Differences among the Anthocyanin Accumulation Patterns and Related Gene Expression Levels in Red Pears
Abstract
1. Introduction
2. Results
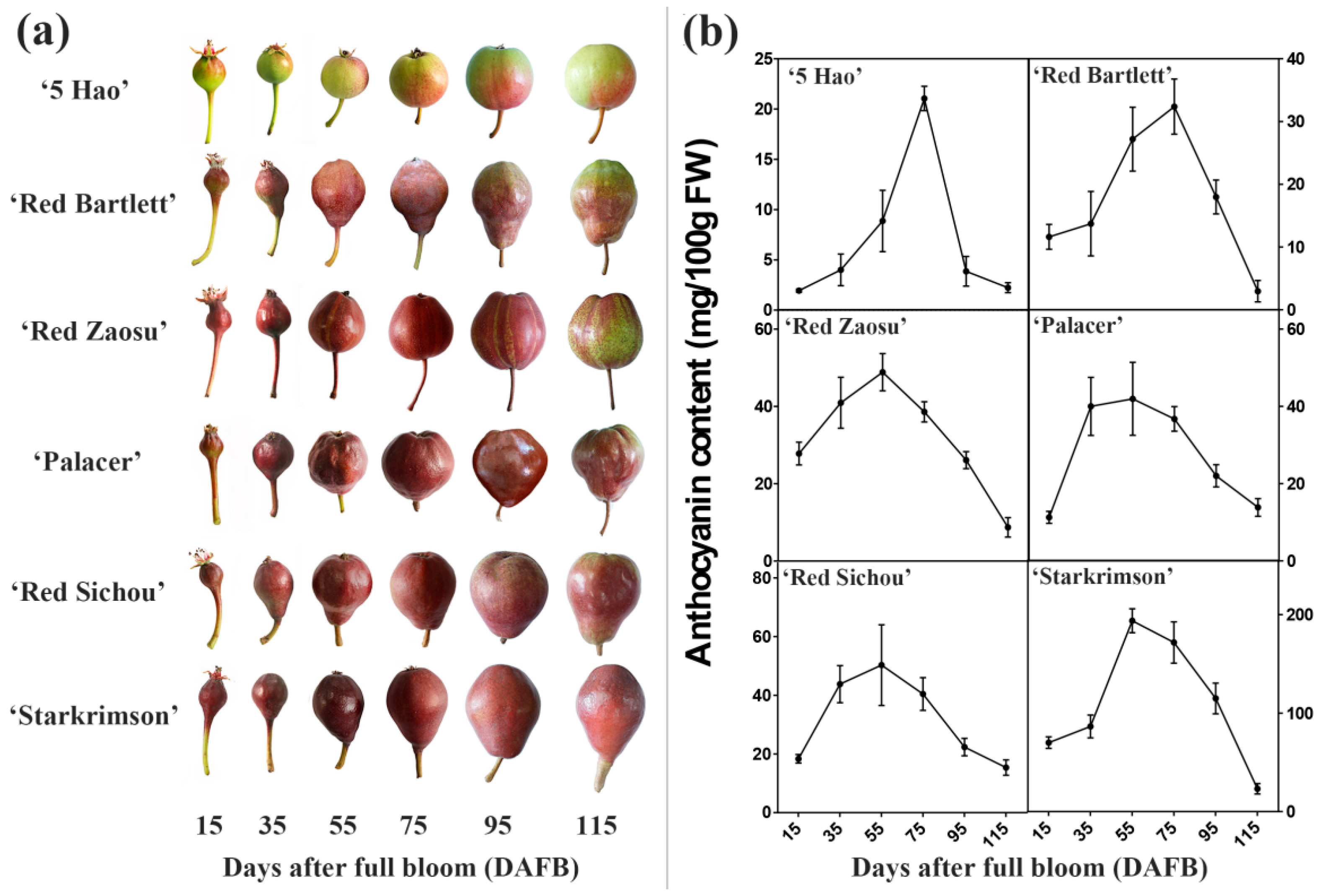
2.1. Total Anthocyanin Accumulation in Different Pear Cultivars
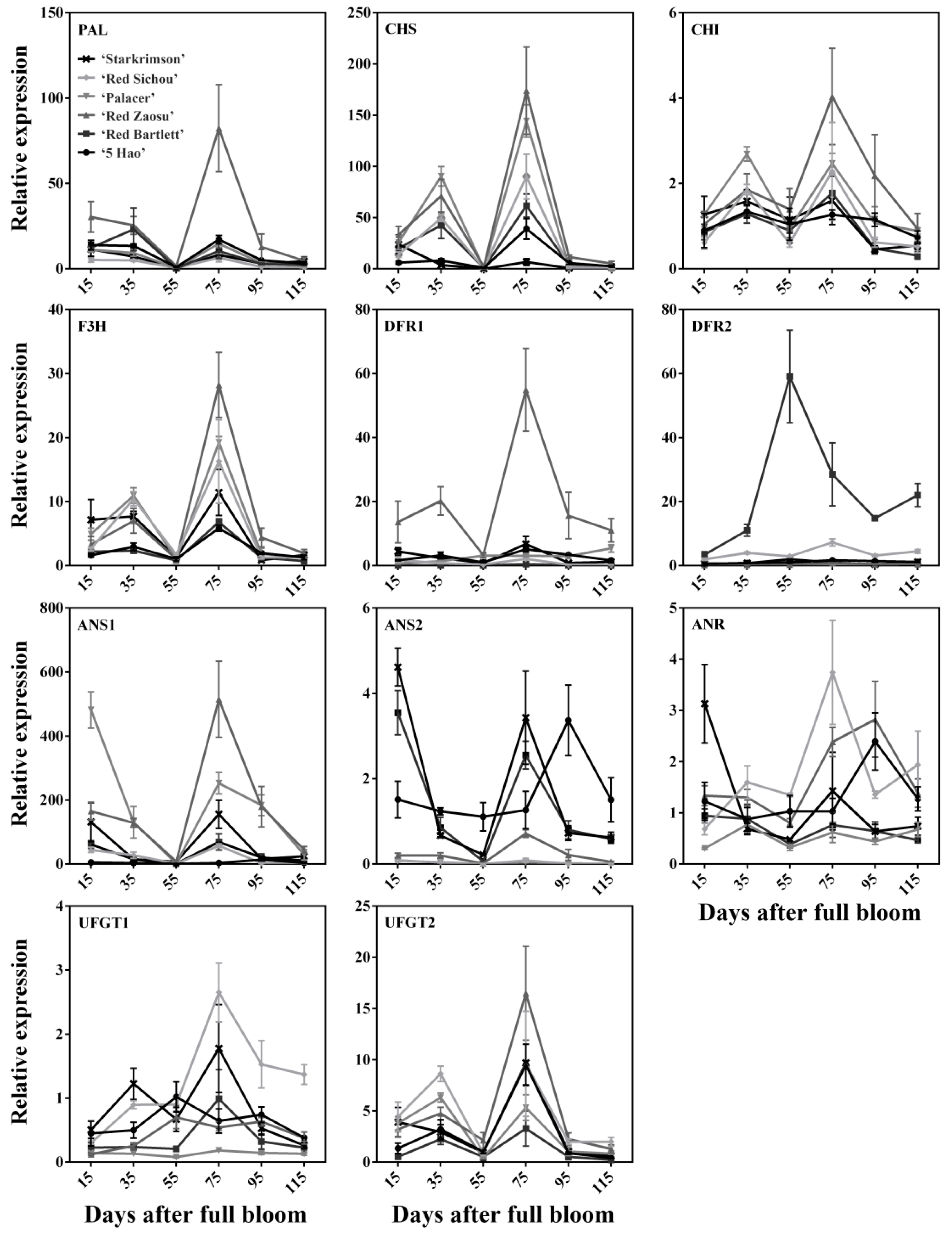
2.2. Expression Levels of Anthocyanin Synthesis-Related Structural Genes
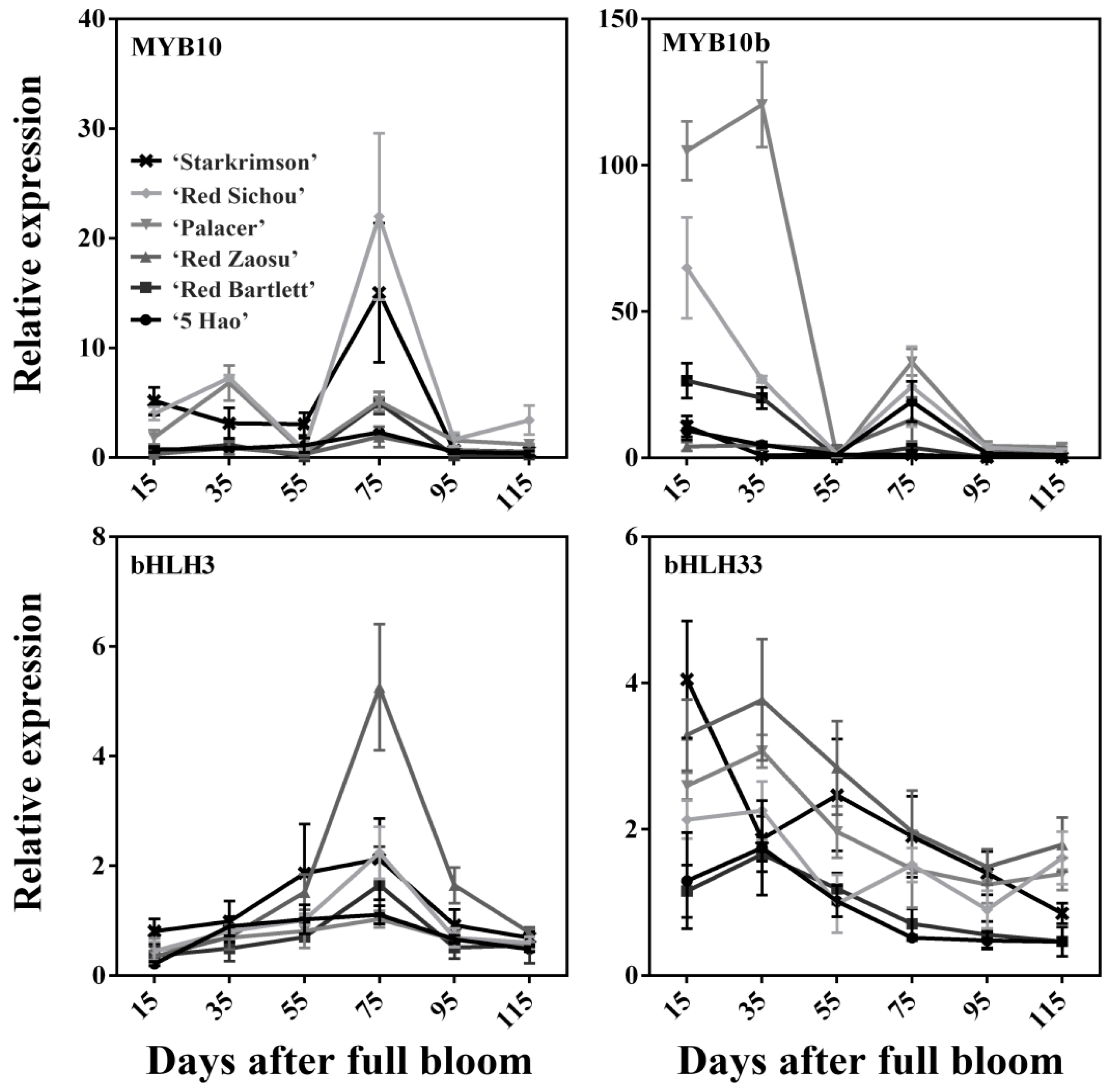
2.3. Expression Levels of Anthocyanin Synthesis-Related Regulatory Genes
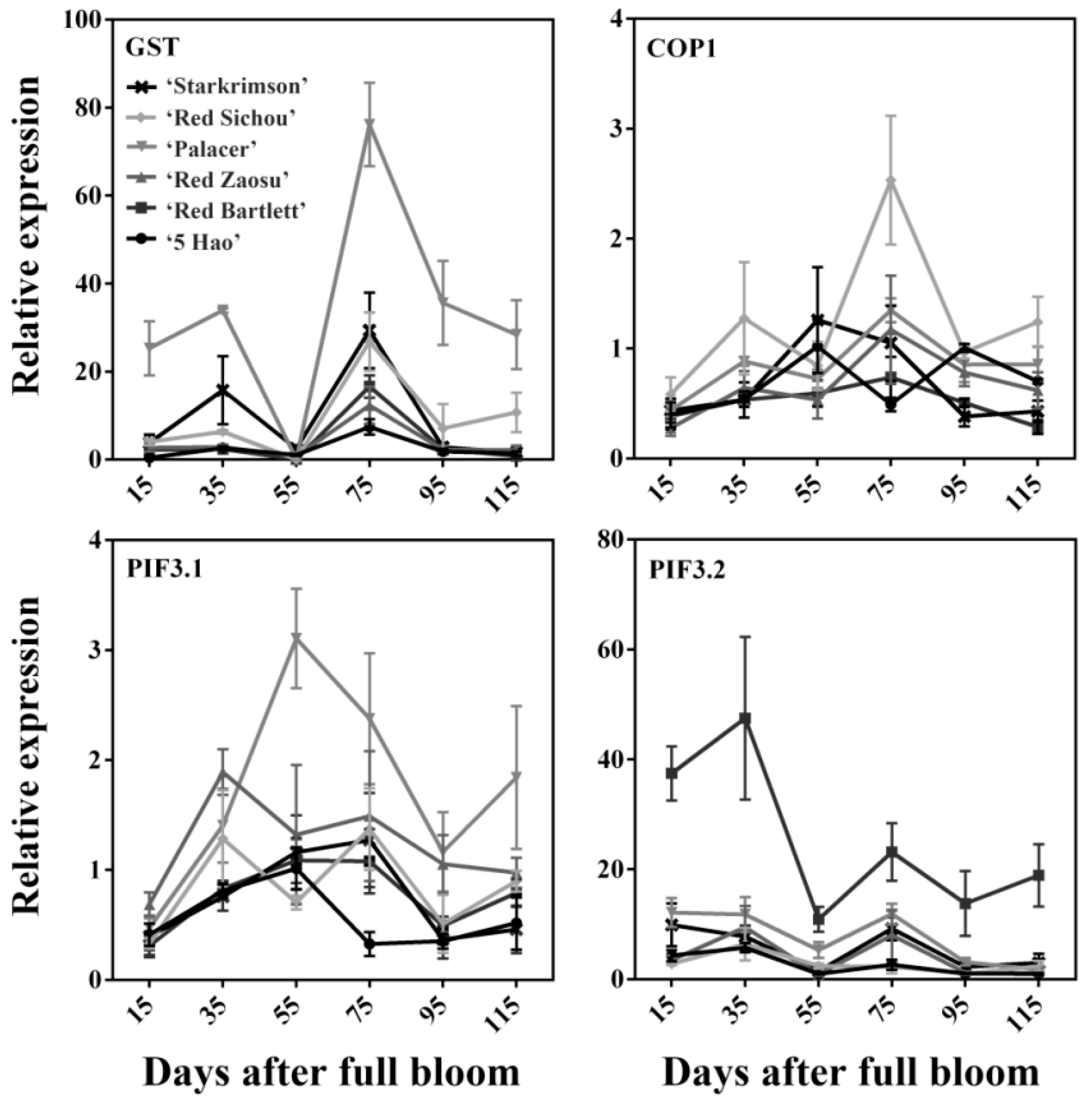
2.4. Expression Levels of Anthocyanins Synthesis-Related Transporter and Light-Responsive Genes
2.5. Correlation Analysis of Anthocyanin Contents and Gene Expression Levels
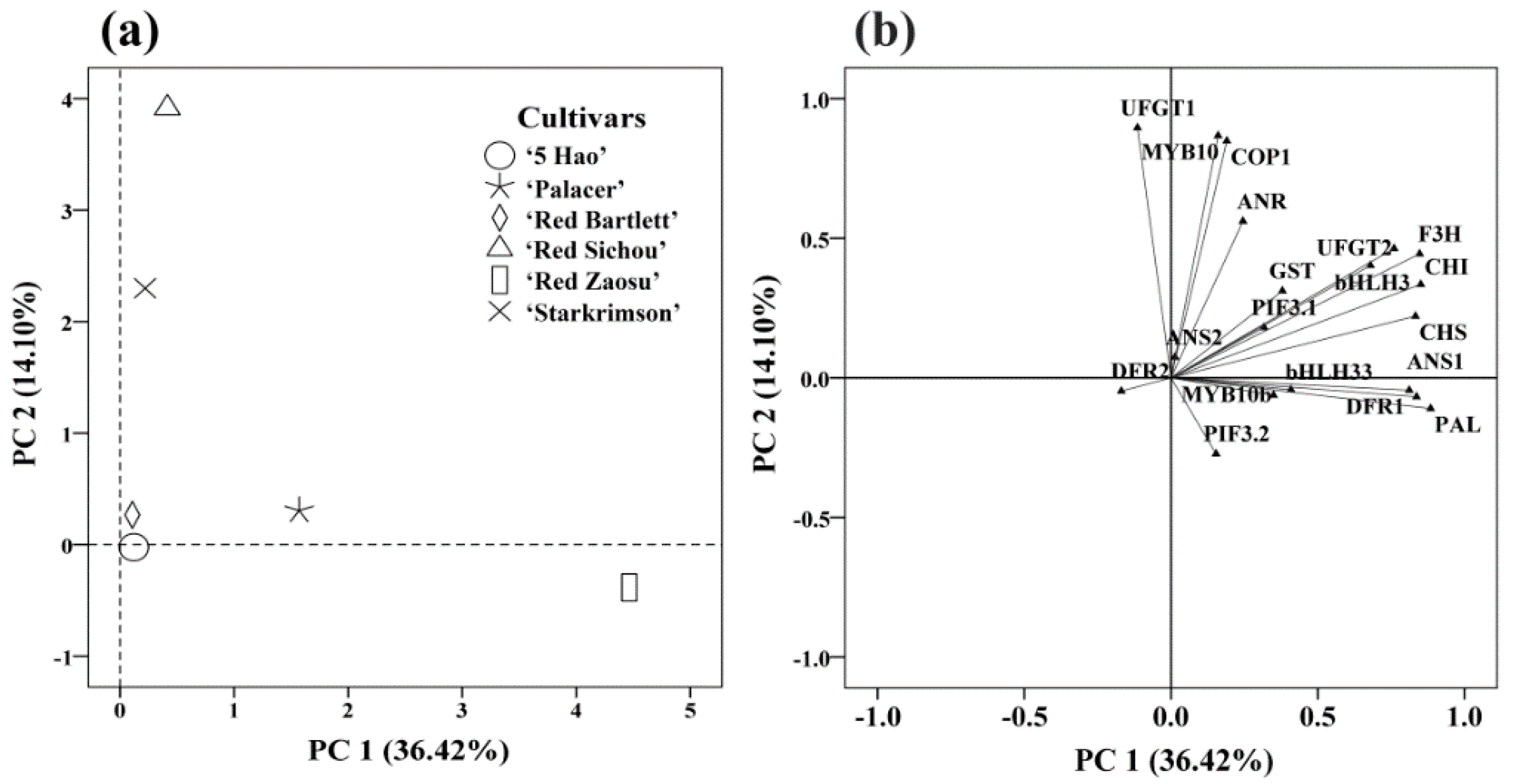
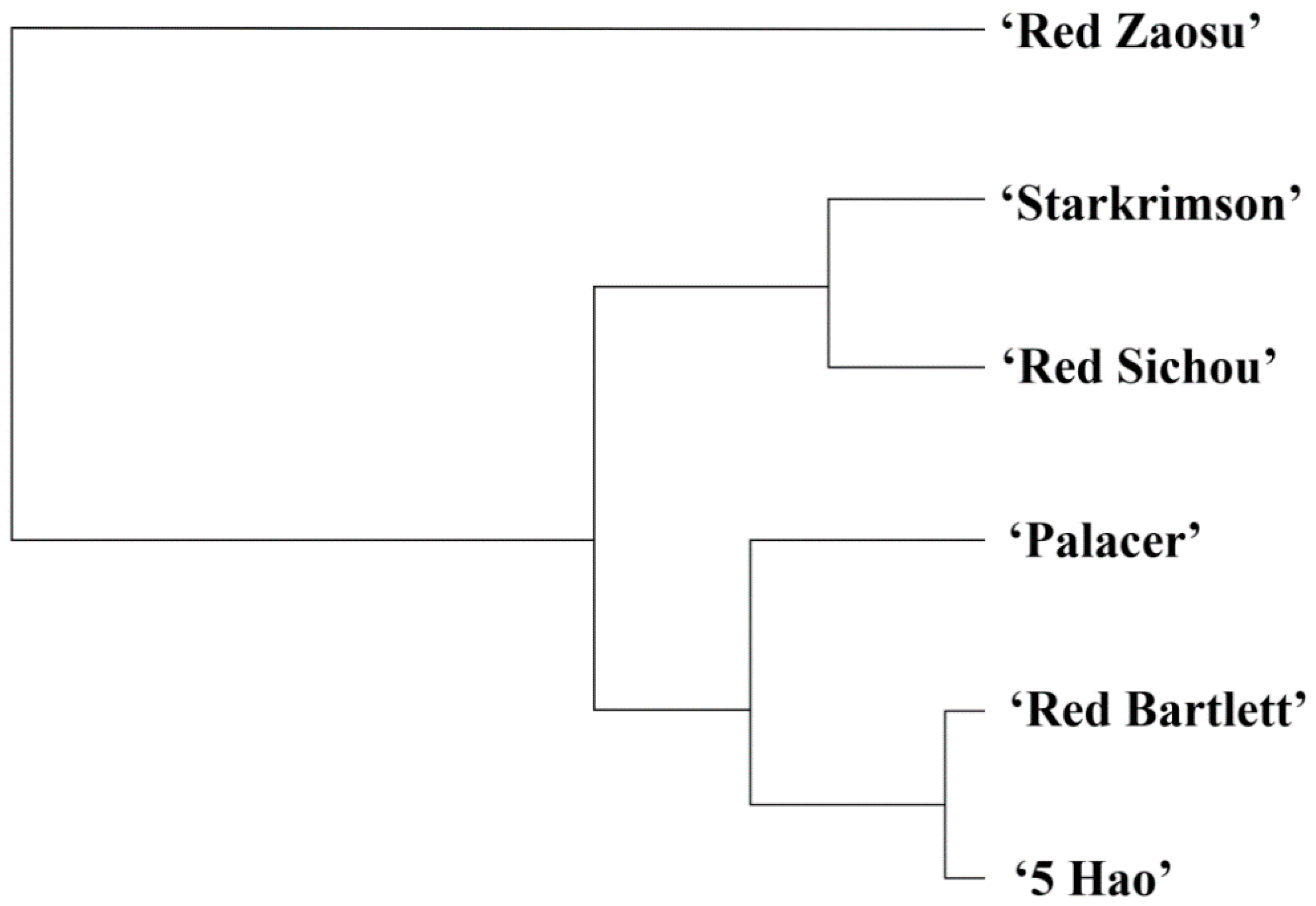
2.6. Classification and Analysis of Important Genes in the Six Red Pear Cultivars
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Experimental Treatments
4.2. Anthocyanin Analysis
4.3. Quantification of Genes Expression
4.4. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Tao, P.; Shu, Q.; Wang, J.J.; Zhang, W.B. Present situation and prospect of research and utilization on red pear germplasm resources. Southwest China J. Agric. Sci. 2004, 17, 409–412. [Google Scholar]
- Dussi, M.C.; Sugar, D.; Wrolstad, R.E. Characterizing and quantifying anthocyanins in red pears and the effect of light quality on fruit color. J. Am. Soc. Hortic. Sci. 1995, 120, 785–789. [Google Scholar] [CrossRef]
- Dayton, D.F. The pattern and inheritance of anthocyanin distribution in red pears. J. Am. Soc. Hortic. Sci. 1996, 89, 110–116. [Google Scholar]
- Xiao, C.; Jiaming, L.I.; Yao, G.F.; Liu, J.; Hu, H.J. Characteristics of components and contents of anthocyanin in peel of red-skinned pear fruits from different species. J. Nanjing Agric. Univ. 2004, 37, 60–66. [Google Scholar]
- Holton, T.A.; Cornish, E.C. Genetics and Biochemistry of Anthocyanin Biosynthesis. Plant Cell 1995, 7, 1071–1083. [Google Scholar] [CrossRef] [PubMed]
- Boss, P.K.; Davies, C.; Robinson, S.P. Analysis of the Expression of Anthocyanin Pathway Genes in Developing Vitis vinifera L. cv Shiraz Grape Berries and the Implications for Pathway Regulation. Plant Physiol. 1996, 111, 1059–1066. [Google Scholar] [CrossRef] [PubMed]
- Grotewold, E. The genetics and biochemistry of floral pigments. Annu. Rev. Plant Biol. 2006, 57, 761–780. [Google Scholar] [CrossRef]
- Bogs, J.; Ebadi, A.; McDavid, D.; Robinson, S.P. Identification of the flavonoid hydroxylases from grapevine and their regulation during fruit development. Plant Physiol. 2006, 140, 279–291. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.P.; Vimolmangkang, S.; Soria-Guerra, R.E.; Rosales-Mendoza, S.; Zheng, D.M.; Lygin, A.V.; Korban, S.S. Ectopic expression of apple F3’H genes contributes to anthocyanin accumulation in the Arabidopsis tt7 mutant grown under nitrogen stress. Plant Physiol. 2010, 153, 806–820. [Google Scholar] [CrossRef] [PubMed]
- Allan, A.C.; Hellens, R.P.; Laing, W.A. MYB transcription factors that colour our fruit. Trends Plant Sci. 2008, 13, 99–102. [Google Scholar] [CrossRef]
- Feller, A.; Machemer, K.; Braun, E.L.; Grotewold, E. Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J. 2011, 66, 94. [Google Scholar] [CrossRef]
- Lai, Y.; Huanxiu, L.I.; Yamagishi, M. A review of target gene specificity of flavonoid R2R3-MYB transcription factors and a discussion of factors contributing to the target gene selectivity. Front. Biol. 2013, 8, 577–598. [Google Scholar] [CrossRef]
- Starkevič, P.; Paukštytė, J.; Kazanavičiūtė, V.; Denkovskienė, E.; Stanys, V.; Bendokas, V. Expression and anthocyanin biosynthesis-modulating potential of sweet cherry (prunus avium L.) MYB10 and bHLH genes. PLoS ONE 2015, 10, e0126991. [Google Scholar]
- Cutuli, B.; Lemanski, C.; Fourquet, A.; Lafontan, B.D.; Giard, S.; Lancrenon, S. Environmental significance of anthocyanins in plant stress responses. Photochem. Photobiol. 2010, 70, 1–9. [Google Scholar]
- Jaakola, L. New insights into the regulation of anthocyanin biosynthesis in fruit. Trends Plant Sci. 2013, 18, 477–483. [Google Scholar] [CrossRef] [PubMed]
- Shaked-Sachray, L.; Weiss, D.; Reuveni, M.; Nissim-Levi, A.; Oren-Shamir, M. Increased anthocyanin accumulation in aster flowers at elevated temperatures due to magnesium treatment. Physiol. Plant. 2002, 114, 559–565. [Google Scholar] [CrossRef] [PubMed]
- Oren-Shamir, M. Does anthocyanin degradation play a significant role in determining pigment concentration in plants? Plant Sci. 2009, 177, 310–316. [Google Scholar] [CrossRef]
- Vaknin, H.; Bar-Akiva, A.; Ovadia, R.; Nissim-Levi, A.; Forer, I.; Weiss, D.; Oren-Shamir, M. Active anthocyanin degradation in brunfelsia calycina(yesterday–today–tomorrow) flowers. Planta 2005, 222, 19–26. [Google Scholar] [CrossRef] [PubMed]
- Zipor, G.; Patrícia, D.; Inês, C.; Shahar, L.; Ovadia, R.; Teper-Bamnolker, P.; Eshel, D.; Levin, Y.; Doron-Faigenboim, A.; Sottomayor, M.; et al. In planta anthocyanin degradation by a vacuolar class III peroxidase in brunfelsia calycina flowers. New Phytol. 2015, 205, 653–665. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.Y.; Mao, K.; Zhao, C.; Zhao, X.Y.; Zhang, H.L.; Shu, H.R.; Hao, Y.J. MdCOP1 Ubiquitin E3 Ligases Interact with MdMYB1 to Regulate Light-Induced Anthocyanin Biosynthesis and Red Fruit Coloration in Apple. Plant Physiol. 2012, 160, 1011–1022. [Google Scholar] [CrossRef]
- Toledoortiz, G.; Huq, E.; Quail, P.H. The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 2003, 15, 1749–1770. [Google Scholar] [CrossRef]
- Shin, J.; Park, E.; Choi, G. PIF3 regulates anthocyanin biosynthesis in an HY5-dependent manner with both factors directly binding anthocyanin biosynthetic gene promoters in Arabidopsis. Plant J. 2007, 49, 981–994. [Google Scholar] [CrossRef] [PubMed]
- Marr, K.A.; Alfenito, M.R.; Lloyd, A.M.; Walbot, V. A glutathione S-transferase involved in vacuolar transfer encoded by the maize gene Bronze-2. Nature 1995, 375, 397–400. [Google Scholar] [CrossRef] [PubMed]
- Mueller, L.A.; Goodman, C.D.; Silady, R.A.; Walbot, V. AN9, a petunia glutathione S-transferase required for anthocyanin sequestration, is a flavonoid-binding protein. Plant Physiol. 2000, 123, 1561–1570. [Google Scholar] [CrossRef] [PubMed]
- Kitamura, S.; Shikazono, N.; Tanaka, A. TRANSPARENT TESTA 19 is involved in the accumulation of both anthocyanins and proanthocyanidins in Arabidopsis. Plant J. 2004, 37, 104–114. [Google Scholar] [CrossRef] [PubMed]
- Conn, S.; Curtin, C.; Bezier, A.; Franco, C.; Zhang, W. Purification, molecular cloning, and characterization of glutathione S-transferases (GSTs) from pigmented Vitis vinifera L. cell suspension cultures as putative anthocyanin transport proteins. J. Exp. Bot. 2008, 59, 3621–3634. [Google Scholar] [CrossRef]
- Kitamura, S.; Akita, Y.; Ishizaka, H.; Narumi, I.; Tanaka, A. Molecular characterization of an anthocyanin-related glutathione S-transferase gene in cyclamen. J. Plant Physiol. 2012, 169, 636–642. [Google Scholar] [CrossRef]
- Fischer, T.C.; Gosch, C.; Pfeiffer, J.; Halbwirth, H.; Halle, C.; Stich, K.; Forkmann, G. Flavonoid genes of pear (Pyrus communis). Trees 2007, 21, 521–529. [Google Scholar] [CrossRef]
- Yu, B.; Zhang, D.; Huang, C.; Qian, M.J.; Zheng, X.Y.; Teng, Y.W.; Su, J.; Shu, Q. Isolation of anthocyanin biosynthetic genes in red Chinese sand pear (Pyrus pyrifolia Nakai) and their expression as affected by organ/tissue, cultivar, bagging and fruit side. Sci. Hortic. 2012, 136, 29–37. [Google Scholar] [CrossRef]
- Yang, Y.N.; Yao, G.F.; Zheng, D.M.; Zhang, S.L.; Wang, C.; Zhang, M.Y.; Wu, J. Expression differences of anthocyanin biosynthesis genes reveal regulation patterns for red pear coloration. Plant Cell Rep. 2015, 34, 189–198. [Google Scholar] [CrossRef]
- Steyn, W.J.; Holcroft, D.M.; Wand, S.J.E.; Jacobs, G. Regulation of pear color development in relation to activity of flavonoid enzymes. J. Am. Soc. Hortic. Sci. 2004, 129, 6–12. [Google Scholar] [CrossRef]
- Feng, S.Q.; Wang, Y.L.; Yang, S.; Xu, Y.T.; Chen, X.S. Anthocyanin biosynthesis in pears is regulated by a R2R3-MYB transcription factor PyMYB10. Planta 2010, 232, 245–255. [Google Scholar] [CrossRef]
- Wang, Z.G.; Meng, D.; Wang, A.D.; Li, T.L.; Jiang, S.L.; Cong, P.H.; Li, T. The methylation of the PcMYB10 promoter is associated with green-skinned sport in Max Red Bartlett pear. Plant Physiol. 2013, 162, 885–896. [Google Scholar] [CrossRef]
- Zhai, R.; Wang, Z.M.; Zhang, S.W.; Meng, G.; Song, L.Y.; Wang, Z.G.; Li, P.; Ma, F.; Xu, L. Two MYB transcription factors regulate flavonoid biosynthesis in pear fruit (Pyrus bretschneideri Rehd.). J. Exp. Bot. 2016, 67, 1275. [Google Scholar] [CrossRef]
- Wang, Z.G.; Du, H.; Zhai, R.; Song, L.Y.; Ma, F.W.; Xu, L.F. Transcriptome Analysis Reveals Candidate Genes Related to Color Fading of Red Bartlett (Pyrus communis L.). Front. Plant Sci. 2017, 8, e0142112. [Google Scholar] [CrossRef]
- Kobayashi, S.; Ishimaru, M.; Ding, C.K.; Yakushiji, H.; Goto, N. Comparison of UDP-glucose: Flavonoid 3-O-glucosyltransferase (UFGT) gene sequences between white grapes (Vitis vinifera) and their sports with red skin. Plant Sci. 2001, 163, 543–550. [Google Scholar] [CrossRef]
- Kim, S.H.; Lee, J.R.; Hong, S.T.; Yoo, Y.K.; An, G.; Kim, S.R. Molecular cloning and analysis of anthocyanin biosynthesis genes preferentially expressed in apple skin. Plant Sci. 2003, 165, 403–413. [Google Scholar] [CrossRef]
- Zhao, Z.C.; Hu, G.B.; Hu, F.C.; Wang, H.C.; Yang, Z.Y.; Lai, B. The udp glucose: Flavonoid-3-o-glucosyltransferase (ufgt) gene regulates anthocyanin biosynthesis in litchi (litchi chinesissonn.) during fruit coloration. Mol. Biol. Rep. 2012, 39, 6409–6415. [Google Scholar] [CrossRef]
- Pierantoni, L.; Dondini, L.; De Franceschi, P.; Musacchi, S.; Winkel, B.S.J.; Sansavini, S. Mapping of an anthocyanin-regulating MYB transcription factor and its expression in red and green pear, Pyrus communis. Plant Physiol. Biochem. 2010, 48, 1020. [Google Scholar] [CrossRef]
- Yang, Y.N.; Zhao, G.; Yue, W.Q.; Zhang, S.L.; Gu, C.; Wu, J. Molecular cloning and gene expression differences of the anthocyanin biosynthesis-related genes in the red/green skin color mutant of pear (Pyrus communis L.). Tree Genet. Genomes 2013, 9, 1351–1360. [Google Scholar] [CrossRef]
- Zhu, Y.F.; Su, J.; Yao, G.F.; Liu, H.N.; Gu, C.; Qin, M.F.; Bai, B.; Cai, S.S.; Wang, G.M.; Wang, R.Z.; et al. Different light-response patterns of coloration and related gene expression in red pears (Pyrus L.). Sci. Hortic. 2018, 229, 240–251. [Google Scholar] [CrossRef]
- Zhai, R.; Liu, X.T.; Feng, W.T.; Chen, S.S.; Xu, L.F.; Wang, Z.G.; Zhang, J.L.; Li, P.M.; Ma, F.W. Different biosynthesis patterns among flavonoid 3-glycosides with distinct effects on accumulation of other flavonoid metabolites in pears (Pyrus bretschneideri Rehd.). PLoS ONE 2014, 9, e91945. [Google Scholar] [CrossRef]
- Qian, M.J.; Sun, Y.W.; Allan, A.C.; Teng, Y.W.; Zhang, D. The red sport of ‘Zaosu’ pear and its red-striped pigmentation pattern are associated with demethylation of the PyMYB10 promoter. Phytochemistry 2014, 107, 16–23. [Google Scholar] [CrossRef]
- Huang, C.H.; Ge, C.L.; Zhang, X.H.; Wu, H.; Qu, X.Y.; Xu, X.B. Expression analysis of structural genes related to anthocyanin synthesis in a mutant of ‘Hongyang’ kiwifruit. J. Fruit Sci. 2014, 31, 169–174. [Google Scholar]
- Chagné, D.; Carlisle, C.M.; Blond, C.; Volz, R.K.; Whitworth, C.J.; Oraguzie, N.C.; Crowhurst, R.N.; Allan, A.C.; Espley, R.V.; Hellens, R.P. Mapping a candidate gene (MdMYB10) for red flesh and foliage colour in apple. BMC Genom. 2007, 8, 212. [Google Scholar] [CrossRef] [PubMed]
- Kui, L.W.; Bolitho, K.; Grafton, K.; Kortstee, A.; Karunairetnam, S.; McGhie, T.K.; Espley, R.V.; Hellens, R.P.; Allan, A.C. An R2R3 MYB transcription factor associated with regulation of the anthocyanin biosynthetic pathway in Rosaceae. BMC Plant Biol. 2010, 10, 50. [Google Scholar]
- Rahim, M.A.; Busatto, N.; Trainotti, L. Regulation of anthocyanin biosynthesis in peach fruits. Planta 2014, 240, 913–929. [Google Scholar] [CrossRef] [PubMed]
- Espley, R.V.; Hellens, R.P.; Putterill, J.; Stevenson, D.E.; Kutty-Amma, S.; Allan, A.C. Colouration in apple fruit is due to the activity of the MYB transcription factor MdMYB10. Plant J. 2007, 49, 414. [Google Scholar] [CrossRef] [PubMed]
- Mori, K.; Sugaya, S.; Gemma, H. Decreased anthocyanin biosynthesis in grape berries grown under elevated night temperature condition. Sci Hortic. 2005, 105, 319–330. [Google Scholar] [CrossRef]
- Li, T.; Jia, K.P.; Lian, H.L.; Yang, X.; Li, L.; Yang, H.Q. Jasmonic acid enhancement of anthocyanin accumulation is dependent on phytochrome a signaling pathway under far-red light in Arabidopsis. Biochem. Biophys. Res. Commun. 2014, 454, 78–83. [Google Scholar] [CrossRef]
- Maier, A.; Schrader, A.; Kokkelink, L.; Falke, C.; Welter, B.; Iniesto, E.; Rubio, V.; Uhrig, J.F.; Hülskamp, M.; Hoecker, U. Light and the E3 ubiquitin ligase COP1/SPA control the protein stability of the MYB transcription factors PAP1 and PAP2 involved in anthocyanin accumulation in Arabidopsis. Plant J. 2013, 74, 638–651. [Google Scholar] [CrossRef]
- Kimura, T.; Sawamura, Y.; Kotobuki, K.; Matsuta, N.; Hayashi, T.; Ban, Y.; Yamamoto, T. Parentage Analysis in Pear Cultivars Characterized by SSR Markers. J. Jpn. Soc. Hortic. Sci. 2003, 72, 182–189. [Google Scholar] [CrossRef]
- Giusti, M.M.; Wrolstad, R.E. Characterization and Measurement of Anthocyanins by UV-Visible Spectroscopy. In Current Protocols in Food Analytical Chemistry; John Wiley & Sons, Inc.: New York, NY, USA, 2001. [Google Scholar]
- Reid, K.E.; Olsson, N.; Schlosser, J.; Peng, F.; Lund, S.T. An optimized grapevine RNA isolation procedure and statistical determination of reference genes for real-time RT-PCR during berry development. BMC Plant Biol. 2006, 6, 27. [Google Scholar] [CrossRef] [PubMed]
| Anthocyanin | 5 Hao | Red Bartlett | Red Zaosu | Palacer | Red Sichou | Starkrimson |
|---|---|---|---|---|---|---|
| PAL | 0.590 * | 0.09 | 0.534 * | 0.178 | 0.571 * | 0.058 |
| CHS | 0.962 ** | 0.779 ** | 0.639 * | 0.611 * | 0.861 ** | −0.102 |
| CHI | 0.382 | 0.759 ** | 0.473 | 0.692 ** | 0.830 ** | 0.416 |
| F3H | 0.943 ** | 0.882 ** | 0.504 | 0.556 * | 0.850 ** | 0.557 * |
| DFR1 | 0.794 ** | 0.759 ** | 0.499 | −0.003 | 0.358 | 0.586 * |
| DFR2 | 0.578 * | 0.451 | 0.005 | 0.033 | 0.648 ** | −0.137 |
| ANS1 | −0.278 | 0.578 * | 0.429 | −0.47 | 0.492 | 0.458 |
| ANS2 | −0.221 | 0.29 | 0.553 * | −0.411 | 0.336 | 0.257 |
| ANR | −0.28 | 0.227 | 0.065 | 0.590 * | 0.523 * | −0.049 |
| UFGT1 | 0.301 | 0.792 ** | −0.073 | 0.231 | 0.449 | 0.754 ** |
| UFGT2 | 0.949 ** | 0.725 ** | 0.513 | 0.459 | 0.775 ** | 0.767 ** |
| MYB10b | 0.809 ** | −0.169 | 0.358 | 0.017 | −0.065 | −0.222 |
| MYB10 | 0.953 ** | 0.875 ** | 0.392 | 0.745 ** | 0.679 ** | 0.725 ** |
| bHLH3 | 0.744 ** | 0.747 ** | 0.311 | 0.639 * | 0.680 ** | 0.751 ** |
| bHLH33 | −0.273 | −0.055 | 0.47 | 0.023 | 0.219 | −0.009 |
| GST | 0.948 ** | 0.908 ** | 0.399 | 0.477 | 0.513 | 0.773 ** |
| COP1 | −0.299 | 0.860 ** | 0.071 | 0.670 ** | 0.278 | 0.716 ** |
| PIF3.1 | −0.294 | 0.488 | 0.684 ** | 0.379 | 0.791 ** | 0.692 ** |
| PIF3.2 | −0.046 | −0.04 | 0.783 ** | 0.143 | 0.481 | 0.297 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, M.; Liu, J.; Song, L.; Li, X.; Cong, L.; Yue, R.; Yang, C.; Liu, Z.; Xu, L.; Wang, Z. Differences among the Anthocyanin Accumulation Patterns and Related Gene Expression Levels in Red Pears. Plants 2019, 8, 100. https://doi.org/10.3390/plants8040100
Wu M, Liu J, Song L, Li X, Cong L, Yue R, Yang C, Liu Z, Xu L, Wang Z. Differences among the Anthocyanin Accumulation Patterns and Related Gene Expression Levels in Red Pears. Plants. 2019; 8(4):100. https://doi.org/10.3390/plants8040100
Chicago/Turabian StyleWu, Meng, Jianlong Liu, Linyan Song, Xieyu Li, Liu Cong, Rongrong Yue, Chengquan Yang, Zhuo Liu, Lingfei Xu, and Zhigang Wang. 2019. "Differences among the Anthocyanin Accumulation Patterns and Related Gene Expression Levels in Red Pears" Plants 8, no. 4: 100. https://doi.org/10.3390/plants8040100
APA StyleWu, M., Liu, J., Song, L., Li, X., Cong, L., Yue, R., Yang, C., Liu, Z., Xu, L., & Wang, Z. (2019). Differences among the Anthocyanin Accumulation Patterns and Related Gene Expression Levels in Red Pears. Plants, 8(4), 100. https://doi.org/10.3390/plants8040100




