Comparative Transcriptome Analysis of Waterlogging-Sensitive and Tolerant Zombi Pea (Vigna vexillata) Reveals Energy Conservation and Root Plasticity Controlling Waterlogging Tolerance
Abstract
1. Introduction
2. Results and Discussion
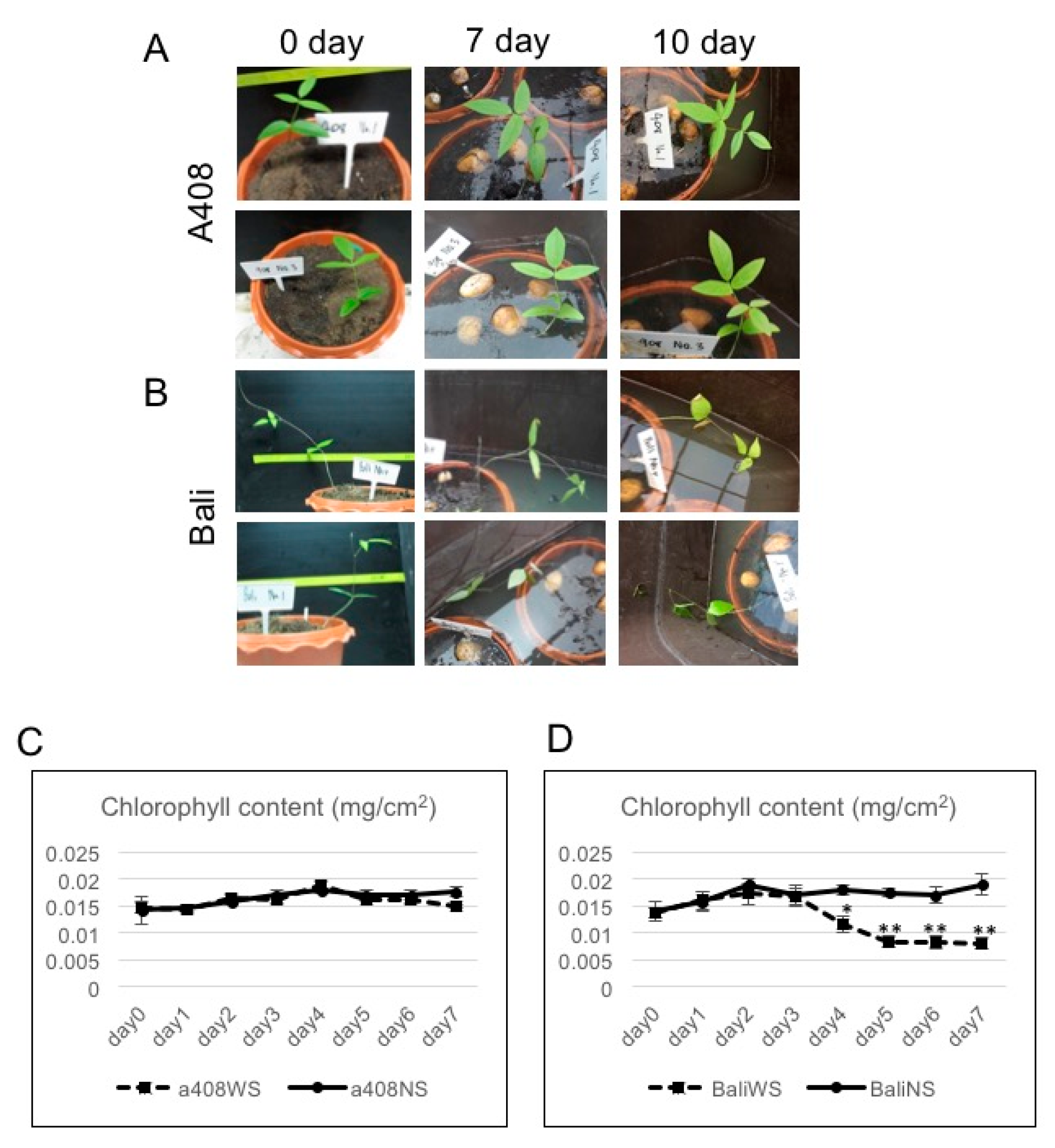
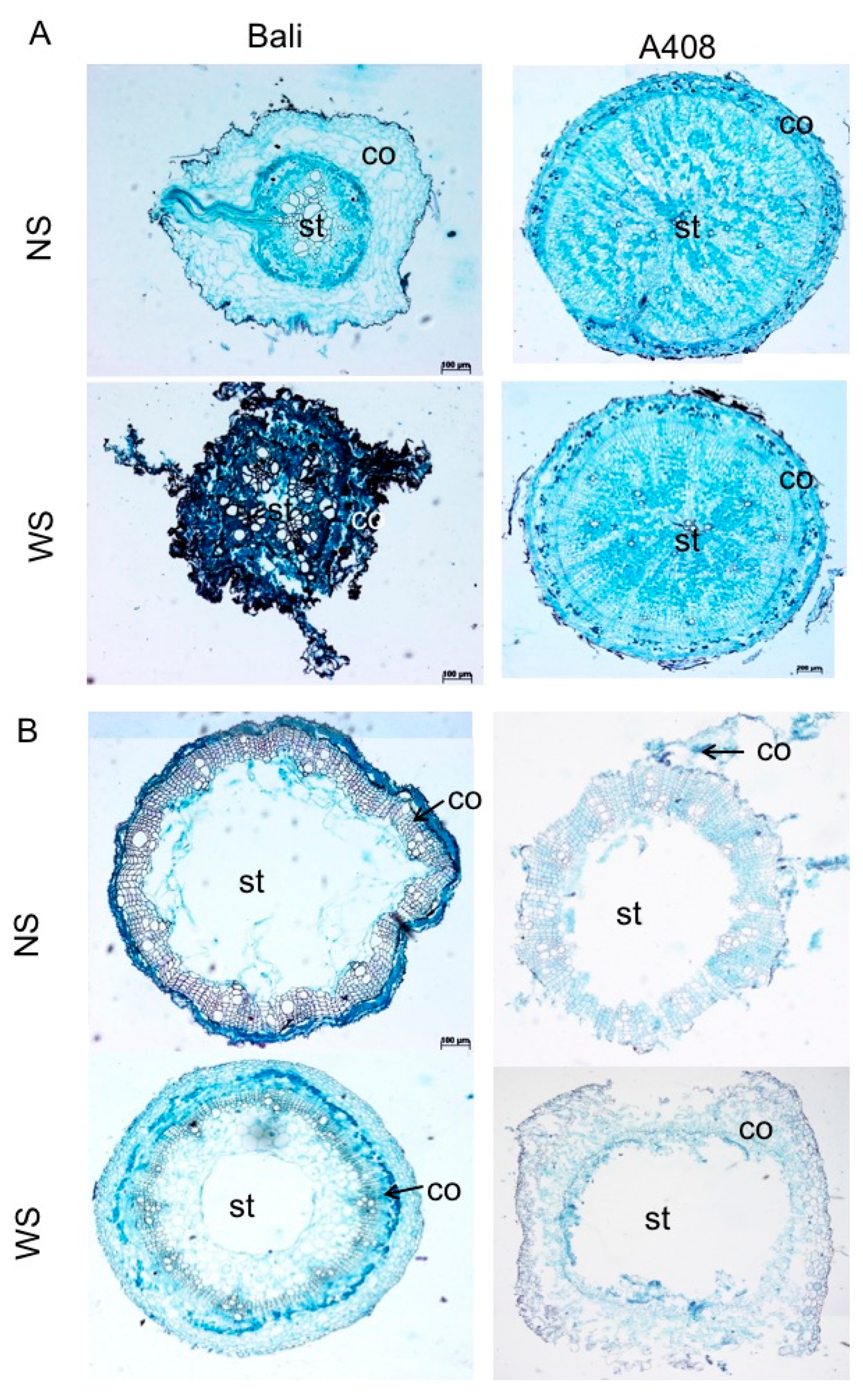
2.1. Anatomical and Morphological Changes of Zombi Pea Varieties Subjected to Waterlogging Stress
2.2. De Novo Transcriptome Analysis
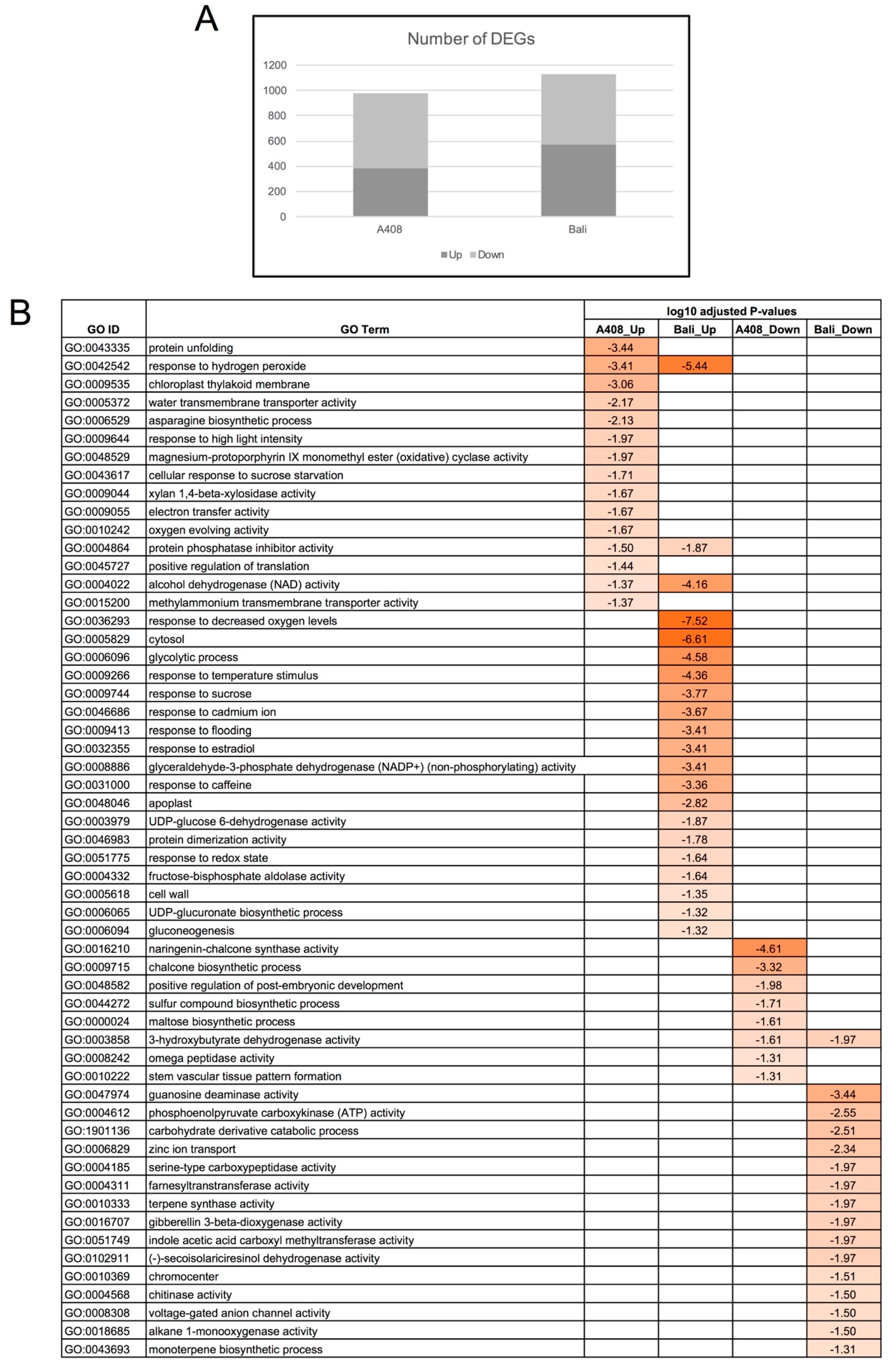
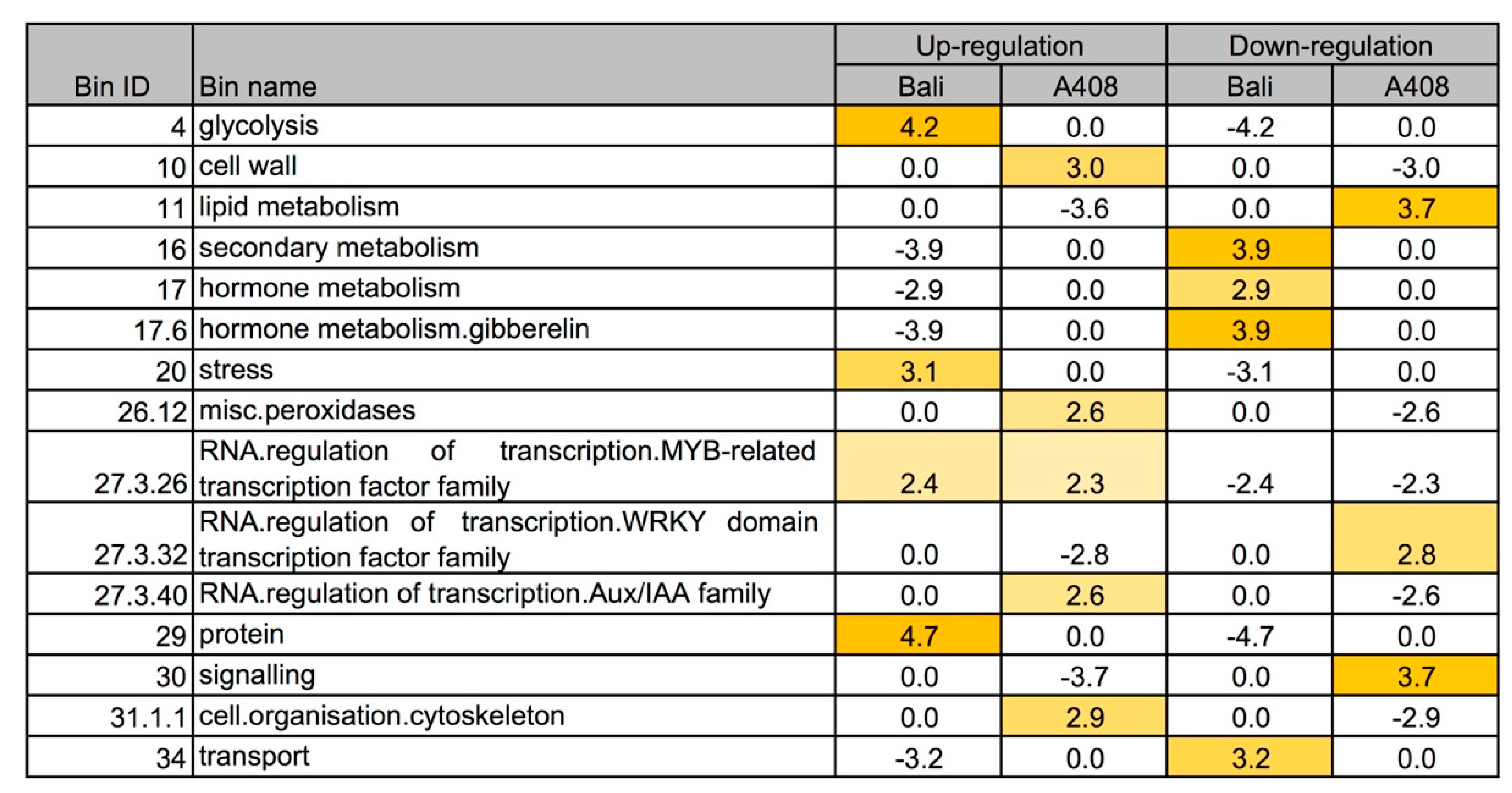
2.3. Differential Gene Expression, Functional Enrichment, and Comparative Transcriptome Analyses
2.4. Waterlogging Resulted in Stronger Induction of Glycolysis and Fermentative Genes in “Bali” than in “A408”
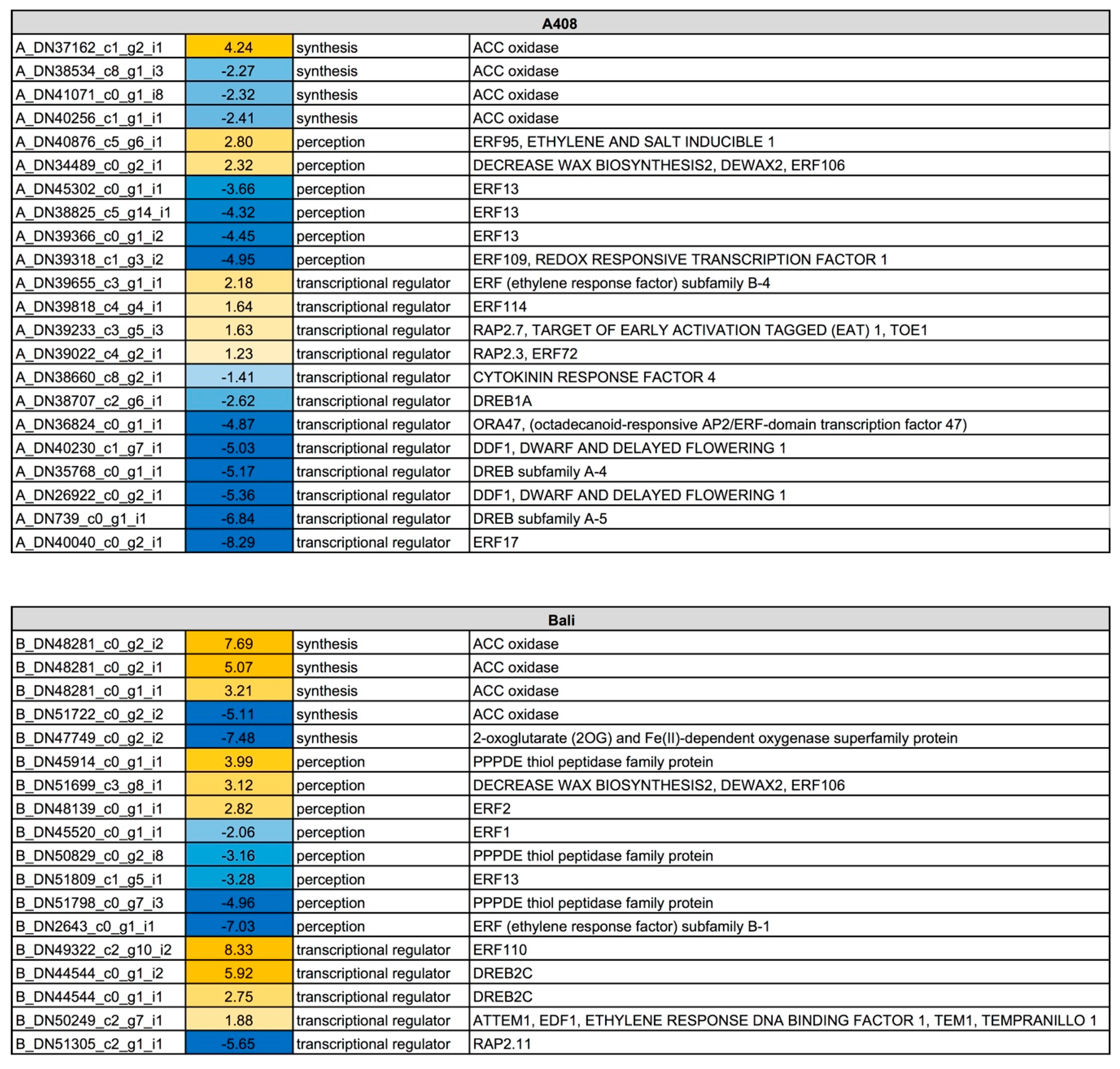
2.5. Waterlogging Resulted in Stronger Induction of Ethylene Synthesis, Perception and Responsive Genes in “Bali” than in “A408”
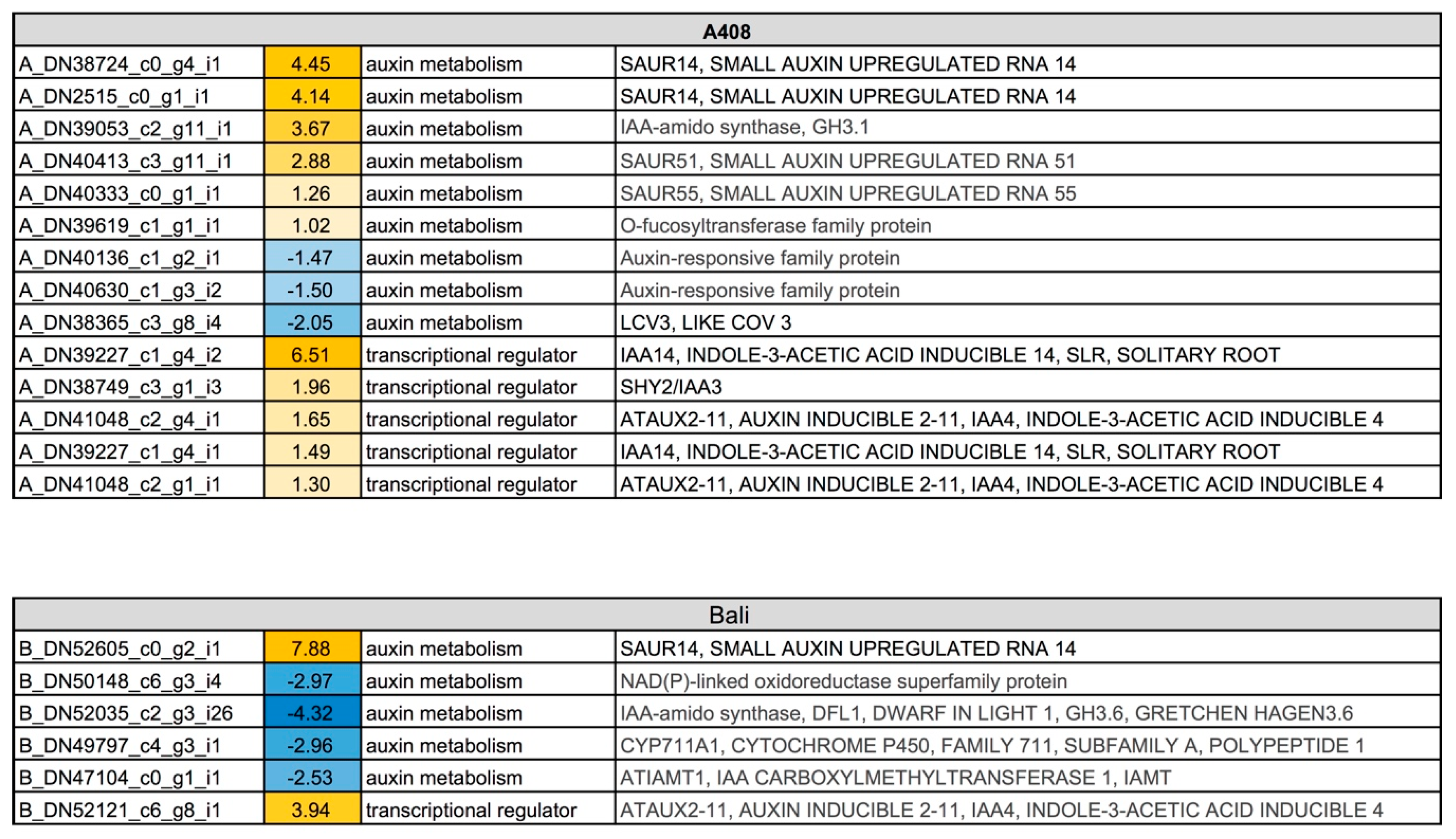
2.6. Auxin Metabolism and Auxin-Regulated Transcription Factor Genes were Predominantly Induced in the Roots of Waterlogging-Tolerant Zombi Pea
2.7. Differential Expression of Abscisic Acid and Gibberellic Acid Metabolic Genes
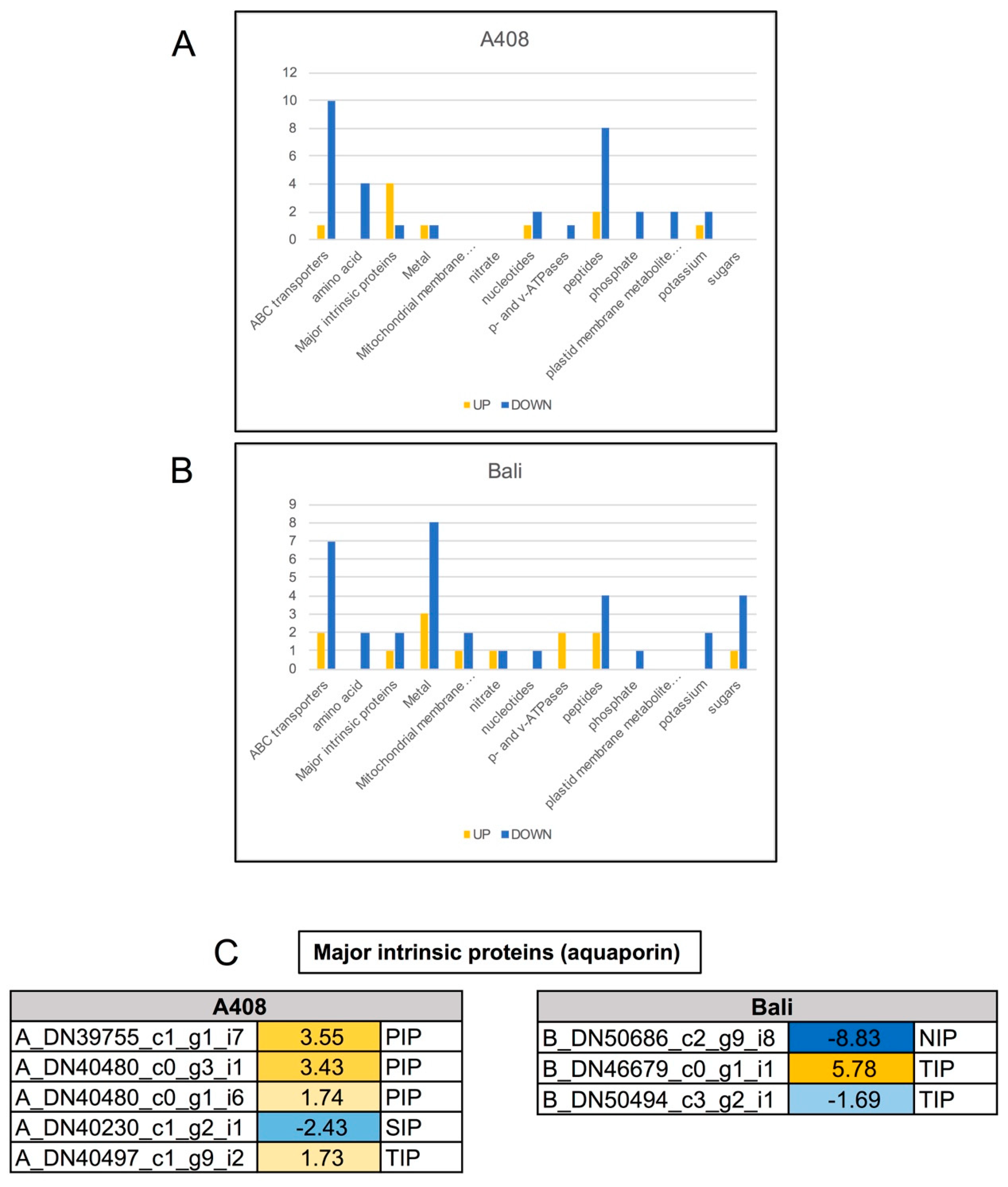
2.8. Differential Expression of Transport Genes
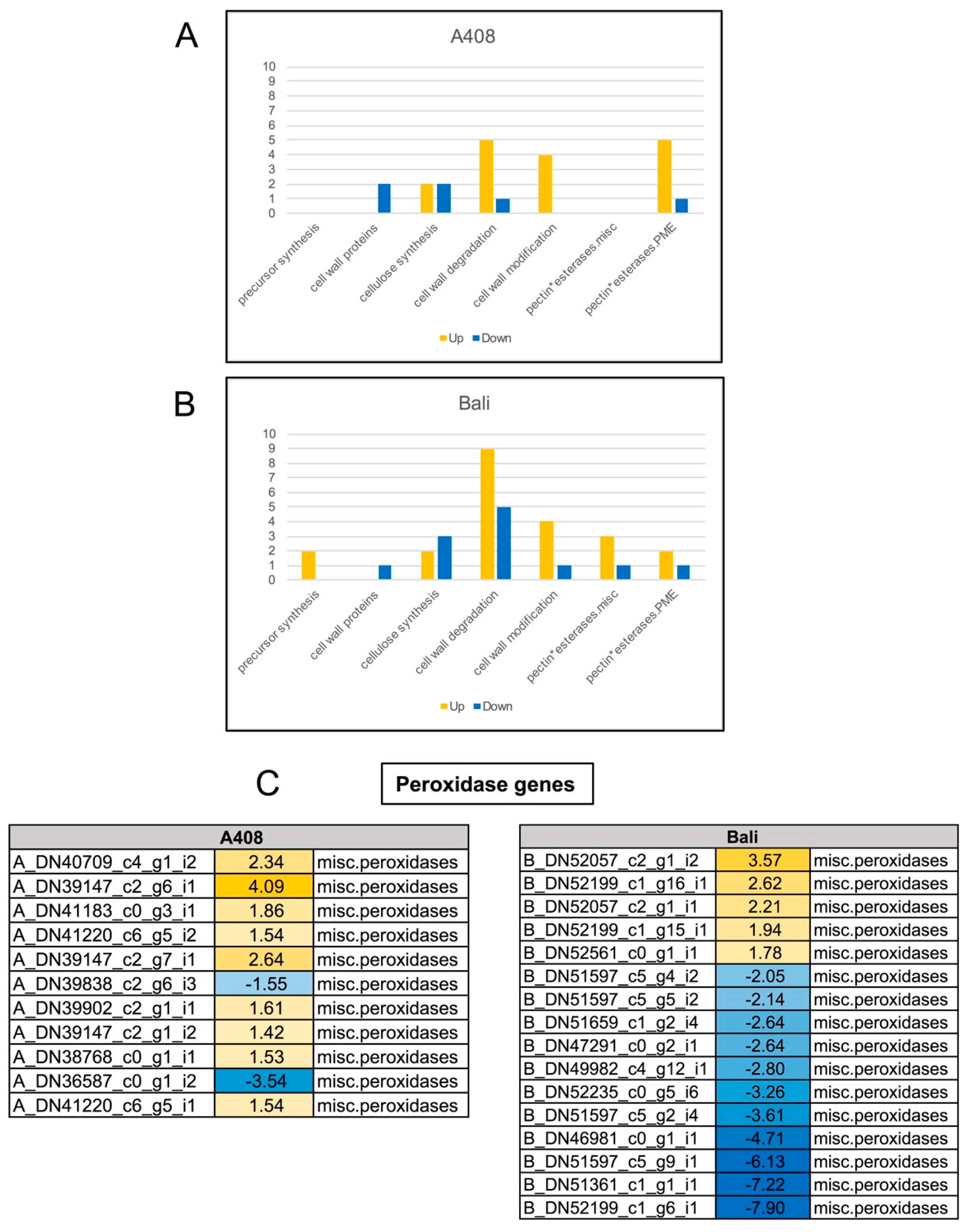
2.9. Differential Expression of Plant Cell Wall-Related Genes
2.10. Validation of Transcriptome Data by Quantitative Real-Time PCR
3. Material and Methods
3.1. Plant Material and Stress Treatment
3.2. Analysis of Leaf Chlorophyll Content
3.3. Root Anatomical and Morphological Analysis
3.4. Analysis of Total Soluble Carbohydrate Content
3.5. RNA Extraction, Library Preparation, and Sequencing
3.6. De Novo Assembly and Annotation
3.7. Differential Gene Expression Analysis
3.8. Quantitative Real-Time PCR
4. Conclusions
Supplementary Materials
Author contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sasidharan, R.; Bailey-Serres, J.; Ashikari, M.; Atwell, B.J.; Colmer, T.D.; Fagerstedt, K.; Fukao, T.; Geigenberger, P.; Hebelstrup, K.H.; Hill, R.D.; et al. Community recommendations on terminology and procedures used in flooding and low oxygen stress research. New Phytol. 2017, 214, 1403–1407. [Google Scholar] [CrossRef] [PubMed]
- Juntawong, P.; Sirikhachornkit, A.; Pimjan, R.; Sonthirod, C.; Sangsrakru, D.; Yoocha, T.; Tangphatsornruang, S.; Srinives, P. Elucidation of the molecular responses to waterlogging in Jatropha roots by transcriptome profiling. Front. Plant Sci. 2014, 5, 658. [Google Scholar] [CrossRef] [PubMed]
- Bailey-Serres, J.; Fukao, T.; Gibbs, D.J.; Holdsworth, M.J.; Lee, S.C.; Licausi, F.; Perata, P.; Voesenek, L.A.; van Dongen, J.T. Making sense of low oxygen sensing. Trends Plant Sci. 2012, 17, 129–138. [Google Scholar] [CrossRef] [PubMed]
- Fukao, T.; Xiong, L. Genetic mechanisms conferring adaptation to submergence and drought in rice: Simple or complex? Curr. Opin. Plant Biol. 2013, 16, 196–204. [Google Scholar] [CrossRef] [PubMed]
- Voesenek, L.A.; Bailey-Serres, J. Flood adaptive traits and processes: An overview. New Phytol. 2015, 206, 57–73. [Google Scholar] [CrossRef]
- Xu, K.; Xu, X.; Fukao, T.; Canlas, P.; Maghirang-Rodriguez, R.; Heuer, S.; Ismail, A.M.; Bailey-Serres, J.; Ronald, P.C.; Mackill, D.J. Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice. Nature 2006, 442, 705–708. [Google Scholar] [CrossRef]
- Hattori, Y.; Nagai, K.; Furukawa, S.; Song, X.J.; Kawano, R.; Sakakibara, H.; Wu, J.; Matsumoto, T.; Yoshimura, A.; Kitano, H.; et al. The ethylene response factors SNORKEL1 and SNORKEL2 allow rice to adapt to deep water. Nature 2009, 460, 1026–1030. [Google Scholar] [CrossRef]
- Rivera-Contreras, I.K.; Zamora-Hernandez, T.; Huerta-Heredia, A.A.; Capataz-Tafur, J.; Barrera-Figueroa, B.E.; Juntawong, P.; Pena-Castro, J.M. Transcriptomic analysis of submergence-tolerant and sensitive Brachypodium distachyon ecotypes reveals oxidative stress as a major tolerance factor. Sci. Rep. 2016, 6, 27686. [Google Scholar] [CrossRef]
- Valliyodan, B.; Van Toai, T.T.; Alves, J.D.; Goulart, P.D.P.; Lee, J.D.; Fritschi, F.B.; Rahman, M.A.; Islam, R.; Shannon, J.G.; Nguyen, H.T. Expression of Root-Related Transcription Factors Associated with Flooding Tolerance of Soybean (Glycine max). Int. J. Mol. Sci. 2014, 15, 17622–17643. [Google Scholar] [CrossRef]
- Jitsuyama, Y. Morphological root responses of soybean to rhizosphere hypoxia reflect waterlogging tolerance. Can. J. Plant Sci. 2015, 95, 999–1005. [Google Scholar] [CrossRef]
- Kim, Y.H.; Hwang, S.J.; Wagas, M.; Khan, A.L.; Lee, J.H.; Lee, J.D.; Nguyen, H.T.; Lee, I.J. Comparative analysis of endogenous hormones level in two soybean (Glycine max L.) lines differing in waterlogging tolerance. Front. Plant Sci. 2015, 6, 714. [Google Scholar] [CrossRef]
- Sakazono, S.; Nagata, T.; Matsuo, R.; Kajihara, S.; Watanabe, M.; Ishimoto, M.; Shimamura, S.; Harada, K.; Takahashi, R.; Mochizuki, T. Variation in Root Development Response to flooding among 92 Soybean Lines during Early Growth Stages. Plant Prod. Sci. 2014, 17, 228–236. [Google Scholar] [CrossRef]
- Cornelious, B.; Chen, P.; Hou, A.; Shi, A.; Shannon, J.G. Yield Potential and Waterlogging Tolerance of Selected Near-Isogenic Lines and Recombinant Inbred Lines from Two Southern Soybean Populations. J. Crop Improv. 2006, 16, 97–111. [Google Scholar] [CrossRef]
- Nguyen, V.T.; Vuong, T.D.; VanToai, T.; Lee, J.D.; Wu, X.; Mian, M.A.R.; Dorrance, A.E.; Shannon, J.G.; Nguyen, H.T. Mapping of Quantitative Trait Loci Associated with Resistance to Phytophthora sojae and Flooding Tolerance in Soybean. Crop Sci. 2012, 52, 2481–2493. [Google Scholar] [CrossRef]
- Rhine, M.D.; Stevens, G.; Shannon, G.; Wrather, A.; Sleper, D. Yield and nutritional responses to waterlogging of soybean cultivars. Irrig. Sci. 2010, 28, 135–142. [Google Scholar] [CrossRef]
- VanToai, T.T.; Beuerlein, A.F.; Schmitthenner, S.K.; St. Martin, S.K. Genetic Variability for Flooding Tolerance in Soybeans. Crop Sci. 1994, 34, 1112–1115. [Google Scholar] [CrossRef]
- Ye, H.; Song, L.; Chen, H.T.; Valliyodan, B.; Cheng, P.; Ali, L.; Vuong, T.; Wu, C.J.; Orlowski, J.; Buckley, B.; et al. A major natural genetic variation associated with root system architecture and plasticity improves waterlogging tolerance and yield in soybean. Plant Cell Environ. 2018, 41, 2169–2182. [Google Scholar] [CrossRef]
- Gibberd, M.R.; Gray, J.D.; Cocks, P.S.; Colmer, T.D. Waterlogging tolerance among a diverse range of Trifolium accessions is related to root porosity, lateral root formation and ‘aerotropic rooting’. Ann. Bot. 2001, 88, 579–589. [Google Scholar] [CrossRef]
- Malik, A.I.; Ailewe, T.I.; Erskine, W. Tolerance of three grain legume species to transient waterlogging. AoB Plants 2015, 7, 40. [Google Scholar] [CrossRef]
- Lewis, G.P. Legumes of the World; Royal Botanic Gardens Kew: London, UK, 2005. [Google Scholar]
- Dachapak, S.; Somta, P.; Poonchaivilaisak, S.; Yimram, T.; Srinives, P. Genetic diversity and structure of the zombi pea (Vigna vexillata (L.) A. Rich) gene pool based on SSR marker analysis. Genetica 2017, 145, 189–200. [Google Scholar] [CrossRef]
- Tomooka, N.; Yoon, M.S.; Doi, K.; Kaga, A.; Vaughan, D. AFLP analysis of diploid species in the genus Vigna subgenus Ceratotropis. Genet. Resour. Crop Evol. 2002, 49, 521–530. [Google Scholar] [CrossRef]
- Tomooka, N.; Naito, K.; Kaga, A.; Sakai, H.; Isemura, T.; Ogiso-Tanaka, E.; Iseki, K.; Takahashi, Y. Evolution, domestication and neo-domestication of the genus Vigna. Plant Genet. Resour. 2014, 12, 168–171. [Google Scholar] [CrossRef]
- Kumar, P.; Pal, M.; Joshi, R.; Sairam, R.K. Yield, growth and physiological responses of mung bean [Vigna radiata (L.) Wilczek] genotypes to waterlogging at vegetative stage. Physiol. Mol. Biol. Plants 2013, 19, 209–220. [Google Scholar] [CrossRef]
- Mustroph, A. Improving Flooding Tolerance of Crop Plants. Agronomy 2018, 8, 160. [Google Scholar] [CrossRef]
- Tomooka, N.; Kaga, A.; Isemura, T.; Vaughan, D. Vigna. In Wild Crop Relatives: Genomic and Breeding Resources: Legume Crops and Forages; Kole, C., Ed.; Springer: Berlin/Heidelberg, Germany, 2011; pp. 291–311. [Google Scholar] [CrossRef]
- Karuniawan, A.; Iswandi, A.; Kale, P.R.; Heinzemann, J.; Gruneberg, W.J. Vigna vexillata (L.) A. Rich. cultivated as a root crop in Bali and Timor. Genet. Resour. Crop Evol. 2006, 53, 213–217. [Google Scholar] [CrossRef]
- Lawn, R.J.; Watkinson, A.R. Habitats, morphological diversity, and distribution of the genus Vigna Savi in Australia. Aust. J. Agric. Res. 2002, 53, 1305–1316. [Google Scholar] [CrossRef]
- Roecklein, J.C.; Leung, P.S. A Profile of Economic Plants; Transaction Publishers: Piscataway/New Brunswick, NJ, USA, 1987. [Google Scholar]
- MIller, I.L.; Williams, W.T. Tolerance of some tropical legumes to six months of stimulated waterlogging. Trop. Glassland 1981, 15, 39–41. [Google Scholar]
- Phuphak, S.; Setter, T.L. Adverse effects of waterlogging on growth of lupin and field pea cultivars. In Proceedings of the 5th Australian Agronomy Conference 1989, the University of Western Australia, Perth, Western Australia, 24–29 September 1989. [Google Scholar]
- Fukao, T.; Barrera-Figueroa, B.E.; Juntawong, P.; Pena-Castro, J.M. Submergence and Waterlogging Stress in Plants: A Review Highlighting Research Opportunities and Understudied Aspects. Front. Plant Sci. 2019, 10, 340. [Google Scholar] [CrossRef]
- Kortz, A.; Hochholdinger, F.; Yu, P. Cell Type-Specific Transcriptomics of Lateral Root Formation and Plasticity. Front. Plant Sci. 2019, 10, 21. [Google Scholar] [CrossRef]
- Armstrong, W. Aeration in Higher Plants. In Advances in Botanical Research; Woolhouse, H.W., Ed.; Academic Press: Cambridge, MA, USA, 1980; Volume 7, pp. 225–332. [Google Scholar]
- Mustroph, A.; Zanetti, M.E.; Jang, C.J.; Holtan, H.E.; Repetti, P.P.; Galbraith, D.W.; Girke, T.; Bailey-Serres, J. Profiling translatomes of discrete cell populations resolves altered cellular priorities during hypoxia in Arabidopsis. Proc. Natl. Acad. Sci. USA 2009, 106, 18843–18848. [Google Scholar] [CrossRef]
- Sasidharan, R.; Mustroph, A.; Boonman, A.; Akman, M.; Ammerlaan, A.M.; Breit, T.; Schranz, M.E.; Voesenek, L.A.; van Tienderen, P.H. Root transcript profiling of two Rorippa species reveals gene clusters associated with extreme submergence tolerance. Plant Physiol. 2013, 163, 1277–1292. [Google Scholar] [CrossRef]
- Cai, X.T.; Xu, P.; Zhao, P.X.; Liu, R.; Yu, L.H.; Xiang, C.B. Arabidopsis ERF109 mediates cross-talk between jasmonic acid and auxin biosynthesis during lateral root formation. Nat. Commun. 2014, 5, 5833. [Google Scholar] [CrossRef]
- Negi, S.; Ivanchenko, M.G.; Muday, G.K. Ethylene regulates lateral root formation and auxin transport in Arabidopsis thaliana. Plant J. 2008, 55, 175–187. [Google Scholar] [CrossRef]
- Muday, G.K.; Rahman, A.; Binder, B.M. Auxin and ethylene: Collaborators or competitors? Trends Plant Sci. 2012, 17, 181–195. [Google Scholar] [CrossRef]
- Lewis, D.R.; Negi, S.; Sukumar, P.; Muday, G.K. Ethylene inhibits lateral root development, increases IAA transport and expression of PIN3 and PIN7 auxin efflux carriers. Development 2011, 138, 3485–3495. [Google Scholar] [CrossRef]
- Abbas, M.; Hernandez-Garcia, J.; Pollmann, S.; Samodelov, S.L.; Kolb, M.; Friml, J.; Hammes, U.Z.; Zurbriggen, M.D.; Blazquez, M.A.; Alabadi, D. Auxin methylation is required for differential growth in Arabidopsis. Proc. Natl. Acad. Sci. USA 2018, 115, 6864–6869. [Google Scholar] [CrossRef]
- Spartz, A.K.; Ren, H.; Park, M.Y.; Grandt, K.N.; Lee, S.H.; Murphy, A.S.; Sussman, M.R.; Overvoorde, P.J.; Gray, W.M. SAUR Inhibition of PP2C-D Phosphatases Activates Plasma Membrane H+-ATPases to Promote Cell Expansion in Arabidopsis. Plant Cell 2014, 26, 2129–2142. [Google Scholar] [CrossRef]
- Paponov, I.A.; Paponov, M.; Teale, W.; Menges, M.; Chakrabortee, S.; Murray, J.A.; Palme, K. Comprehensive transcriptome analysis of auxin responses in Arabidopsis. Mol. Plant 2008, 1, 321–337. [Google Scholar] [CrossRef]
- van Mourik, H.; van Dijk, A.D.J.; Stortenbeker, N.; Angenent, G.C.; Bemer, M. Divergent regulation of Arabidopsis SAUR genes: A focus on the SAUR10-clade. BMC Plant Biol. 2017, 17, 245. [Google Scholar] [CrossRef]
- Abel, S.; Nguyen, M.D.; Theologis, A. The PS-IAA4/5-like family of early auxin-inducible mRNAs in Arabidopsis thaliana. J. Mol. Biol. 1995, 251, 533–549. [Google Scholar] [CrossRef]
- Tiwari, S.B.; Wang, X.J.; Hagen, G.; Guilfoyle, T.J. AUX/IAA proteins are active repressors, and their stability and activity are modulated by auxin. Plant Cell 2001, 13, 2809–2822. [Google Scholar] [CrossRef]
- Ulmasov, T.; Murfett, J.; Hagen, G.; Guilfoyle, T.J. Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 1997, 9, 1963–1971. [Google Scholar] [CrossRef]
- Goh, T.; Joi, S.; Mimura, T.; Fukaki, H. The establishment of asymmetry in Arabidopsis lateral root founder cells is regulated by LBD16/ASL18 and related LBD/ASL proteins. Development 2012, 139, 883–893. [Google Scholar] [CrossRef]
- Cutler, S.; Ghassemian, M.; Bonetta, D.; Cooney, S.; McCourt, P. A protein farnesyl transferase involved in abscisic acid signal transduction in Arabidopsis. Science 1996, 273, 1239–1241. [Google Scholar] [CrossRef]
- Wang, Y.; Ying, J.; Kuzma, M.; Chalifoux, M.; Sample, A.; McArthur, C.; Uchacz, T.; Sarvas, C.; Wan, J.; Dennis, D.T.; et al. Molecular tailoring of farnesylation for plant drought tolerance and yield protection. Plant J. 2005, 43, 413–424. [Google Scholar] [CrossRef]
- Ruppel, N.J.; Kropp, K.N.; Davis, P.A.; Martin, A.E.; Luesse, D.R.; Hangarter, R.P. Mutations in GERANYLGERANYL DIPHOSPHATE SYNTHASE 1 affect chloroplast development in Arabidopsis thaliana (Brassicaceae). Am. J. Bot. 2013, 100, 2074–2084. [Google Scholar] [CrossRef]
- Mitchum, M.G.; Yamaguchi, S.; Hanada, A.; Kuwahara, A.; Yoshioka, Y.; Kato, T.; Tabata, S.; Kamiya, Y.; Sun, T.P. Distinct and overlapping roles of two gibberellin 3-oxidases in Arabidopsis development. Plant J. 2006, 45, 804–818. [Google Scholar] [CrossRef]
- Chaumont, F.; Tyerman, S.D. Aquaporins: Highly regulated channels controlling plant water relations. Plant Physiol. 2014, 164, 1600–1618. [Google Scholar] [CrossRef]
- Hachez, C.; Laloux, T.; Reinhardt, H.; Cavez, D.; Degand, H.; Grefen, C.; De Rycke, R.; Inze, D.; Blatt, M.R.; Russinova, E.; et al. Arabidopsis SNAREs SYP61 and SYP121 coordinate the trafficking of plasma membrane aquaporin PIP2;7 to modulate the cell membrane water permeability. Plant Cell 2014, 26, 3132–3147. [Google Scholar] [CrossRef]
- Ridley, B.L.; O’Neill, M.A.; Mohnen, D.A. Pectins: Structure, biosynthesis, and oligogalacturonide-related signaling. Phytochemistry 2001, 57, 929–967. [Google Scholar] [CrossRef]
- Sobry, S.; Havelange, A.; Van Cutsem, P. Immunocytochemistry of pectins in shoot apical meristems: Consequences for intercellular adhesion. Protoplasma 2005, 225, 15–22. [Google Scholar] [CrossRef]
- Wen, F.; Zhu, Y.; Hawes, M.C. Effect of pectin methylesterase gene expression on pea root development. Plant Cell 1999, 11, 1129–1140. [Google Scholar] [CrossRef]
- Jin, J.; Hewezi, T.; Baum, T.J. Arabidopsis peroxidase AtPRX53 influences cell elongation and susceptibility to Heterodera schachtii. Plant Signal. Behav. 2011, 6, 1778–1786. [Google Scholar] [CrossRef]
- Johansen, D.A. Plant Microtechnique; McGraw-Hill Book, Co.: New York, NY, USA, 1940. [Google Scholar]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef]
- Sirikhachornkit, A.; Suttangkakul, A.; Vuttipongchaikij, S.; Juntawong, P. De novo transcriptome analysis and gene expression profiling of an oleaginous microalga Scenedesmus acutus TISTR8540 during nitrogen deprivation-induced lipid accumulation. Sci. Rep. 2018, 8, 3668. [Google Scholar] [CrossRef]
- Horan, K.; Jang, C.; Bailey-Serres, J.; Mittler, R.; Shelton, C.; Harper, J.F.; Zhu, J.K.; Cushman, J.C.; Gollery, M.; Girke, T. Annotating genes of known and unknown function by large-scale coexpression analysis. Plant Physiol. 2008, 147, 41–57. [Google Scholar] [CrossRef]
- Usadel, B.; Poree, F.; Nagel, A.; Lohse, M.; Czedik-Eysenberg, A.; Stitt, M. A guide to using MapMan to visualize and compare Omics data in plants: A case study in the crop species, Maize. Plant Cell Environ. 2009, 32, 1211–1229. [Google Scholar] [CrossRef]
- Xu, L.; Dong, Z.; Fang, L.; Luo, Y.; Wei, Z.; Guo, H.; Zhang, G.; Gu, Y.Q.; Coleman-Derr, D.; Xia, Q.; et al. OrthoVenn2: A web server for whole-genome comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res. 2019, 47, W52–W58. [Google Scholar] [CrossRef]
- Liu, Y.; Wei, G.; Xia, Y.; Liu, X.; Tang, J.; Lu, Y.; Lan, H.; Zhang, S.; Li, C.; Cao, M. Comparative transcriptome analysis reveals that tricarboxylic acid cycle-related genes are associated with maize CMS-C fertility restoration. BMC Plant Biol. 2018, 18, 190. [Google Scholar] [CrossRef]
- Zhao, H.; Basu, U.; Kebede, B.; Qu, C.; Li, J.; Rahman, H. Fine mapping of the major QTL for seed coat color in Brassica rapa var. Yellow Sarson by use of NIL populations and transcriptome sequencing for identification of the candidate genes. PLoS ONE 2019, 14, e0209982. [Google Scholar] [CrossRef]


© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Butsayawarapat, P.; Juntawong, P.; Khamsuk, O.; Somta, P. Comparative Transcriptome Analysis of Waterlogging-Sensitive and Tolerant Zombi Pea (Vigna vexillata) Reveals Energy Conservation and Root Plasticity Controlling Waterlogging Tolerance. Plants 2019, 8, 264. https://doi.org/10.3390/plants8080264
Butsayawarapat P, Juntawong P, Khamsuk O, Somta P. Comparative Transcriptome Analysis of Waterlogging-Sensitive and Tolerant Zombi Pea (Vigna vexillata) Reveals Energy Conservation and Root Plasticity Controlling Waterlogging Tolerance. Plants. 2019; 8(8):264. https://doi.org/10.3390/plants8080264
Chicago/Turabian StyleButsayawarapat, Pimprapai, Piyada Juntawong, Ornusa Khamsuk, and Prakit Somta. 2019. "Comparative Transcriptome Analysis of Waterlogging-Sensitive and Tolerant Zombi Pea (Vigna vexillata) Reveals Energy Conservation and Root Plasticity Controlling Waterlogging Tolerance" Plants 8, no. 8: 264. https://doi.org/10.3390/plants8080264
APA StyleButsayawarapat, P., Juntawong, P., Khamsuk, O., & Somta, P. (2019). Comparative Transcriptome Analysis of Waterlogging-Sensitive and Tolerant Zombi Pea (Vigna vexillata) Reveals Energy Conservation and Root Plasticity Controlling Waterlogging Tolerance. Plants, 8(8), 264. https://doi.org/10.3390/plants8080264




