Google Scholar]
- Xu, L.; Wang, Y.; Zhai, L.L.; Xu, Y.Y.; Wang, L.J.; Zhu, X.W.; Gong, Y.Q.; Yu, R.G.; Cecilia, L.; Liu, L.W. Genome-wide identification and characterization of cadmium-responsive microRNAs and their target genes in radish (Raphanus sativus L.) roots. J. Exp. Bot. 2013, 64, 4271–4287. [Google Scholar] [CrossRef]
- Khaldun, A.B.; Huang, W.; Liao, S.; Lv, H.; Wang, Y. Identification of microRNAs and target genes in the fruit and shoot tip of Lycium chinense: A traditional Chinese medicinal plant. PLoS ONE 2015, 10, e0116334. [Google Scholar] [CrossRef]
- Friedländer, M.R.; Mackowiak, S.D.; Na, L.; Wei, C.; Nikolaus, R. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2012, 40, 37–52. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef]
- Wen, M.; Shen, Y.; Shi, S.; Tang, T. miREvo: An integrative microRNA evolutionary analysis platform for next-generation sequencing experiments. BMC Bioinform. 2012, 13, 140. [Google Scholar] [CrossRef]
- Meyers, B.C.; Axtell, M.J.; Bartel, B.; Bartel, D.P.; Baulcombe, D.; Bowman, J.L.; Cao, X.; Carrington, J.C.; Chen, X.; Green, P.J.; et al. Criteria for annotation of plant MicroRNAs. Plant Cell 2008, 20, 3186–3190. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Leti, F.; Malenica, I.; Doshi, M.; Courtright, A.; Van Keuren-Jensen, K.; Legendre, C.; Still, C.D.; Gerhard, G.S.; DiStefano, J.K. High-throughput sequencing reveals altered expression of hepatic microRNAs in nonalcoholic fatty liver disease–related fibrosis. Transl. Res. 2015, 166, 304–314. [Google Scholar] [CrossRef] [PubMed]
- Unger, L.; Jagannathan, V.; Pacholewska, A.; Leeb, T.; Gerber, V. Differences in miRNA differential expression in whole blood between horses with sarcoid regression and progression. J. Vet. Intern. Med. 2019, 33, 241–250. [Google Scholar] [CrossRef] [PubMed]
- Hamzeiy, H.; Allmer, J.; Yousef, M. Computational methods for microRNA target prediction. Methods Mol. Biol. 2014, 1107, 207–221. [Google Scholar] [PubMed]
- Wu, H.J.; Ma, Y.K.; Chen, T.; Wang, M.; Wang, X.J. PsRobot: A web-based plant small RNA meta-analysis toolbox. Nucleic Acids Res. 2012, 40, W22–W28. [Google Scholar] [CrossRef] [PubMed]
- Allantaz, F.; Cheng, D.T.; Bergauer, T.; Ravindran, P.; Rossier, M.F.; Ebeling, M.; Badi, L.; Reis, B.; Bitter, H.; D’Asaro, M.; et al. Expression profiling of human immune cell subsets identifies miRNA-mRNA regulatory relationships correlated with cell type specific expression. PLoS ONE 2012, 7, e29979. [Google Scholar] [CrossRef]

| Category | Raw Reads | N% > 10% | Low Quality Reads | 5′ Adapter Contamine | 3′ Adapter Null | Poly (A/T/G/C) | Unique Reads | Clean Reads |
|---|---|---|---|---|---|---|---|---|
| CS-1 | 13,869,156 (100.00%) | 564 (0.00%) | 19,225 (0.14%) | 33,490 (0.24%) | 224,816 (1.62%) | 12,005 (0.09%) | 1,315,057 | 13,579,056 (97.91%) |
| CS-2 | 16,819,324 (100.00%) | 639 (0.00%) | 21,867 (0.13%) | 15,406 (0.09%) | 964,063 (5.73%) | 7615 (0.05%) | 1,376,042 | 15,809,734 (94.00%) |
| CS-3 | 14,212,954 (100.00%) | 543 (0.00%) | 16,330 (0.11%) | 14,962 (0.11%) | 398,880 (2.81%) | 7654 (0.05%) | 1,156,054 | 13,774,585 (96.92%) |
| LS-1 | 14,552,453 (100.00%) | 527 (0.00%) | 18,159 (0.12%) | 13,581 (0.09%) | 702,302 (4.83%) | 4291 (0.03%) | 993,154 | 13,813,593 (94.92%) |
| LS-2 | 12,328,793 (100.00%) | 451 (0.00%) | 19,864 (0.16%) | 13,884 (0.11%) | 179,427 (1.46%) | 7466 (0.06%) | 1,442,730 | 12,107,701 (98.21%) |
| LS-3 | 13,908,660 (100.00%) | 504 (0.00%) | 15,636 (0.11%) | 11,487 (0.08%) | 290,553 (2.09%) | 4276 (0.03%) | 818,193 | 13,586,204 (97.68%) |
| MS-1 | 12,887,720 (100.00%) | 535 (0.00%) | 19,878 (0.15%) | 8991 (0.07%) | 270,670 (2.10%) | 3690 (0.03%) | 1,014,692 | 12,583,956 (97.64%) |
| MS-2 | 17,338,149 (100.00%) | 680 (0.00%) | 23,377 (0.13%) | 17,416 (0.10%) | 490,306 (2.83%) | 12,336 (0.07%) | 2,026,130 | 16,794,034 (96.86%) |
| MS-3 | 14,594,879 (100.00%) | 532 (0.00%) | 15,790 (0.11%) | 13,876 (0.10%) | 743,198 (5.09%) | 5511 (0.04%) | 1,145,458 | 13,815,972 (94.66%) |
| Read Types | Total | Sequences Mapped to Genome | Known miRNA | Novel miRNA | Ribosomal RNA | Transfer RNA | Small Nuclear RNA | Small Nucleolar RNA | Trans-Acting Small Interfering RNAs | Unannotated |
|---|---|---|---|---|---|---|---|---|---|---|
| CS-1 | 9,127,130 | 8,474,172 (92.85%) | 12,039 (0.14%) | 1244 (0.01%) | 1,245,902 (14.70%) | 10 (0.00%) | 2883 (0.03%) | 6680 (0.08%) | 641 (0.01%) | 7,204,773 (85.02%) |
| CS-2 | 8,007,781 | 7,381,145 (92.17%) | 61,754 (0.84%) | 4490 (0.06%) | 1,644,956 (22.29%) | 15 (0.00%) | 3419 (0.05%) | 5609 (0.08%) | 3583 (0.05%) | 5,657,319 (76.65%) |
| CS-3 | 9,375,007 | 8,975,019 (95.73%) | 20,397 (0.23%) | 1423 (0.02%) | 1,497,323 (16.68%) | 5 (0.00%) | 2433 (0.03%) | 3034 (0.03%) | 1038 (0.01%) | 7,449,366 (83.00%) |
| LS-1 | 7,270,911 | 6,886,984 (94.72%) | 16,112 (0.23%) | 1515 (0.02%) | 1,366,452 (19.84%) | 9 (0.00%) | 3193 (0.05%) | 5135 (0.07%) | 1646 (0.02%) | 5,492,922 (79.76%) |
| LS-2 | 8,520,845 | 7,737,807 (90.81%) | 32,024 (0.41%) | 3086 (0.04%) | 1,356,215 (17.53%) | 9 (0.00%) | 2832 (0.04%) | 7026 (0.09%) | 2159 (0.03%) | 6,334,456 (81.86%) |
| LS-3 | 9,018,168 | 8,779,523 (97.35%) | 7119 (0.08%) | 604 (0.01%) | 1,482,050 (16.88%) | 6 (0.00%) | 2286 (0.03%) | 3677 (0.04%) | 419 (0.00%) | 7,283,362 (82.96%) |
| MS-1 | 8,981,640 | 8,581,120 (95.54%) | 5456 (0.06%) | 138 (0.00%) | 1,452,970 (16.93%) | 7 (0.00%) | 2749 (0.03%) | 1702 (0.02%) | 121 (0.00%) | 7,117,977 (82.95%) |
| MS-2 | 10,925,787 | 9,849,558 (90.15%) | 112,529 (1.14%) | 4902 (0.05%) | 1,888,880 (19.18%) | 10 (0.00%) | 4224 (0.04%) | 5038 (0.05%) | 6472 (0.07%) | 7,827,503 (79.47%) |
| MS-3 | 6,096,367 | 5,522,031 (90.58%) | 51,689 (0.94%) | 2498 (0.05%) | 927,311 (16.79%) | 9 (0.00%) | 1767 (0.03%) | 4458 (0.08%) | 3196 (0.06%) | 4,531,103 (82.06%) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, H.; Liu, X.; Yang, X.; Wu, H.; Zhu, J.; Zhang, H. miRNA–mRNA Integrated Analysis Reveals Roles for miRNAs in a Typical Halophyte, Reaumuria soongorica, during Seed Germination under Salt Stress. Plants 2020, 9, 351. https://doi.org/10.3390/plants9030351
Zhang H, Liu X, Yang X, Wu H, Zhu J, Zhang H. miRNA–mRNA Integrated Analysis Reveals Roles for miRNAs in a Typical Halophyte, Reaumuria soongorica, during Seed Germination under Salt Stress. Plants. 2020; 9(3):351. https://doi.org/10.3390/plants9030351
Chicago/Turabian StyleZhang, Huilong, Xiaowei Liu, Xiuyan Yang, Haiwen Wu, Jianfeng Zhu, and Huaxin Zhang. 2020. "miRNA–mRNA Integrated Analysis Reveals Roles for miRNAs in a Typical Halophyte, Reaumuria soongorica, during Seed Germination under Salt Stress" Plants 9, no. 3: 351. https://doi.org/10.3390/plants9030351
APA StyleZhang, H., Liu, X., Yang, X., Wu, H., Zhu, J., & Zhang, H. (2020). miRNA–mRNA Integrated Analysis Reveals Roles for miRNAs in a Typical Halophyte, Reaumuria soongorica, during Seed Germination under Salt Stress. Plants, 9(3), 351. https://doi.org/10.3390/plants9030351






