One-Step Reverse-Transcription Digital PCR for Reliable Quantification of Different Pepino Mosaic Virus Genotypes
Abstract
1. Introduction
2. Results and Discussion
2.1. Optimisation of the Reverse-Transcription Digital Polymerase Chain Reaction (RT-dPCR) Assays on the QX100/200 Platform
2.2. Optimisation of the RT-dPCR Assays on the Naica Platform
2.3. Validation of the RT-dPCR Assays
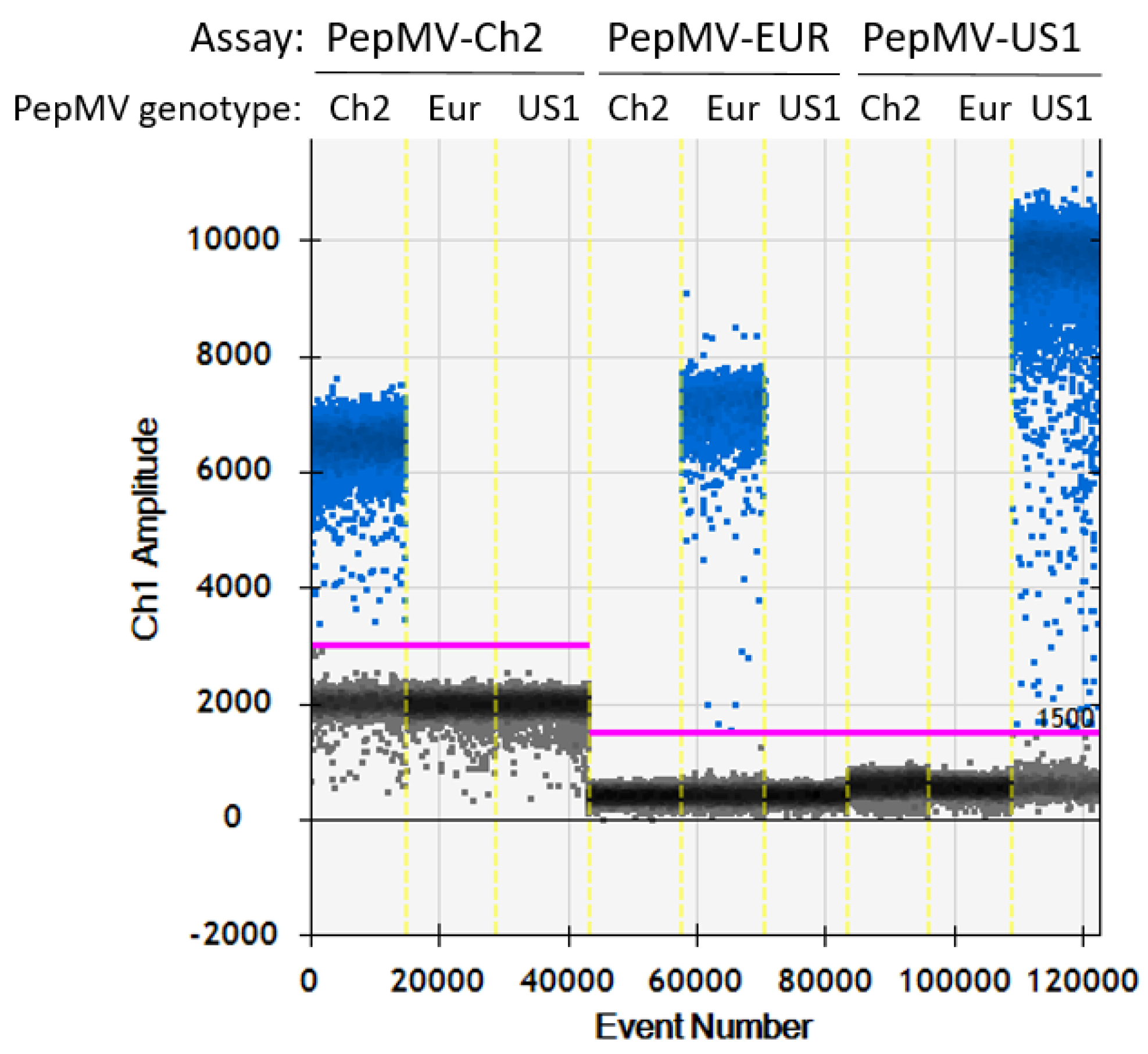
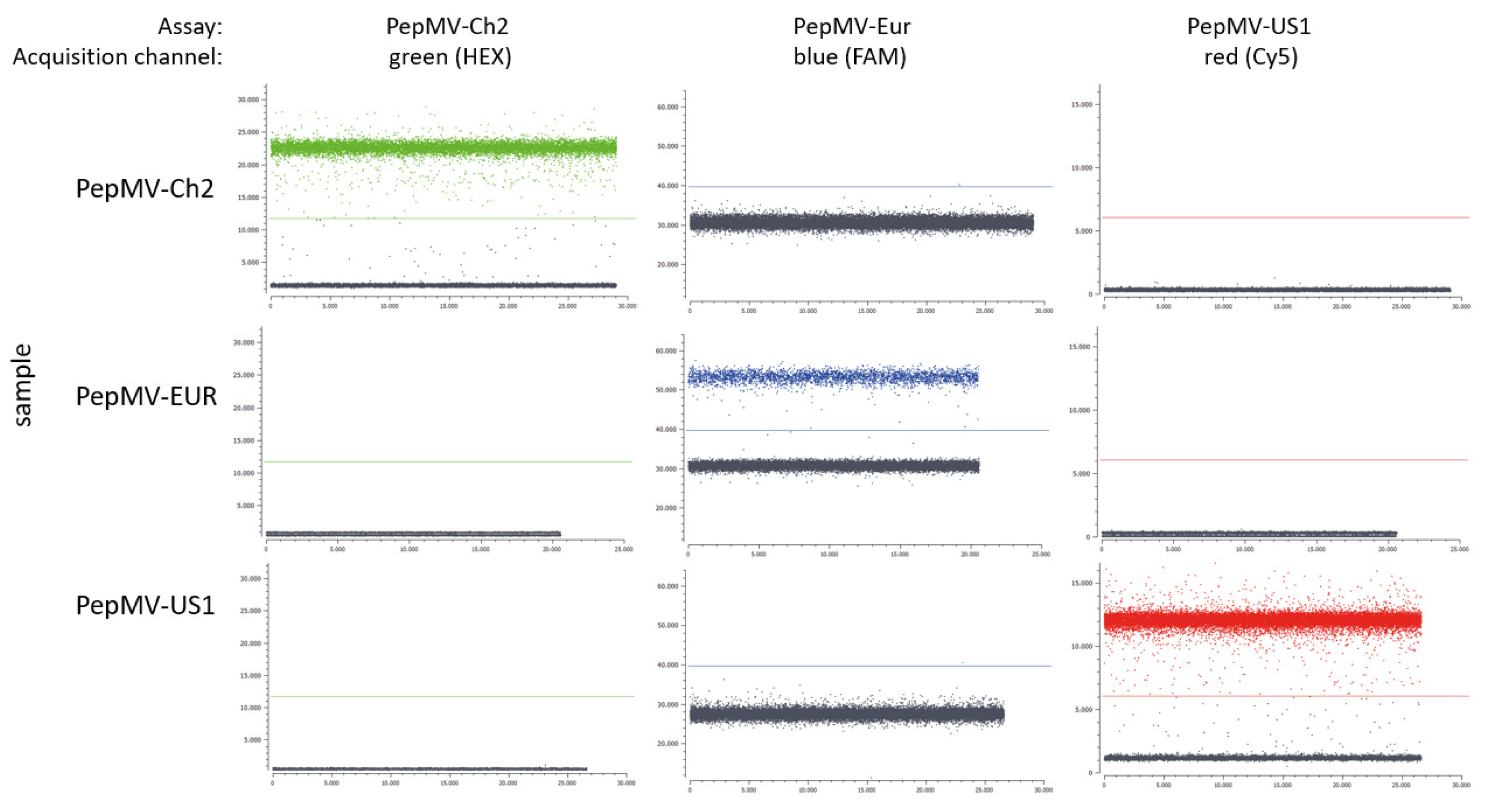
2.3.1. Specificity, Repeatability and Reproducibility of the RT-dPCR Assays
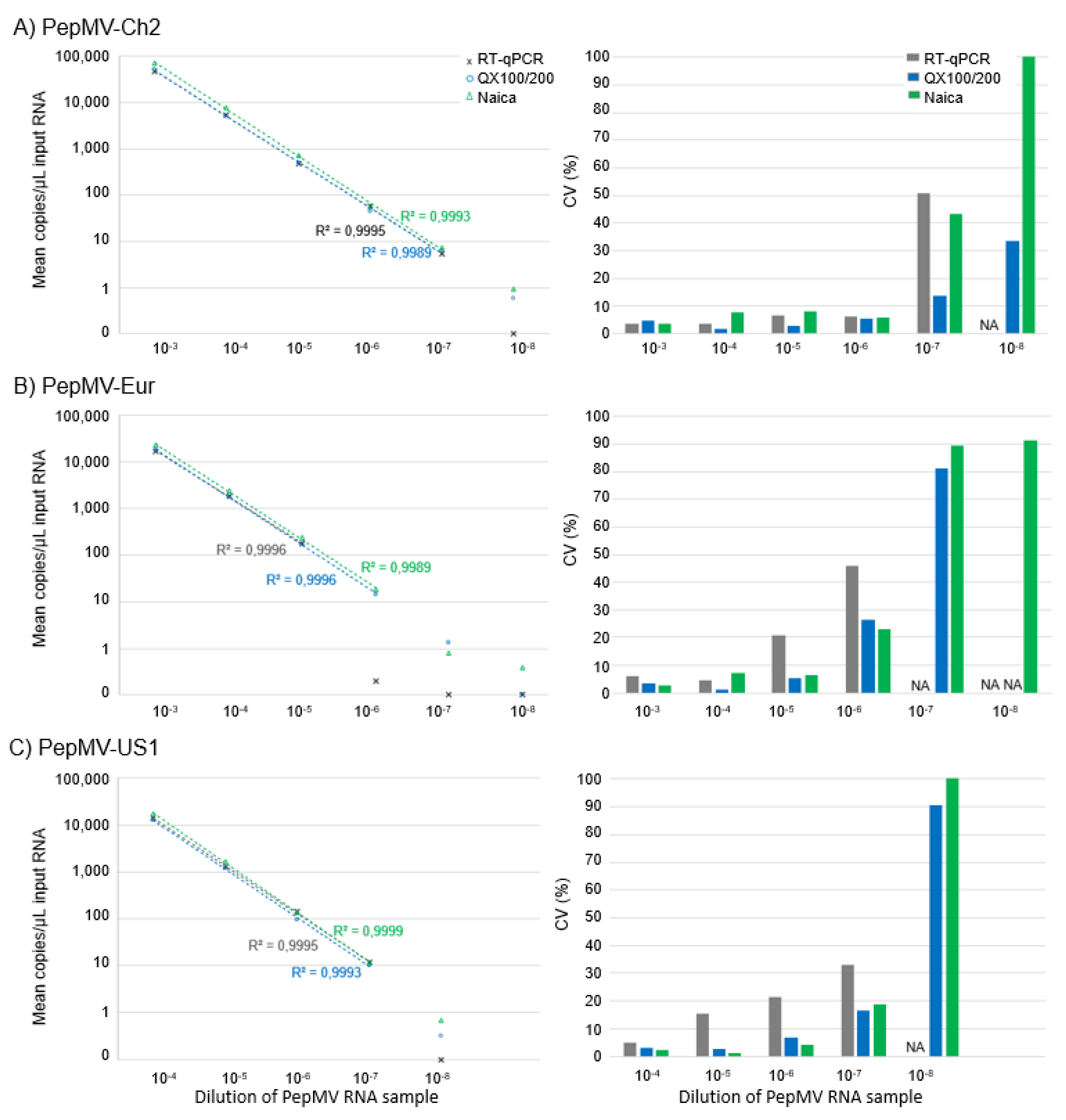
2.3.2. Sensitivity and Linear Range for Different PepMV Genotypes by RT-dPCR Compared to Reverse-Transcription Real-Time Quantitative Polymerase Chain Reaction (RT-qPCR)
3. Materials and Methods
3.1. Samples and Sample Preparation
3.2. Assays Used
3.3. RT-qPCR
3.4. Optimisation of RT-dPCR: QX100/200
3.5. Optimisation of RT-dPCR: Naica
3.6. Validation of RT-dPCR Assays for Quantification of Different PepMV Genotypes
3.7. Data Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jones, R.A.C.; Koenig, R.; Lesemann, D.E. Pepino mosaic virus, a new potexvirus from pepino (Solanum muricatum). Ann. Appl. Biol. 1980, 94, 61–68. [Google Scholar] [CrossRef]
- Aguilar, J.; Hernández-Gallardo, M.; Cenis, J.; Lacasa, A.; Aranda, M.A. Complete sequence of the pepino mosaic virus RNA genome. Arch. Virol. 2002, 147, 2009–2015. [Google Scholar] [CrossRef] [PubMed]
- van der Vlugt, R.A.A.; Stijer, C.C.M.M.; Verhoeven, J.; Lesemann, D.E. First report of pepino mosaic virus on tomato. Plant Dis. 2000, 84, 103. [Google Scholar] [CrossRef] [PubMed]
- Minicka, J.; Hasiów-Jaroszewska, B.; Borodynko-Filas, N.; Pospieszny, H.; Hanssen, I.M. Rapid evolutionary dynamics of the pepino mosaic virus—Status and future perspectives. J. Plant Prot. Res. 2016, 56, 337–345. [Google Scholar] [CrossRef]
- Roggero, P.; Masenga, V.; Lenzi, R.; Coghe, F.; Ena, S.; Winter, S. First report of pepino mosaic virus in tomato in Italy. Plant Pathol. 2001, 50, 798–800. [Google Scholar] [CrossRef]
- Spence, N.J.; Basham, J.; Mumford, R.A.; Hayman, G.; Edmondson, R.; Jones, D.R. Effect of pepino mosaic virus on the yield and quality of glasshouse-grown tomatoes in the UK. Plant Pathol. 2006, 55, 595–606. [Google Scholar] [CrossRef]
- Hanssen, I.M.; Paeleman, A.; Vandewoestijne, E.; Van Bergen, L.; Bragard, C.; Lievens, B.; Vanachter, A.C.R.C.; Thomma, B.P.H.J. Pepino mosaic virus isolates and differential symptomatology in tomato. Plant Pathol. 2009, 58, 450–460. [Google Scholar] [CrossRef]
- Hasiów-Jaroszewska, B.; Paeleman, A.; Ortega-Parra, N.; Borodynko, N.; Minicka, J.; Czerwoniec, A.; Thomma, B.P.; Hanssen, I.M. Ratio of mutated versus wild-type coat protein sequences in pepino mosaic virus determines the nature and severity of yellowing symptoms on tomato plants. Mol. Plant Pathol. 2013, 14, 923–933. [Google Scholar] [CrossRef]
- Domingo, E.; Sheldon, J.; Perales, C. Viral quasispecies evolution. Microbiol. Mol. Biol. Rev. 2012, 76, 159–216. [Google Scholar] [CrossRef]
- Ling, K.S.; Wechter, P.; Jordan, R. Development of a one-step immunocapture real-time TaqMan RT-PCR assay for the broad spectrum detection of pepino mosaic virus. J. Virol. Methods 2007, 144, 65–72. [Google Scholar] [CrossRef]
- Maroon-Lango, C.J.; Guaragna, M.A.; Jordan, R.L.; Hammond, J.; Bandla, M.; Marquardt, S.K. Two unique isolates of pepino mosaic virus from a limited source of pooled tomato tissue are distinct from a third (European-like) US isolate. Arch. Virol. 2005, 150, 1187–1201. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Pérez, M.G.; Pagán, I.; Aragón-Caballero, L.; Cáceres, F.; Fraile, A.; García-Arenal, F. Ecological and genetic determinants of pepino mosaic virus emergence. J. Virol. 2014, 88, 3359–3368. [Google Scholar] [CrossRef] [PubMed]
- French, C.J.; Dubeau, C.; Bunckle, A.; Ferguson, G.; Haesevoets, R.; Bouthillier, M.; Bernardy, M.G. Overview of pepino mosaic virus research. Can. J. Plant Pathol. 2008, 30, 373–374. [Google Scholar]
- Ling, K.S.; Li, R.G.; Bledsoe, M. Pepino mosaic virus genotype shift in North America and development of a loop-mediated isothermal amplification for rapid genotype identification. Virol. J. 2013, 10, 117. [Google Scholar] [CrossRef] [PubMed]
- Hanssen, I.M.; Paeleman, A.; Wittemans, L.; Goen, K.; Lievens, B.; Bragard, C.; Vanachter, A.C.R.C.; Thomma, B.P.H.J. Genetic characterization of pepino mosaic virus isolates from Belgian greenhouse tomatoes reveals genetic recombination. Eur. J. Plant Pathol. 2008, 121, 131–146. [Google Scholar] [CrossRef]
- Ling, K.S. Molecular characterization of two pepino mosaic virus variants from imported tomato seed reveals high levels of sequence identity between Chilean and US isolates. Virus Genes 2007, 34, 1–8. [Google Scholar] [CrossRef]
- Gómez, P.; Sempere, R.N.; Aranda, M.A. Pepino mosaic virus and tomato torrado virus: Emerging viruses affecting tomato crops in the Mediterranean basin. Adv. Virus Res. 2012, 84, 505–532. [Google Scholar] [CrossRef]
- Gómez, P.; Sempere, R.N.; Elena, S.F.; Aranda, M.A. Mixed infections of pepino mosaic virus strains modulate the evolutionary dynamics of this emergent virus. J. Virol. 2009, 83, 12378–12387. [Google Scholar] [CrossRef]
- Hanssen, I.M.; Mumford, R.; Blystad, D.R.; Cortez, I.; Hasiów-Jaroszewska, B.; Hristova, D.; Pagán, I.; Pereira, A.M.; Peters, J.; Pospieszny, H.; et al. Seed transmission of pepino mosaic virus in tomato. Eur. J. Plant Pathol. 2010, 126, 145–152. [Google Scholar] [CrossRef]
- Mehle, N.; Gutierrez-Aguirre, I.; Prezelj, N.; Delić, D.; Vidic, U.; Ravnikar, M. Survival and transmission of potato virus Y, pepino mosaic virus, and potato spindle tuber viroid in water. Appl. Environ. Microb. 2014, 80, 1455–1462. [Google Scholar] [CrossRef]
- Ortega-Parra, N.; Hasiów-Jaroszewska, B.; Borodynko, N.; Paele-man, A.; Hanssen, I.M. Single nucleotide polymorphisms in the coat protein of PepMV are responsible for yellowing pathotypes in tomato crops. In Proceedings of the 13th International Plant Virus Epidemiology Sympo-Sium, Avignon, France, 6–10 June 2016. [Google Scholar]
- Gutiérrez-Aguirre, I.; Mehle, N.; Delić, D.; Gruden, K.; Mumford, R.; Ravnikar, M. Real-time quantitative PCR based sensitive detection and strain discrimination of pepino mosaic virus. J. Virol. Methods 2009, 162, 46–55. [Google Scholar] [CrossRef] [PubMed]
- Vogelstein, B.; Kinzler, K.W. Digital PCR. Proc. Natl. Acad. Sci. USA 1999, 96, 9236–9241. [Google Scholar] [CrossRef] [PubMed]
- Rački, N.; Dreo, T.; Gutierrez-Aguirre, I.; Blejec, A.; Ravnikar, M. Reverse transcriptase droplet digital PCR shows high resilience to PCR inhibitors from plant, soil and water samples. Plant Methods 2014, 10, 42. [Google Scholar] [CrossRef] [PubMed]
- Bogožalec Košir, A.; Demšar, T.; Štebih, D.; Žel, J.; Milavec, M. Digital PCR as an effective tool for GMO quantification in complex matrices. Food Chem. 2019, 294, 73–78. [Google Scholar] [CrossRef]
- Whale, A.S.; Cowen, S.; Foy, C.A.; Huggett, J.F. Methods for applying accurate digital PCR analysis on low copy DNA samples. PLoS ONE 2013, 8, e58177. [Google Scholar] [CrossRef]
- Baker, M. Digital PCR hits its stride. Nat. Methods 2012, 9, 541–544. [Google Scholar] [CrossRef]
- Vallone, P.M.; Butler, J.M. AutoDimer: A screening tool for primer-dimer and hairpin structures. Biotechniques 2004, 37, 226–231. [Google Scholar] [CrossRef]
- Lievens, A.; Jacchia, S.; Kagkli, D.; Savini, C.; Querci, M. Measuring digital PCR quality: Performance parameters and their optimization. PLoS ONE 2016, 11, e0153317. [Google Scholar] [CrossRef]
- Mehle, N.; Dobnik, D.; Ravnikar, M.; Pompe Novak, M. Validated reverse transcription droplet digital PCR serves as a higher order method for absolute quantification of potato virus Y strains. Anal. Bioanal. Chem. 2018, 410, 3815–3825. [Google Scholar] [CrossRef]
- Pinheiro, L.B.; Coleman, V.A.; Hindson, C.M.; Herrmann, J.; Hindson, B.J.; Bhat, S.; Emslie, K.R. Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal. Chem. 2012, 84, 1003–1011. [Google Scholar] [CrossRef]
- Dagata, J.A.; Farkas, N.; Kramer, J.A. Method for Measuring the Volume of Nominally 100 μm Diameter Spherical Water-In-Oil Emulsion Droplets; NIST Special Publication 260-184; National Institute of Standards and Technology (NIST): Gaithersburg, MD, USA, 2016; 23p. [Google Scholar] [CrossRef]
- Bogožalec Košir, A.; Divieto, C.; Pavšič, J.; Pavarelli, S.; Dobnik, D.; Dreo, T.; Bellotti, R.; Sassi, M.P.; Žel, J. Droplet volume variability as a critical factor for accuracy of absolute quantification using droplet digital PCR. Anal. Bioanal. Chem. 2017, 409, 6689–6697. [Google Scholar] [CrossRef] [PubMed]
- Emslie, K.R.; McLaughlin, J.L.; Griffiths, K.; Forbes-Smith, M.; Pinheiro, L.B.; Burke, D.G. Droplet volume variability and impact on digital PCR copy number concentration measurements. Anal. Chem. 2019, 91, 4124–4131. [Google Scholar] [CrossRef] [PubMed]
- Madic, J.; Zocevic, A.; Senlis, V.; Fradet, E.; Andre, B.; Muller, S.; Dangla, R.; Droniou, M.E. Three-color crystal digital PCR. Biomol. Detect. Quantif. 2016, 10, 34–46. [Google Scholar] [CrossRef]
- European and Mediterranean Plant Protection Organisation. Specific requirements for laboratories preparing accreditation for a plant pest diagnostic activity PM7/98 (2). EPPO Bull. 2014, 44, 117–147. [Google Scholar] [CrossRef]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A. Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays. J. Mol. Endocrinol. 2000, 25, 169–193. [Google Scholar] [CrossRef]
| Genotype | Simplex | Duplex | Simplex vs. Duplex | |||
|---|---|---|---|---|---|---|
| Input RNA (mean; copies/µL) | CV (%) | Input RNA (mean; copies/µL) | CV (%) | Input RNA (Δ; copies/µL) | Δ Duplex (%Simplex) | |
| Mix a | 20,573 | 1.1 | 22,144 | 3.8 | 1571 | 7.6 |
| Ch2 | 7021 | 1.4 | 7292 | 7.9 | 271 | 3.9 |
| Eur | 2398 | 3.1 | 2253 | 7.1 | 145 | 6.0 |
| US1 | 16,717 | 1.6 | 17,027 | 2.3 | 310 | 1.9 |
| Sample | Runs (n) | Replicates (n) | QX100/200 | Naica | ||
|---|---|---|---|---|---|---|
| Input RNA (mean; copies/µL) | CV (%) | Input RNA (mean; copies/µL) | CV (%) | |||
| PepMV-Ch2-M | 3 | 8 | 5483 | 7.9 | 7552 | 5.6 |
| PepMV-Eur-M | 3 | 8 | 1616 | 8.2 | 2165 | 6.1 |
| PepMV-US1-M | 3 | 8 | 12,905 | 4.1 | 16,896 | 3.0 |
| PepMV-Ch2-L | 3 | 8 | 51 | 14.3 | 71 | 20.0 |
| PepMV-Eur-L | 3 | 7–8 a | 14 | 21.2 | 21 | 21.8 |
| PepMV-US1-L | 3 | 6–8 b | 110 | 13.9 | 152 | 15.1 |
| PepMV-Ch2-zero | 3 | 8 | 0 | NA | 0 c | NA |
| PepMV-Eur-zero | 3 | 5 | 0 | NA | 0 c | NA |
| PepMV-US1-zero | 3 | 5 | 0 | NA | 0 | NA |
| Sample | RNA Dilution | RT-qPCR Cq | Input RNA (copies/µL) | ||
|---|---|---|---|---|---|
| RT-qPCR a | QX100/200 b | Naica b | |||
| PepMV-Ch2 | 10−3 | 21.1 ± 0.1 | 46,737 ± 1648 | 51,907 ± 2337 e | 71,818 ± 2502 |
| 10−4 | 24.4 ± 0.1 | 5350 ± 186 | 5031 ± 90 | 7495 ± 588 | |
| 10−5 | 28.2 ± 0.1 | 465 ± 30 | 500 ± 14 | 727 ± 58 | |
| 10−6 | 31.5 ± 0.1 | 58 ± 3.5 | 44 ± 2.4 | 57 ± 3.4 | |
| 10−7 | 35.4 ± 1.0 | 5 ± 2.7 | 6 ± 0.9 | 7 ± 3.1 | |
| 10−8 | Negative c | 0 c | 1 ± 0.2 | 1 ± 1.4 e | |
| PepMV-Eur | 10−3 | 22.4 ± 0.1 | 16,485 ± 1005 | 17,254 ± 603 | 21,777 ± 610 |
| 10−4 | 25.6 ± 0.1 | 1822 ± 86 | 1713 ± 23 | 2315 ± 164 | |
| 10−5 | 29.1 ± 0.3 | 170 ± 35 | 175 ± 9 | 235 ± 15 | |
| 10−6 | 38.7 ± 0.8 | 0 ± 0.1 | 14 ± 3.8 | 18 ± 4.2 | |
| 10−7 | Negative | 0 | 1 ± 1.1 | 1 ± 0.7 d | |
| 10−8 | Negative | 0 | 0 c | 0 ± 0.4 d | |
| PepMV-US1 | 10−3 | 20.5 ± 0.1 | 109,074 ± 8848 | NA | 115,505 ± 19,153 e |
| 10−4 | 23.8 ± 0.1 | 13,888 ± 676 | 12,495 ± 384 | 17,500 ± 396 | |
| 10−5 | 27.6 ± 0.2 | 1288 ± 196 | 1215 ± 33 | 1616 ± 20 | |
| 10−6 | 31.1 ± 0.4 | 142 ± 30 | 94 ± 6.4 | 135 ± 6 | |
| 10−7 | 35.1 ± 0.6 | 12 ± 3.9 | 10 ± 1.7 | 12 ± 2.3 | |
| 10−8 | Negative | 0 | 0 ± 0.3d | 1 ± 0.8 d | |
| PepMV-Ch2 (in RNA from healthy plants) | 10−4 | 25.1 ± 0.1 | 4297 ± 184 | 4608 ± 175 | 7273 ± 127 |
| 10−5 | 28.4 ± 0.1 | 440 ± 34 | 459 ± 6 | 716 ± 15 e | |
| 10−6 | 31.8 ± 0.2 | 45 ± 5.0 | 44 ± 5.8 | 70 ± 2.2 | |
| 10−7 | 34.8 ± 0.4 | 6 ± 1.6 | 5 ± 1.2 | 9 ± 0.8 e | |
| 10−8 | 37.8 ± 0.6 d | 1 ± 0.4 d | 0 | 1 ± 0.9 | |
| 10−9 | Negative | 0 | 0 ± 0.4 d | 1 ± 0.5 d | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mehle, N.; Gregur, L.; Bogožalec Košir, A.; Dobnik, D. One-Step Reverse-Transcription Digital PCR for Reliable Quantification of Different Pepino Mosaic Virus Genotypes. Plants 2020, 9, 326. https://doi.org/10.3390/plants9030326
Mehle N, Gregur L, Bogožalec Košir A, Dobnik D. One-Step Reverse-Transcription Digital PCR for Reliable Quantification of Different Pepino Mosaic Virus Genotypes. Plants. 2020; 9(3):326. https://doi.org/10.3390/plants9030326
Chicago/Turabian StyleMehle, Nataša, Larisa Gregur, Alexandra Bogožalec Košir, and David Dobnik. 2020. "One-Step Reverse-Transcription Digital PCR for Reliable Quantification of Different Pepino Mosaic Virus Genotypes" Plants 9, no. 3: 326. https://doi.org/10.3390/plants9030326
APA StyleMehle, N., Gregur, L., Bogožalec Košir, A., & Dobnik, D. (2020). One-Step Reverse-Transcription Digital PCR for Reliable Quantification of Different Pepino Mosaic Virus Genotypes. Plants, 9(3), 326. https://doi.org/10.3390/plants9030326






