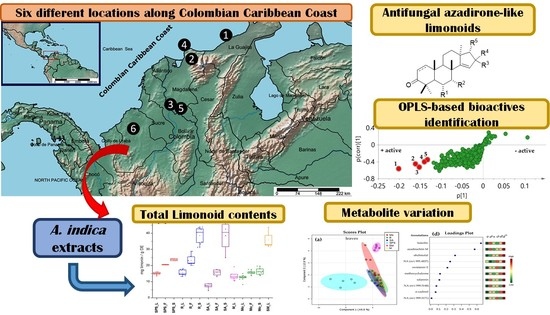
Chemical and Antifungal Variability of Several Accessions of Azadirachta indica A. Juss. from Six Locations Across the Colombian Caribbean Coast: Identification of Antifungal Azadirone Limonoids
Abstract
1. Introduction
2. Results and Discussion
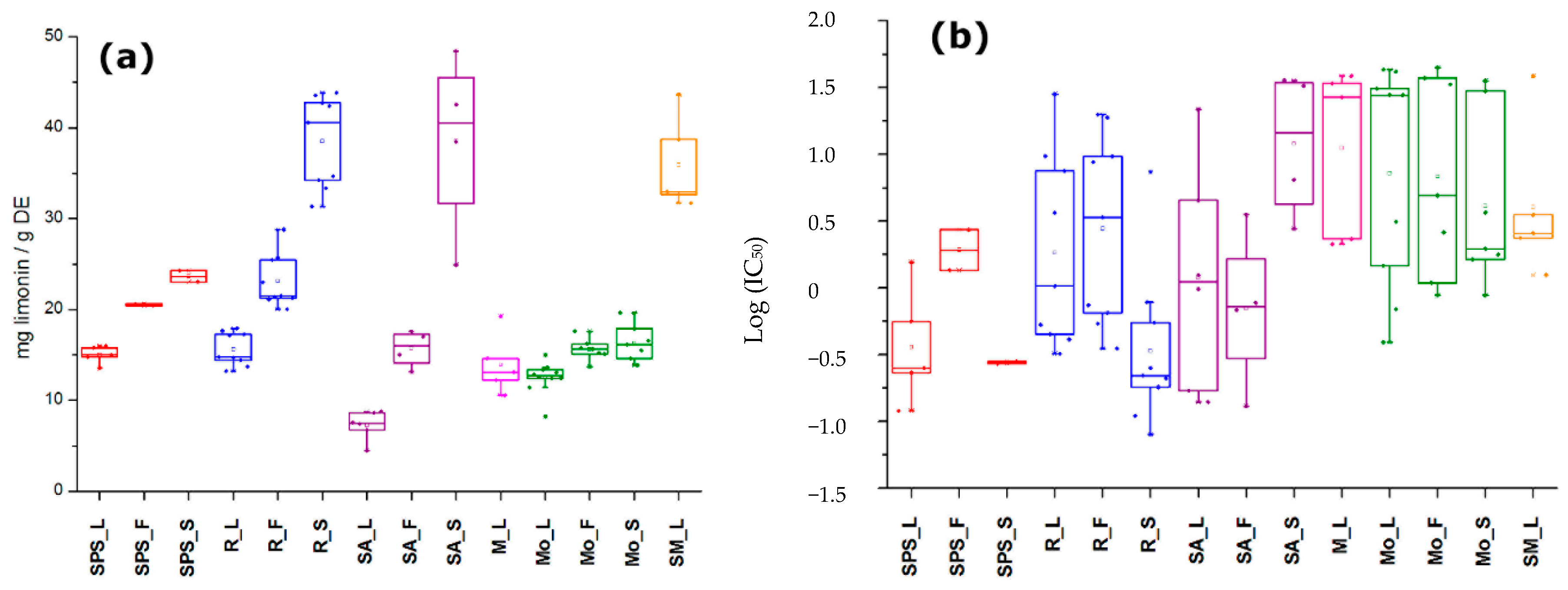
2.1. Total Limonoid Contents and Antifungal Activity of A. indica-Derived Extracts
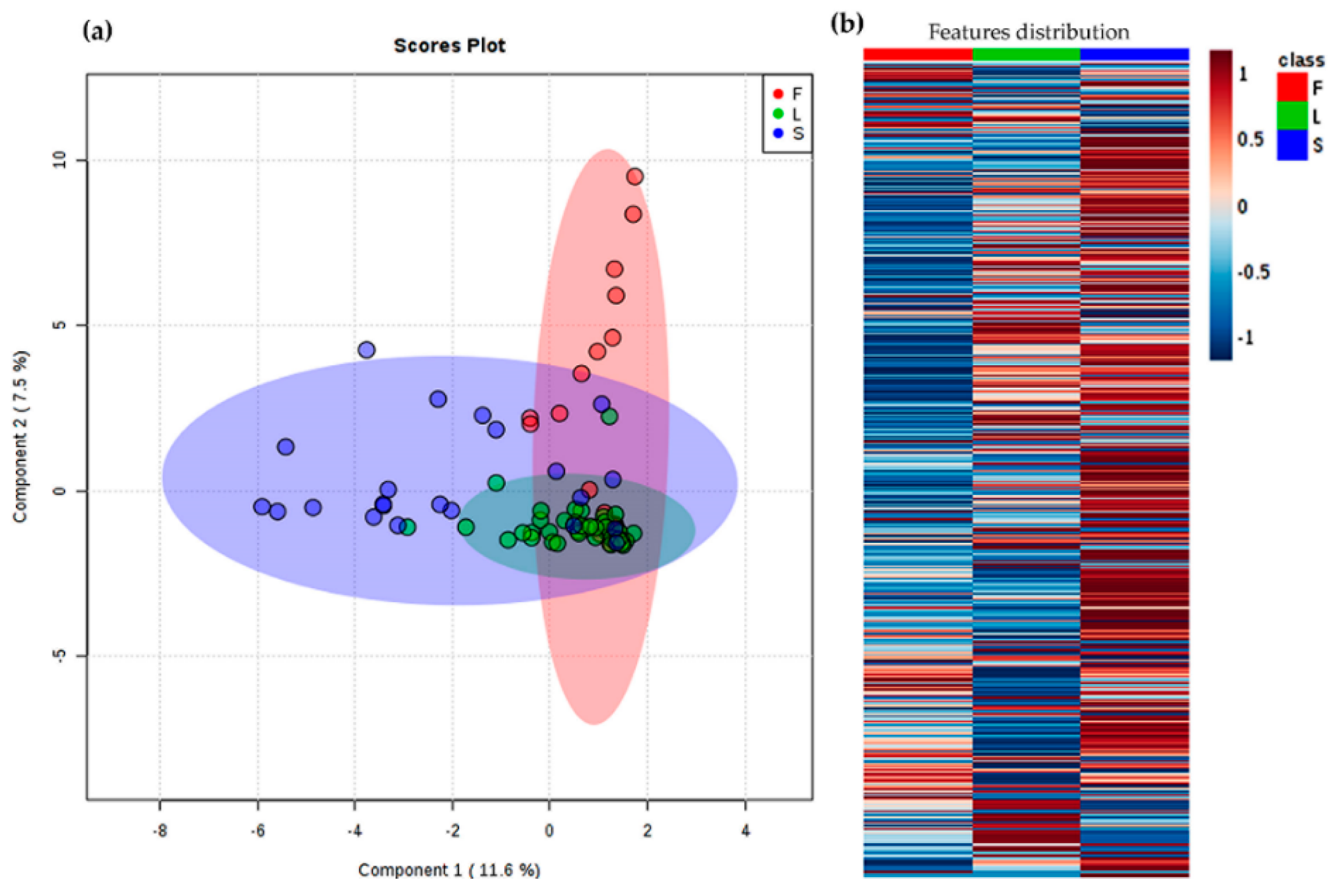
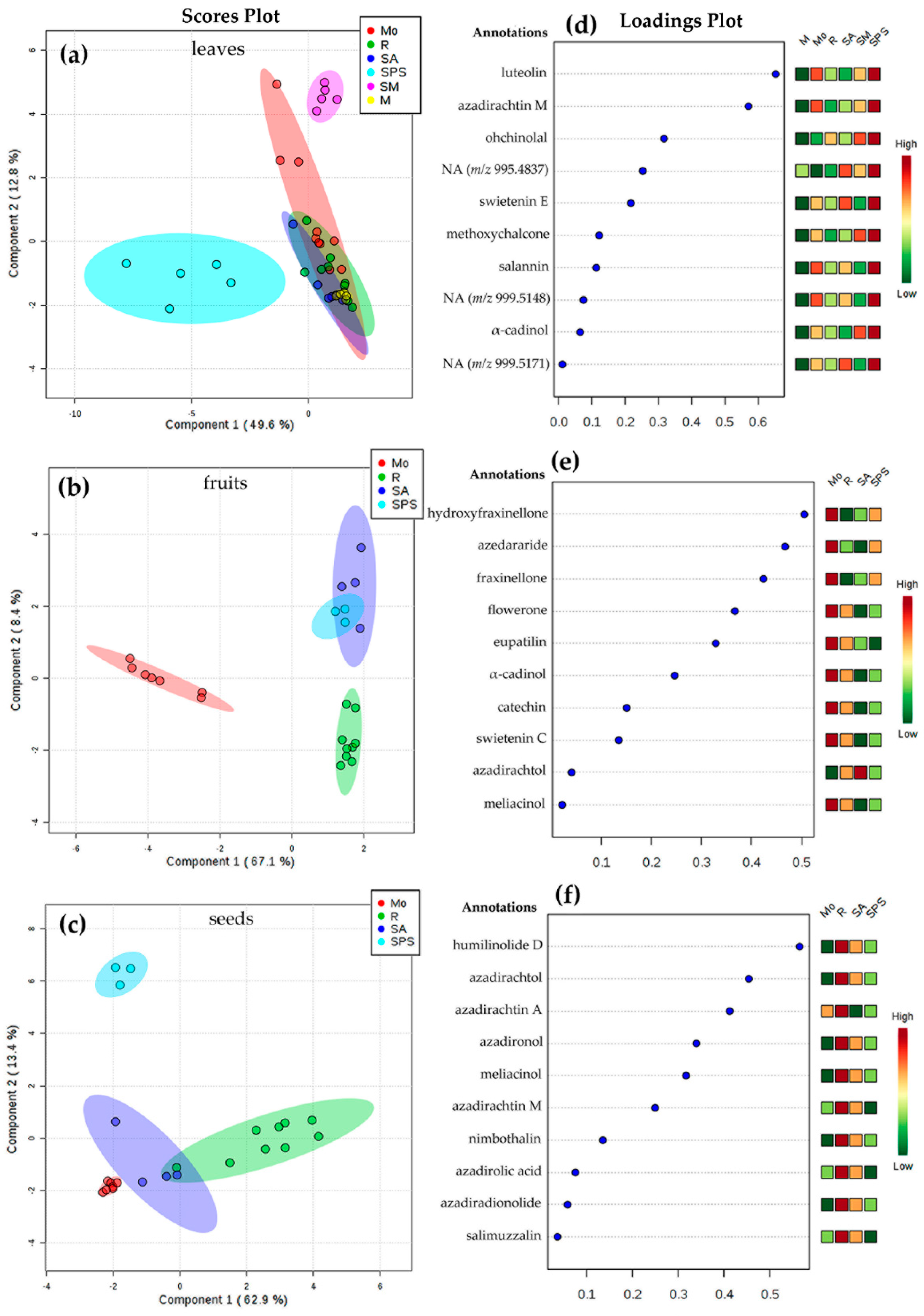
2.2. Chemical Variability through Metabolic Profiling of Crude Extracts of A. indica
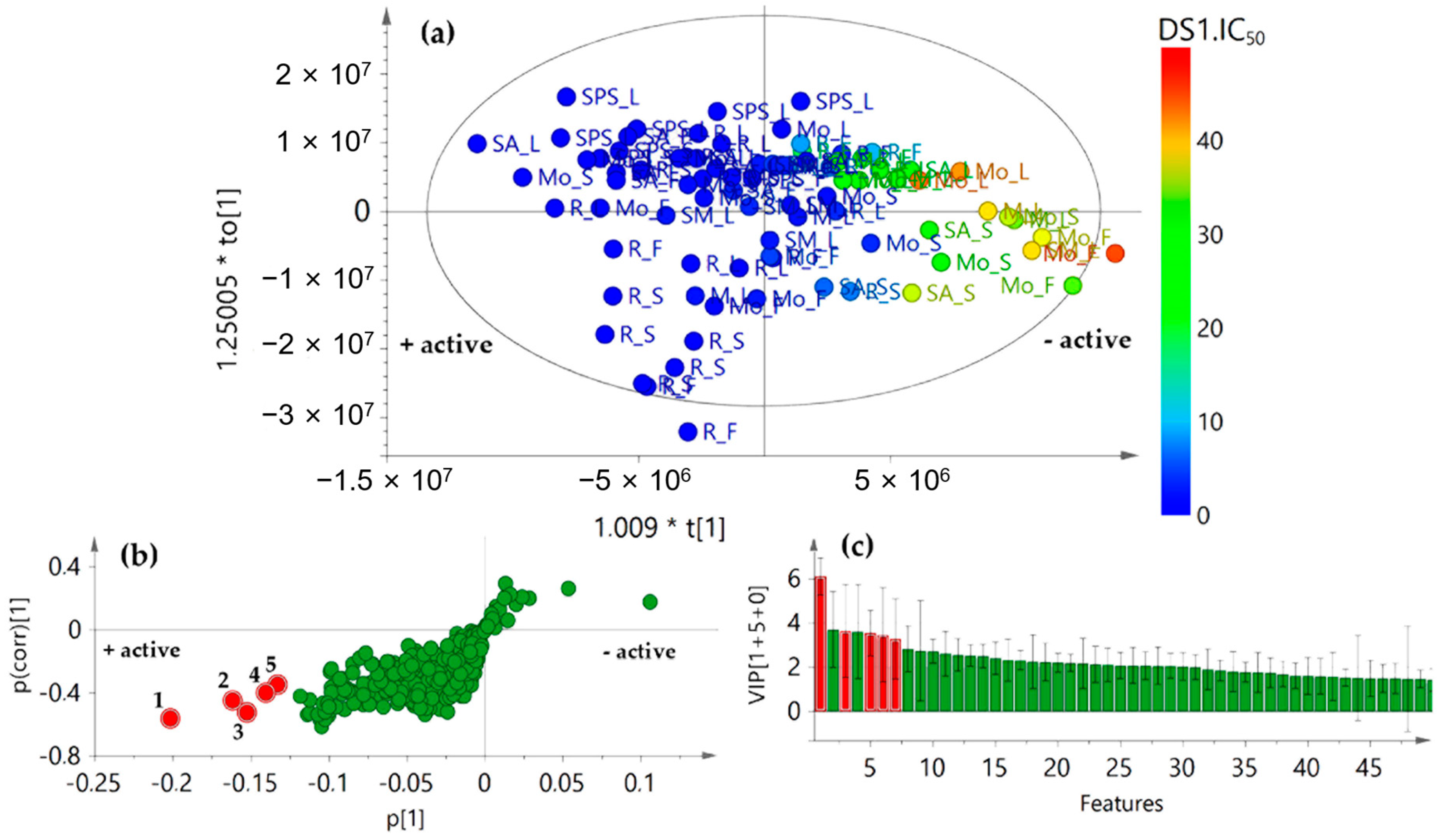
2.3. Integration of Chemical and Antifungal Activity Datasets for Detecting Bioactives
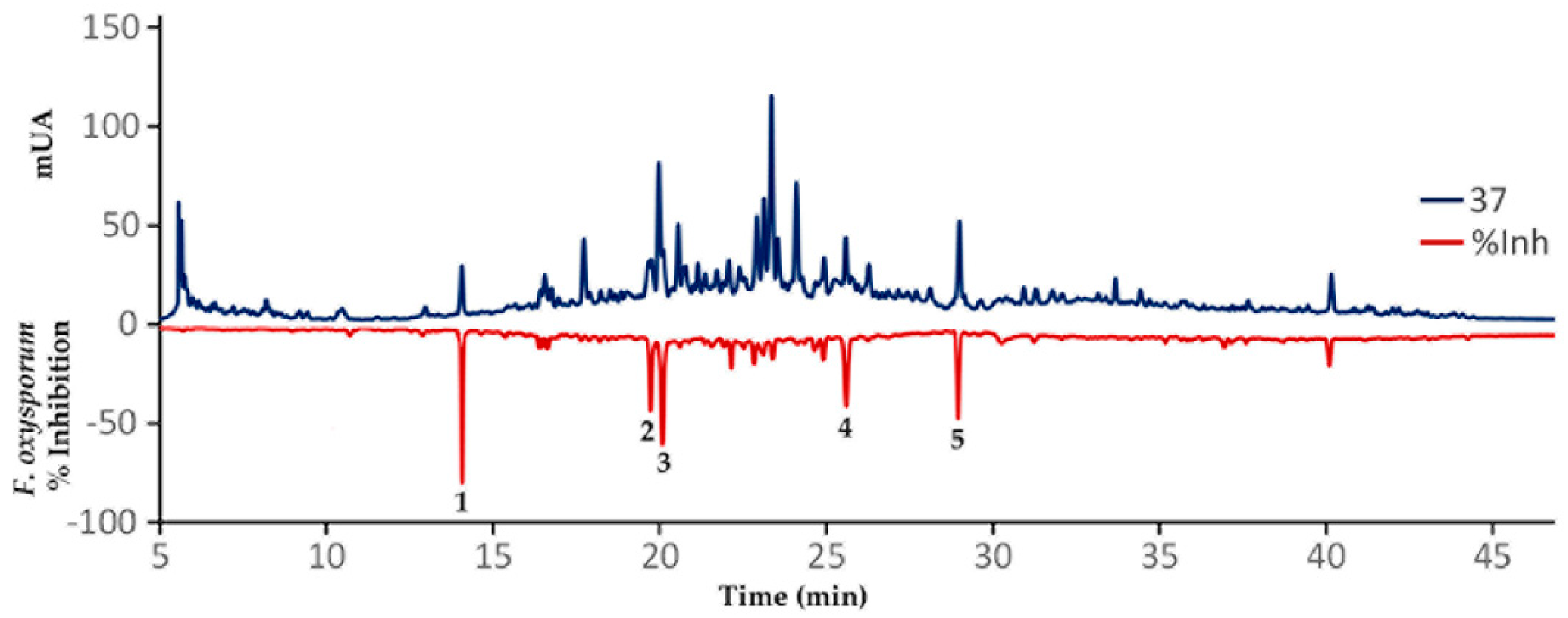
2.4. Validation of the OPLS Model by Direct Identification of Bioactives after High Performance Liquid Chromatography (HPLC)-Based Microfractionation
3. Materials and Methods
3.1. Plant Collection
3.2. Preparation of Crude Ethanolic Extracts
3.3. Measurement of Total Limonoid Contents (TLiC)
3.4. Analysis of the Effect of A. indica Extracts on Viability and Germination of F. oxysporum conidia
3.5. Liquid Chromatography Coupled to Mass Spectrometry (LC-MS) Analysis and Peak Annotations
3.6. Microfractionation of Extract 37 (SA_L_1) by Analytical-Scale High Performance Liquid Chromatography (HPLC)
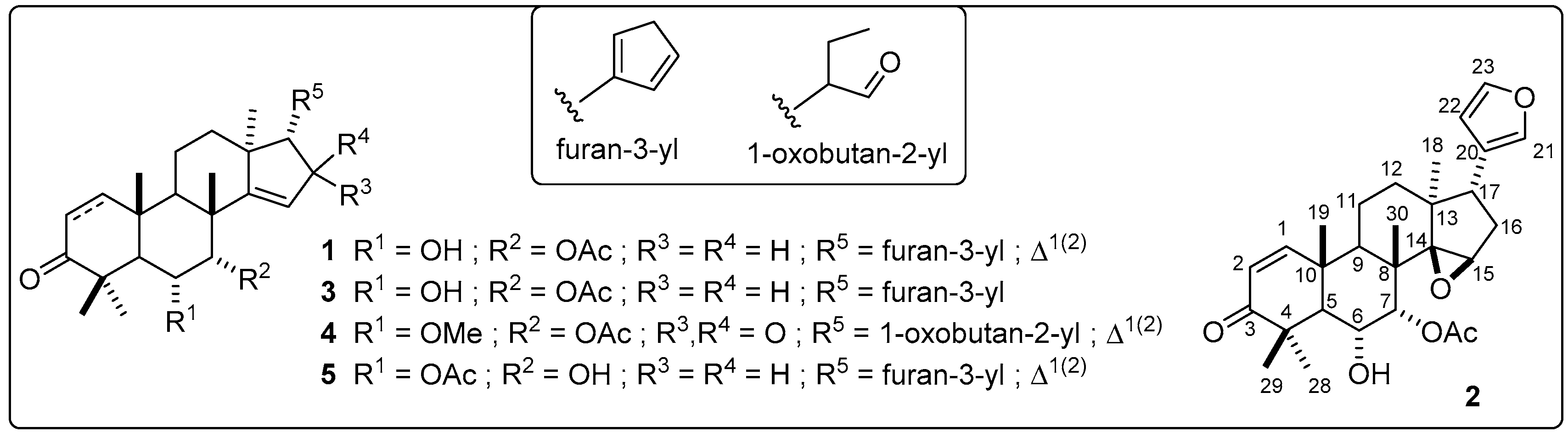
3.7. Purification and Identification of Most-Active Compounds from Extract 37 by Semipreparative-Scale HPLC
3.8. Statistical Analyses
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
| Location | Accession | Coordinates |
|---|---|---|
| San Pedro de la Sierra (SPS) | 1 | 10°53′6.31″ N 74° 6′28.44″ W |
| 2 | 10°53′40.66″ N 74° 5′56.10″ W | |
| 3 | 10°53′41.15″ N 74° 5′53.93″ W | |
| 4 | 10°53′20.87″ N 74° 6′16.43″ W | |
| 5 | 10°52′53.63″ N 74° 9′9.93″ W | |
| Santa Ana (SA) | 1 | 9°19′19.10″ N 74°34′46.31″ W |
| 2 | 9°19′39.14″ N 74°34′48.64″ W | |
| 3 | 9°19′39.49″ N 74°34′47.71″ W | |
| 4 | 9°19′39.62″ N 74°34′47.48″ W | |
| 5 | 9°19′39.71″ N 74°34′47.29″ W | |
| 6 | 9°19′13.63″ N 74°34′23.03″ W | |
| Riohacha (R) | 1 | 11°31′28.03″ N 72°54′7.55″ W |
| 2 | 11°31′27.97″ N 72°54′7.90″ W | |
| 3 | 11°31′28.84″ N 72°54′8.95″ W | |
| 4 | 11°31′30.62″ N 72°54′13.34″ W | |
| 5 | 11°31′30.18″ N 72°54′13.74″ W | |
| 6 | 11°31′29.21″ N 72°54′15.56″ W | |
| 7 | 11°31′29.97″ N 72°54′16.63″ W | |
| 8 | 11°31′29.68″ N 72°54′18.33″ W | |
| 9 | 11°31′26.78″ N 72°54′11.65″ W | |
| Mompox (M) | 1 | 9°14′36.17″ N 74°25′37.73″ W |
| 2 | 9°14′36.06″ N 74°25′37.68″ W | |
| 3 | 9°14′28.10″ N 74°25′22.49″ W | |
| 4 | 9°14′29.11″ N 74°25′24.05″ W | |
| 5 | 9°14′17.76″ N 74°25′15.06″ W | |
| Santa Marta (SM) | 1 | 11°13′26.45″ N 74°11′6.14″ W |
| 2 | 11°13′26.29″ N 74°11′6.63″ W | |
| 3 | 11°13′26.16″ N 74°11′5.90″ W | |
| 4 | 11°13′26.07″ N 74°11′6.15″ W | |
| 5 | 11°13′25.92″ N 74°11′6.57″ W | |
| Montería (Mo) | 1 | 8°45′44.71″ N 75°52′39.16″ W |
| 2 | 8°45′50.04″ N 75°52′43.17″ W | |
| 3 | 8°45′50.12″ N 75°52′43.08″ W | |
| 4 | 8°45′50.20″ N 75°52′43.00″ W | |
| 5 | 8°45′50.27″ N 75°52′42.96″ W | |
| 6 | 8°45′5.16″ N 75°53′27.34″ W | |
| 7 | 8°45′5.34″ N 75°53′27.24″ W | |
| 8 | 8°45′3.99″ N 75°53′27.19″ W | |
| 9 | 8°45′3.98″ N 75°53′26.99″ W | |
| 10 | 8°45′6.67″ N 75°53′28.20″ W |
| No | Extract a | TLiC b | No | Extract a | TLiC b | No | Extract a | TLiC b | No | Extract a | TLiC b |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | SPS_L_1 | 13.5 ± 0.5 | 22 | R_F_4 | 25.7 ± 1.5 | 43 | SA_F_2 | 17.6 ± 2.9 | 64 | Mo_L_9 | 12.4 ± 0.7 |
| 2 | SPS_L_2 | 14.8 ± 0.8 | 23 | R_F_5 | 21.1 ± 2.1 | 44 | SA_F_4 | 15.1 ± 1.8 | 65 | Mo_L_10 | 11.4 ± 0.8 |
| 3 | SPS_L_3 | 16.0 ± 0.6 | 24 | R_F_6 | 25.5 ± 1.5 | 45 | SA_F_5 | 13.1 ± 1.5 | 66 | Mo_F_1 | 13.7 ± 3.8 |
| 4 | SPS_L_4 | 15.8 ± 0.3 | 25 | R_F_7 | 28.8 ± 2.4 | 46 | SA_F_6 | 17.0 ± 2.7 | 67 | Mo_F_2 | 15.6 ± 0.7 |
| 5 | SPS_L_5 | 15.0 ± 1.0 | 26 | R_F_8 | 21.3 ± 2.1 | 47 | SA_S_2 | 48.5 ± 1.7 | 68 | Mo_F_3 | 16.2 ± 0.9 |
| 6 | SPS_F_4 | 20.4 ± 0.3 | 27 | R_F_9 | 23 ± 1.1.8 | 48 | SA_S_4 | 24.9 ± 4.2 | 69 | Mo_F_4 | 15.2 ± 0.4 |
| 7 | SPS_F_5 | 20.6 ± 1.4 | 28 | R_S_1 | 42.8 ± 3.2 | 49 | SA_S_5 | 38.5 ± 9.4 | 70 | Mo_F_5 | 15.7 ± 0.4 |
| 8 | SPS_S_4 | 23.0 ± 0.4 | 29 | R_S_2 | 33.4 ± 1.3 | 50 | SA_S_6 | 42.5 ± 5.8 | 71 | Mo_F_6 | 15.1 ± 1.1 |
| 9 | SPS_S_5 | 24.3 ± 0.7 | 30 | R_S_3 | 43.6 ± 1.0 | 51 | M_L_1 | 19.2 ± 0.7 | 72 | Mo_F_7 | 17.6 ± 2.9 |
| 10 | R_L_1 | 17.9 ± 1.6 | 31 | R_S_4 | 34.2 ± 1.5 | 52 | M_L_2 | 10.5 ± 0.8 | 73 | Mo_S_1 | 13.9 ± 3.0 |
| 11 | R_L_2 | 14.7 ± 3.0 | 32 | R_S_5 | 34.7 ± 6.1 | 53 | M_L_3 | 12.2 ± 0.8 | 74 | Mo_S_2 | 17.9 ± 4.2 |
| 12 | R_L_3 | 13.2 ± 2.6 | 33 | R_S_6 | 31.3 ± 2.2 | 54 | M_L_4 | 13.1 ± 1.5 | 75 | Mo_S_3 | 16.1 ± 1.3 |
| 13 | R_L_4 | 14.4 ± 2.1 | 34 | R_S_7 | 42.4 ± 2.4 | 55 | M_L_5 | 14.6 ± 7.3 | 76 | Mo_S_4 | 15.5 ± 1.3 |
| 14 | R_L_5 | 17.1 ± 1.4 | 35 | R_S_8 | 40.6 ± 2.7 | 56 | Mo_L_1 | 13.7 ± 1.8 | 77 | Mo_S_5 | 14.6 ± 4.4 |
| 15 | R_L_6 | 14.8 ± 1.8 | 36 | R_S_9 | 43.9 ± 1.9 | 57 | Mo_L_2 | 13.4 ± 2.4 | 78 | Mo_S_6 | 19.6 ± 1.8 |
| 16 | R_L_7 | 13.7 ± 1.7 | 37 | SA_L_1 | 6.7 ± 0.5 | 58 | Mo_L_3 | 12.4 ± 1.0 | 79 | Mo_S_7 | 16.5 ± 1.4 |
| 17 | R_L_8 | 17.3 ± 0.5 | 38 | SA_L_2 | 4.5 ± 0.4 | 59 | Mo_L_4 | 15.0 ± 0.3 | 80 | SM_L_1 | 32.6 ± 2.4 |
| 18 | R_L_9 | 17.6 ± 0.7 | 39 | SA_L_3 | 7.4 ± 0.6 | 60 | Mo_L_5 | 12.5 ± 0.4 | 81 | SM_L_2 | 33.0 ± 2.3 |
| 19 | R_F_1 | 21.5 ± 1.0 | 40 | SA_L_4 | 8.7 ± 0.8 | 61 | Mo_L_6 | 13.1 ± 0.9 | 82 | SM_L_3 | 31.7 ± 1.0 |
| 20 | R_F_2 | 21.4 ± 1.2 | 41 | SA_L_5 | 7.6 ± 0.8 | 62 | Mo_L_7 | 8.2 ± 0.5 | 83 | SM_L_4 | 43.6 ± 4.3 |
| 21 | R_F_3 | 20.0 ± 1.1 | 42 | SA_L_6 | 8.8 ± 0.5 | 63 | Mo_L_8 | 12.9 ± 0.6 | 84 | SM_L_5 | 38.7 ± 2.2 |
| No | Extract a | IC50 c | No | Extract a | IC50 c | No | Extract a | IC50 c | No | Extract a | IC50 c |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | SPS_L_1 | 0.56 ± 0.04 | 22 | R_F_4 | 9.63 ± 0.58 | 43 | SA_F_2 | 3.53 ± 0.28 | 64 | Mo_L_9 | 0.39 ± 0.05 |
| 2 | SPS_L_2 | 1.56 ± 0.12 | 23 | R_F_5 | 19.8 ± 0.99 | 44 | SA_F_4 | 0.77 ± 0.03 | 65 | Mo_L_10 | 0.16 ± 0.01 |
| 3 | SPS_L_3 | 0.23 ± 0.02 | 24 | R_F_6 | 0.65 ± 0.03 | 45 | SA_F_5 | 0.13 ± 0.01 | 66 | Mo_F_1 | 44.8 ± 1.79 |
| 4 | SPS_L_4 | 0.12 ± 0.01 | 25 | R_F_7 | 0.54 ± 0.04 | 46 | SA_F_6 | 0.68 ± 0.05 | 67 | Mo_F_2 | 4.91 ± 0.39 |
| 5 | SPS_L_5 | 0.25 ± 0.03 | 26 | R_F_8 | 0.35 ± 0.04 | 47 | SA_S_2 | 2.78 ± 0.25 | 68 | Mo_F_3 | 37.1 ± 3.71 |
| 6 | SPS_F_4 | 2.72 ± 0.27 | 27 | R_F_9 | 0.74 ± 0.03 | 48 | SA_S_4 | 32.6 ± 3.26 | 69 | Mo_F_4 | 2.60 ± 0.16 |
| 7 | SPS_F_5 | 1.35 ± 0.08 | 28 | R_S_1 | 0.25 ± 0.02 | 49 | SA_S_5 | 6.44 ± 0.52 | 70 | Mo_F_5 | 33.4 ± 4.01 |
| 8 | SPS_S_4 | 0.28 ± 0.03 | 29 | R_S_2 | 0.22 ± 0.01 | 50 | SA_S_6 | 35.8 ± 2.15 | 71 | Mo_F_6 | 0.88 ± 0.04 |
| 9 | SPS_S_5 | 0.27 ± 0.03 | 30 | R_S_3 | 0.18 ± 0.01 | 51 | M_L_1 | 2.32 ± 0.09 | 72 | Mo_F_7 | 1.09 ± 0.12 |
| 10 | R_L_1 | 0.32 ± 0.02 | 31 | R_S_4 | 0.11 ± 0.01 | 52 | M_L_2 | 26.8 ± 2.14 | 73 | Mo_S_1 | 1.95 ± 0.14 |
| 11 | R_L_2 | 3.64 ± 0.44 | 32 | R_S_5 | 7.35 ± 0.29 | 53 | M_L_3 | 2.12 ± 0.08 | 74 | Mo_S_2 | 1.63 ± 0.15 |
| 12 | R_L_3 | 1.03 ± 0.10 | 33 | R_S_6 | 0.08 ± 0.01 | 54 | M_L_4 | 38.6 ± 2.32 | 75 | Mo_S_3 | 1.77 ± 0.09 |
| 13 | R_L_4 | 0.45 ± 0.05 | 34 | R_S_7 | 0.55 ± 0.06 | 55 | M_L_5 | 33.9 ± 3.39 | 76 | Mo_S_4 | 29.8 ± 2.68 |
| 14 | R_L_5 | 9.67 ± 0.77 | 35 | R_S_8 | 0.21 ± 0.02 | 56 | Mo_L_1 | 41.6 ± 3.74 | 77 | Mo_S_5 | 3.67 ± 0.18 |
| 15 | R_L_6 | 28.2 ± 2.5 | 36 | R_S_9 | 0.78 ± 0.05 | 57 | Mo_L_2 | 27.9 ± 1.40 | 78 | Mo_S_6 | 35.6 ± 4.27 |
| 16 | R_L_7 | 7.52 ± 0.83 | 37 | SA_L_1 | 0.14 ± 0.01 | 58 | Mo_L_3 | 3.12 ± 0.22 | 79 | Mo_S_7 | 0.88 ± 0.04 |
| 17 | R_L_8 | 0.41 ± 0.04 | 38 | SA_L_2 | 0.17 ± 0.02 | 59 | Mo_L_4 | 31.2 ± 3.74 | 80 | SM_L_1 | 2.56 ± 0.23 |
| 18 | R_L_9 | 0.53 ± 0.03 | 39 | SA_L_3 | 1.25 ± 0.08 | 60 | Mo_L_5 | 42.9 ± 5.15 | 81 | SM_L_2 | 38.7 ± 4.26 |
| 19 | R_F_1 | 18.8 ± 1.32 | 40 | SA_L_4 | 0.98 ± 0.11 | 61 | Mo_L_6 | 27.8 ± 1.95 | 82 | SM_L_3 | 2.36 ± 0.21 |
| 20 | R_F_2 | 3.37 ± 0.40 | 41 | SA_L_5 | 21.6 ± 1.30 | 62 | Mo_L_7 | 1.46 ± 0.13 | 83 | SM_L_4 | 1.25 ± 0.10 |
| 21 | R_F_3 | 8.74 ± 1.05 | 42 | SA_L_6 | 4.51 ± 0.18 | 63 | Mo_L_8 | 0.69 ± 0.08 | 84 | SM_L_5 | 3.54 ± 0.28 |
| Leaves | Fruits | Seeds | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| No a | m/z [M+H]+ | tR (min) | MF b | Annotation | Error | No a | m/z [M+H]+ | tR (min) | MF b | Annotation | Error | No a | m/z [M+H]+ | tR (min) | MF b | Annotation | Error |
| 1 | 287.0563 | 18.2 | C15H11O6+ | luteolin | −2.79 | 1 | 249.1133 | 11.8 | C14H17O4+ | hydroxyfraxinellone | −2.79 | 1 | 587.2479 | 27.6 | C31H39O11+ | humilinolide D | 2.21 |
| 2 | 635.2712 | 18.9 | C32H43O13+ | azadirachtin M | −1.42 | 2 | 261.1137 | 8.1 | C15H17O4+ | azedararide | −1.42 | 2 | 527.3379 | 23.7 | C32H47O6+ | azadirachtol | −1.33 |
| 3 | 613.3021 | 13.7 | C34H45O10+ | ohchinolal | −1.47 | 3 | 233.1168 | 9.9 | C14H17O3+ | fraxinellone | −1.47 | 3 | 721.2716 | 29.5 | C35H45O16+ | azadirachtin A | −1.25 |
| 4 | 995.4837 | 18.5 | C50H75O20+ | NAc | 1.41 | 4 | 357.1325 | 16.4 | C20H21O6+ | flowerone | 1.41 | 4 | 523.3041 | 20.9 | C32H43O6+ | azadironol | 3.44 |
| 5 | 571.2919 | 19.9 | C32H43O9+ | swietenin E | −2.10 | 5 | 345.0986 | 10.4 | C18H17O7+ | eupatilin | −2.10 | 5 | 555.2963 | 17.6 | C32H43O8+ | meliacinol | −1.08 |
| 6 | 239.1081 | 14.5 | C16H15O2+ | methoxychalcone | −3.76 | 6 | 223.2069 | 21.9 | C15H27O+ | α-cadinol | −3.76 | 6 | 635.2711 | 18.9 | C32H43O13+ | azadirachtin M | −1.26 |
| 7 | 597.3069 | 24.8 | C34H45O9+ | salannin | −1.00 | 7 | 291.0851 | 5.81 | C15H15O6+ | catechin | −1.00 | 7 | 369.2049 | 26.1 | C23H29O4+ | nimbothalin | 4.33 |
| 8 | 999.5148 | 22.5 | C50H79O20+ | NAc | 1.60 | 8 | 557.2766 | 34.1 | C31H41O9+ | swietenin C | 1.60 | 8 | 499.3069 | 26.9 | C30H43O6+ | azadirolic acid | −2.00 |
| 9 | 223.2071 | 21.9 | C15H27O+ | α-cadinol | −4.48 | 9 | 527.3361 | 23.7 | C32H47O6+ | azadirachtol | −4.48 | 9 | 467.2421 | 34.8 | C28H35O6+ | azadiradionolide | 2.57 |
| 10 | 999.5171 | 21.3 | C50H79O20+ | NAc | −0.70 | 10 | 555.2971 | 17.6 | C32H43O8+ | meliacinol | −0.70 | 10 | 513.2859 | 37.4 | C30H41O7+ | salimuzzalin | −1.36 |
| 1 | 2 | 3 | 4 | 5 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| No | Type | δC | δH, mult, J (Hz) | Type | δC | δH, mult, J (Hz) | Type | δC | δH, mult, J (Hz) | Type | δC | δH, mult, J (Hz) | Type | δC | δH, mult, J (Hz) |
| C-1 | CH | 157.1 | 7.14, d, 10.2 | CH | 157.4 | 7.10, d, 9.8 | CH2 | 33.5 | 1.41–1.79, m | CH | 157.3 | 7.05, d, 10.2 | CH | 157.9 | 7.12, d, 10.3 |
| C-2 | CH | 125.9 | 5.92, d, 10.2 | CH | 126.1 | 5.95, d, 9.8 | CH2 | 38.3 | 2.18-2.41, m | CH | 126.8 | 5.89, d, 10.2 | CH | 126.3 | 5.93, d, 10.3 |
| C-3 | C | 206.1 | C | 205.9 | C | 209.7 | C | 204.1 | C | 205.7 | |||||
| C-4 | C | 40.4 | C | 40.5 | C | 41.2 | C | 43.4 | C | 40.8 | |||||
| C-5 | CH | 49.8 | 2.20, d, 11.5 | CH | 50.2 | 2.23, d, 11.8 | CH | 47.1 | 2.07, d, 11.0 | CH | 48.6 | 2.13, d, 11.31 | CH | 49.6 | 2.76, d, 12.2 |
| C-6 | CH | 68.2 | 4.39, dd, 11.5, 2.4 | CH | 68.6 | 4.34, dd, 11.8, 2.5 | CH | 68.8 | 4.17, dd, 11.0, 3.1 | CH | 67.5 | 4.25, dd, 2.7, 11.1 | CH | 76.4 | 5.44, dd, 12.2, 2.6 |
| C-7 | CH | 79.0 | 5.34, d, 2.4 | CH | 77.5 | 4.95, d, 2.7 | CH | 79.2 | 5.24, d, 3.1 | CH | 82.3 | 5.29, d, 2.7 | CH | 68.4 | 4.06, d, 2.6 |
| C-8 | C | 45.4 | C | 42.7 | C | 43.5 | C | 45.6 | C | 45.2 | |||||
| C-9 | CH | 37.3 | 2.15–2.28, m | CH | 39.3 | 2.50–2.67, m | CH | 34.3 | 2.60–2.75, m | CH | 38.6 | 2.22, dd, 14.1, 5.1 | CH | 37.3 | 2.31, d, 12.6, 6.4 |
| C-10 | C | 43.4 | C | 41.7 | C | 38.5 | C | 42.1 | C | 43.0 | |||||
| C-11 | CH2 | 16.6 | 1.95–2.04, m (Ha) 1.70–1.81, m (Hb) | CH2 | 16.4 | 1.90–2.02 m (Ha) 1.71–1.87, m (Hb) | CH2 | 16.5 | 1.40–1.89, m | CH2 | 16.5 | 2.35–2.50, m (Ha) 2.05–2.21, m (Hb) | CH2 | 16.3 | 2.21–2.38, m (Ha) 2.02–2.29, m (Hb) |
| C-12 | CH2 | 32.5 | 1.84–1.92, m (Ha) 2.26–2.33, m (Hb) | CH2 | 29.1 | 1.71–1.87, m (Ha) 2.40–2.42, m (Hb) | CH2 | 31.7 | 1.44–1.89, m | CH2 | 32.9 | 2.38–2.48, m (Ha) 2.79–2.85, m (Hb) | CH2 | 33.5 | 2.02–2.29, m (Ha) 2.36–2.43, m (Hb) |
| C-13 | C | 47.3 | C | 45.5 | C | 47.1 | C | 59.5 | C | 46.9 | |||||
| C-14 | C | 158.6 | C | 73.0 | C | 158.6 | C | 193.4 | C | 158.5 | |||||
| C-15 | CH | 119.4 | 5.45, dd, 2.7, 1.9 | CH | 57.3 | 3.48–3.55, m | CH | 119.7 | 5.35, m | CH | 123.9 | 5.75, s | CH | 119.0 | 5.56, dd, 3.5, 1.7 |
| C-16 | CH2 | 34.2 | 1.68–1.75, m (Ha) 2.32–2.41, m (Hb) | CH2 | 32.2 | 1.59–1.65, m (Ha) 2.10–2.31, m (Hb) | CH2 | 33.1 | 1.44–1.89, m | C | 208.0 | CH2 | 34.5 | 1.95–2.01, m (Ha) 2.41, ddd, 14.4, 7.2, 3.5 (Hb) | |
| C-17 | CH | 51.8 | 2.84, dd, 10.6, 7.1 | CH | 39.4 | 2.66, dd, 3.6, 2.9 | CH | 51.4 | CH | 56.4 | 3.20, dd, 8.1, 1.7 | CH | 51.6 | 2.44, dd, 7.2, 3.5 | |
| C-18 | CH3 | 20.4 | 0.85, s | CH3 | 21.8 | 0.95, s | CH3 | 20.7 | 0.77, s | CH3 | 18.4 | 1.08, s | CH3 | 20.6 | 0.84, s |
| C-19 | CH3 | 20.1 | 1.15, s | CH3 | 20.4 | 1.11, s | CH3 | 16.6 | 0.81, s | CH3 | 16.1 | 1.17, s | CH3 | 14.0 | 1.14, s |
| C-20 | C | 124.3 | C | 123.8 | C | 124.5 | CH | 143.6 | 3.33, m | C | 124.5 | ||||
| C-21 | CH | 142.8 | 7.25, dd, 1.7, 1.0 | CH | 142.8 | 7.36, dd, 1.6, 0.8 | CH | 142.6 | 7.35, m | C | 203.1 | 9.78, d, 7.5 | CH | 142.6 | 7.26, dd, 1.7, 0.8 |
| C-22 | CH | 111.1 | 6.32, dd, 1.7, 1.0 | CH | 111.3 | 6.19, dd, 1.6, 0.8 | CH | 111.7 | 6.17, m | CH2 | 32.1 | 2.02-2.34, m | CH | 111.2 | 6.25, dd, 1.7, 0.8 |
| C-23 | CH | 139.4 | 7.38, t, 1.7 | CH | 139.6 | 7.16, t, 1.6 | CH | 139.5 | 7.19, m | CH3 | 21.9 | 1.24, t, 8.2 | CH | 139.4 | 7.37, t, 1.7 |
| C-28 | CH3 | 31.6 | 1.27, s | CH3 | 31.8 | 1.19, s | CH3 | 31.6 | 1.132, s | CH3 | 25.0 | 1.22, s | CH3 | 26.9 | 1.19, s |
| C-29 | CH3 | 20.6 | 1.30, s | CH3 | 19.4 | 1.32, s | CH3 | 19.5 | 1.21, s | CH3 | 21.3 | 1.25, s | CH3 | 19.5 | 1.25, s |
| C-30 | CH3 | 26.8 | 1.39, s | CH3 | 21.8 | 1.39, s | CH3 | 26.2 | 1.25, s | CH3 | 25.7 | 1.26, s | CH3 | 20.1 | 1.32, s |
| COCH3 | C | 171.8 | C | 171.6 | C | 172.2 | C | 169.2 | C | 172.3 | |||||
| COCH3 | CH3 | 21.1 | 2.05, s | CH3 | 21.4 | 2.13, s | CH3 | 21.5 | 2.01, s | CH3 | 19.7 | 1.95, s | CH3 | 21.5 | 2.00, s |
| OCH3 | - | - | - | CH3 | 3.50, s | - | |||||||||
References
- Kumar, V.S.; Navaratnam, V. Neem (Azadirachta indica): Prehistory to contemporary medicinal uses to humankind. Asian Pac. J. Trop. Biomed. 2013, 3, 505–514. [Google Scholar] [CrossRef]
- Koriem, K.M.M. Review on pharmacological and toxicologyical effects of oleum azadirachti oil. Asian Pac. J. Trop. Biomed. 2013, 3, 834–840. [Google Scholar] [CrossRef]
- Gupta, S.C.; Prasad, S.; Tyagi, A.K.; Kunnumakkara, A.B.; Aggarwal, B.B. Neem (Azadirachta indica): An indian traditional panacea with modern molecular basis. Phytomedicine 2017, 34, 14–20. [Google Scholar] [CrossRef] [PubMed]
- Saleem, S.; Muhammad, G.; Hussain, M.A.; Bukhari, S.N.A. A comprehensive review of phytochemical profile, bioactives for pharmaceuticals, and pharmacological attributes of Azadirachta indica. Phyther. Res. 2018, 32, 1241–1272. [Google Scholar] [CrossRef] [PubMed]
- Tan, Q.-G.; Luo, X.-D. Meliaceous limonoids: Chemistry and biological activities. Chem. Rev. 2011, 111, 7437–7522. [Google Scholar] [CrossRef]
- Gualdani, R.; Cavalluzzi, M.M.; Lentini, G.; Habtemariam, S. The chemistry and pharmacology of citrus limonoids. Molecules 2016, 21, 1530. [Google Scholar] [CrossRef]
- Benelli, G.; Canale, A.; Toniolo, C.; Higuchi, A.; Murugan, K.; Pavela, R.; Nicoletti, M. Neem (Azadirachta indica): Towards the ideal insecticide? Nat. Prod. Res. 2017, 31, 369–386. [Google Scholar] [CrossRef]
- Mahmoud, D.A.; Hassanein, N.M.; Youssef, K.A.; Abou Zeid, M.A. Antifungal activity of different neem leaf extracts and the nimonol against some important human pathogens. Braz. J. Microbiol. 2011, 42, 1007–1016. [Google Scholar] [CrossRef]
- Niaz, I.; Kazmi, S.A.R. Effect of neem seed coat oil fractions on stored grain fungi. Int. J. Biol. Biotechnol. 2005, 2, 705–706. [Google Scholar]
- Moslem, M.A.; El-Kholie, E.M. Effect of Neem (Azardirachta indica A. Juss) seeds and leaves extract on some plant pathogenic fungi. Pak. J. Biol. Sci. 2009, 12, 1045–1048. [Google Scholar] [CrossRef]
- Dubey, R.C.; Kumar, R. Efficacy of azadirachtin and fungicides on growth and survival of sclerotia of Macrophomina phaseolina causing charcoal rot of soybean. Indian Phytopathol. 2003, 56, 216–217. [Google Scholar]
- Suresh, G.; Narasimhan, N.S.; Masilamani, S.; Partho, P.D.; Gopalakrishnan, G. Antifungal fractions and compounds from uncrushed Green Leaves of Azadirachta indica. Phytoparasitica 1997, 25, 33–39. [Google Scholar] [CrossRef]
- Poli, A.; Bertetti, D.; Rapetti, S.; Gullino, M.L.; Garibaldi, A. Characterization and identification of Colombian isolates of Fusarium oxysporum f. sp. Dianthi. J. Plant Pathol. 2013, 95, 255–263. [Google Scholar]
- Radwanski, S.A.; Wickens, G.E. Vegetative fallows and potential value of the Neem tree (Azadirachta indica) in the tropics. Econ. Bot. 1981, 35, 398–414. [Google Scholar] [CrossRef]
- Florez-Florez, C.P.; Leon-Pelaez, J.D.; Osorio-Vega, N.W.; Restrepo-Llano, M.F. Nutrient dynamics in forest plantations of Azadirachta indica (Meliaceae) established for restoration of degraded lands in Colombia. Rev. Biol. Trop. 2013, 61, 515–529. (In Spanish) [Google Scholar] [CrossRef][Green Version]
- Restrepo, M.F.; Florez, C.P.; Osorio, N.W.; León, J.D. Passive and active restoration strategies to activate soil biogeochemical nutrient cycles in a degraded tropical dry land. ISRN Soil Sci. 2013, 2013, 461984. [Google Scholar] [CrossRef]
- Barliza, J.C.; Rodríguez, O.B.; Peláez, J.D.L.; Chávez, L.F. Planted forests for open coal mine spoils rehabilitation in Colombian drylands: Contributions of fine litterfall through an age chronosequence. Ecol. Eng. 2019, 138, 180–187. [Google Scholar] [CrossRef]
- Isah, T. Stress and defense responses in plant secondary metabolites production. Biol. Res. 2019, 52, 39. [Google Scholar] [CrossRef]
- Yang, L.; Wen, K.-S.; Ruan, X.; Zhao, Y.-X.; Wei, F.; Wang, Q. Response of plant secondary metabolites to Environmental Factors. Molecules 2018, 23, 762. [Google Scholar] [CrossRef]
- Kroymann, J. Natural diversity and adaptation in plant secondary metabolism. Curr. Opin. Plant Biol. 2011, 14, 246–251. [Google Scholar] [CrossRef]
- Ghisoni, S.; Lucini, L.; Rocchetti, G.; Chiodelli, G.; Farinelli, D.; Tombesi, S.; Trevisan, M. Untargeted metabolomics with multivariate analysis to discriminate hazelnut (Corylus avellana L.) cultivars and their geographical origin. J. Sci. Food Agric. 2019. [Google Scholar] [CrossRef] [PubMed]
- Erban, A.; Fehrle, I.; Martinez-Seidel, F.; Brigante, F.; Más, A.L.; Baroni, V.; Wunderlin, D.; Kopka, J. Discovery of food identity markers by metabolomics and machine learning technology. Sci. Rep. 2019, 9, 9697. [Google Scholar] [CrossRef] [PubMed]
- Lim, D.K.; Mo, C.; Lee, J.H.; Long, N.P.; Dong, Z.; Li, J.; Lim, J.; Kwon, S.W. The integration of multi-platform MS-based metabolomics and multivariate analysis for the geographical origin discrimination of Oryza sativa L. J. Food Drug Anal. 2018, 26, 769–777. [Google Scholar] [CrossRef] [PubMed]
- Sidhu, O.P.; Kumar, V.; Behl, H.M. Variability in Neem (Azadirachta indica) with respect to azadirachtin content. J. Agric. Food Chem. 2003, 51, 910–915. [Google Scholar] [CrossRef]
- Rangel-Buitrago, N.G.; Anfuso, G.; Williams, A.T. Coastal erosion along the Caribbean coast of Colombia: Magnitudes, causes and management. Ocean Coast. Manag. 2015, 114, 129–144. [Google Scholar] [CrossRef]
- Dai, J.; Yaylayan, V.A.; Vijaya Raghavan, G.S.; Parè, J.R.; Liu, Z. Multivariate calibration for the determination of total azadirachtin-related limonoids and simple terpenoids in neem extracts using vanillin assay. J. Agric. Food Chem. 2001, 49, 1169–1174. [Google Scholar] [CrossRef]
- Datta, A.; Grün, I.U.; Kwasniewski, M.T.; Fernando, L.N. Comparison of two adsorbent based de-bittering procedures for Neem (Azadirachta indica A. Juss) tea- effect on polyphenols, anti-oxidant capacity, color and volatile profile. Plant Foods Hum. Nutr. 2017, 72, 88–95. [Google Scholar] [CrossRef]
- Pluskal, T.; Castillo, S.; Villar-Briones, A.; Orešič, M. MZmine 2: Modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinform. 2010, 11, 395. [Google Scholar] [CrossRef]
- Salek, R.; Steinbeck, C.; Viant, M.R.; Goodacre, R.; Dunn, W.B. The role of reporting standards for metabolite annotation and identification in metabolomic studies. Gigascience 2013, 2, 13. [Google Scholar] [CrossRef]
- Worley, B.; Powers, R. Multivariate analysis in metabolomics. Curr. Metab. 2013, 1, 92–107. [Google Scholar]
- Haldar, S.; Phapale, P.B.; Kolet, S.P.; Thulasiram, H.V. Expedient preparative isolation, quantification and characterization of limonoids from Neem fruits. Anal. Methods 2013, 5, 5386–5391. [Google Scholar] [CrossRef]
- Pais, A.L.; Li, X.; Xiang, Q.-Y. Discovering variation of secondary metabolite diversity and its relationship with disease resistance in Cornus florida L. Ecol. Evol. 2018, 8, 5619–5636. [Google Scholar] [CrossRef] [PubMed]
- Nanduri, S.; Thunuguntla, S.S.R.; Nyavanandi, V.K.; Kasu, S.; Kumar, P.M.; Ram, P.S.; Rajagopal, S.; Kumar, R.A.; Deevi, D.S.; Rajagopalan, R.; et al. Biological investigation and structure–activity relationship studies on azadirone from Azadirachta indica A. Juss. Bioorg. Med. Chem. Lett. 2003, 13, 4111–4115. [Google Scholar] [CrossRef] [PubMed]
- Tundis, R.; Loizzo, M.R.; Menichini, F. An overview on chemical aspects and potential health benefits of limonoids and their derivatives. Crit. Rev. Food Sci. Nutr. 2014, 54, 225–250. [Google Scholar] [CrossRef]
- Schmidt, J.S.; Nyberg, N.T.; Staerk, D. Assessment of constituents in Allium by multivariate data analysis, high-resolution α-glucosidase inhibition assay and HPLC-SPE-NMR. Food Chem. 2014, 161, 192–198. [Google Scholar] [CrossRef]
- Liu, Y.; Nielsen, M.; Staerk, D.; Jäger, A.K. High-resolution bacterial growth inhibition profiling combined with HPLC–HRMS–SPE–NMR for identification of antibacterial constituents in Chinese plants used to treat snakebites. J. Ethnopharmacol. 2014, 155, 1276–1283. [Google Scholar] [CrossRef]
- Narender, T.; Khaliq, T. Shweta 13C NMR spectroscopy of D and B, D-ring seco-limonoids of Meliaceae family. Nat. Prod. Res. 2008, 22, 763–800. [Google Scholar] [CrossRef]
- Suresh, G.; Narasimhan, N.S.; Palani, N. Structure of nimonol from fresh whole green leaves of Azadirachta indica. Phytochemistry 1997, 45, 807–810. [Google Scholar] [CrossRef]
- Govindachari, T.R.; Malathi, R.; Gopalakrishnan, G.; Suresh, G.; Rajan, S.S. Isolation of a new tetranortriterpenoid from the uncrushed green leaves of Azadirachta indica. Phytochemistry 1999, 52, 1117–1119. [Google Scholar] [CrossRef]
- Siddiqui, B.S.; Afshan, F.; Gulzar, T.; Hanif, M. Tetracyclic triterpenoids from the leaves of Azadirachta indica. Phytochemistry 2004, 65, 2363–2367. [Google Scholar] [CrossRef]
- Siddiqui, B.S.; Afshan, F.; Ghiasuddin; Faizi, S.; Naqvi, S.N.H.; Tariq, R.M. Two insecticidal tetranortriterpenoids from Azadirachta indica. Phytochemistry 2000, 53, 371–376. [Google Scholar] [CrossRef]
- Breksa, A.P.; Ibarra, P. Colorimetric method for the estimation of total limonoid aglycones and glucoside contents in citrus juices. J. Agric. Food Chem. 2007, 55, 5013–5017. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S.; Pathak, N.; Srivastava, P. Identification of limiting factors for the optimum growth of Fusarium oxysporum in liquid medium. Toxicol. Int. 2011, 18, 111–116. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Wang, J.; Gao, Q.; Ma, X.; Wang, Y.; Zhang, Y.; Xun, H.; Yao, X.; Tang, F. Simultaneous determination of five azadirachtins in the seed and leaf extracts of Azadirachta indica by automated online solid-phase extraction coupled with LC-Q-TOF-MS. Chem. Cent. J. 2018, 12, 85. [Google Scholar] [CrossRef]
- Haldar, S.; Mulani, F.A.; Aarthy, T.; Dandekar, D.S.; Thulasiram, H.V. Expedient preparative isolation and tandem mass spectrometric characterization of C-seco triterpenoids from Neem oil. J. Chromatogr. A 2014, 1366, 1–14. [Google Scholar] [CrossRef]
- Chong, J.; Wishart, D.S.; Xia, J. Using MetaboAnalyst 4.0 for comprehensive and integrative metabolomics data analysis. Curr. Protoc. Bioinform. 2019, 68, e86. [Google Scholar] [CrossRef]
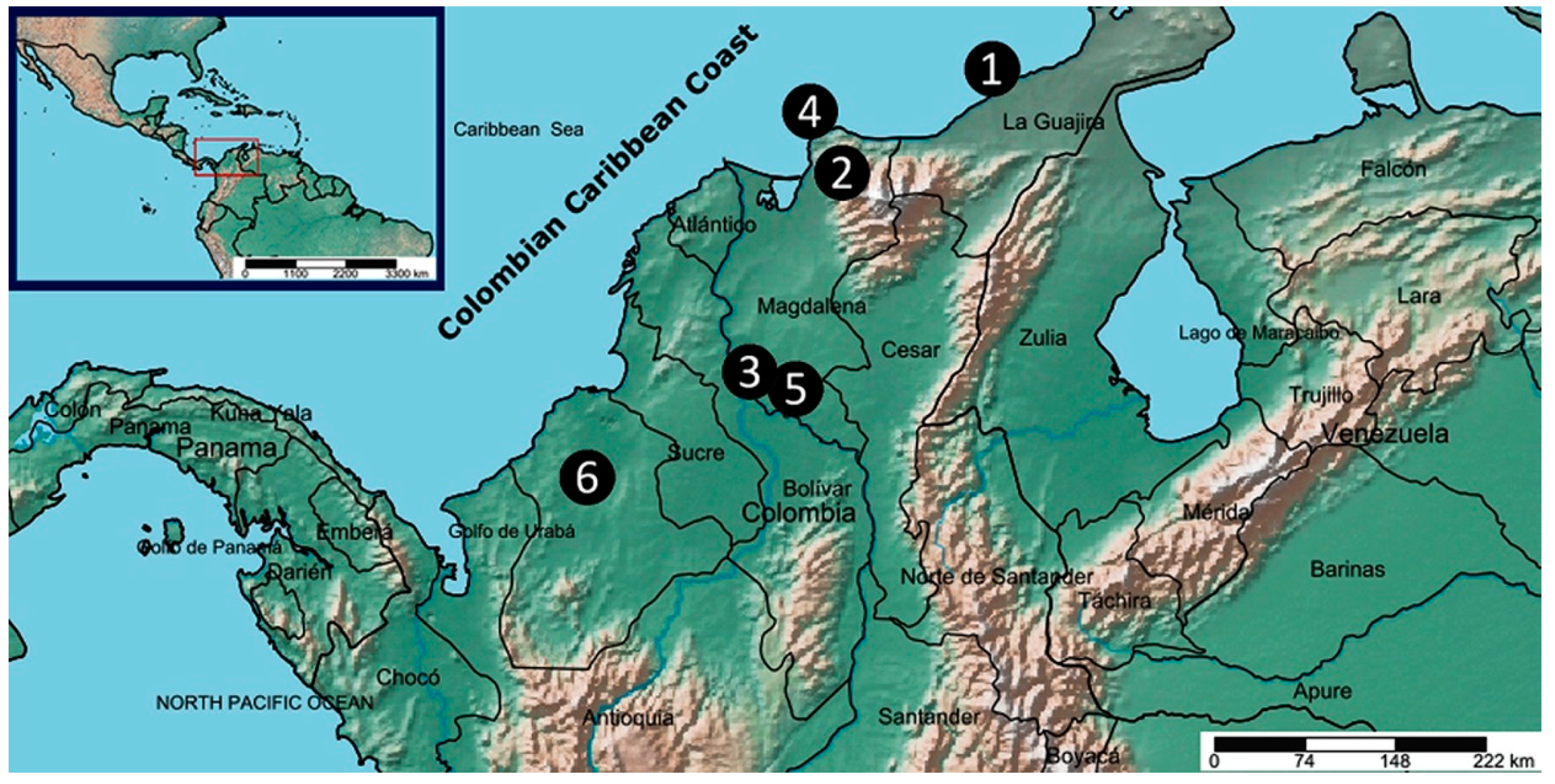
| Location | Abb a | Colombian Caribbean Areas | A (m) b | Climate conditions | Samples | |||
|---|---|---|---|---|---|---|---|---|
| KGC c | T (˚C) d | RF (mm) e | N f | PP g | ||||
| Riohacha | R | La Guajira Peninsula | 5 | Bsh | 28.3 | 588 | 9 | L,F,S |
| San Pedro de la Sierra | SPS | The Sierra Nevada | 1378 | Am | 18.9 | 2721 | 5 | L,F,S |
| Santa Ana | SA | Magdalena River | 25 | Aw | 24.0 | 1468 | 6 | L,F,S |
| Santa Marta | SM | The Sierra Nevada | 9 | Bsh | 28.3 | 512 | 5 | L |
| Mompox | M | Magdalena River | 16 | Aw | 27.5 | 1573 | 10 | L |
| Montería | Mo | Sinú Belt | 20 | Aw | 27.4 | 1225 | 5 | L,F,S |
| No a | m/z [M+H]+ | tR (min) | MF b | Annotation | Error |
|---|---|---|---|---|---|
| 1 | 453.2649 | 14.1 | C28H37O6+ | hydroxyazadirone isomer 1 | −1.76 |
| 2 | 469.2585 | 19.4 | C28H37O13+ | epoxyhydroxyazadirone | 1.07 |
| 3 | 455.2791 | 20.5 | C28H39O10+ | dehydrohydroxyazadirone | 1.32 |
| 4 | 485.2915 | 25.3 | C29H41O20+ | didehydrooxohydroxyazadirone | −2.47 |
| 5 | 453.2632 | 28.8 | C28H37O9+ | hydroxyazadirone isomer 2 | 1.99 |
| Compounds | 1 | 2 | 3 | 4 | 5 |
|---|---|---|---|---|---|
| IC50 a | 1.48 ± 0.11 | 6.42 ± 0.26 | 15.17 ± 0.22 | 4.57 ± 0.23 | 10.07 ± 0.31 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Álvarez-Caballero, J.M.; Coy-Barrera, E. Chemical and Antifungal Variability of Several Accessions of Azadirachta indica A. Juss. from Six Locations Across the Colombian Caribbean Coast: Identification of Antifungal Azadirone Limonoids. Plants 2019, 8, 555. https://doi.org/10.3390/plants8120555
Álvarez-Caballero JM, Coy-Barrera E. Chemical and Antifungal Variability of Several Accessions of Azadirachta indica A. Juss. from Six Locations Across the Colombian Caribbean Coast: Identification of Antifungal Azadirone Limonoids. Plants. 2019; 8(12):555. https://doi.org/10.3390/plants8120555
Chicago/Turabian StyleÁlvarez-Caballero, Juan Manuel, and Ericsson Coy-Barrera. 2019. "Chemical and Antifungal Variability of Several Accessions of Azadirachta indica A. Juss. from Six Locations Across the Colombian Caribbean Coast: Identification of Antifungal Azadirone Limonoids" Plants 8, no. 12: 555. https://doi.org/10.3390/plants8120555
APA StyleÁlvarez-Caballero, J. M., & Coy-Barrera, E. (2019). Chemical and Antifungal Variability of Several Accessions of Azadirachta indica A. Juss. from Six Locations Across the Colombian Caribbean Coast: Identification of Antifungal Azadirone Limonoids. Plants, 8(12), 555. https://doi.org/10.3390/plants8120555