Tomato Phenotypic Diversity Determined by Combined Approaches of Conventional and High-Throughput Tomato Analyzer Phenotyping
Abstract
1. Introduction
2. Results
2.1. Field Evaluation Using Conventional Descriptors
2.1.1. Phenotyping of Vegetative and Reproductive Traits
Vegetative Traits
Fruit Color and Size
Traits Related to Fruit Form
Fruit Yield and Productivity
2.2. TA Descriptor Characterization
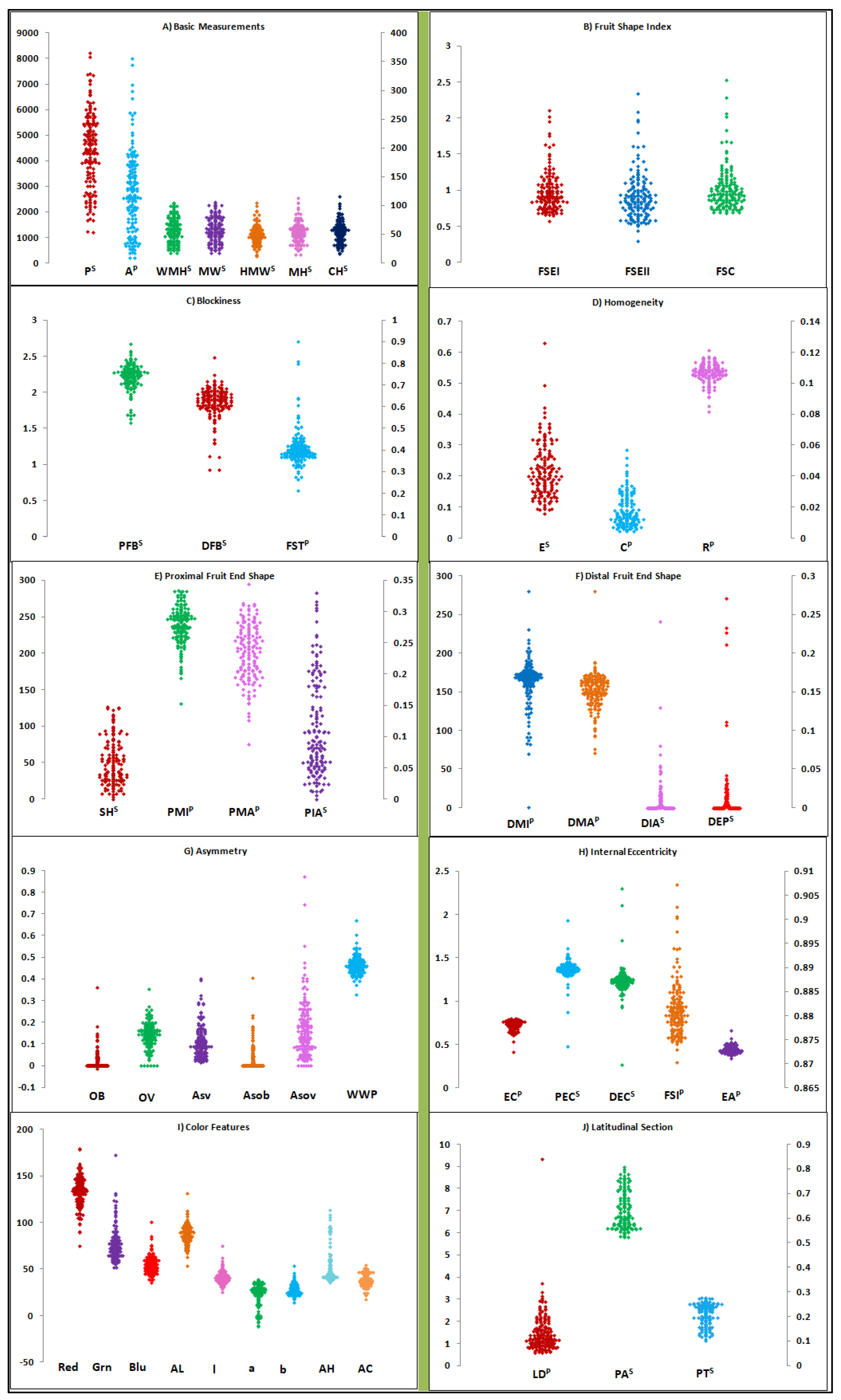
2.2.1. Fruit Trait Variation between and within Varietal Groups
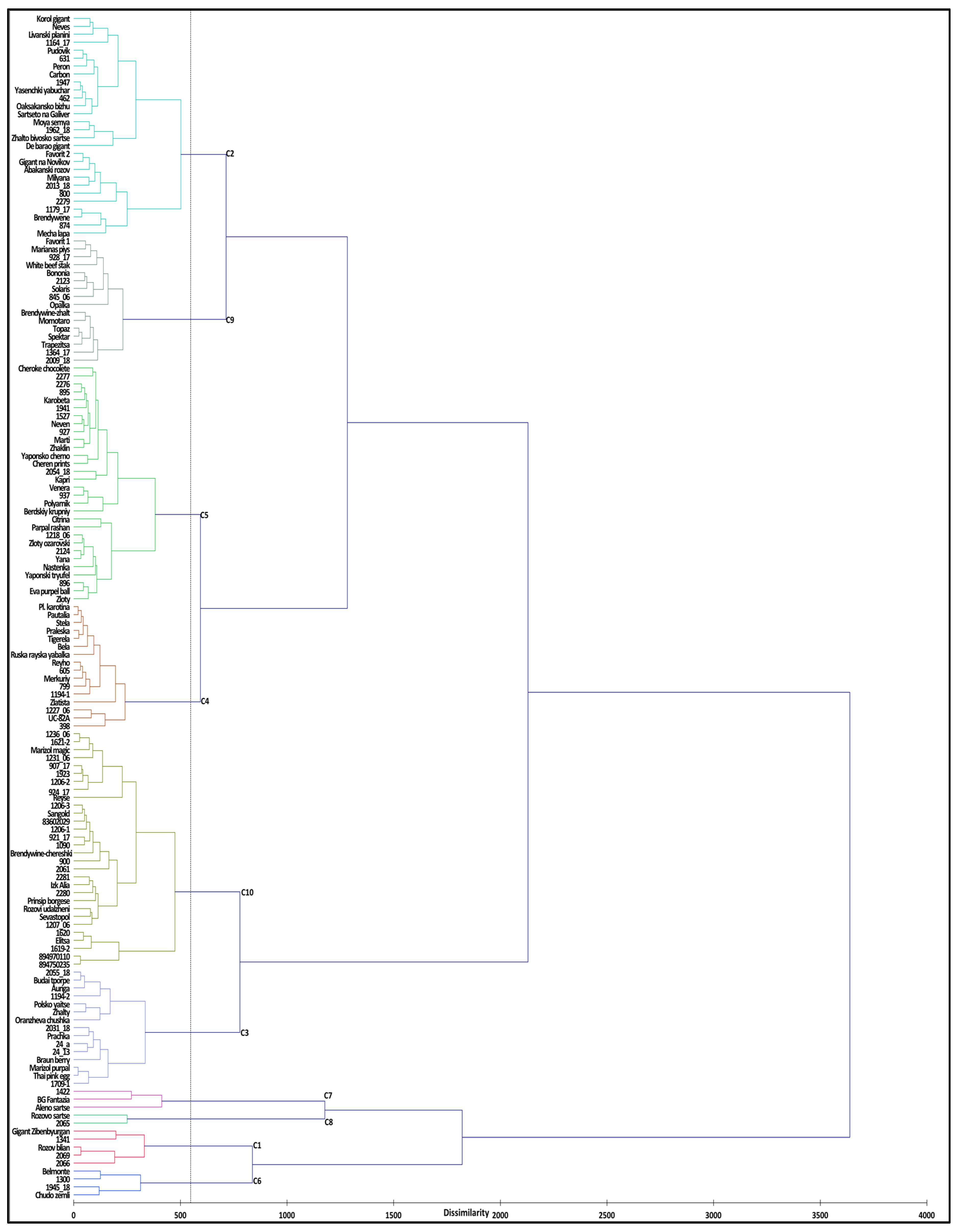
2.2.2. Cluster Analysis
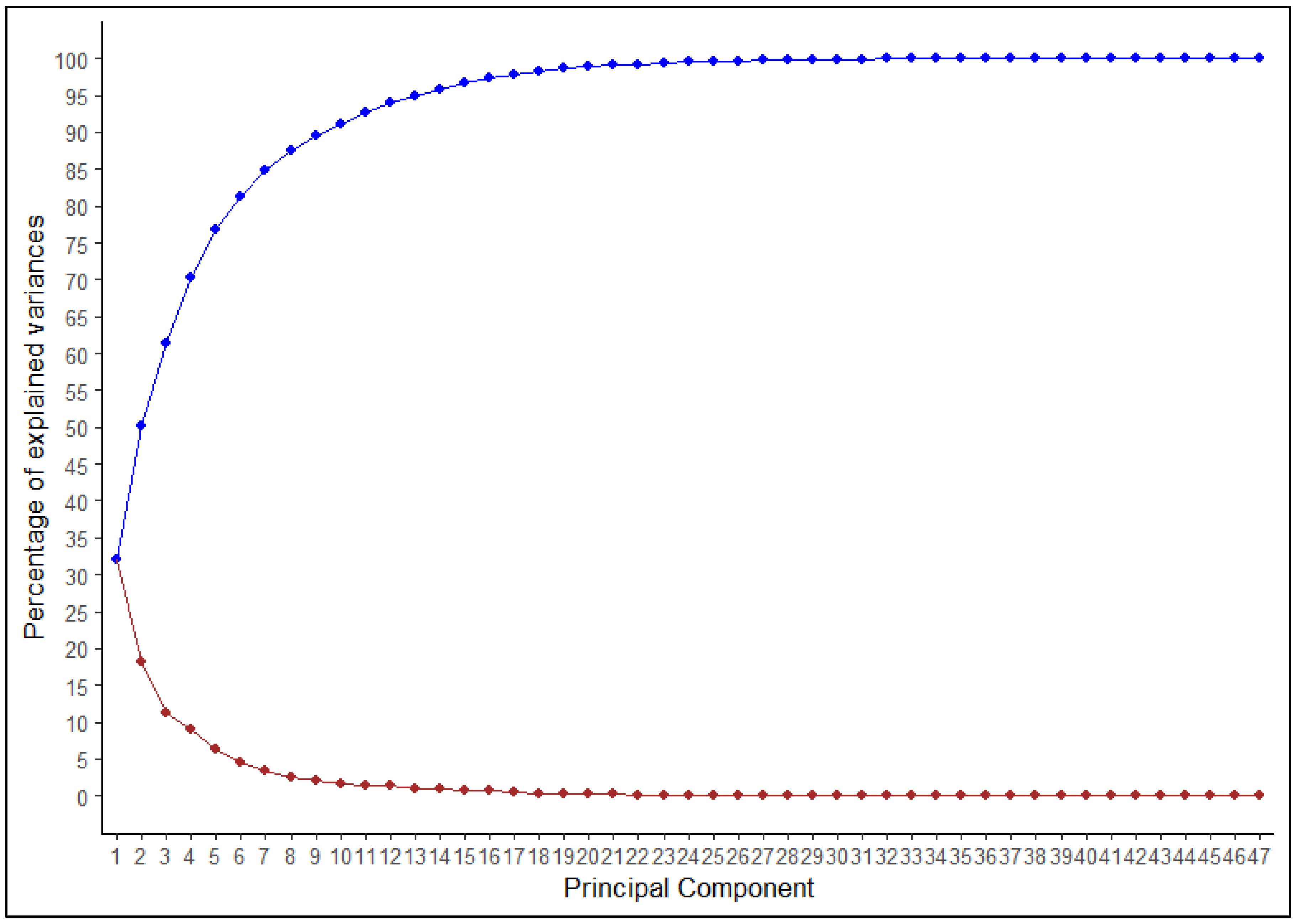
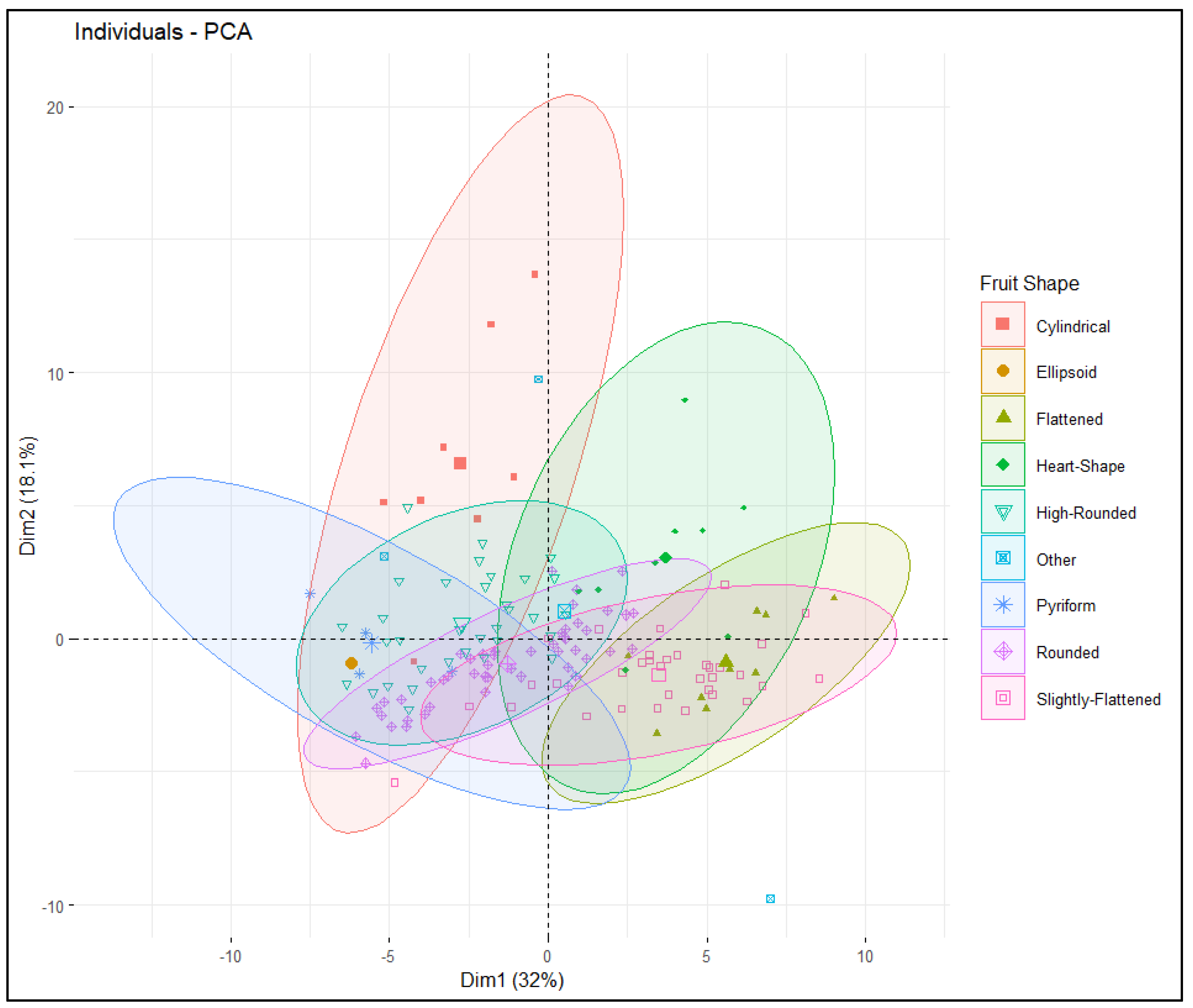
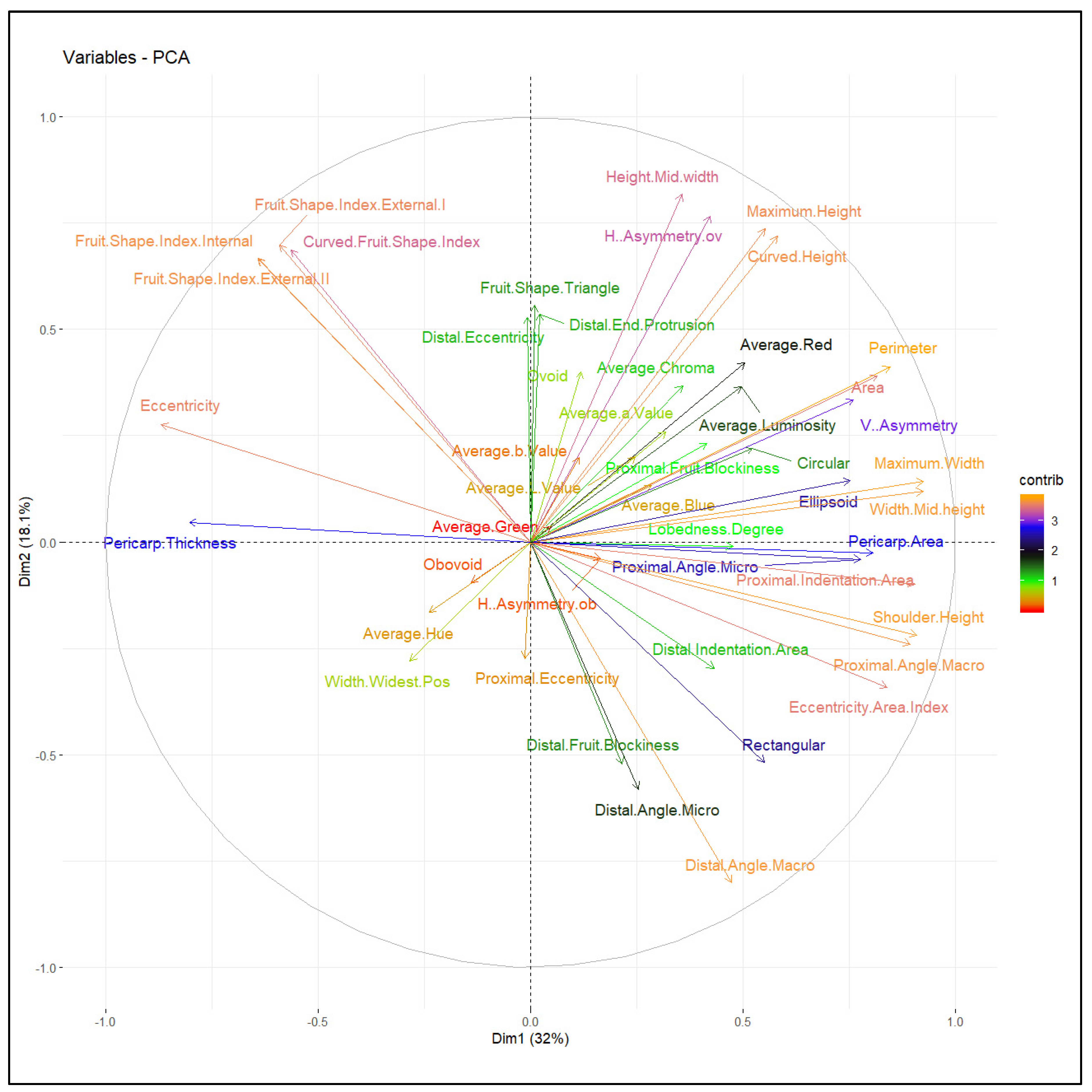
2.2.3. Multivariate Analysis
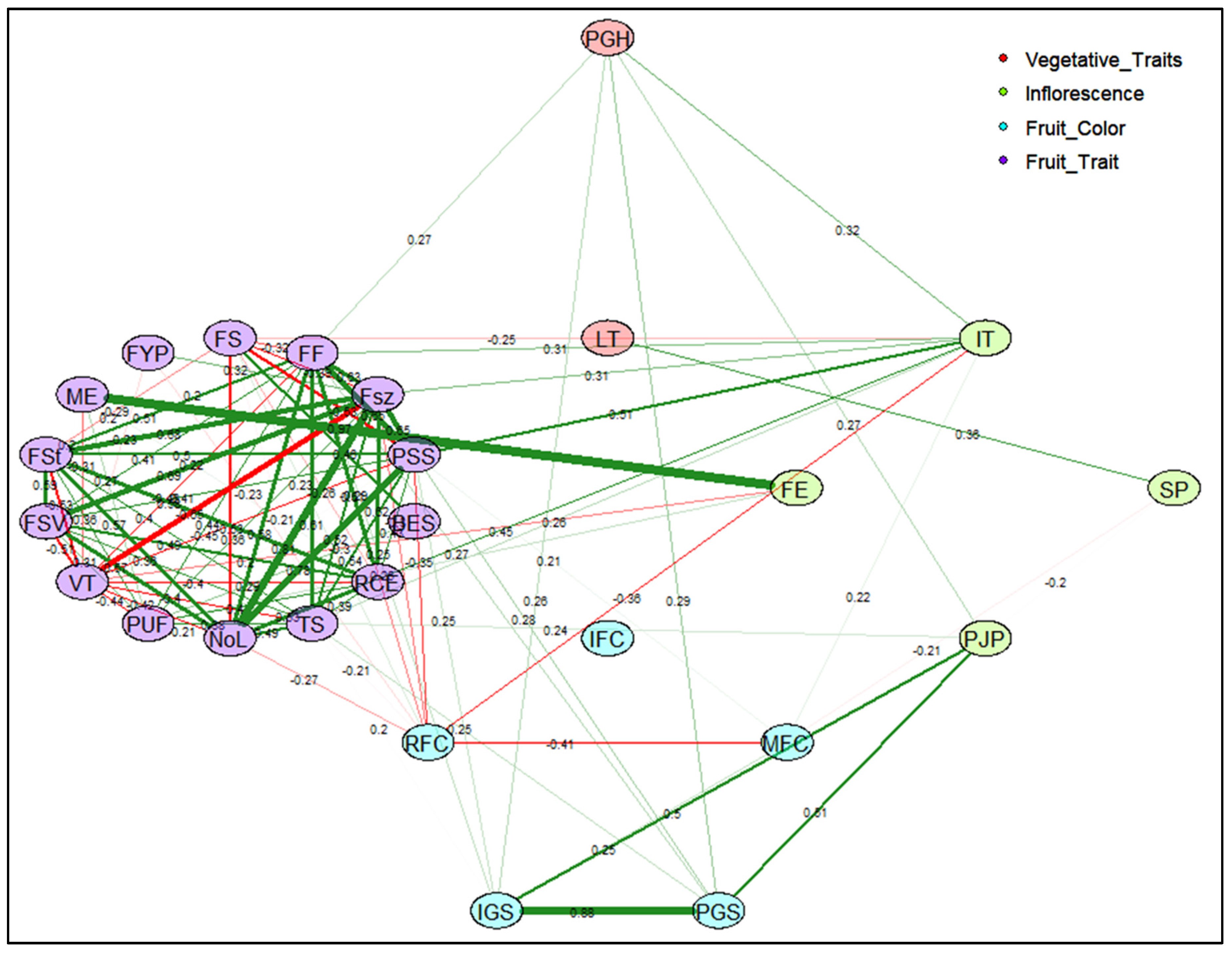
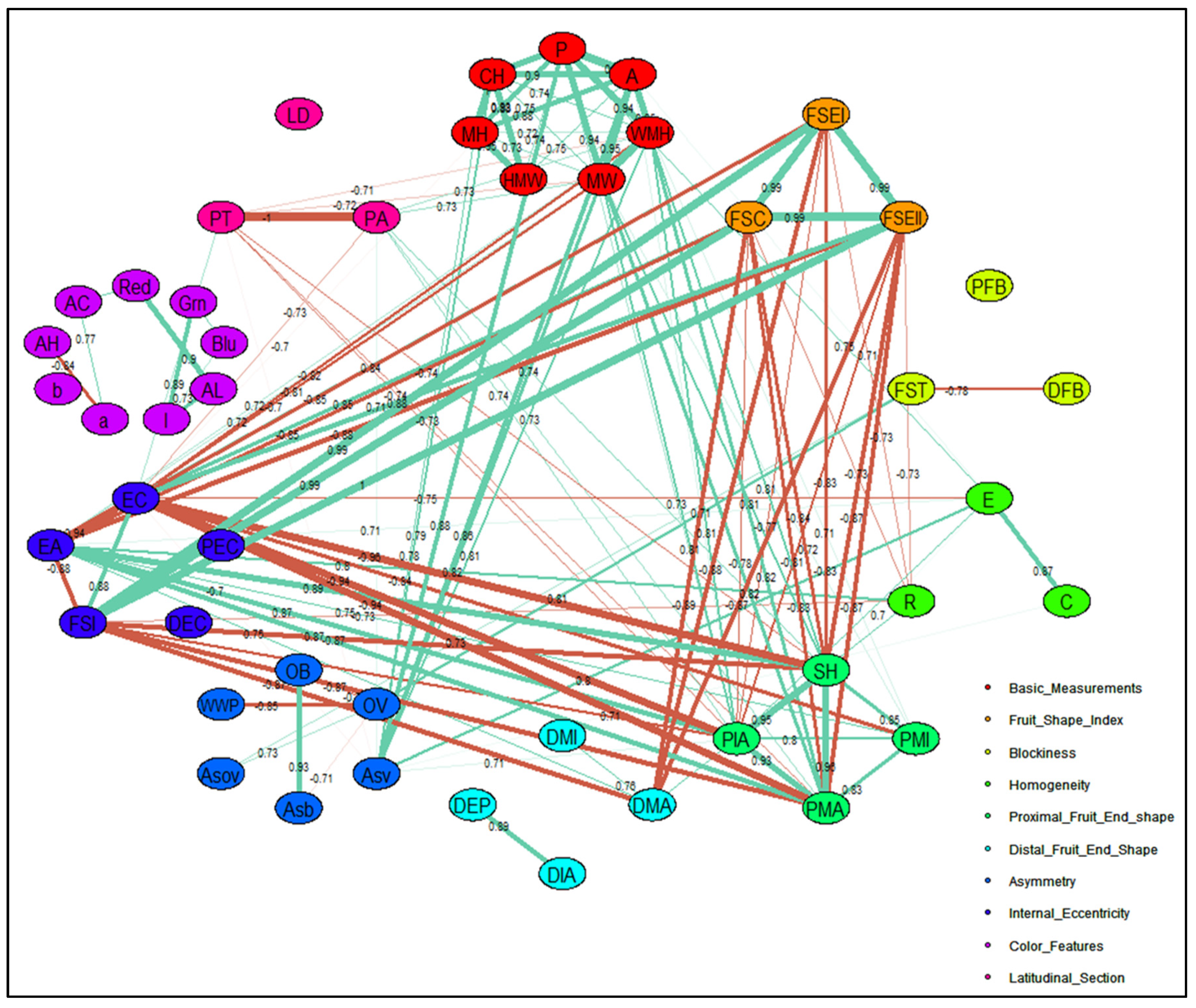
2.2.4. Correlation Network
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Seed Germination, Transplanting, and Plant Growth
4.3. Morphological Characterization
4.3.1. Conventional Descriptor Characterization
4.3.2. Tomato Analyzer Descriptor Characterization
4.4. Statistical Analyses
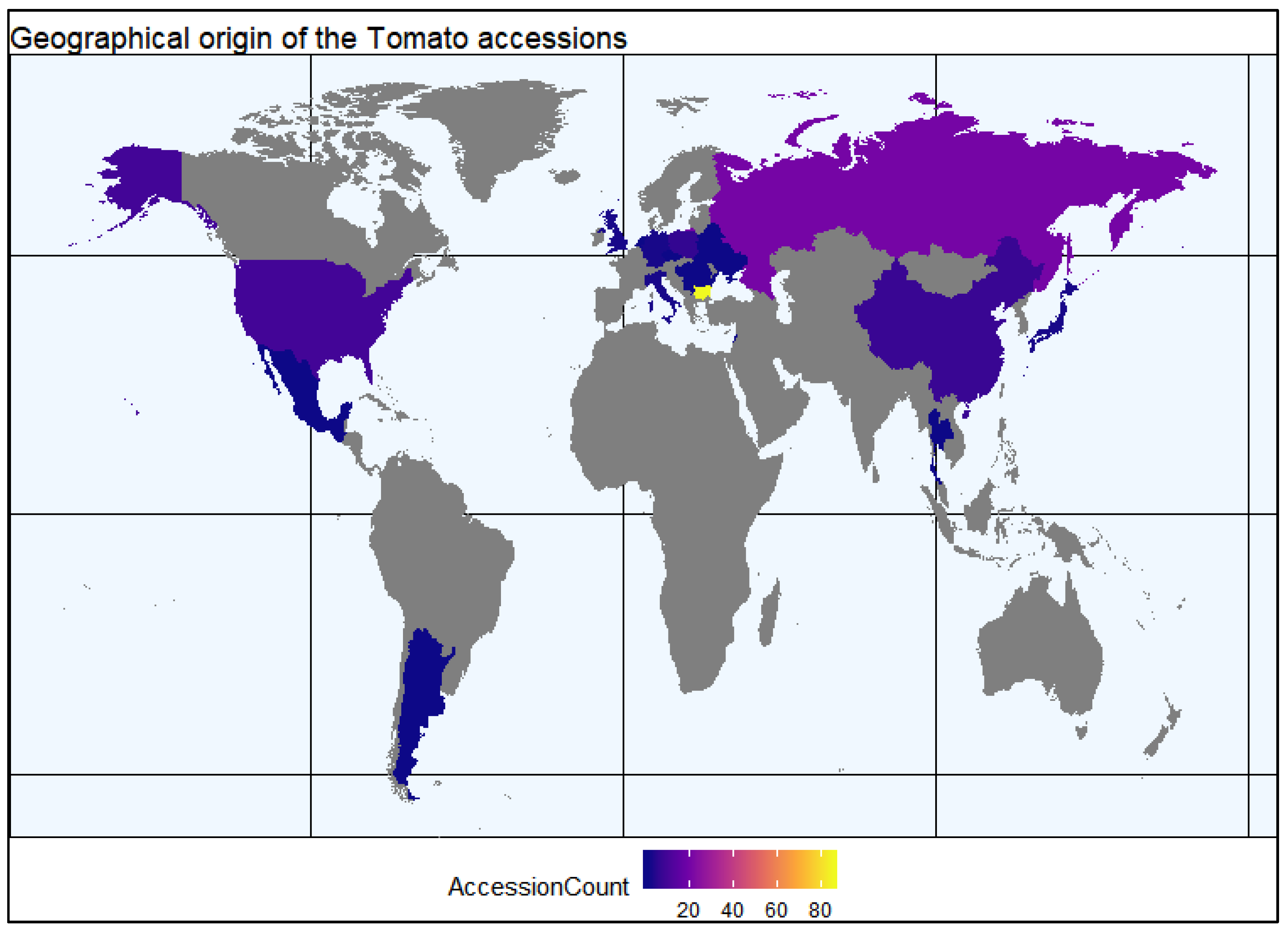
4.4.1. Construction of World Map
4.4.2. Analysis of Variance (ANOVA)
4.4.3. Hierarchical Cluster Analysis (HCA) and Factor Analysis
4.4.4. Multivariate Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mohan, V.; Gupta, S.; Thomas, S.; Mickey, H.; Charakana, C.; Chauhan, V.S.; Sharma, K.; Kumar, R.; Tyagi, K.; Sarma, S.; et al. Tomato fruits show wide phenomic diversity but fruit developmental genes show low genomic diversity. PLoS ONE 2016, 11, e0152907. [Google Scholar] [CrossRef] [PubMed]
- FAOSTAT. 2017. Available online: http://www.fao.org/faostat/en/#data/QC (accessed on 13 January 2020).
- Bergougnoux, V. The history of tomato: From domestication to biopharming. Biotechnol. Adv. 2014, 32, 170–189. [Google Scholar] [CrossRef] [PubMed]
- United States Department of Agriculture (USDA) Food Data Central. 2019. Available online: https://fdc.nal.usda.gov/fdc-app.html#/food-details/342502/nutrients (accessed on 13 January 2020).
- Beecher, G.R. Nutrient content of tomatoes and tomato products. Exp. Biol. Med. 1998, 218, 98–100. [Google Scholar] [CrossRef] [PubMed]
- Laquatra, I.; Yeung, D.L.; Storey, M.; Forshee, R. Health benefits of lycopene in tomatoes. Conference summary. Nutr. Today 2005, 40, 29–36. [Google Scholar]
- Asprelli, P.D.; Sance, M.; Insani, E.M.; Asis, R.; Vale, E.M.; Carrari, F.; Galmarini, C.R.; Peralta, I.E. Agronomic performance and fruit nutritional quality of an Andean tomato collection. Acta Hortic. 2017, 197–204. [Google Scholar] [CrossRef]
- Peralta, I.E.; Spooner, D.M. Classification of wild tomatoes: A review. Kurtziana 2000, 28, 145–154. [Google Scholar]
- Baldina, S.; Picarela, M.E.; Troise, A.D.; Pucci, A.; Ruggieri, V.; Ferracane, R.; Barone, A.; Fogliano, V.; Mazzucato, A. Metabolite profiling of Italian tomato landraces with different fruit types. Front. Plant Sci. 2016, 7, 1–13. [Google Scholar] [CrossRef]
- García-Martínez, S.; Corrado, G.; Ruiz, J.J.; Rao, R. Diversity and structure of a simple of traditional Italian and Spanish tomato accessions. Genet. Resour. Crop Evol. 2013, 60, 789–798. [Google Scholar] [CrossRef]
- Andreakis, N.; Giordano, I.; Pentangelo, A.; Foliano, V.; Graziani, G.; Monti, L.M.; Rao, R. DNA fingerprinting and quality traits of corbarino cherry-like tomato landraces. J. Agric. Food Chem. 2004, 52, 3366–3371. [Google Scholar] [CrossRef]
- Esquinas-Alcazar, J.; Nuez Viñals, F.; Rodriguez del Rincon, A.; Tello, J.; Cuartero, J.; Segura, B. Situación taxonómica, domesticación y difusión del tomate. In El Cultivo del Tomate; Nuez Viñals, F., Ed.; MundiPrensa: Madrid, Spain, 1995; pp. 15–43. [Google Scholar]
- Danailov, Z. Breeding and seed production of tomato (Solanum lycopersicum L.). In History, Methods, Achievements, Trends; Academic Publishing House “Prof. Marin Drinov”: Sofia, Bulgaria, 2012; pp. 1–265. [Google Scholar]
- Fernie, A.R.; Tadmor, Y.; Zamir, D. Natural genetic variation for improving crop quality. Curr. Opin. Plant Biol. 2006, 9, 196–202. [Google Scholar] [CrossRef]
- Passam, H.C.; Karapanos, I.C.; Bebeli, P.J.; Savvas, D. A review of recent research on tomato nutrition, breeding and post-harvest technology with reference to fruit quality. Eur. J. Plant Sci. Biotech. 2007, 1, 1–21. [Google Scholar]
- Takač, A.; Gvozdenović, Đ.; Gvozdanović-Varga, J.; Vasić, M.; Bugarski, D. Characteristics of old tomato cultivars. Nat. Montenegrina Podgor. 2005, 4, 83–91. [Google Scholar]
- Terzopoulos, P.J.; Bebeli, P.J. DNA and morphological diversity of selected Greek tomato (Solanum lycopersicum L.) landraces. Sci. Hortic. 2008, 116, 353–361. [Google Scholar] [CrossRef]
- Atanasov, A.; Todorovska, E.; Rusanov, K.; Atanasov, I. Molecular markers for characterization of genetic resources. In Proceedings of the Jubilee Scientific Session “120 Years Agricultural Sciences in Sadovo”, Sadovo-Plovdiv, Bulgaria, 21–22 May 2002; pp. 13–18. [Google Scholar]
- Krasteva, L.; Heikov, S.; Valchevaa, N. Evaluation and Management of Genetic Resources in Solanaceae; Scientific Books Agrarian University: Plovdiv, LIV, Bulgaria, 2009; pp. 19–24. [Google Scholar]
- Ganeva, D.; Pevicharova, G.; Danailov, Z. Characterization of local tomato germplasm in relation to breeding. Genetika 2014, 229–233. [Google Scholar]
- IPGRI. Descriptors for Tomato Lycopersicum spp.; International Plant Genetic Resources Institute: Rome, Italy, 1996. [Google Scholar]
- Mazzucato, A.; Papa, R.; Bitocchi, E.; Mosconi, P.; Nanni, L.; Negri, V.; Picarella, M.E.; Siligato, F.; Soressi, G.P.; Tiranti, B.; et al. Genetic diversity, structure and marker-trait associations in a collection of Italian tomato (Solanum lycopersicum L.) landraces. Theor. Appl. Genet. 2008, 116, 657–669. [Google Scholar] [CrossRef] [PubMed]
- Al-Aysh, F.; Kutma, H.; Al- Zouabi, A. Genetic variation, heritability and interrelationships of some important characteristics in Syrian tomato landraces (Solanum lycopersicum L.). Acad. Arena 2012, 4, 1–5. [Google Scholar]
- Osei, M.K.; Bonsu, K.O.; Agyeman, A.; Choi, H.S. Genetic diversity of tomato germplasm in Ghana using morphological characters. Internat. J. Plant Soil Sci. 2014, 3, 220–231. [Google Scholar] [CrossRef]
- Henareh, M.; Dursun, A.; Mandoulakani, A.B. Genetic diversity in tomato landraces collected from Turkey and Iran revealed by morphological characters. Acta. Sci. Pol. Hortorum Cultus 2015, 14, 87–96. [Google Scholar]
- Díez, M.J.; Nuez, F. Tomato. In Handbook of Plant Breeding: Vegetables II; Prohens, J., Nuez, F., Eds.; Springer: New York, NY, USA, 2008; pp. 249–323. [Google Scholar]
- Riggi, E.; Patanè, C.; Cosentino, S.L. Contenuto di composti antiossidanti in ecotipi di pomodoro da serbo del Meridione d’Italia. Italus Hortus 2006, 13, 779–782. [Google Scholar]
- Figàs, M.R.; Prohens, J.; Raigón, M.D.; Fernández-de-Córdova, P.; Fita, A.; Soler, S. Characterization of a collection of local varieties of tomato (Solanum lycopersicum L.) using conventional descriptors and the high-throughput phenomics tool Tomato Analyzer. Genet. Resour. Crop Evol. 2015, 62, 189–204. [Google Scholar] [CrossRef]
- Sim, S.C.; Robbins, M.D.; Chilcott, C.; Zhu, T.; Francis, D.M. Oligonucleotide array discovery of polymorphisms in cultivated tomato (Solanum lycopersicum L.) reveals patterns of SNP variation associated with breeding. BMC Genom. 2009, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Tripodi, P.; Massa, D.; Venezia, A.; Cardi, T. Sensing Technologies for Precision Phenotyping in Vegetable Crops: Current Status and Future Challenges. Agronomy 2018, 8, 57. [Google Scholar] [CrossRef]
- Humplik, J.F.; Lazar, D.; Husickova, A.; Spichal, L. Automated phenotyping of plant shoots using imaging methods for analysis of plant stress responses—A review. Plant Methods 2015, 11, 29. [Google Scholar] [CrossRef] [PubMed]
- Gur, A.; Zamir, D. Unused natural variation can lift yield barriers in plant breeding. PLoS Biol. 2004, 2, e245. [Google Scholar] [CrossRef]
- Brewer, M.T.; Lang, L.; Fujimura, K.; Dujmovic, N.; Gray, S.; van der Knaap, E. Development of a controlled vocabulary and software application to analyze fruit shape variation in tomato and other plant species. Plant Physiol. 2006, 141, 15–25. [Google Scholar] [CrossRef]
- Darrigues, A.; Hall, J.; van der Knaap, E.; Francis, D.M.; Dujmovic, N.; Gray, S. Tomato Analyzer-color test: A new tool for efficient digital phenotyping. J. Am. Soc. Hortic. Sci 2008, 133, 579–586. [Google Scholar] [CrossRef]
- Tripodi, P.; Greco, B. Large scale phenotyping provides insight into the diversity of vegetative and reproductive organs in a wide collection of wild and domesticated peppers (Capsicum spp.). Plants 2018, 7, 103. [Google Scholar] [CrossRef]
- Zamir, D. Where have all the crop phenotypes gone. PLoS Biol 2013, 11, e1001595. [Google Scholar] [CrossRef]
- Mazzucato, A.; Ficcadenti, N.; Caioni, M.; Mosconi, P.; Piccinini, E.; Sanampudi, V.R.R.; Sestili, S.; Ferrari, V. Genetic diversity and distinctiveness in tomato (Solanum lycopersicum L.) landraces: The Italian case study of ‘A pera Abruzzese’. Sci. Hortic. 2010, 125, 55–62. [Google Scholar] [CrossRef]
- Rodríguez, G.R.; Mu˜noz, S.; Anderson, C.; Sim, S.C.; Michel, A.; Causse, M.; McSpadden Gardener, B.B.; Francis, D.M.; van der Knaap, E. Distribution of SUN, OVATELC, and FAS in the tomato germplasm and the relationship to fruit shape diversity. Plant Physiol. 2011, 156, 275–285. [Google Scholar] [CrossRef]
- Van der Knaap, E.; Chakrabarti, M.; Chu, Y.H.; Clevenger, J.P.; Illa-Berenguer, E.; Huang, Z.; Keyhaninejad, N.; Mu, Q.; Sun, L.; Wang, Y.; et al. What lies beyond the eye: The molecular mechanisms regulating tomato fruit weight and shape. Front. Plant Sci. 2014, 5, 227. [Google Scholar] [CrossRef] [PubMed]
- Chusreeaeom, K.; Ariizumi, T.; Asamizu, E.; Okabe, Y.; Shirasawa, K.; Ezura, H. A novel tomato mutant, Solanum lycopersicum elongated fruit1 (Slelf1), exhibits an elongated fruit shape caused by increased cell layers in the proximal region of the ovary. Mol. Genet. Genom. 2014, 289, 399–409. [Google Scholar] [CrossRef] [PubMed]
- Tanksley, S.D. The genetic, developmental, and molecular bases of fruit size and shape variation in tomato. Plant Cell 2004, 16, S181–S189. [Google Scholar] [CrossRef] [PubMed]
- Gonzalo, M.J.; van der Knaap, E. A comparative analysis into the genetic bases of morphology in tomato varieties exhibiting elongated fruit shape. Theor. Appl. Genet. 2008, 116, 647–656. [Google Scholar] [CrossRef]
- Casals, J.; Bosch, L.; Casan˜as, F.; Cebolla, J.; Nuez, F. Montgrí, a cultivar within the Montserrat type. Hort. Sci. 2011, 45, 1885–1886. [Google Scholar] [CrossRef]
- Blanca, J.; Montero-Pau, J.; Sauvage, C.; Bauchet, G.; Illa, E.; Díez, M.J.; Francis, D.M.; Causse, M.; van der Knaap, E.; Cañizares, J. Genomic variation in tomato, from wild ancestors to contemporary breeding accessions. BMC Genom. 2015, 16, 257. [Google Scholar] [CrossRef]
- Rao, R.; Corrado, G.; Bianchi, M.; Di Mauro, A. (GATA) DNA fingerprinting identifies morphologically characterized ‘San Marzano’ tomato plants. Plant Breed. 2006, 125, 173–176. [Google Scholar] [CrossRef]
- Mercati, F.; Longo, C.; Poma, D.; Araniti, F.; Lupini, A.; Mammano, M.M.; Fiore, M.C.; Abenavoli, M.R.; Sunseri, F. Genetic variation of an Italian long shelf-life tomato (Solanum lycopersicum L.) collection by using SSR and morphological fruit traits. Genet. Resour. Crop. Evol. 2014, 62, 721–732. [Google Scholar] [CrossRef]
- García-Martínez, S.; Andreani, L.; Garcia-Gusano, M.; Geuna, F.; Ruiz, J.J. Evaluation of amplified fragment length polymorphism and simple sequence repeats for tomato germplasm fingerprinting: Utility for grouping closely related traditional cultivars. Genome 2006, 49, 648–656. [Google Scholar] [CrossRef]
- Colonna, V.; D’Agostino, N.; Garrison, E.; Meisner, J.; Albrechtsen, A.; Facchiano, A.; Cardi, T.; Tripodi, P. Genomic diversity and novel genome-wide association with fruit morphology in Capsicum from 746k polymorphic sites. Sci. Rep. 2019, 9, 10067. [Google Scholar] [CrossRef]
- Nankar, A.N.; Tringovska, I.; Grozeva, S.; Todorova, V.; Kostova, D. Application of high-throughput phenotyping tool Tomato Analyzer to characterize Balkan Capsicum fruit diversity. Sci. Hortic. 2019, 260, 108862. [Google Scholar] [CrossRef]
- Hurtado, M.; Vilanova, S.; Plazas, M.; Gramazio, P.; Javier Herraiz, F.; Andujar, I.; Prohens, J. Phenomics of fruit shape in eggplant (Solanum melongena L.) using Tomato Analyzer software. Sci. Hortic. 2013, 164, 625–632. [Google Scholar] [CrossRef]
- Cebolla-Cornejo, J.; Rosello’, S.; Nuez, F. Phenotypic and genetic diversity of Spanish tomato landraces. Sci. Hortic. 2013, 162, 150–164. [Google Scholar] [CrossRef]
- Gonzalo, M.J.; Brewer, M.T.; Anderson, C.; Sullivan, D.; Gray, S.; van der Knaap, E. Tomato fruit shape analysis using morphometric and morphology attributes implemented in Tomato Analyzer software program. J. Am. Soc. Hortic. Sci. 2009, 134, 77–87. [Google Scholar] [CrossRef]
- Chaim, A.B.; Paran, I.; Grube, R.C.; Jahn, M.; van Wijk, R.; Peleman, J. QTL mapping of fruit-related traits in pepper (Capsicum annuum). Theor Appl Genet. Appl. Genet. 2001, 102, 1016–1028. [Google Scholar] [CrossRef]
- Xue-Jun, C.; Rong, F.; Kun-Hua, Z.; Nan-Sheng, M.; Chang-Lin, H. Major gene plus polygene inheritance analysis of fruit traits in pepper. Acta. Bot. Sin. 2012, 32, 246–251. [Google Scholar]
- Pereira-Dias, L.; Vilanova, S.; Fita, A.; Prohens, J.; Rodríguez-Burruezo, A. Genetic diversity, population structure, and relationships in a collection of pepper (Capsicum spp.) landraces from the Spanish center of diversity revealed by genotyping-by-sequencing (GBS). Hortic. Res. 2019, 6, 54. [Google Scholar] [CrossRef]
- Rodríguez, G.R.; Strecker, J.; Brewer, M.; Gonzalo, M.J.; Anderson, C.; Lang, L.; Sullivan, D.; Wagner, E.; Strecker, B.; Drushal, R.; et al. Tomato Analyzer Version 3 User Manual. 2010. Available online: https://vanderknaaplab.uga.edu/files/Tomato_Analyzer_3.0_Manual.pdf (accessed on 13 January 2020).
- Strecker, J.; Rodríguez, G.R.; Njanji, I.; Thomas, J.; Jack, A.; Darrigues, A.; Hall, N.; Dujmovic, S.; Gray, E.; van der Knaap, E.; et al. Tomato Analyzer Color Test Manual Version 3. 2010. Available online: https://vanderknaaplab.uga.edu/files/Color_Test_3.0_Manual.pdf (accessed on 13 January 2020).

| Category | Sub-Category | No. of Accessions | Accessions Showing Variation | % Variation | ||
|---|---|---|---|---|---|---|
| I | Architecture | 1. Plant growth habit | Dwarf | 4 | 61 | 40.7 |
| Determinate | 36 | |||||
| Semi-Determinate | 21 | |||||
| Indeterminate | 89 | |||||
| II | Leaf | 2. Leaf type | Dwarf | 4 | 17 | 11.3 |
| Potato Type | 11 | |||||
| Standard | 133 | |||||
| Peruvianum | 2 | |||||
| III | Inflorescence | 3. Inflorescence Type | Uniparous | 70 | 80 | 53.3 |
| Both | 40 | |||||
| Multiparous | 40 | |||||
| 4. Style Position | Inserted | 0 | 2 | 1.3 | ||
| Same as Stamen | 148 | |||||
| Slightly Exerted | 2 | |||||
| Highly Exerted | 0 | |||||
| 5. Presence of Jointless Pedicel | Absent | 37 | 37 | 24.7 | ||
| Present | 113 | |||||
| 6. Flowering Earliness | Early | 8 | 15 | 10 | ||
| Medium | 135 | |||||
| Late | 7 | |||||
| IV | Fruit | 7. Fruit size | Very Small | 18 | 99 | 66 |
| Small | 24 | |||||
| Medium | 51 | |||||
| Large | 40 | |||||
| Very Large | 17 | |||||
| 8. Fruit Shape | Flattened | 9 | 101 | 67.3 | ||
| Slightly Flattened | 34 | |||||
| Rounded | 49 | |||||
| High Rounded | 33 | |||||
| Heart Shape | 9 | |||||
| Cylindrical | 8 | |||||
| Pyriform | 4 | |||||
| Ellipsoid | 1 | |||||
| Other | 3 | |||||
| 9. Immature Fruit color | Greenish White | 2 | 10 | 7.0 | ||
| Light Green | 7 | |||||
| Green | 140 | |||||
| Dark Green | 1 | |||||
| 10. Mature Fruit color | Green | 2 | 81 | 54 | ||
| Yellow | 8 | |||||
| Orange | 18 | |||||
| Red | 69 | |||||
| Pink | 34 | |||||
| Orange Red | 3 | |||||
| Brown | 8 | |||||
| Other | 8 | |||||
| 11. Ripened Fruit Skin Color | Colorless | 37 | 37 | 24.7 | ||
| Yellow | 113 | |||||
| 12. Presence of Greenback | Present | 54 | 54 | 36 | ||
| Absent | 96 | |||||
| 13. Intensity of Greenback | Absent | 54 | 92 | 61.3 | ||
| Slight | 25 | |||||
| Intermediate | 58 | |||||
| Strong | 13 | |||||
| 14. Fruit Size Variation | Uniform | 53 | 86 | 57.3 | ||
| Slight | 64 | |||||
| Medium | 32 | |||||
| High | 1 | |||||
| 15. Fruit Setting | Low | 91 | 91 | 60.7 | ||
| Intermediate | 52 | |||||
| High | 7 | |||||
| Very High | 0 | |||||
| 16. Varietal Type | Salad | 18 | 80 | 53.3 | ||
| Beef | 9 | |||||
| Roma | 20 | |||||
| Processing Salad | 70 | |||||
| Cherry | 32 | |||||
| Pear | 1 | |||||
| 17. Fruit Yield per Plant | Very Low | 11 | 70 | 46.7 | ||
| Low | 31 | |||||
| Medium | 80 | |||||
| High | 25 | |||||
| Very High | 3 | |||||
| 18. Ribbing at Calix End | Absent | 104 | 46 | 30.7 | ||
| Slight | 12 | |||||
| Medium | 32 | |||||
| Strong | 2 | |||||
| 19. Blossom End Shape | Flat | 125 | 25 | 16.7 | ||
| Indented | 24 | |||||
| Unknown | 1 | |||||
| 20. Maturity Earliness | Early | 8 | 14 | 9.0 | ||
| Medium | 136 | |||||
| Late | 6 |
| Descriptor (Unit) | Code # | Across Varietal Types | |||||
|---|---|---|---|---|---|---|---|
| Descriptive Statistics | ANOVA | ||||||
| F-Value | Sum of Squares (%) | ||||||
| Mean | Range | CV | Accession | Accession | Residual | ||
| Fruit Weight (gm) | 132.4 | 1.23–576.6 | 33.7 | 70.25 *** | 206,808 | 26,695 | |
| Fruits per Plant | 29.9 | 4.0–267.7 | 25.4 | 67.30 *** | 5802 | 150 | |
| Productivity (gm) | 1858.5 | 99.7–5888.0 | 34.8 | 7.23 *** | 4,517,679 | 1,169,403 | |
| Basic Measurements: | |||||||
| Perimeter (mm) | P | 191.91 | 53.52–365.8 | 9.79 | 43.34 *** | 22,821.37 | 1374.82 |
| Area (mm2) | A | 2617.43 | 202.1–8002.9 | 19.54 | 32.50 *** | 12,670,602 | 1,017,690 |
| Width Mid-Height (mm) | WMH | 56.05 | 16.87–104.7 | 11.69 | 38.13 *** | 2443.20 | 167.26 |
| Maximum Width (mm) | MW | 56.63 | 16.9–105.4 | 11.56 | 38.63 *** | 2466.26 | 166.68 |
| Height mid-Width (mm) | HMW | 48,11 | 11.5–105.2 | 8.28 | 53.07 *** | 1255.66 | 61.767 |
| Maximum Height (mm) | MH | 53.15 | 14.1–112.6 | 7.99 | 58.27 *** | 1567.98 | 70.251 |
| Curved Height (mm) | CH | 54.41 | 16.2–115.7 | 8.63 | 48.03 *** | 1578.42 | 85.8002 |
| Fruit Shape Index: | |||||||
| Fruit Shape Index External I | FSEI | 0.995 | 0.57–2.11 | 8.25 | 40.58 *** | 0.4073 | 0.0262 |
| Fruit Shape Index External II | FSEII | 0.932 | 0.30–2.34 | 10.81 | 36.63 *** | 0.5544 | 0.0395 |
| Curved Fruit Shape Index | FSC | 1.04 | 0.69–2.53 | 9.36 | 34.86 *** | 0.4885 | 0.0365 |
| Blockiness: | |||||||
| Proximal Fruit Blockiness | PFB | 0.737 | 0.525–0.890 | 7.51 | 4.06 *** | 0.0186 | 0.0119 |
| Distal Fruit Blockiness | DFB | 0.619 | 0.309–0.826 | 8.78 | 6.72 *** | 0.0296 | 0.0115 |
| Fruit Shape Triangle | FST | 1.23 | 0.63–2.70 | 15.51 | 6.76 *** | 0.3662 | 0.1413 |
| Homogeneity: | |||||||
| Ellipsoid | E | 0.042 | 0.016–0.126 | 20.55 | 12.97 *** | 0.00145 | 0.00029 |
| Circular | C | 0.084 | 0.023–0.285 | 27.54 | 16.52 *** | 0.0134 | 0.00212 |
| Rectangular | R | 0.534 | 0.408–0.607 | 4.51 | 5.88 *** | 0.0051 | 0.00225 |
| Proximal Fruit End Shape: | |||||||
| Shoulder Height | SH | 0.057 | 0.0–0.15 | 32.99 | 12.92 *** | 0.00676 | 0.00137 |
| Proximal Angle Micro (Degrees) | PMI | 235.98 | 130.9–285.8 | 8.83 | 5.73 *** | 3707.77 | 1689.38 |
| Proximal Angle Macro (Degrees) | PMA | 198.18 | 75.6–295.2 | 8.67 | 17.24 *** | 7592.9 | 1149.96 |
| Proximal Indentation Area | PIA | 0.107 | 0.0–0.33 | 45.53 | 8.73 *** | 0.0309 | 0.0093 |
| Distal Fruit End Shape: | |||||||
| Distal Angle Micro (Degrees) | DMI | 163.46 | 0.50–280.2 | 15.44 | 4.15 *** | 3944.70 | 2480.03 |
| Distal Angle Macro (Degrees) | DMA | 150.33 | 70.53–280.4 | 7.75 | 12.58 *** | 2541.33 | 527.51 |
| Distal Indentation Area | DIA | 0.007 | 0.0- 0.24 | 265.85 | 4.28 *** | 0.0022 | 0.0014 |
| Distal End Protrusion | DEP | 0.013 | 0.0–0.27 | 271.32 | 4.72 *** | 0.0086 | 0.0048 |
| Asymmetry: | |||||||
| Obovoid | OB | 0.02 | −0.02–0.36 | 180.65 | 5.54 *** | 0.0108 | 0.0051 |
| Ovoid | OV | 0.141 | 0.0–0.35 | 42.06 | 3.57 *** | 0.0187 | 0.0137 |
| V. Asymmetry | Asv | 0.101 | 0.012–0.399 | 57.98 | 5.23 *** | 0.0269 | 0.0134 |
| H. Asymmetry. Ob | Asob | 0.024 | 0.0- 0.41 | 231.6 | 3.47 *** | 0.0161 | 0.0121 |
| H. Asymmetry. Ov | Asov | 0.166 | 0.00–0.87 | 53.83 | 7.62 *** | 0.0909 | 0.0311 |
| Width Widest Pos | WWP | 0.461 | 0.33–0.67 | 8.97 | 3.51 *** | 0.0089 | 0.0066 |
| Internal Eccentricity: | |||||||
| Eccentricity | EC | 0.728 | 0.414–0.796 | 3.74 | 12.80 *** | 0.0141 | 0.0029 |
| Proximal Eccentricity | PEC | 0.889 | 0.87–0.90 | 0.32 | 1.71 *** | 0.00002 | 0.00003 |
| Distal Eccentricity | DEC | 0.887 | 0.87–0.91 | 0.39 | 2.19 *** | 0.00004 | 0.00005 |
| Fruit Shape Index Internal | FSI | 0.933 | 0.29–2.34 | 10.85 | 36.56 *** | 0.5577 | 0.0393 |
| Eccentricity Area Index | EA | 0.437 | 0.34–0.66 | 5.22 | 11.02 *** | 0.0085 | 0.00202 |
| Average Color Values: | |||||||
| Red | Red | 133.16 | 74.99–179.3 | 4.89 | 21.15 *** | 1341.43 | 165.61 |
| Green | Green | 75.91 | 51.49–172.4 | 8.42 | 28.60 *** | 1742.58 | 159.09 |
| Blue | Blue | 54.31 | 35.62–100.37 | 8.67 | 13.84 *** | 456.65 | 86.14 |
| Luminosity | AL | 88.26 | 53.26–131.46 | 5.50 | 15.26 *** | 535.91 | 91.67 |
| L | l | 41.47 | 35.58–35.71 | 6.07 | 21.71 *** | 205.05 | 24.66 |
| a | a | 23.81 | −11.7–38.85 | 11.46 | 58.79 *** | 652.03 | 28.95 |
| B | b | 27.58 | 13.6–53.49 | 6.12 | 40.54 *** | 172.03 | 11.08 |
| Hue | AH | 50.59 | 35.2–113.2 | 6.89 | 86.77 *** | 1572.6 | 47.31 |
| Chroma | AC | 37.95 | 17.1–54.7 | 6.67 | 24.01 *** | 229.4 | 24.95 |
| Latitudinal Section: | |||||||
| Lobedness Degree | LD | 1.43 | 1.68–0.93 | 42.93 | 5.47 *** | 3.055 | 1.357 |
| Pericarp Area | PA | 0.62 | 0.52–0.81 | 4.11 | 23.3 *** | 0.022 | 0.0023 |
| Pericarp Thickness | PT | 0.21 | 0.10–0.28 | 7.39 | 23.1 *** | 0.0086 | 0.0009 |
| TA Descriptor Category | Factor | Correlation between the Factor and Variable | |
|---|---|---|---|
| Positive | Negative | ||
| Size | 1 | Basic measurement (size) | Eccentricity |
| Shape | 2 | Fruit shape index external I | Proximal/distal fruit end shape |
| Fruit shape index external II | Fruit shape index internal | ||
| Curved fruit shape index | |||
| H. asymmetry. ov | |||
| 5 | Average chroma | Fruit shape triangle | |
| Circular | |||
| 6 | Pericarp area | Average chroma | |
| 7 | Proximal fruit blockiness | ||
| Lobedness degree | Distal fruit blockiness | ||
| Rectangular | |||
| 8 | Height mid-width | Distal end protrusion | |
| Color | 3 | Green | a |
| l | |||
| 4 | a | Green | |
| l | |||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nankar, A.N.; Tringovska, I.; Grozeva, S.; Ganeva, D.; Kostova, D. Tomato Phenotypic Diversity Determined by Combined Approaches of Conventional and High-Throughput Tomato Analyzer Phenotyping. Plants 2020, 9, 197. https://doi.org/10.3390/plants9020197
Nankar AN, Tringovska I, Grozeva S, Ganeva D, Kostova D. Tomato Phenotypic Diversity Determined by Combined Approaches of Conventional and High-Throughput Tomato Analyzer Phenotyping. Plants. 2020; 9(2):197. https://doi.org/10.3390/plants9020197
Chicago/Turabian StyleNankar, Amol N., Ivanka Tringovska, Stanislava Grozeva, Daniela Ganeva, and Dimitrina Kostova. 2020. "Tomato Phenotypic Diversity Determined by Combined Approaches of Conventional and High-Throughput Tomato Analyzer Phenotyping" Plants 9, no. 2: 197. https://doi.org/10.3390/plants9020197
APA StyleNankar, A. N., Tringovska, I., Grozeva, S., Ganeva, D., & Kostova, D. (2020). Tomato Phenotypic Diversity Determined by Combined Approaches of Conventional and High-Throughput Tomato Analyzer Phenotyping. Plants, 9(2), 197. https://doi.org/10.3390/plants9020197





