Abstract
Because they are sessile organisms, plants need rapid and finely tuned signaling pathways to adapt to adverse environments, including salt stress. In this study, we identified a gene named Arabidopsis thaliana stress-induced BTB protein 1 (AtSIBP1), which encodes a nucleus protein with a BTB domain in its C-terminal side and is induced by salt and other stresses. The expression of the β-glucuronidase (GUS) gene driven by the AtSIBP1 promoter was found to be significantly induced in the presence of NaCl. The sibp1 mutant that lost AtSIBP1 function was found to be highly sensitive to salt stress and more vulnerable to salt stress than the wild type WT, while the overexpression of AtSIBP1 transgenic plants exhibited more tolerance to salt stress. According to the DAB staining, the sibp1 mutant accumulated more reactive oxygen species (ROS) than the WT and AtSIBP1 overexpression plants after salt stress. In addition, the expression levels of stress-induced marker genes in AtSIBP1 overexpression plants were markedly higher than those in the WT and sibp1 mutant plants. Therefore, our results demonstrate that AtSIBP1 was a positive regulator in salinity responses in Arabidopsis.
1. Introduction
In nature, plants must cope with different kinds of abiotic stress, such as high temperatures, cold, salt, drought and osmotic stresses; salt stress is one of the most common and severe stresses [1]. High salinity causes retarded plant growth (even death) and reduces agricultural productivity on more than 20% of the cultivated land worldwide [2]. The unfavorable effects of salt on plants are consequences of both water deficits that result in osmotic stress and excess sodium ions on critical biochemical process [3,4]. In order to deal with unfavorable environmental signals, plants show specific changes in gene expression, metabolism, and physiology in response to different environmental stress conditions [5].
In Arabidopsis, there are more than 1400 genes that are predicted to encode E3 ubiquitin ligase enzymes [6], which can be largely divided into two groups based on their structure: single-subunit E3 ligases and multi-subunit E3 ligases [7]. Among the multi-subunit E3 ligases, cullin-RING E3 ligases (CRLs) have been best characterized to date. Five cullins (cullin 1/2/3a/3b/4) have been identified as components of the CRLs [7,8]. The cullin3a/3b ligase has been identified in the genome of all eukaryotes, and it interacts with ‘‘Bric-a-brac, Tramtrack and broad (BTB)/complex/pox virus and zinc finger’’ (POZ) (hereafter called BTB) domain proteins [9]. The BTB domain, found in eukaryotes and some viruses, is a highly conserved protein motif with a length of around 116 amino acids that folds into five β-sheets and six α-helices [10]. A BTB domain-containing protein acts as the substrate receptor of cullin 3a/3b and facilitates the assembly of cullin 3a/3b into E3 ligase complexes [11]. The BTB superfamily is divided into multiple groups according to their structure, such as BTB-ZF (BTB zinc-finger), BBK (BTB-BACK-kelch), BBP (BTB-BACKPHR), BTB-ankyrin, all of which contain the BTB domain and other domains, while Skp1 (S-phase kinase associated protein 1) and ElonginC contain only BTB domain [12,13].
BTB plays an important role in plant growth, development, stress resistance, protein ubiquitination and degradation, cytoskeletal composition, ion channel, and the regulation of the cell cycle [14,15]. Previous reports have indicated that the Arabidopsis NPR1 (non-expressor of pathogenesis related genes 1) protein that contains the BTB domain is a key regulator at the intersection of the ISR (induced systemic resistance) and SAR (systemic acquired resistance) signaling pathways [16,17,18]. Many BTB protein family members in plants are involved in the regulation of hormone-mediated plant-related signaling such as SA (salicylic acid), JA (jasmonic acid), ABA (abscisic acid), GA3 (gibberellin 3). AHT1 (ABA-HYPERSENSITIVE BTB/POZ PROTEIN 1), a BTB-containing protein, negatively regulates the ABA-mediated inhibition of germination in Arabidopsis [19]. Six Arabidopsis BTB/POZ-MATH (meprin and TRAF homology) proteins (BPMs) have been reported to participate in ABA responses such as germination and stomatal closing [9]. ATHB6, a homeobox-leucine zipper transcription factor, acts as a negative regulator in ABA and is a target of BPMs for ubiquitination and degradation [19]. The disruption of BT2 (BTB and TAZ(Transcription Adaptor putative Zinc finger) domain protein 2) leads to the enhancement of the ABA-mediated inhibition of germination [9]. Therefore, the BTB-containing protein family is involved in many biological processes in plants including abiotic and biotic stress responses.
Here, we found a novel BTB domain-containing gene called AtSIBP1 (Arabidopsis thaliana stress-induced BTB protein 1) that belongs to the Skp1 or ElonginC families. The expression of AtSIBP1 was induced by salt and other stresses. The overexpression of AtSIBP1 exhibited an enhanced tolerance to NaCl, while the sibp1 mutant was sensitive to NaCl. The expression levels of stress-responsive genes were significantly increased in AtSIBP1 overexpression lines. Our analysis demonstrates that AtSIBP1 acts as a positive regulator in salt stress responses.
2. Results
2.1. Expression Pattern of AtSIBP1
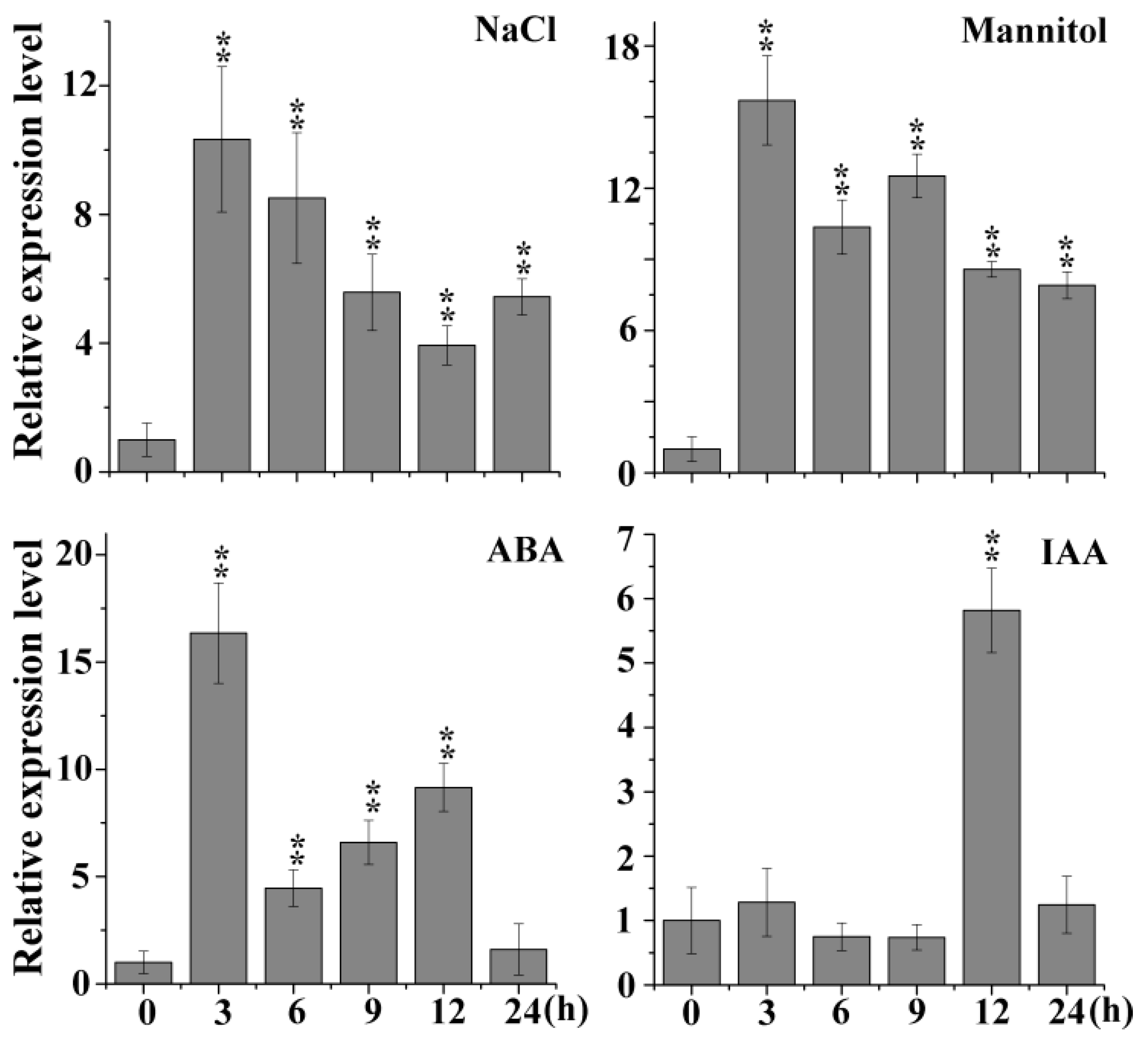
Most BTB family members are involved in abiotic and biotic stress responses. In the Arabidopsis eFP (electronic Fluorescent Pictograph) browser, we noticed a novel gene (At1g55760) that is predicted to respond to salt and other stresses and is also a BTB-containing protein (supplementary Figures S1 and S2). Subsequently, the transcripts of At1g55760 were detected by qRT-PCR under different treatments, and the results showed that the expression levels of At1g55760 were significantly induced under NaCl, mannitol, ABA and indole-3-acetic acid (IAA) treatment (Figure 1). Therefore, we named it AtSIBP1 (Arabidopsis thaliana stress-induced BTB protein 1). In this study, we focused on the functions of AtSIBP1 in response to the salt stress.
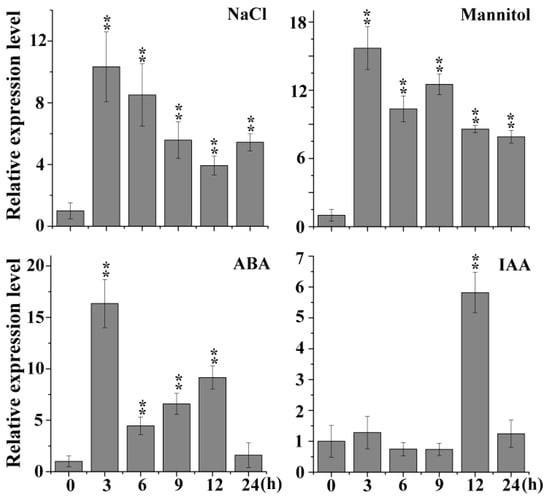
Figure 1.
Expression profiles of AtSIBP1 under stress treatment. An analysis of the expression levels of AtSIBP1 under abscisic acid (ABA), indole-3-acetic acid (IAA) and abiotic stresses. Eight-day-old wild type (WT) Arabidopsis seedlings were treated with 50 μM of ABA, 10 μM of IAA, 200 mM of NaCl and 300 mM of mannitol. The total RNA was extracted at the indicated times. The expression level of AtSIBP1 was monitored by real-time PCR. ACTIN2 was used as the internal control. Values are the means (± SD) of three individual experiments, and asterisks indicate significant differences from the WT using the unpaired Student’s t-test (* p < 0.05; ** p < 0.01).
2.2. Identification of Arabidopsis sibp1 Mutant and AtSIBP1 Overexpression Lines
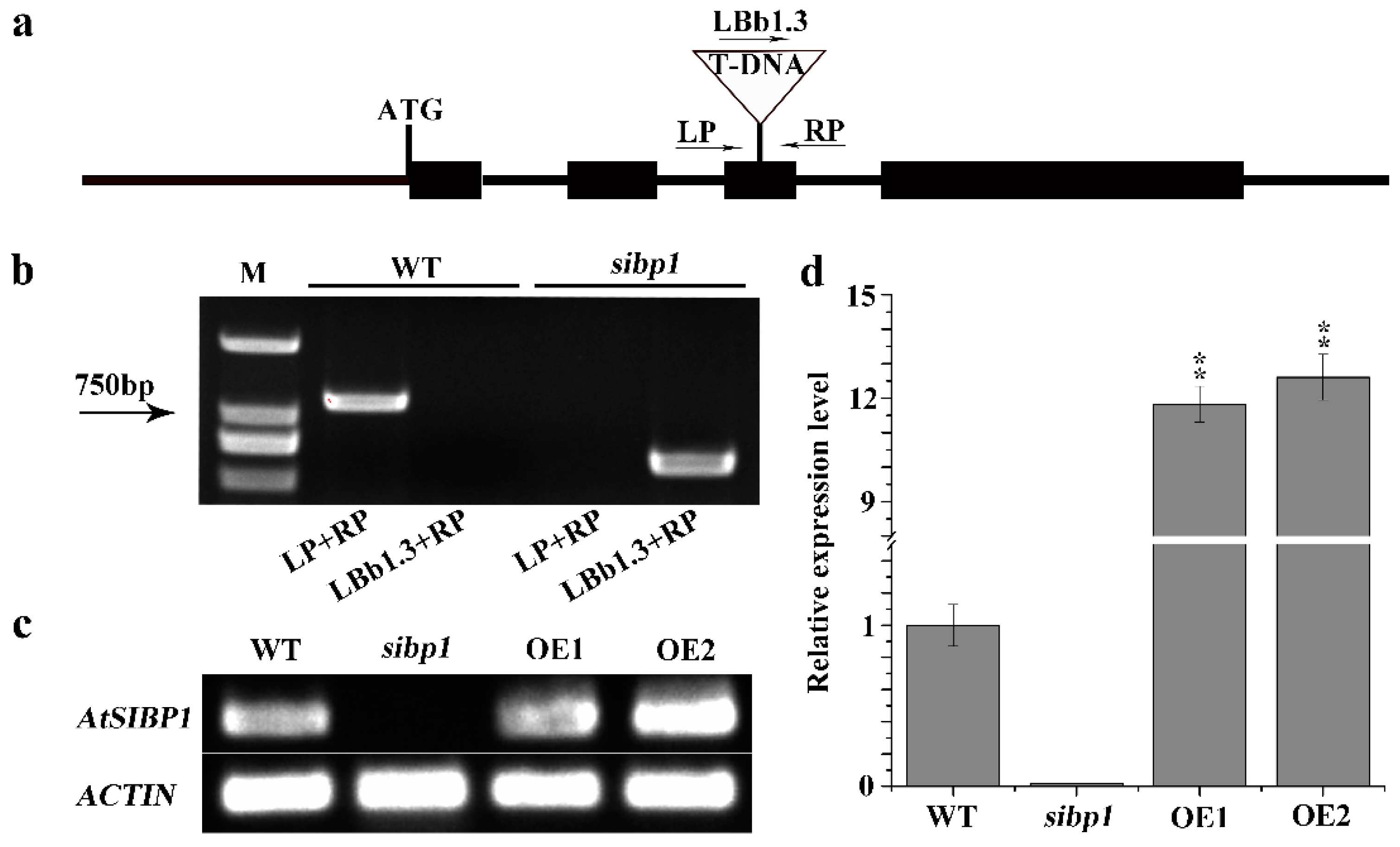
To investigate the function of AtSIBP1 in the response of Arabidopsis to salt stress, we isolated a homozygous AtSIBP1 T-DNA insertion mutant (SALK_075267) and generated two independent AtSIBP1-overexpression lines (abbreviated as OE1 and OE2). The mutant and the insertion site of AtSIBP1 was confirmed by PCR and sequencing, respectively (Figure 2a,b). A single copy of T-DNA was inserted into the third exon of AtSIBP1, and no full-length AtSIBP1 transcript was detected in the sibp1 mutant by RT-PCR analysis, suggesting that AtSIBP1 is disrupted in sibp1 mutant plants (Figure 2b). The result of semi-quantitative RT-PCR further confirmed that the transcripts of AtSIBP1 were totally disrupted in the sibp1 mutant, while the overexpression of AtSIBP1 plants increased 11.8 fold in OE1 and 12.6 fold in OE2 compared to the wild type (WT) (Figure 2c,d).
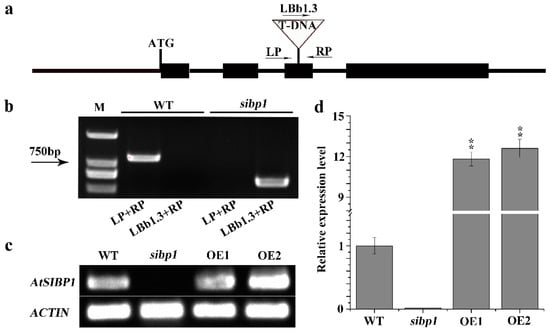
Figure 2.
Identification of the sibp1 T-DNA insertion mutant and construction of AtSIBP1-overexpression plants. (a) Structure of Arabidopsis AtSIBP1 and T-DNA insertion site in the sibp1 mutant (SALK_075267). (b) Molecular analysis of sibp1 and the WT, primers LP, RP and LBb1.3 (Left, Right genomic primer and the left T-DNA border primer) were used to target the flanking sequences of the T-DNA; M represents molecular marker. (c) Semi-quantitative RT-PCR analysis of AtSIBP1 expression in the WT and the sibp1 mutant. (d) qRT-PCR analysis of AtSIBP1 expression in the WT, the sibp1 mutant and the AtSIBP1-overexpressing lines OE1 and OE2. ACTIN2 was used as internal control for both semi-quantitative RT-PCR and qRT-PCR.
2.3. Tissue Expression Pattern of AtSIBP1 and Subcellular Localization of AtSIBP1-eGFP Fusion Protein
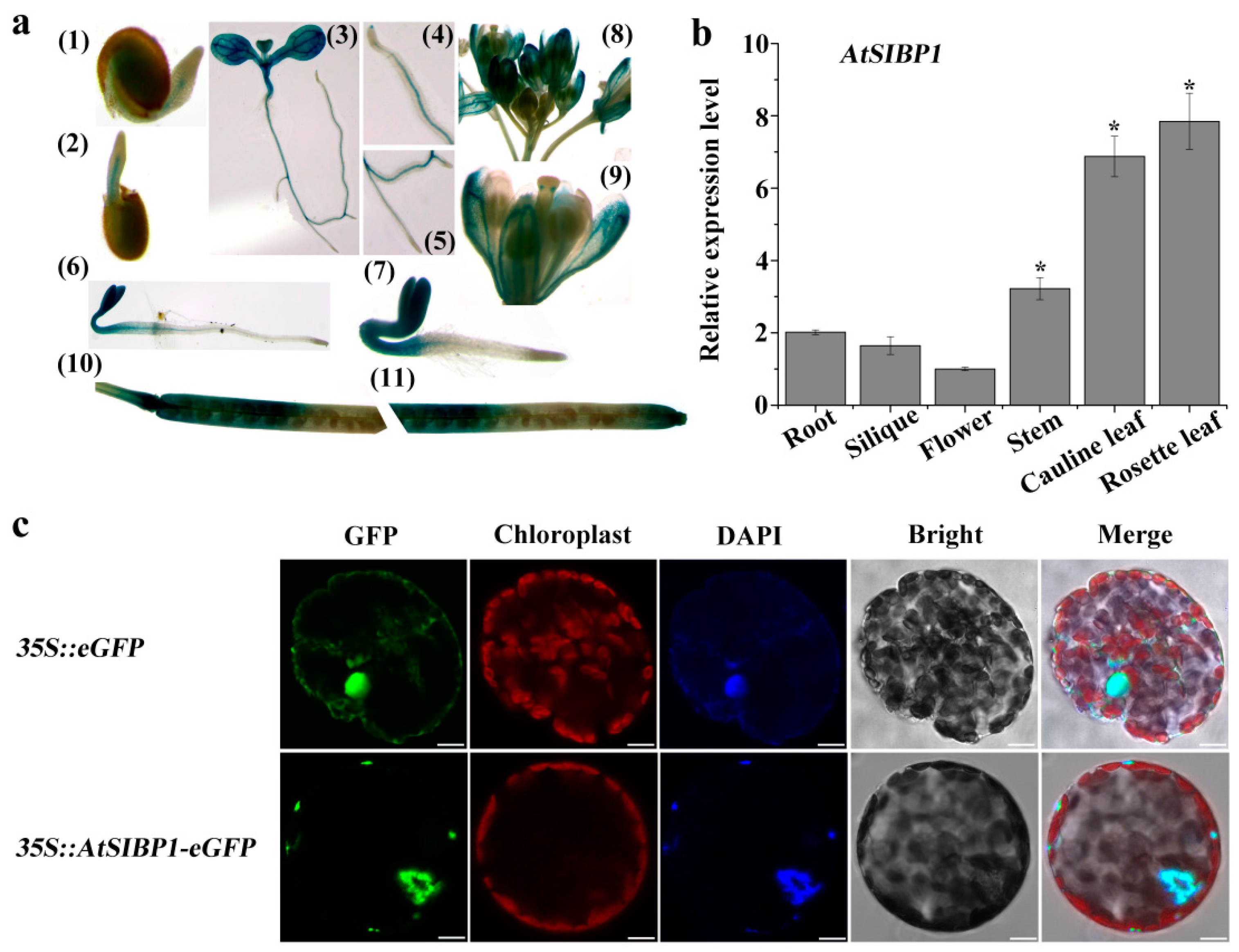
To further confirm the expression pattern of AtSIBP1, the tissue-specific expression of AtSIBP1 was first analyzed. Transgenic lines carrying proAtSIBP1::GUS (β-glucuronidase) were generated, and the T3 homozygous seeds were used for the subsequent staining experiments. Histochemical staining showed that GUS activity could be monitored at all developmental stages (Figure 3a). The GUS staining was first detected in the emerging radicles of the germinated seeds 24h after planting (Figure 3a (1)) and became more obvious in two- and three-day-old seedlings (Figure 3a (2,6,7)). After seven days of growth, GUS activity was found mainly in the leaf and vascular tissues, including the root tips and the lateral roots (Figure 3a (3,4,5)). In mature plants, GUS activity was observed in the generative organs and siliques (Figure 3a (8,9,10,11)). Subsequently, the mRNA of AtSIBP1 was detected in different organs of Arabidopsis, and the results showed that AtSIBP1 was expressed in all organs; the rosette leaves had the highest expression while the flowers had the lowest expression (Figure 3b). Hence, our results indicate that AtSIBP1 is expressed throughout the whole life cycle of Arabidopsis.
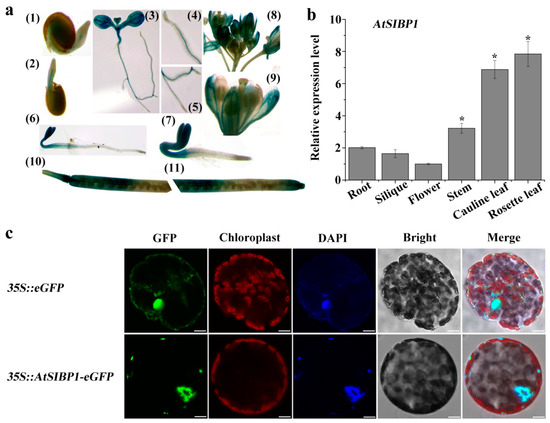
Figure 3.
Tissue-specific expression and subcellular localization of AtSIBP1. (a) Histochemical staining of proAtSIBP1::GUS (β-glucuronidase) T3 homozygous plants. AtSIBP1 promoter activity was detected in the (1–3) germinating seedlings at day one (1), day two (2), day seven (3), the root tip of the seven-day-old seedlings (4), the lateral root of the seven-day-old seedlings(5), the three-day-old etiolated seedlings (6), the three-day-old light-grown seedlings (7), inflorescence (8), the opened flowers (9), and the siliques (10,11). (b) qRT-PCR analysis of AtSIBP1 expression in different organs. The value for the flower was set to 1. ACTIN2 was used as an internal control. Values are the means (± SD) of three individual experiments, and asterisks indicate significant differences from the WT using the unpaired Student’s t-test (* p < 0.05; ** p < 0.01). (c) Subcellular localization of AtSIBP1-eGFP in tobacco mesophyll protoplasts. At 16 hours after transfection, fluorescence was imaged by confocal microscope. The 35S: eGFP was used as a control. The same cell was stained with DAPI (4’, 6’-diamidino-2-phenylindole) to show its nucleus. Bars = 10 μm.
In order to identify the subcellular localization of AtSIBP1, a 35S::AtSIBP1-eGFP construct was generated and transiently transformed into tobacco mesophyll protoplasts. The green fluorescent signal of the positive control was detected in the nucleus and cytoplasm, while the signal of AtSIBP1 was only detected in the nucleus (Figure 3c). DAPI (4’, 6’-diamidino-2-phenylindole) dye was used to stain the nucleus, and the results further confirmed that AtSIBP1 was localized in the nucleus (Figure 3c).
2.4. AtSIBP1 Acts as a Positive Regulator in Response to Salt Stress
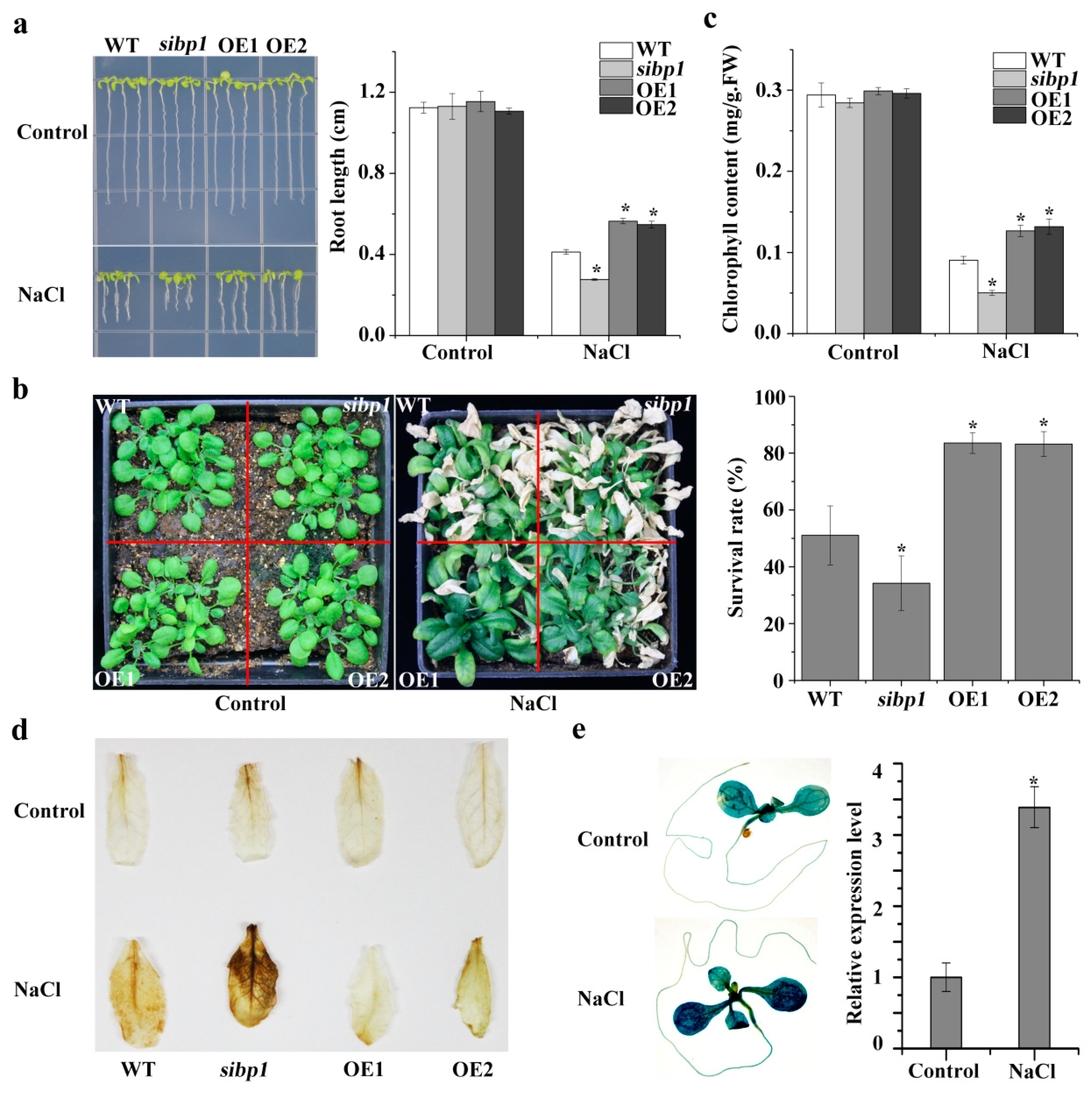
To gain insights into the physiological function of AtSIBP1 in salt stress, phenotypic experiments were executed. A root growth assay demonstrated that there was no difference among plants under normal conditions (Figure 4a). When plants were exposed to salt stress, the overexpression of AtSIBP1 plants showed a reduced sensitivity, while the sibp1 mutant exhibited an increased sensitivity to salt stress compared to the WT (Figure 4a). To further characterize the function of AtSIBP1 in salt tolerance, the plants grown in soil were also investigated. The overexpression of AtSIBP1 transgenic plants consistently showed a higher survival rate, while the sibp1 mutant had a lower survival rate than that of the WT after the treatment with NaCl (Figure 4b). The analysis of chlorophyll contents revealed that the OE1 and OE2 plants had higher chlorophyll contents and that of the sibp1 mutant was lower than that of the WT after being treated with 200 mM of NaCl (Figure 4c).
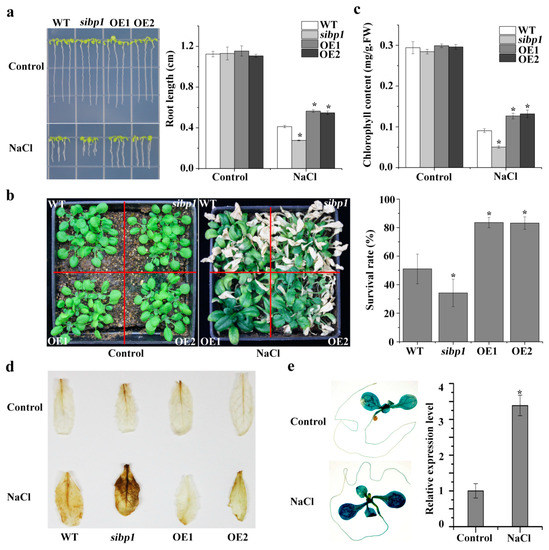
Figure 4.
AtSIBP1 is involved in salt stress response in Arabidopsis thaliana. (a) Root length of the WT, the sibp1 mutant, and the overexpression of AtSIBP1 planted two and a half days on an MS (Murashige and Skoog) medium following a seven-day vertical culture on a 1/2 MS medium supplemented with or without 160 mM of NaCl. (b) Phenotype and survival rate of the 16-day-old WT, sibp1 and AtSIBP1-overexpressing lines before and after NaCl treatment. (c) Chlorophyll content of the WT, sibp1, OE1, and OE2 before and after salt stress. (d) DAB staining assay of the rosette leaves of the WT, sibp1, OE1, and OE2 with or without salt stress. (e) Histochemical staining and expression level of the GUS gene in proAtSIBP1::GUS T3 homozygous seedlings with or without salt stress. ACTIN2 was used as an internal control. Values are the means (± SD) of three individual experiments, and asterisks indicate significant differences from the untreated using the unpaired Student’s t-test (* p < 0.05; ** p < 0.01).
When plants suffer biotic or abiotic stress, the level of reactive oxygen species (ROS) sharply increases and then impairs plant growth [20]. The DAB staining showed that the sibp1 mutant plants accumulated more ROS (stained deeper), while the OE lines accumulated less ROS after salt stress, and there was no difference in plants under normal conditions (Figure 4d). In addition, the GUS staining of the Arabidopsis plants expressing proAtSIBP1::GUS also showed that the expression of the GUS reporter gene was significantly induced by salt stress (Figure 4e). An analysis by qPCR also revealed an increased level of GUS transcripts in proAtSIBP1::GUS transgenic Arabidopsis under the treatment with NaCl (Figure 4e). Taken together, our results indicate that AtSIBP1 plays a positive regulator in the salt stress responses of plants.
2.5. Disruption or Overexpressing of AtSIBP1 Altered the Expression of Stress Induced Genes
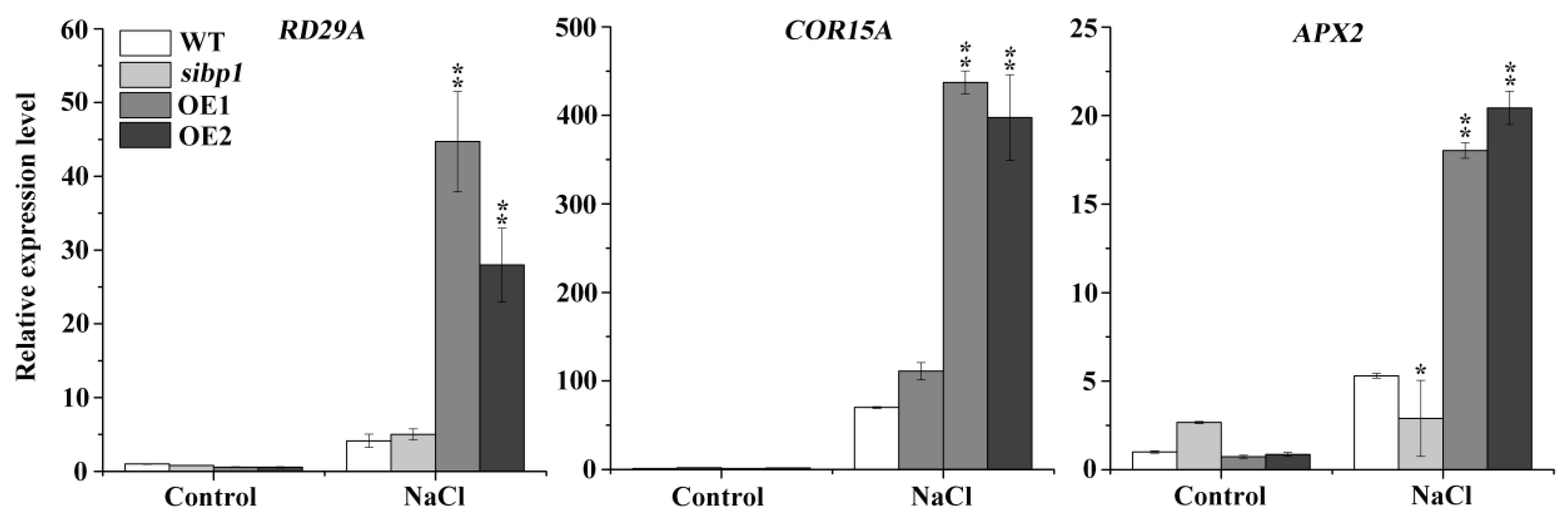
To further elucidate the role of the AtSIBP1 response to salt stress in Arabidopsis, eight-day-old seedlings were treated with 200 mM of NaCl for six hours, and then the transcript levels of the stress-inducible genes were analyzed, including the responsive to desiccation 29A (RD29A), cold-regulated 15A (COR15A), and ascorbate peroxidase 2 (APX2) genes. The expression levels of them were significantly increased in the overexpression of AtSIBP1 transgenic plants, while the expression level of APX2 was decreased in the sibp1 mutant compared to the WT after the treatment with salt stress (Figure 5). In summary, AtSIBP1 plays a positive role in plants’ response to salt stress through the up-regulation of the expressions of the stress-inducible genes RD29A, COR15A and APX2.
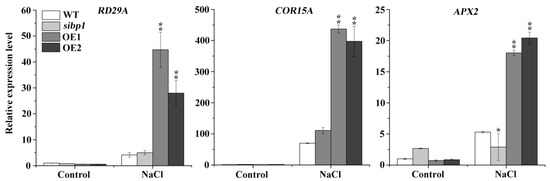
Figure 5.
Transcriptional expression levels of stress-inducible genes. Eight-day-old seedlings of the WT, sibp1, OE1,and OE2 were treated with 200 mM of NaCl for six hours, and then the transcription levels of RD29A, COR15A and APX2 were tested by qRT-PCR. Data represent the means (± SD) from three independent experiments, and asterisks indicate significant differences from the WT using the unpaired Student’s t-test (* p < 0.05; ** p < 0.01).
3. Discussion
In Arabidopsis thaliana, there are about 150 BTB proteins that play important roles in the processes of plant growth, development, and stress responses [14,15]. The BTB proteins contain a conserved BTB protein–protein interaction motif and are highly diverse in Arabidopsis [21]. In this study, AtSIBP1 was found to be a typical BTB protein containing one BTB domain and acted as a potential substrate receptor for CRL3 (cullin3-RING E3 ligases) (Supplementary Figure S2).
The analysis of data from the publicly available Arabidopsis microarray database and the results of qRT-PCR indicated that AtSIBP1 was induced by salt stress (Figure 1). In addition, the histochemical staining assay revealed that the GUS reporter gene driven by the AtSIBP1 promoter was expressed in the whole process of the life cycle and was induced by NaCl, suggesting that AtSIBP1 has an important role in the response to salt stress of plants (Figure 3 and Figure 4e). The root system of plants is one of the most sensitive organs for sensing the availability of water and nutrients or other adverse soil conditions [22,23]. The analysis of root length, seedlings and the content of chlorophyll showed that the sibp1 mutant was sensitive to NaCl, while the overexpression of AtSIBP1 transgenic plants were insensitive to NaCl (Figure 4). The mutation of AHT1, one BTB-domain protein, resulted in plants that were sensitive to salinity [19]. The overexpression of AtNPR1, first the BTB-domain protein cloned in Arabidopsis, has been found to have a negative effect on dehydration and salt stresses in rice [24]. However, the overexpression of the Malus hupehensis MhNPR1 gene has been shown to increase tolerance to salt and osmotic stress by inducing the expression of pathogenesis-related genes (PRs) and osmotic-stress related genes in transgenic tobacco [25]. Abiotic stresses generally increase the production rate of ROS, which may react with proteins, lipids and deoxyribonucleic acid and then cause oxidative damage and impair the normal functions of plant cells [26]. Our DAB staining indicated that the amount of ROS was significantly reduced in the AtSIBP1 overexpression lines and was accumulated in the sibp1 mutant (Figure 4d), indicating that AtSIBP1 is involved in the depression of ROS accumulation in response to salinity.
The nuclear localization of the BTB protein may be a prerequisite for its activation and the regulation of plant immune response [27]. Here, we found that AtSIBP1 was localized in the nucleus (Figure 3c); thus, we speculate that the degraded target protein is a nucleus-localization regulator in the salt signaling pathway. Our work analyzed the expression levels of salt stress responsive genes, and our results demonstrated that the overexpression of AtSIBP1 led to the induction of the expressions of RD29A, COR15A and APX2 (Figure 5). Previous reports have indicated that the RD29A and COR15A promoters contain several DREs (dehydration-responsive elements) and ABREs (abscisic acid-responsive elements), which have the potential to confer abiotic stress resistance. In our study, salt stress induced the expression of RD29A and COR15A in AtSIBP1-overexpressions to increase the tolerance of salinity [28,29]. APX2, an ascorbate peroxidase, is one of the most important antioxidant enzymes in plants and detoxifies hydrogen peroxide by using ascorbate as a reductant [30]. The overexpression of AtSIBP1 was found to result in a higher expression level of APX2, indicating an improved antioxidant capacity of plants during salt stress. Therefore, we conclude that the overexpression of AtSIBP1 leads to increased tolerance of plants to high salinity through the up-regulation of APX2 expression, and then mediates the reduction of ROS accumulation.
The functions of BTB proteins in biotic stress have been highly clarified, but their roles in abiotic stress are far from elucidated. In this study, we demonstrated that AtSIBP1 positively regulated the responses of plants to salt stress. AtSIBP1, as a potential substrate receptor for the CRL3 complex, is involved in the tolerance to salt stress. Thus, we infer that its degraded target protein may be a negative regulator in the salt signaling pathway. However, the substrates of AtSIBP1 have yet to be identified, and the details how AtSIBP1 works in response to salinity and other stresses need to be further explored in Arabidopsis.
4. Materials and Methods
4.1. Plant Material and Growth Conditions
All Arabidopsis used in this study were of the Columbia (Col-0) ecotype. The T-DNA insertion mutant At1g55760 (SALK_075267) was obtained from the Arabidopsis Biological Resource Center (ABRC). All plants were grown in pots containing a mixture of vermiculite and soil (1:3, v/v) in a greenhouse at 22 °C under 70% relative humidity with 16 h light/8 h dark photoperiod. All seeds were sterilized with NaClO (0.5%, v/v) for 15min, were washed with sterile water six times, and then stored at 4 °C for 3 d to break dormancy. Surface-sterilized seeds were sown on a Murashige and Skoog (MS) medium containing 2% (w/v) sucrose and 0.8% (w/v) agar, pH 5.8.
4.2. Generation of Transgenic Plants
To generate the AtSIBP1-overexpressing and the proAtSIBP1::GUS constructs, the full length CDS (Coding sequence) (990bp) and the promoter of AtSIBP1 (1521bp) were amplified and cloned into pCAMBIA1302 and pCAMBIA1301, respectively. The constructed plasmids were transformed into Agrobacterium tumefaciens (GV3101) and then infiltrated into the WT (wild type) using the floral dip method [31]. The seeds of the transgenic plants were screened on an MS medium supplemented with hygromycin. The mRNA levels of AtSIBP1 were identified with a qRT-PCR assay. The third generation homozygous seeds of the transgenic plants (T3) were used for further analysis. The primers that were used in this assay were listed in Supplementary Table S1.
4.3. Subcellular Localization Assay
The CDS of AtSIBP1 was cloned into pBI221 containing a 35S promoter for a green fluorescence protein (eGFP) by homologous recombination. The constructed plasmid 35S::AtSIBP1-eGFP and the control of 35S::eGFP were transfected into mesophyll protoplasts from tobacco leaves using the PEG-calcium (polyethylene glycol-calcium) transfection method, as described in [32]. The pBI221-eGFP vector was used as a control. To confirm the location of AtSIBP1, 4’, 6’-diamidino-2-phenylindole (DAPI) was used to stain the nucleus of the cell. After transfection at 22 °C for 16 h in the dark, the fusion fluorescence proteins were observed and captured using a confocal laser-scanning microscope [33] (Leica TCSSP5 II system, Leica, Germany).
4.4. Phenotype Analysis
For the root elongation assay, the seeds were sown on an MS medium for vertical incubation for two and a half days and then transferred to new ½ MS media supplemented with or without NaCl. The root length was determined 1 week after transfer by using the Image J software. For salt treatment, after 16 d, plants grew under normal conditions were irrigated with a 200 mM NaCl solution applied at the bottom of the pots. The morphological phenotypes and survival rates of plants were recorded after 15 d of NaCl treatment. The measurement of chlorophyll content was determined as described [34].
4.5. GUS and DAB Staining
The histochemical GUS staining of the homozygous T3 transgenic lines harboring proAtSIBP1::GUS was done as described [35]. For DAB (3, 3’-Diaminobenzidine tetrahydrochloride) staining, five Arabidopsis rosette leaves treated with NaCl were placed in 5 mL tubes supplemented with a 3 mL DAB staining solution (1 mg/mL DAB, Sigma-Aldrich, adjusted to pH 3.8 with HCl), then vacuumed for 5 min. After 6 h of incubation in the dark with gentle shaking, the leaves were immersed in a bleaching solution (ethanol, acetic acid, and glycerol = 3:1:1) and boiled for 20 min to decolorize the leaves (except for the deep brown polymerization product that was produced by the reaction of DAB with H2O2), and then the images were captured.
4.6. Analysis of Gene Expression
Eight-day-old Arabidopsis seedlings were used for gene expression level identification with or without NaCl treatment. The total RNA was extracted from plants with RNAiso Plus reagents (Takara) according to the manufacturer’s instruction. The cDNA was synthesized using 2 μg of RNA by a Prime Script RT reagent Kit with a gDNA Eraser (Takara). For semi-quantitative RT-PCR, the semi-quantitative primers were used to test the expression of AtSIBP1. For the qRT-PCR analysis, the TB Green Premix Ex TaqII kit and an Applied Biosystems 7500 real-time PCR system were used for reaction according to the manufacturer’s instruction. ACTIN2 was used as the internal control. All primers are shown in Supplementary Table S1.
4.7. Sequence Analysis of AtSIBP1
The putative protein structure of AtSIBP1 was obtained from SMART (SIMPLE MODULAR ARCHITECTURE RESEARCH TOOL) and NCBI (National Center for Biotechnology Information). Multiple alignment results were edited in the DNAMAN8.0 program. The phylogenetic analysis was performed with the MEGA7.0 program.
4.8. Statistical Analysis
The data are represented as means ± SD (Standard Deviation) (n = 3). Statistical analysis was performed using Student’s t-test. The values of p < 0.05 were considered to be significant, and Values of p < 0.01 were considered as more significant.
Supplementary Materials
The following are available online at https://www.mdpi.com/2223-7747/8/12/573/s1. (Figure S1: The development map of At1g55760 in Arabidopsis eFP browse, Figure S2: Domain organization and phylogenetic analysis of At1g55760, Table S1: Primers used in the Paper)
Author Contributions
X.W. designed and wrote the research. L.P. and X.W. performed the experiments. J.X. collected the plant material. J.W. and X.L. (Xiaoyi Li) supervised the experiments. Y.Y. and X.L. (Xufeng Li) were project administration.
Funding
This study was funded by NSFC31870240 (Y.Y.) and by the National transgenic project 2016ZX08009003-002 (to X.L. (Xufeng Li)).
Conflicts of Interest
The authors declare no competing financial interests.
References
- Zhu, J.K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273. [Google Scholar] [CrossRef] [PubMed]
- Ganie, S.A.; Molla, K.A.; Henry, R.J.; Bhat, K.V.; Mondal, T.K. Advances in understanding salt tolerance in rice. Theor. Appl. Genet. 2019, 132, 851–870. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.-K. Regulation of ion homeostasis under salt stress. Curr. Opin. Plant Biol. 2003, 6, 441–445. [Google Scholar] [CrossRef]
- Tada, Y. The HKT Transporter Gene from Arabidopsis, AtHKT1; 1, Is Dominantly Expressed in Shoot Vascular Tissue and Root Tips and Is Mild Salt Stress-Responsive. Plants 2019, 8, 204. [Google Scholar] [CrossRef]
- Zhu, J.; Fu, X.; Koo, Y.D.; Zhu, J.K.; Jenney, F.E.; Adams, M.W.; Zhu, Y.; Shi, H.; Yun, D.-J.; Hasegawa, P.M.; et al. An enhancer mutant of Arabidopsis salt overly sensitive 3 mediates both ion homeostasis and the oxidative stress response. Mol. Cell. Biol. 2007, 27, 5214–5224. [Google Scholar] [CrossRef] [PubMed]
- Vierstra, R. The ubiquitin/26S proteasome pathway, the complex last chapter in the life of many plant proteins. Trends Plant Sci. 2003, 8, 135–142. [Google Scholar] [CrossRef]
- Lee, J.H.; Kim, W.T. Regulation of abiotic stress signal transduction by E3 ubiquitin ligases in Arabidopsis. Mol. Cells 2011, 31, 201–208. [Google Scholar] [CrossRef]
- Petroski, M.D.; Deshaies, R.J. Function and regulation of cullin-RING ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 2005, 6, 9–20. [Google Scholar] [CrossRef]
- Lechner, E.; Leonhardt, N.; Eisler, H.; Parmentier, Y.; Alioua, M.; Jacquet, H.; Leung, J.; Genschik, P. MATH/BTB CRL3 receptors target the homeodomain-leucine zipper ATHB6 to modulate abscisic acid signaling. Dev. Cell 2011, 21, 1116–1128. [Google Scholar] [CrossRef]
- Weber, H.; Bernhardt, A.; Dieterle, M.; Hano, P.; Mutlu, A.; Estelle, M.; Genschik, P.; Hellmann, H. Arabidopsis AtCUL3a and AtCUL3b form complexes with members of the BTB/POZ-MATH protein family. Plant Physiol. 2005, 137, 83–93. [Google Scholar] [CrossRef]
- Thomann, A.; Dieterle, M.; Genschik, P. Plant CULLIN-based E3s: Phytohormones come first. FEBS Lett. 2005, 579, 3239–3245. [Google Scholar] [CrossRef] [PubMed]
- Stogios, P.J.; Downs, G.S.; Jauhal, J.J.; Nandra, S.K.; Privé, G. Sequence and structural analysis of BTB domain proteins. Genome Biol. 2005, 6, R82. [Google Scholar] [CrossRef] [PubMed]
- Dow, M.R.; Mains, P.E. Genetic and Molecular Characterization of the Caenorhabditis elegans Gene, mel-26, a Postmeiotic Negative Regulator of MEI-1, a Meiotic-Specific Spindle Component. Genetics 1998, 150, 119–128. [Google Scholar] [PubMed]
- Bomont, P.; Cavalier, L.; Blondeau, F.; Hamida, C.B.; Belal, S.; Tazir, M.; Demir, E.; Topaloglu, H.; Korinthenberg, R.; Tüysüz, B.; et al. The gene encoding gigaxonin, a new member of the cytoskeletal BTB/kelch repeat family, is mutated in giant axonal neuropathy. Nat. Genet. 2000, 26, 370. [Google Scholar] [CrossRef] [PubMed]
- Collins, T.; Stone, J.R.; Williams, A.J. All in the family: The BTB/POZ, KRAB, and SCAN domains. Mol. Cell. Biol. 2001, 21, 3609–3615. [Google Scholar] [CrossRef] [PubMed]
- Spoel, S.H.; Koornneef, A.; Claessens, S.M.; Korzelius, J.P.; Van Pelt, J.A.; Mueller, M.J.; Buchala, A.J.; Métraux, J.-P.; Brown, R.; Kazan, K.; et al. NPR1 Modulates Cross-Talk between Salicylate- and Jasmonate-Dependent Defense Pathways through a Novel Function in the Cytosol. Plant Cell 2003, 15, 760–770. [Google Scholar] [CrossRef]
- Dong, X. NPR1, all things considered. Curr. Opin. Plant Biol. 2004, 7, 547–552. [Google Scholar] [CrossRef]
- Pieterse, C.M.; Van Loon, L.C. NPR1: The spider in the web of induced resistance signaling pathways. Curr. Opin. Plant Biol. 2004, 7, 456–464. [Google Scholar] [CrossRef]
- Kim, H.; Kim, S.H.; Seo, D.H.; Chung, S.; Kim, S.W.; Lee, J.S.; Kim, W.T.; Lee, J.-H. ABA-HYPERSENSITIVE BTB/POZ PROTEIN 1 functions as a negative regulator in ABA-mediated inhibition of germination in Arabidopsis. Plant Mol. Biol. 2016, 90, 303–315. [Google Scholar] [CrossRef]
- Huang, K.; Peng, L.; Liu, Y.; Yao, R.; Liu, Z.; Li, X.; Yang, Y.; Wang, J. Arabidopsis calcium-dependent protein kinase AtCPK1 plays a positive role in salt/drought-stress response. Biochem. Biophys. Res. Commun. 2018, 498, 92–98. [Google Scholar] [CrossRef]
- Cao, H.G.J.; Clarke, J.D.; Volko, S.; Dong, X. The Arabidopsis NPR1 gene that controls systemic acquired resistance encodes a novel protein containing ankyrin repeats. Cell 1997, 88, 57–63. [Google Scholar] [CrossRef]
- Larbat, R.; Le Bot, J.; Bourgaud, F.; Robin, C.; Adamowicz, S. Organ-specific responses of tomato growth and phenolic metabolism to nitrate limitation. Plant Biol. 2012, 14, 760–769. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Huang, B. Root proteomic responses to heat stress in two Agrostis grass species contrasting in heat tolerance. J. Exp. Bot. 2008, 59, 4183–4194. [Google Scholar] [CrossRef] [PubMed]
- Malnoy, M.; Jin, Q.; Borejsza-Wysocka, E.E.; He, S.Y.; Aldwinckle, H.S. Overexpression of the apple MpNPR1 gene confers increased disease resistance in Malus x domestica. Mol. Plant-Microbe Interact. 2007, 12, 1568–1580. [Google Scholar] [CrossRef]
- Zhang, J.Y.; Qu, S.C.; Qiao, Y.S.; Zhang, Z.; Guo, Z.R. Overexpression of the Malus hupehensis MhNPR1 gene increased tolerance to salt and osmotic stress in transgenic tobacco. Mol. Biol. Rep. 2014, 41, 1553–1561. [Google Scholar] [CrossRef]
- Apel, K.; Hirt, H. Reactive oxygen species: Metabolism, oxidative stress, and signal transduction. Annu. Rev. Plant Biol. 2004, 55, 373–399. [Google Scholar] [CrossRef]
- Canet, J.V.; Dobón, A.; Fajmonová, J.; Tornero, P. The BLADE-ON-PETIOLE genes of Arabidopsis are essential for resistance induced by methyl jasmonate. BMC Plant Biol. 2012, 12, 199. [Google Scholar] [CrossRef]
- Hu, Y.; Chen, L.; Wang, H.; Zhang, L.; Wang, F.; Yu, D. Arabidopsis transcription factor WRKY8 functions antagonistically with its interacting partner VQ9 to modulate salinity stress tolerance. Plant J. Cell Mol. Biol. 2013, 74, 730–745. [Google Scholar] [CrossRef]
- Wang, X.; Hou, C.; Zheng, K.; Li, Q.; Chen, S.; Wang, S. Overexpression of ERF96, a small ethylene response factor gene, enhances salt tolerance in Arabidopsis. Biol. Plant. 2017, 61, 693–701. [Google Scholar] [CrossRef]
- Wang, X.; Huang, W.; Yang, Z.; Liu, J.; Huang, B. Transcriptional regulation of heat shock proteins and ascorbate peroxidase by CtHsfA2b from African bermudagrass conferring heat tolerance in Arabidopsis. Sci. Rep. 2016, 6, 28021. [Google Scholar] [CrossRef]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Yoo, S.D.; Cho, Y.H.; Sheen, J. Arabidopsis mesophyll protoplasts: A versatile cell system for transient gene expression analysis. Nat. Protoc. 2007, 2, 1565–1572. [Google Scholar] [CrossRef] [PubMed]
- Byun, M.Y.; Lee, J.; Cui, L.H.; Kang, Y.; Oh, T.K.; Park, H.; Lee, H.; Kim, W.T. Constitutive expression of DaCBF7, an Antarctic vascular plant Deschampsia antarctica CBF homolog, resulted in improved cold tolerance in transgenic rice plants. Plant Sci. Int. J. Exp. Plant Biol. 2015, 236, 61–74. [Google Scholar] [CrossRef] [PubMed]
- Lichtenthaler, H.K. Chlorophylls and carotenoids: Pigments of photosynthetic biomembranes. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 1987; p. 48. [Google Scholar]
- Liu, D.; Gong, Q.; Ma, Y.; Li, P.; Li, J.; Yang, S.; Yuan, L.; Yu, Y.; Pan, D.; Xu, F.; et al. cpSecA, a thylakoid protein translocase subunit, is essential for photosynthetic development in Arabidopsis. J. Exp. Bot. 2010, 61, 1655–1669. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).