Phloem-Conducting Cells in Haustoria of the Root-Parasitic Plant Phelipanche aegyptiaca Retain Nuclei and Are Not Mature Sieve Elements
Abstract
1. Introduction
2. Results
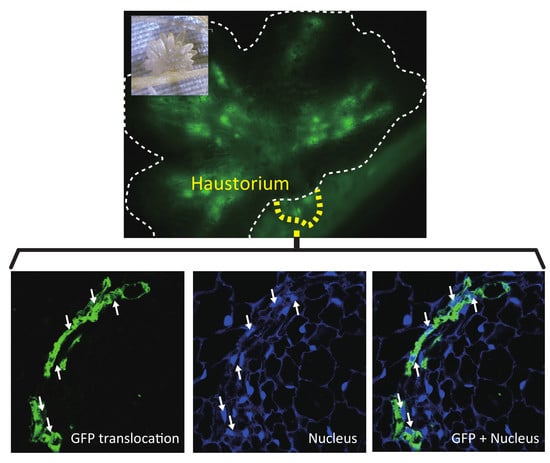
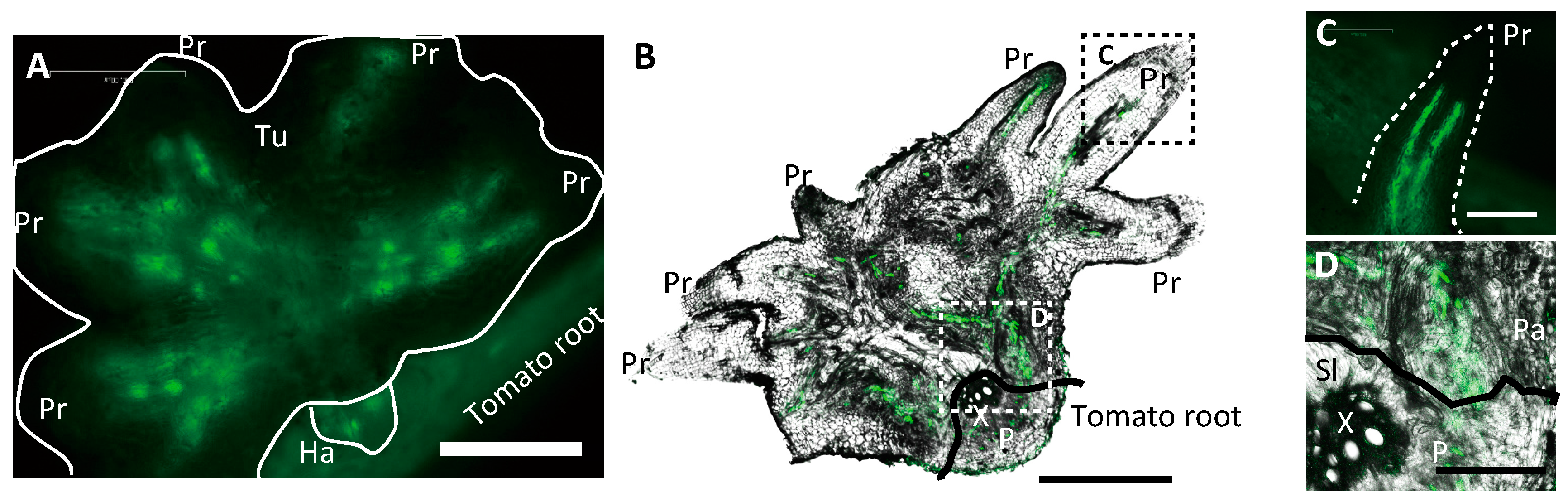
2.1. GFP Expressed in Phloem of Host Plant Visualized a Symplasmic Pathway of a Parasitic Plant
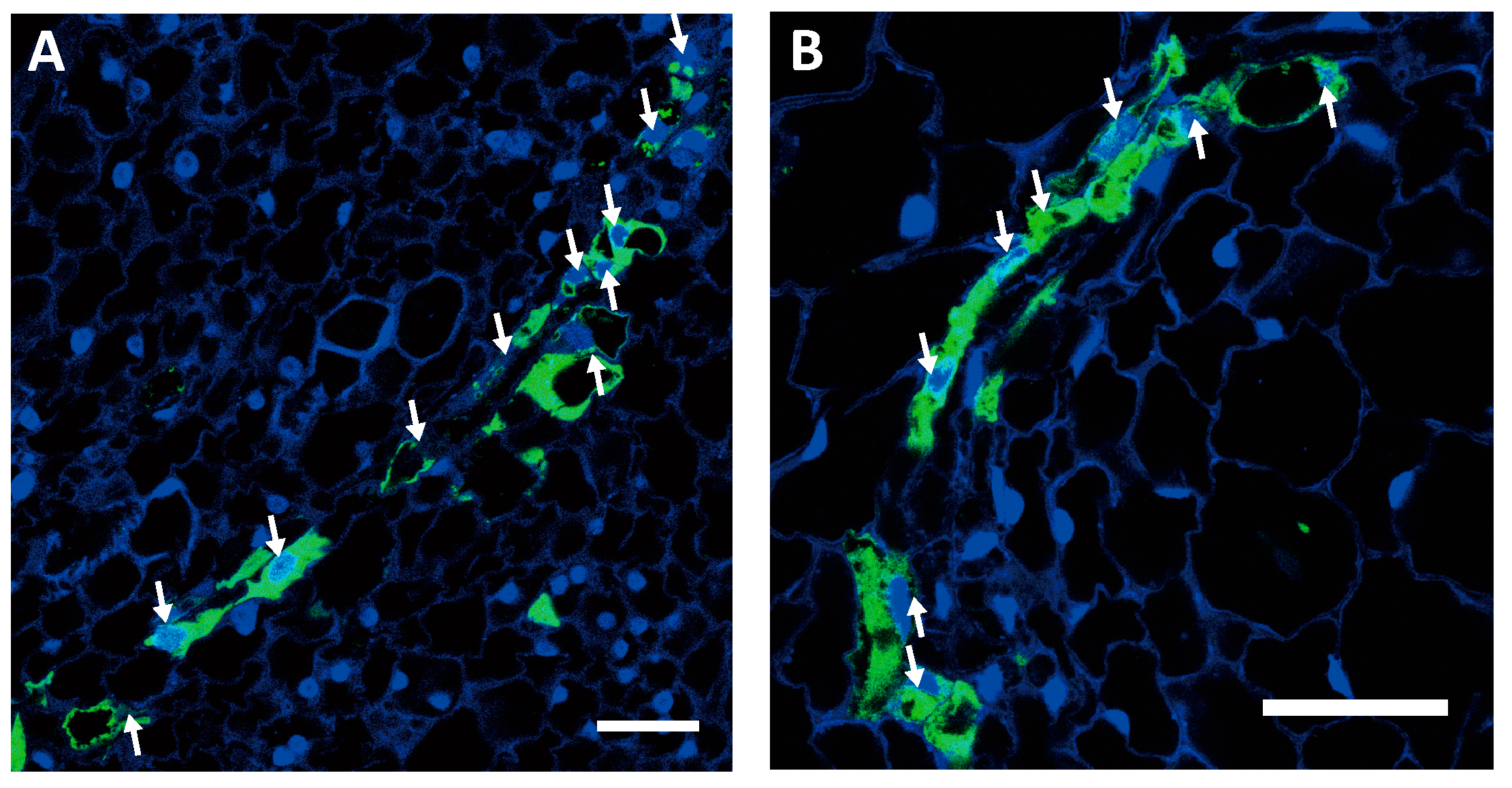
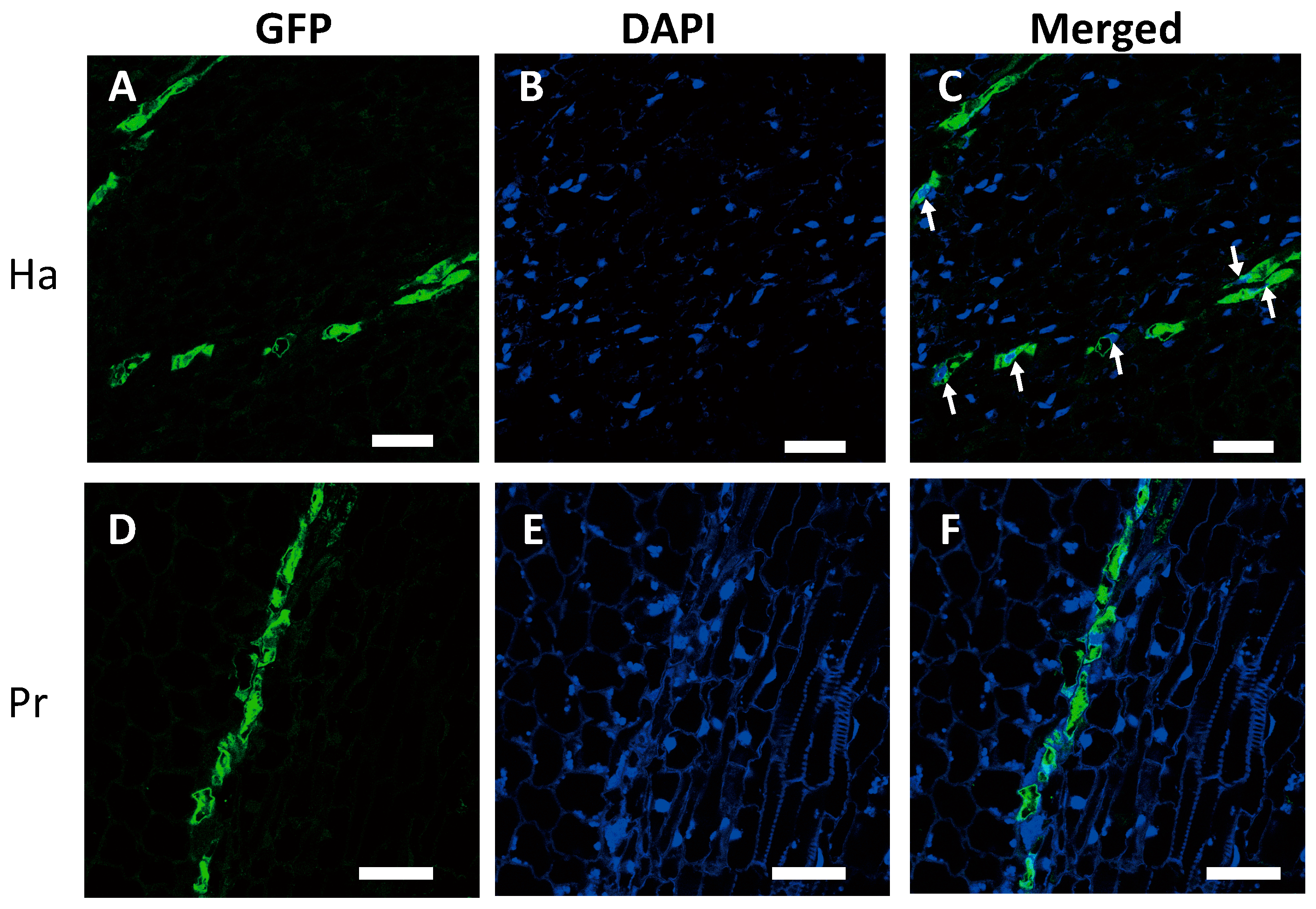
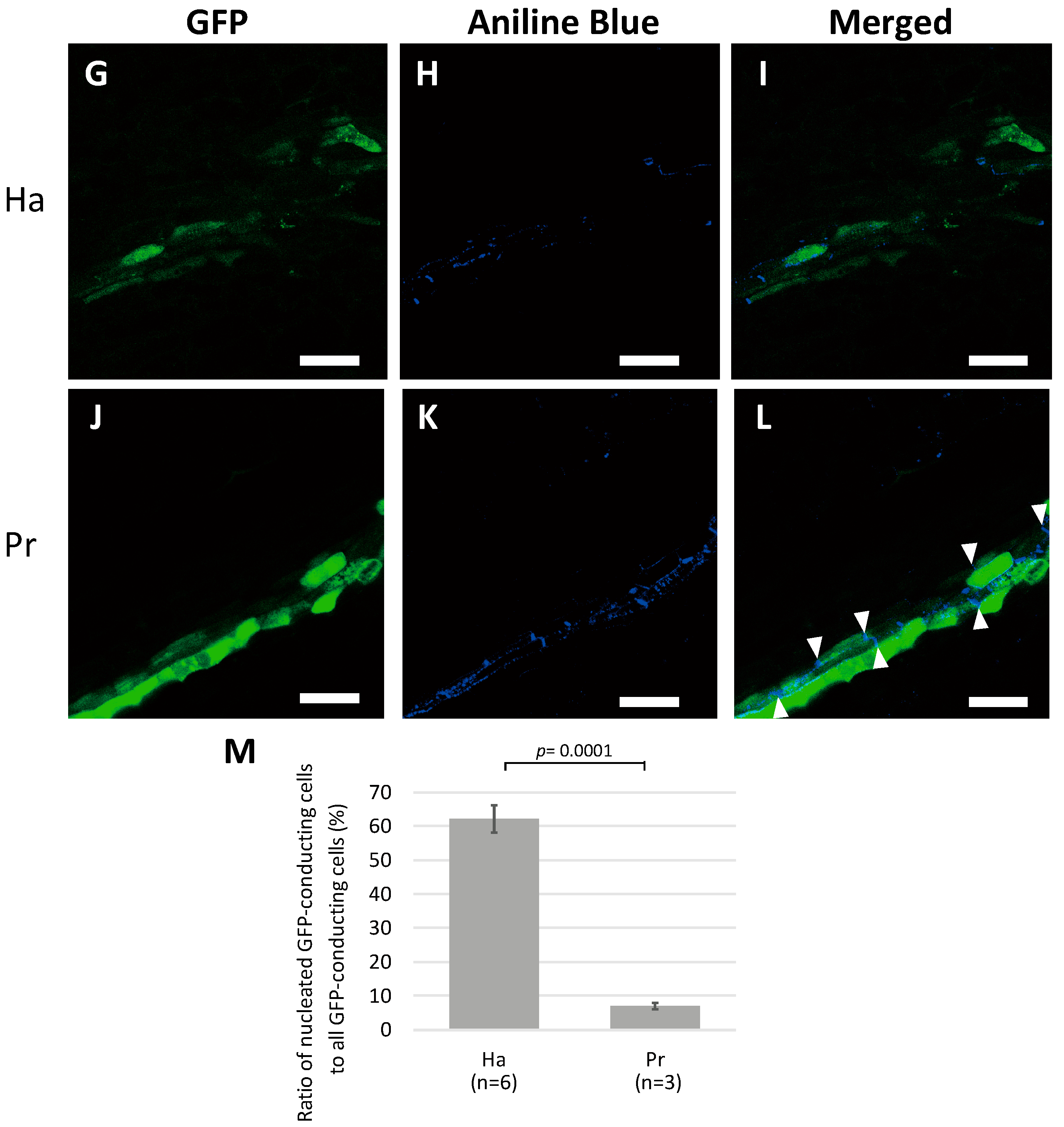
2.2. Nuclei Were Present in GFP-Conducting Cells
2.3. GFP-Conducting Cells in Haustoria Were Not Mature SEs
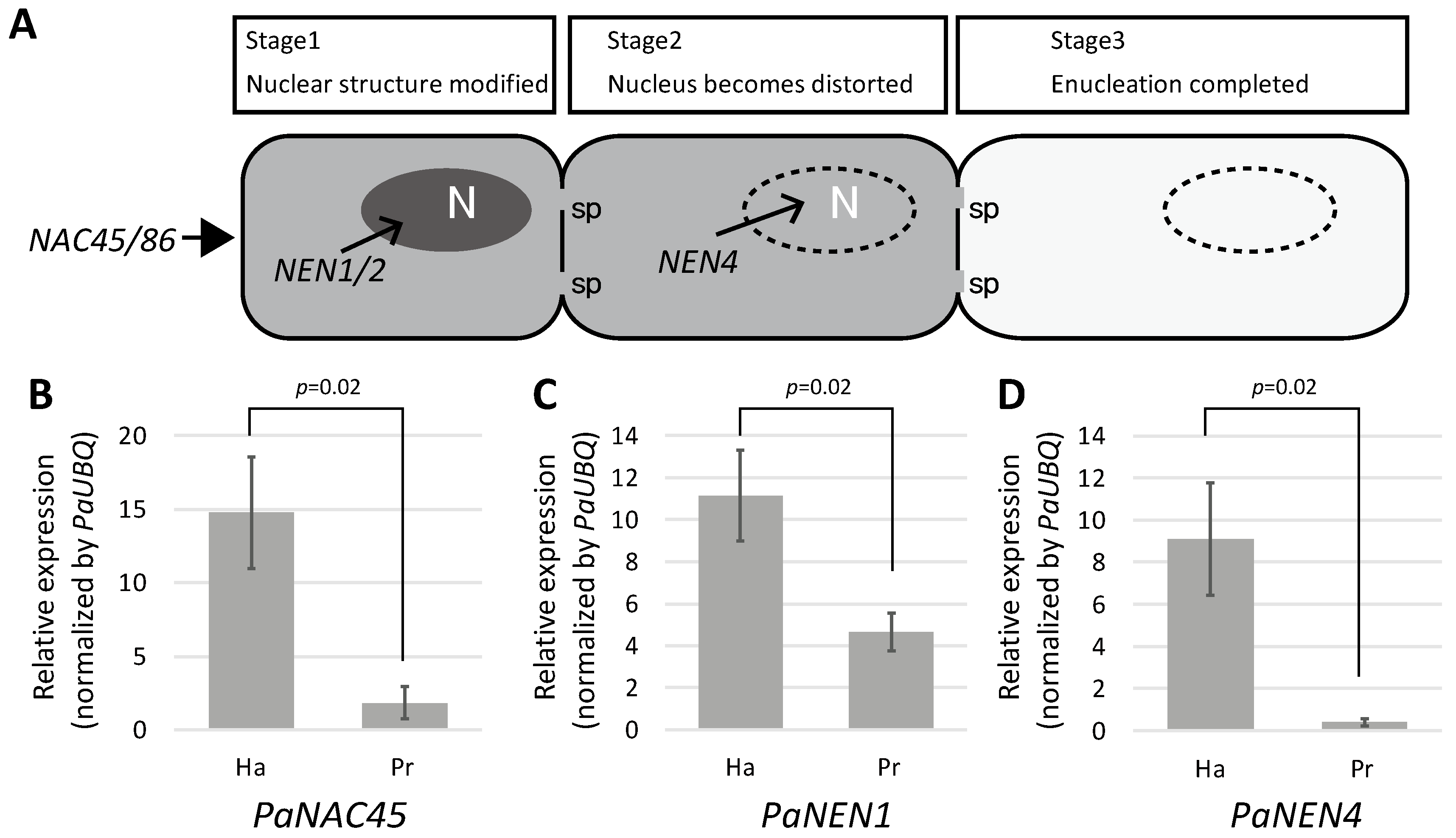
2.4. SE Differentiation-Related Genes Were Expressed in Haustoria
3. Discussion
3.1. Haustorial Phloem-Conducting Cells Are Not Mature SEs
3.2. Why Do Haustorial Phloem-Conducting Cells Fail to Undergo Nuclear Degradation?
3.3. Retention of Nuclei Is Independent of Other SE Differentiation Processes
4. Materials and Methods
4.1. Plant Materials
4.2. Cloning Complementary DNA (cDNA) of PaNEN1, PaNEN4, and PaNAC45
4.3. Real Time PCR
4.4. Preparation and Imaging of Agarose-Embedded Sections
4.5. Immunostaining
4.6. Confocal Laser-Scanning Microscopy
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| BR | brassinosteroid |
| CC | companion cell |
| CHER1 | CHOLINE TRANSPORTER-LIKE 1 |
| DAPI | 4′,6-diamidino-2-phenylindole |
| PBS | phosphate-buffered saline |
| qPCR | quantitative polymerase chain reaction |
| SE | sieve element |
| SEL | size exclusion limit |
| SUC2 | SUCROSE-PROTON SYMPORTER 2 |
References
- Parker, C. Observations on the current status of Orobanche and Striga problems worldwide. Pest Manag. Sci. 2009, 65, 453–459. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Aparicio, M.; Reboud, X.; Gibot-Leclerc, S. Broomrape weeds. Underground mechanisms of parasitism and associated strategies for their control: A review. Front. Plant Sci. 2016, 7, 135. [Google Scholar] [CrossRef] [PubMed]
- Dörr, I. How Striga parasitizes its host: A TEM and SEM study. Ann. Bot. 1997, 79, 463–472. [Google Scholar] [CrossRef]
- Bar-Nun, N.; Sachs, T.; Mayer, A.M. A role for IAA in the infection of Arabidopsis thaliana by Orobanche aegyptiaca. Ann. Bot. 2008, 101, 261–265. [Google Scholar] [CrossRef] [PubMed]
- Truscott, F.H. On the regeneration of new shoots from isolated dodder haustoria. Am. J. Bot. 1958, 45, 169–177. [Google Scholar] [CrossRef]
- Dörr, I. Sieve elements in haustoria of parasitic angiosperms. In Sieve Elements: Comparative Structure, Induction and Development; Behnke, H.-D., Sjolund, R.D., Eds.; Springer: Berlin, Germany, 1990; pp. 239–256. [Google Scholar]
- Haupt, S.; Oparka, K.J.; Sauer, N.; Neumann, S. Macromolecular trafficking between Nicotiana tabacum and the holoparasite Cuscuta reflexa. J. Exp. Bot. 2001, 52, 173–177. [Google Scholar] [CrossRef] [PubMed]
- Birschwilks, M.; Haupt, S.; Hofius, D.; Neumann, S. Transfer of phloem-mobile substances from the host plants to the holoparasite Cuscuta sp. J. Exp. Bot. 2006, 57, 911–921. [Google Scholar] [CrossRef] [PubMed]
- Péron, T.; Candat, A.; Montiel, G.; Veronesi, C.; Macherel, D.; Delavault, P.; Simier, P. New insights into phloem unloading and expression of sucrose transporters in vegetative sinks of the parasitic plant Phelipanche ramosa L. (Pomel). Front. Plant Sci. 2016, 7, 2048. [Google Scholar] [CrossRef] [PubMed]
- Aly, R.; Hamamouch, N.; Abu-Nassar, J.; Wolf, S.; Joel, D.M.; Eizenberg, H.; Kaisler, E.; Cramer, C.; Gal-On, A.; Westwood, J.H. Movement of protein and macromolecules between host plants and the parasitic weed Phelipanche aegyptiaca Pers. Plant Cell Rep. 2011, 30, 2233–2241. [Google Scholar] [CrossRef] [PubMed]
- Spallek, T.; Melnyk, C.W.; Wakatake, T.; Zhang, J.; Sakamoto, Y.; Kiba, T.; Yoshida, S.; Matsunaga, S.; Sakakibara, H.; Shirasu, K. Interspecies hormonal control of host root morphology by parasitic plants. Proc. Natl. Acad. Sci. USA 2017, 114, 5283–5288. [Google Scholar] [CrossRef] [PubMed]
- Furuta, K.M.; Yadav, S.R.; Lehesranta, S.; Belevich, I.; Miyashima, S.; Heo, J.O.; Vatén, A.; Lindgren, O.; De Rybel, B.; Van Isterdael, G.; et al. Plant development. Arabidopsis NAC45/86 direct sieve element morphogenesis culminating in enucleation. Science 2014, 345, 933–937. [Google Scholar] [CrossRef] [PubMed]
- Stadler, R.; Wright, K.M.; Lauterbach, C.; Amon, G.; Gahrtz, M.; Feuerstein, A.; Oparka, K.J.; Sauer, N. Expression of GFP-fusions in Arabidopsis companion cells reveals non-specific protein trafficking into sieve elements and identifies a novel post-phloem domain in roots. Plant J. 2005, 41, 319–331. [Google Scholar] [CrossRef] [PubMed]
- Imlau, A.; Truernit, E.; Sauer, N. Cell-to-cell and long-distance trafficking of the green fluorescent protein in the phloem and symplastic unloading of the protein into sink tissues. Plant Cell 1999, 11, 309–322. [Google Scholar] [CrossRef] [PubMed]
- Esau, K. Development and structure of the phloem tissue. II. Bot. Rev. 1950, 16, 67–114. [Google Scholar] [CrossRef]
- Evert, R.F. Esau’s Plant Anatomy: Meristems, Cells and Tissues of the Plant Body: Their Structure, Function and Development, 3rd ed.; Wiley: Hoboken, NJ, USA, 2006; pp. 357–405. [Google Scholar]
- Wallner, E.S.; López-Salmerón, V.; Belevich, I.; Poschet, G.; Jung, I.; Grünwald, K.; Sevilem, I.; Jokitalo, E.; Hell, R.; Helariutta, Y.; et al. Strigolactone- and karrikin-independent SMXL proteins are central regulators of phloem formation. Curr. Biol. 2017, 27, 1241–1247. [Google Scholar] [CrossRef] [PubMed]
- Bonke, M.; Thitamadee, S.; Mähönen, A.P.; Hauser, M.T.; Helariutta, Y. APL regulates vascular tissue identity in Arabidopsis. Nature 2003, 426, 181–186. [Google Scholar] [CrossRef] [PubMed]
- Xie, B.; Hong, Z. Unplugging the callose plug from sieve pores. Plant Signal. Behav. 2011, 6, 491–493. [Google Scholar] [CrossRef] [PubMed]
- Kondo, Y.; Nurani, A.M.; Saito, C.; Ichihashi, Y.; Saito, M.; Yamazaki, K.; Mitsuda, N.; Ohme-Takagi, M.; Fukuda, H. Vascular cell induction culture system using Arabidopsis leaves (VISUAL) reveals the sequential differentiation of sieve element-like cells. Plant Cell 2016, 28, 1250–1262. [Google Scholar] [CrossRef] [PubMed]
- Heo, J.O.; Blob, B.; Helariutta, Y. Differentiation of conductive cells: A matter of life and death. Curr. Opin. Plant Biol. 2017, 35, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.H.; Breda, A.; Hardtke, C.S. Brassinosteroid signaling directs formative cell divisions and protophloem differentiation in Arabidopsis root meristems. Development 2017, 144, 272–280. [Google Scholar] [CrossRef] [PubMed]
- Anne, P.; Azzopardi, M.; Gissot, L.; Beaubiat, S.; Hématy, K.; Palauqui, J.C. OCTOPUS negatively regulates BIN2 to control phloem differentiation in Arabidopsis thaliana. Curr. Biol. 2015, 25, 2584–2590. [Google Scholar] [CrossRef] [PubMed]
- Truernit, E.; Bauby, H.; Belcram, K.; Barthélémy, J.; Palauqui, J.C. OCTOPUS, a polarly localised membrane-associated protein, regulates phloem differentiation entry in Arabidopsis thaliana. Development 2012, 139, 1306–1315. [Google Scholar] [CrossRef] [PubMed]
- Ruiz Sola, M.A.; Coiro, M.; Crivelli, S.; Zeeman, S.C.; Schmidt Kjølner Hansen, S.; Truernit, E. OCTOPUS-LIKE 2, a novel player in Arabidopsis root and vascular development, reveals a key role for OCTOPUS family genes in root metaphloem sieve tube differentiation. New Phytol. 2017, 216, 1191–1204. [Google Scholar] [CrossRef] [PubMed]
- Enari, M.; Sakahira, H.; Yokoyama, H.; Okawa, K.; Iwamatsu, A.; Nagata, S. A caspase-activated DNase that degrades DNA during apoptosis, and its inhibitor ICAD. Nature 1998, 391, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Woo, E.J.; Kim, Y.G.; Kim, M.S.; Han, W.D.; Shin, S.; Robinson, H.; Park, S.Y.; Oh, B.H. Structural mechanism for inactivation and activation of CAD/DFF40 in the apoptotic pathway. Mol. Cell 2004, 14, 531–539. [Google Scholar] [CrossRef]
- Dettmer, J.; Ursache, R.; Campilho, A.; Miyashima, S.; Belevich, I.; O’Regan, S.; Mullendore, D.L.; Yadav, S.R.; Lanz, C.; Beverina, L.; et al. CHOLINE TRANSPORTER-LIKE1 is required for sieve plate development to mediate long-distance cell-to-cell communication. Nat. Commun. 2014, 5, 4276. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Yu, J.; Li, D.; Zhang, X. Sinbase: An integrated database to study genomics, genetics and comparative genomics in Sesamum indicum. Plant Cell Physiol. 2015, 56, e2. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schaffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- González-Verdejo, C.I.; Die, J.V.; Nadal, S.; Jiménez-Marín, A.; Moreno, M.T.; Román, B. Selection of housekeeping genes for normalization by real-time RT-PCR: Analysis of Or-MYB1 gene expression in Orobanche ramosa development. Anal. Biochem. 2008, 379, 176–181. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Aparicio, M.; Huang, K.; Wafula, E.K.; Honaas, L.A.; Wickett, N.J.; Timko, M.P.; Depamphilis, C.W.; Yoder, J.I.; Westwood, J.H. Application of qRT-PCR and RNA-Seq analysis for the identification of housekeeping genes useful for normalization of gene expression values during Striga hermonthica development. Mol. Biol. Rep. 2013, 40, 3395–3407. [Google Scholar] [CrossRef] [PubMed]
- Hozumi, A.; Bera, S.; Fujiwara, D.; Obayashi, T.; Yokoyama, R.; Nishitani, K.; Aoki, K. Arabinogalactan proteins accumulate in the cell walls of searching hyphae of the stem parasitic plants, Cuscuta campestris and Cuscuta japonica. Plant Cell Physiol. 2017, 58, 1868–1877. [Google Scholar] [CrossRef] [PubMed]
- Ruzin, S.E. Sectioning and mounting. In Plant Microtechnique and Microscopy; Oxford University Press: Oxford, UK, 1999; pp. 73–85. [Google Scholar]

| Primer ID | Sequence (5′ → 3′) |
|---|---|
| PaNEN1 forward | ACAAGCAAGCTATACATAACCGTG |
| PaNEN1 reverse | TAACAGCACCAAAGTTAAATCAAACC |
| PaNEN1 qPCR forward | ACAAGCAAGCTATACATAACCGTG |
| PaNEN1 qPCR reverse | TATTCAGGAAATAAAGAATAATGGGAGC |
| PaNEN4 forward | ACAGATGTAGTGTATGCTAGGATTACC |
| PaNEN4 reverse | ATAACGACACCATCTTACAAAGTTTGAAC |
| PaNEN4 qPCR forward | AGGAAGAACTCATTCTTGCTAACAAC |
| PaNEN4 qPCR reverse | TGACATTGTCAGTTGTAACATACTG |
| PaNAC45 forward | ATTTCAAAAAAGGACCAGGAGACTGG |
| PaNAC45 reverse | AGTTCATTGGCATTGGATATTCAGAAG |
| PaNAC45 qPCR forward | ATCAATGGCCGCAAAATCGATC |
| PaNAC45 qPCR reverse | TACCATTCAAGATCTTTGCTTGGC |
| PaUBQ1 qPCR forward | ATCACTGCCTGATTATCAGACGC |
| PaUBQ1 qPCR reverse | TAGGGAATCAACTCGATTATGGCG |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ekawa, M.; Aoki, K. Phloem-Conducting Cells in Haustoria of the Root-Parasitic Plant Phelipanche aegyptiaca Retain Nuclei and Are Not Mature Sieve Elements. Plants 2017, 6, 60. https://doi.org/10.3390/plants6040060
Ekawa M, Aoki K. Phloem-Conducting Cells in Haustoria of the Root-Parasitic Plant Phelipanche aegyptiaca Retain Nuclei and Are Not Mature Sieve Elements. Plants. 2017; 6(4):60. https://doi.org/10.3390/plants6040060
Chicago/Turabian StyleEkawa, Minako, and Koh Aoki. 2017. "Phloem-Conducting Cells in Haustoria of the Root-Parasitic Plant Phelipanche aegyptiaca Retain Nuclei and Are Not Mature Sieve Elements" Plants 6, no. 4: 60. https://doi.org/10.3390/plants6040060
APA StyleEkawa, M., & Aoki, K. (2017). Phloem-Conducting Cells in Haustoria of the Root-Parasitic Plant Phelipanche aegyptiaca Retain Nuclei and Are Not Mature Sieve Elements. Plants, 6(4), 60. https://doi.org/10.3390/plants6040060