A Spotlight on Rad52 in Cyanidiophytina (Rhodophyta): A Relic in Algal Heritage
Abstract
1. Introduction
2. Results and Discussion
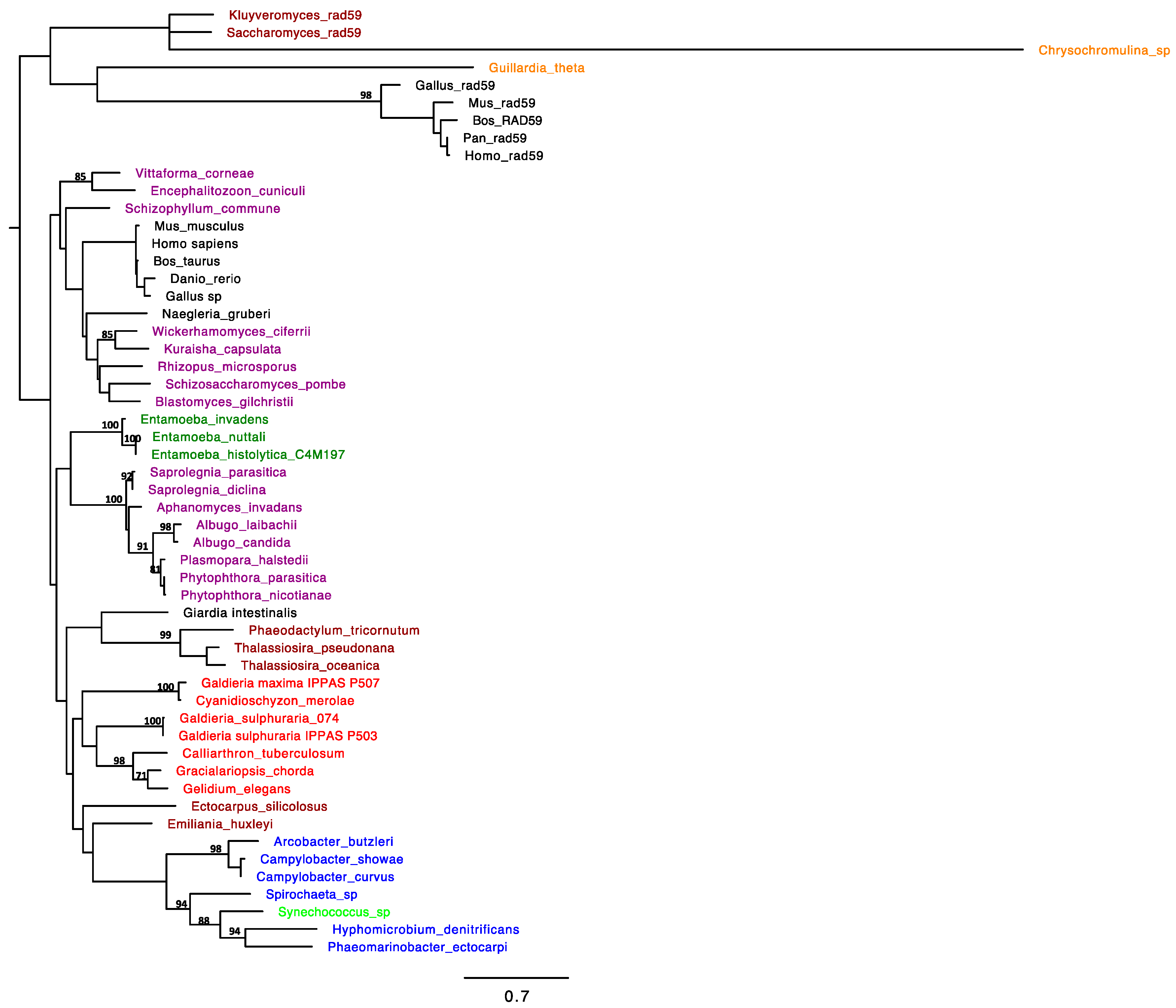
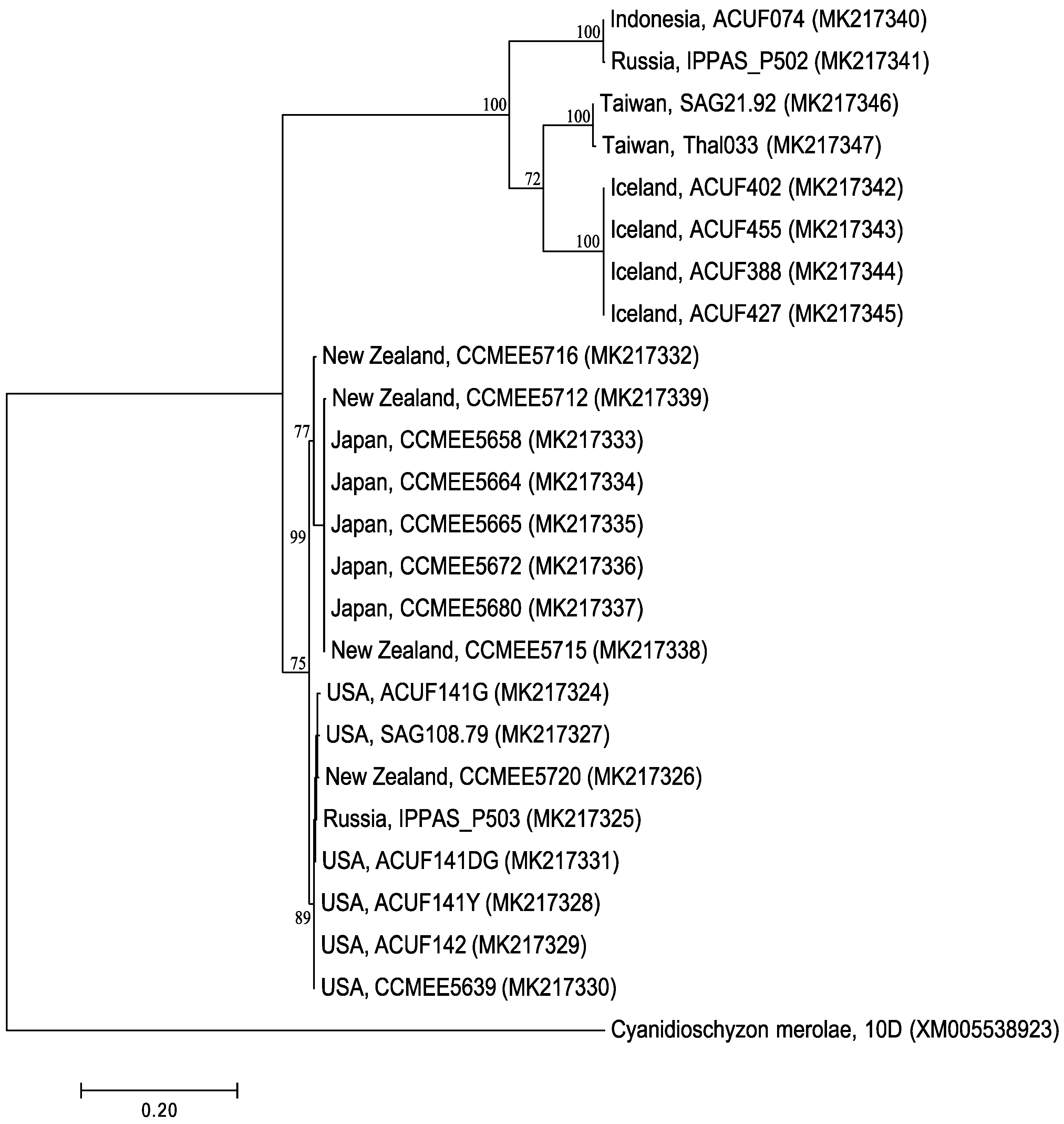
2.1. RAD52 Origin and Distribution
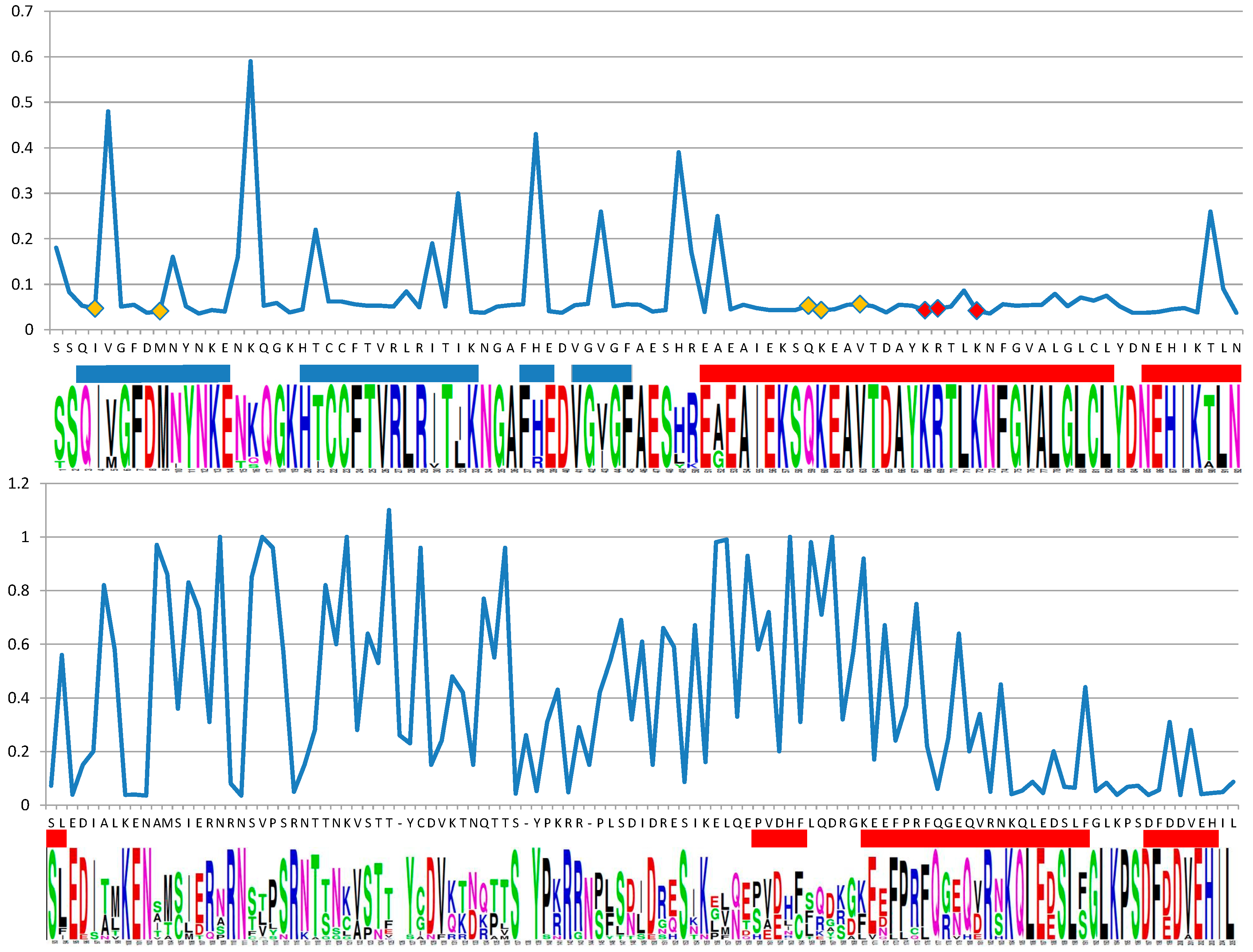
2.2. Support for Functional Homology of RAD52 Protein in Cyanidiophytina
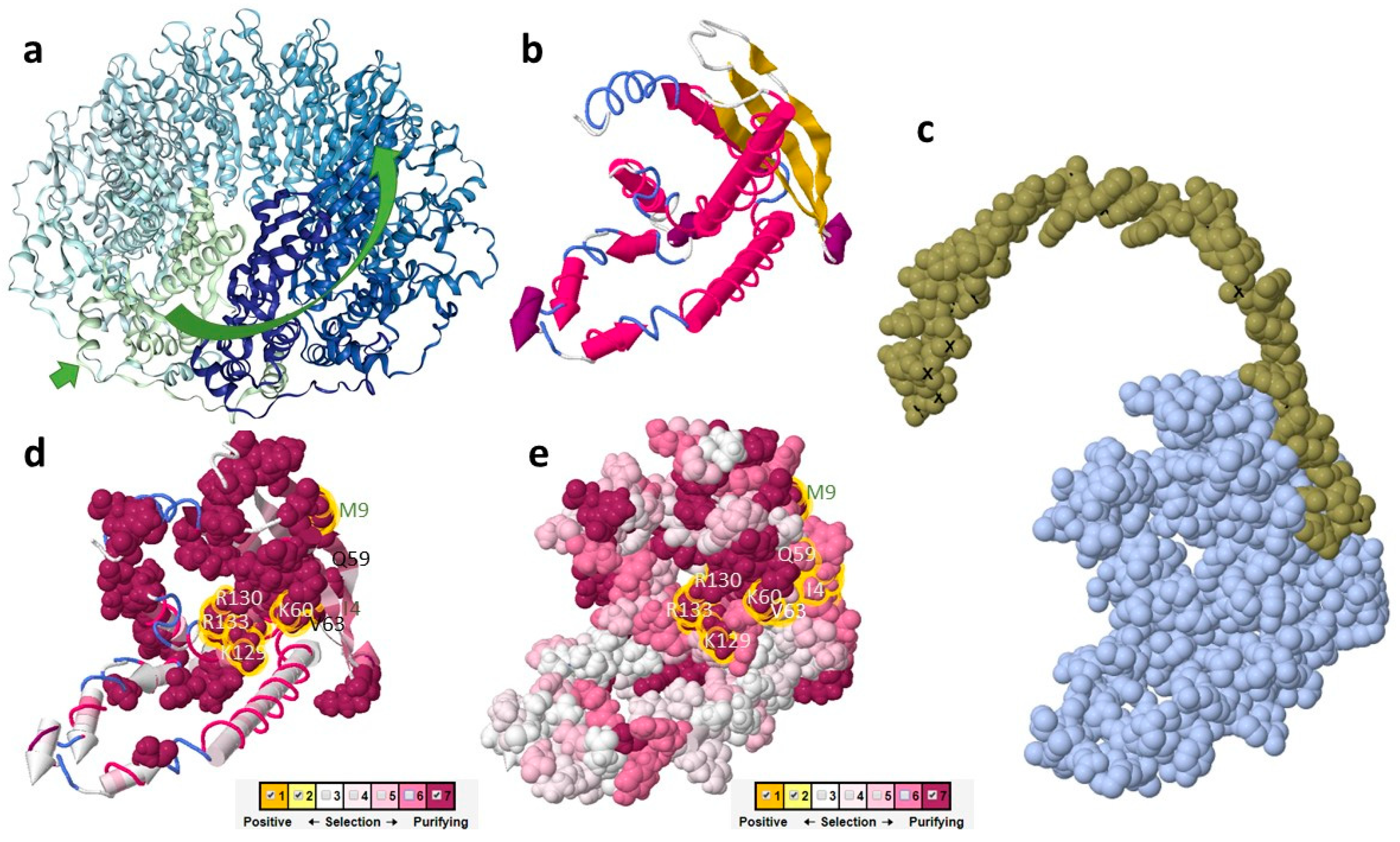
2.3. The Putative Role of RAD52 Protein in Cyanidiophytina
3. Material and Methods
3.1. Bioinformatics and Phylogenetic Analysis
3.2. In Silico Protein Structure Analysis
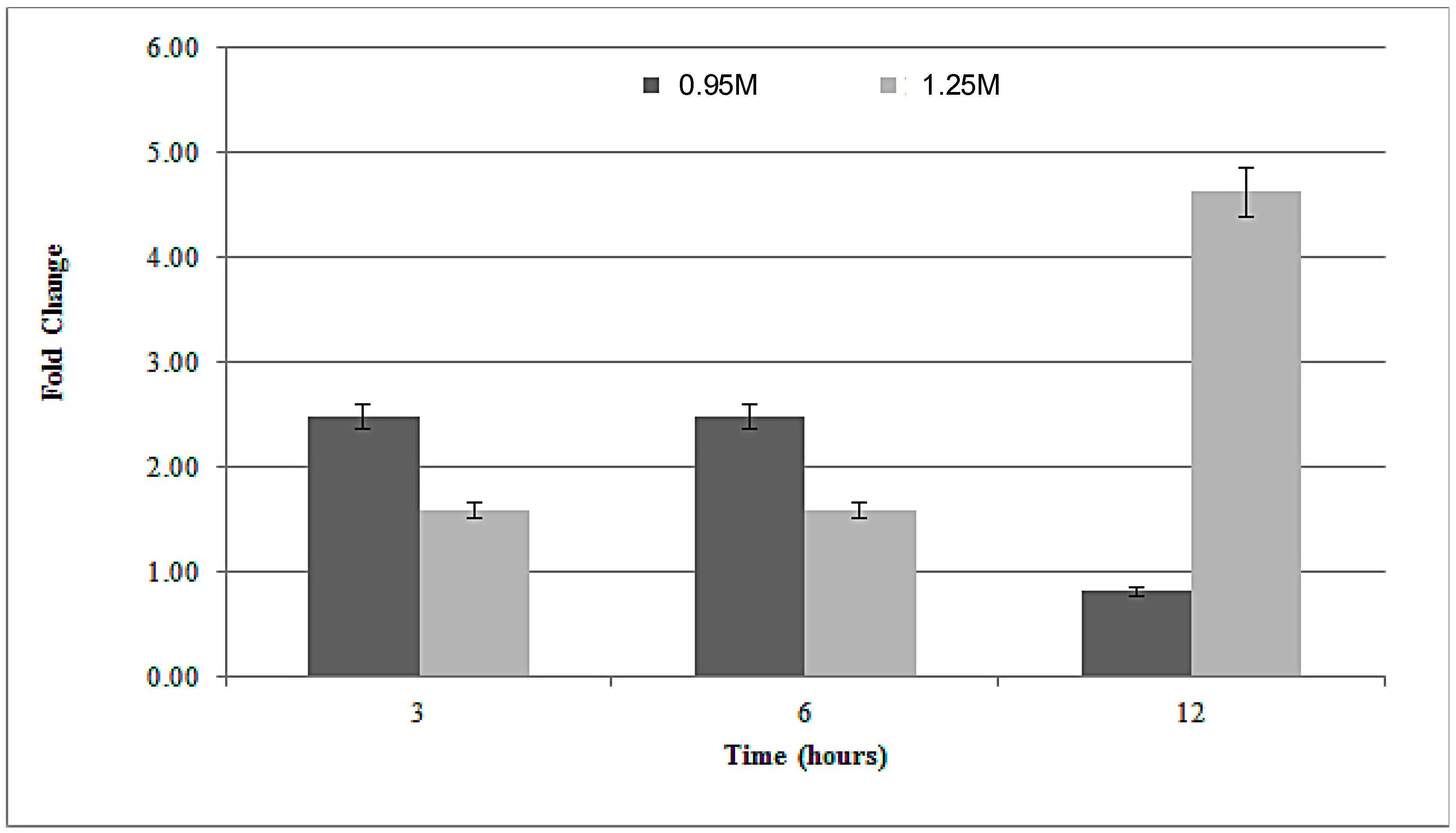
3.3. Rad52 Gene Expression under Salt Stress
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Ciniglia, C.; Yoon, H.S.; Pollio, A.; Pinto, G.; Bhattacharya, D. Hidden biodiversity of the extremophilic Cyanidiales red algae. Mol. Ecol. 2004, 13, 1827–1838. [Google Scholar] [CrossRef] [PubMed]
- Pinto, G.; Ciniglia, C.; Cascone, C.; Pollio, A. Species Composition of Cyanidiales Assemblages in Pisciarelli (Campi Flegrei, Italy) and Description of Galdieria Phlegrea SP. NOV. In Cellular Origin, Life in Extreme Habitats and Astrobiology; Springer: Dordrecht, The Nederland, 2007; pp. 487–502. [Google Scholar]
- Ciniglia, C.; Yang, E.C.; Pollio, A.; Pinto, G.; Iovinella, M.; Vitale, L.; Yoon, H.S. Cyanidiophyceae in Iceland: Plastid rbc L gene elucidates origin and dispersal of extremophilic Galdieria sulphuraria and G. maxima (Galdieriaceae, Rhodophyta). Phycologia 2014, 53, 542–551. [Google Scholar] [CrossRef]
- Cennamo, P.; Ciniglia, C. The algal diversity in the phlegrean fields (campania, italy) archeological districts: An overview. J. Urban Plan. Landsc. Environ. Des. 2017, 2, 97–106. [Google Scholar]
- Iovinella, M.; Eren, A.; Pinto, G.; Pollio, A.; Davis, S.J.; Cennamo, P.; Ciniglia, C. Cryptic dispersal of Cyanidiophytina (Rhodophyta) in non-acidic environments from Turkey. Extremophiles 2018, 22, 713–723. [Google Scholar] [CrossRef] [PubMed]
- Eren, A.; Iovinella, M.; Yoon, H.S.; Cennamo, P.; de Stefano, M.; de Castro, O.; Ciniglia, C. Genetic structure of Galdieria populations from Iceland. Polar Biol. 2018, 41, 1681–1691. [Google Scholar] [CrossRef]
- Yang, E.C.; Boo, S.M.; Bhattacharya, D.; Saunders, G.W.; Knoll, A.H.; Fredericq, S.; Graf, L.; Yoon, H.S. Divergence time estimates and the evolution of major lineages in the florideophyte red algae. Sci. Rep. 2016, 6, 21361. [Google Scholar] [CrossRef] [PubMed]
- Müller, K.M.; Oliveira, M.C.; Sheath, R.G.; Bhattacharya, D. Ribosomal DNA phylogeny of the Bangiophycidae (Rhodophyta) and the origin of secondary plastids. Am. J. Bot. 2001, 88, 1390–1400. [Google Scholar] [CrossRef]
- Yoon, H.; Hackett, J.; Pinto, G.; Bhattacharya, D. The single, ancient origin of chromist plastids. J. Phycol. 2002, 38, 40. [Google Scholar] [CrossRef]
- Gross, J.; Bhattacharya, D. Uniting sex and eukaryote origins in an emerging oxygenic world. Biol. Direct 2010, 5, 53. [Google Scholar] [CrossRef]
- Argueso, J.L.; Westmoreland, J.; Mieczkowski, P.A.; Gawel, M.; Petes, T.D.; Resnick, M.A. Double-strand breaks associated with repetitive DNA can reshape the genome. Proc. Natl. Acad. Sci. USA 2008, 105, 11845–11850. [Google Scholar] [CrossRef]
- Symington, L.S. Role of RAD52 Epistasis Group Genes in Homologous Recombination and Double-Strand Break Repair. Microbiol. Mol. Biol. Rev. 2002, 66, 1092–2172. [Google Scholar] [CrossRef]
- Mortensen, U.H.; Bendixen, C.; Sunjevaric, I.; Rothstein, R. DNA strand annealing is promoted by the yeast Rad52 protein. Proc. Natl. Acad. Sci. USA 1996, 93, 10729–10734. [Google Scholar] [CrossRef] [PubMed]
- Ciniglia, C.; Sansone, C.; Panzella, L.; Napolitano, A.; D’Ischia, M. Effects of walnut husk washing waters and their phenolic constituents on horticultural species. Environ. Sci. Pollut. Res. 2012, 19, 3299–3306. [Google Scholar] [CrossRef] [PubMed]
- Iyer, L.M.; Koonin, E.V.; Aravind, L. Classification and evolutionary history of the single-strand annealing proteins, RecT, Redβ, ERF and RAD52. BMC Genom. 2002, 3, 8. [Google Scholar]
- Malik, S.B.; Pightling, A.W.; Stefaniak, L.M.; Schurko, A.M.; Logsdon, J.M. An expanded inventory of conserved meiotic genes provides evidence for sex in Trichomonas vaginalis. PLoS ONE 2008, 3, e2879. [Google Scholar] [CrossRef]
- Nair, A.; Agarwal, R.; Kant Chittela, R. Biochemical characterization of plant Rad52 protein from rice (Oryza sativa). Plant Physiol. Biochem. 2016, 106, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Aravind, L.; Walker, D.R.; Koonin, E.V. Conserved domains in DNA repair proteins and evolution of repair systems. Nucleic Acids Res. 1999, 27, 1223–1242. [Google Scholar] [CrossRef] [PubMed]
- Kreitman, M. Methods to detect selection in populations with applications to the human. Annu. Rev. Genom. Hum. Genet. 2000, 1, 539–559. [Google Scholar] [CrossRef]
- MacCallum, C.; Hill, E. Being Positive about Selection. PLoS Biol. 2006, 4, e87. [Google Scholar] [CrossRef]
- Roth, C.; Liberles, D.A. A systematic search for positive selection in higher plants (Embryophytes). BMC Plant Biol. 2006, 6, 12. [Google Scholar] [CrossRef]
- Keeling, P.J. The endosymbiotic origin, diversification and fate of plastids. Philos. Trans. R. Soc. B Biol. Sci. 2010, 365, 729–748. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, M.C.; Bhattacharya, D. Phylogeny of the bangiophycidae (Rhodophyta) and the secondary endosymbiotic origin of algal plastids. J. Phycol. 2000, 36, 52. [Google Scholar] [CrossRef]
- Samach, A.; Melamed-Bessudo, C.; Avivi-Ragolski, N.; Pietrokovski, S.; Levy, A.A. Identification of Plant RAD52 Homologs and Characterization of the Arabidopsis thaliana RAD52-Like Genes. Plant Cell 2011, 23, 4266–4279. [Google Scholar] [CrossRef] [PubMed]
- Kagawa, W.; Kurumizaka, H.; Ishitani, R.; Fukai, S.; Nureki, O.; Shibata, T.; Yokoyama, S. Crystal Structure of the Homologous-Pairing Domain from the Human Rad52 Recombinase in the Undecameric Form. Mol. Cell 2002, 10, 359–371. [Google Scholar] [CrossRef]
- Singleton, M.R.; Wentzell, L.M.; Liu, Y.; West, S.C.; Wigley, D.B. Structure of the single-strand annealing domain of human RAD52 protein. Proc. Natl. Acad. Sci. USA 2002, 99, 13492–13497. [Google Scholar] [CrossRef] [PubMed]
- Roy, S. Maintenance of genome stability in plants: Repairing DNA double strand breaks and chromatin structure stability. Front. Plant Sci. 2014, 5, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Graur, D.; Li, W.H. Fundamental of Molecular Evolution; Sinauer Associate Inc.: Sunderland, MA, USA, 2000. [Google Scholar]
- Schönknecht, G.; Chen, W.; Ternes, C.M.; Barbier, G.G.; Shrestha, R.P.; Stanke, M.; Bräutigam, A.; Baker, B.J.; Banfield, J.F.; Garavito, R.M.; et al. Gene Transfer from Bacteria and Archaea Facilitated Evolution of an Extremophilic Eukaryote. Science 2013, 339, 1207–1210. [Google Scholar] [CrossRef]
- Matsuzaki, M.; Misumi, O.; Shin-i, T.; Maruyama, S.; Takahara, M.; Miyagishima, S.; Mori, T.; Nishida, K.; Yagisawa, F.; Nishida, K.; et al. Genome sequence of the ultrasmall unicellular red alga Cyanidioschyzon merolae 10D. Nature 2004, 428, 653–657. [Google Scholar] [CrossRef]
- Davis, A.M.; Iovinella, M.; James, S.; Robshaw, T.; Dodson, J.H.; Herrero-Davila, L.; Clark, J.H.; Agapiou, M.; McQueen-Mason, S.; Pinto, G.; et al. Using MinION nanopore sequencing to generate a de novo eukaryotic draft genome: Preliminary physiological and genomic description of the extremophilic red alga Galdieria sulphuraria strain SAG 107.79. bioRXiv 2017. [Google Scholar] [CrossRef]
- Willing, E.-M.; Rawat, V.; Mandáková, T.; Maumus, F.; James, G.V.; Nordström, K.J.V.; Becker, C.; Warthmann, N.; Chica, C.; Szarzynska, B.; et al. Genome expansion of Arabis alpina linked with retrotransposition and reduced symmetric DNA methylation. Nat. Plants 2015, 1, 14023. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Qiu, H.; Price, D.C.; Weber, A.P.M.; Reeb, V.; Yang, E.C.; Lee, J.M.; Kim, S.Y.; Yoon, H.S.; Bhattacharya, D. Adaptation through horizontal gene transfer in the cryptoendolithic red alga Galdieria phlegrea. Curr. Biol. 2013, 23, R865–R866. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; Mcgettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed]
- Stamatakis, A.; Hoover, P.; Rougemont, J. A rapid bootstrap algorithm for the RAxML web servers. Syst. Biol. 2008, 57, 758–771. [Google Scholar] [CrossRef]
- Hasegawa, M.; Yano, T.; Kishino, H. A new molecular clock of mitochondrial DNA and the evolution of hominoids. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 1984, 60, 95–98. [Google Scholar] [CrossRef]
- Nielsen, R.; Yang, Z. Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 1998, 148, 929–936. [Google Scholar]
- Yang, Z.; Bielawski, J.R. Statistical methods for detecting molecular adaptation. Trends Ecol. Evol. 2000, 15, 496–503. [Google Scholar] [CrossRef]
- Stern, A.; Doron-Faigenboim, A.; Erez, E.; Martz, E.; Bacharach, E.; Pupko, T. Selecton 2007: Advanced models for detecting positive and purifying selection using a Bayesian inference approach. Nucleic Acids Res. 2007, 35, W506–W511. [Google Scholar] [CrossRef]
- Allen, M.M.; Stainer, R.Y. Selective isolation of blue-green algae from water and soil. J. Gen. Microbiol. 1968, 51, 203–209. [Google Scholar] [CrossRef]
- Carfagna, S.; Bottone, C.; Cataletto, P.R.; Petriccione, M.; Pinto, G.; Salbitani, G.; Vona, V.; Pollio, A.; Ciniglia, C. Impact of sulfur starvation in autotrophic and heterotrophic cultures of the Extremophilic Microalga Galdieria Phlegrea (Cyanidiophyceae). Plant Cell Physiol. 2016, 57, 1890–1898. [Google Scholar] [CrossRef] [PubMed]
- Radonić, A.; Thulke, S.; Mackay, I.M.; Landt, O.; Siegert, W.; Nitsche, A. Guideline to reference gene selection for quantitative real-time PCR. Biochem. Biophys. Res. Commun. 2004, 313, 856–862. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Del Mondo, A.; Iovinella, M.; Petriccione, M.; Nunziata, A.; Davis, S.J.; Cioppa, D.; Ciniglia, C. A Spotlight on Rad52 in Cyanidiophytina (Rhodophyta): A Relic in Algal Heritage. Plants 2019, 8, 46. https://doi.org/10.3390/plants8020046
Del Mondo A, Iovinella M, Petriccione M, Nunziata A, Davis SJ, Cioppa D, Ciniglia C. A Spotlight on Rad52 in Cyanidiophytina (Rhodophyta): A Relic in Algal Heritage. Plants. 2019; 8(2):46. https://doi.org/10.3390/plants8020046
Chicago/Turabian StyleDel Mondo, Angelo, Manuela Iovinella, Milena Petriccione, Angelina Nunziata, Seth J. Davis, Diana Cioppa, and Claudia Ciniglia. 2019. "A Spotlight on Rad52 in Cyanidiophytina (Rhodophyta): A Relic in Algal Heritage" Plants 8, no. 2: 46. https://doi.org/10.3390/plants8020046
APA StyleDel Mondo, A., Iovinella, M., Petriccione, M., Nunziata, A., Davis, S. J., Cioppa, D., & Ciniglia, C. (2019). A Spotlight on Rad52 in Cyanidiophytina (Rhodophyta): A Relic in Algal Heritage. Plants, 8(2), 46. https://doi.org/10.3390/plants8020046






