Molecular Mechanisms of Phosphorus Metabolism and Transport during Leaf Senescence
Abstract
:1. Introduction
Nutrient Remobilization during Leaf Senescence
| Species (Common Name) | Phosphorus Resorption Efficiency (%) | Source |
|---|---|---|
| Acacia truncata (angle-leafed wattle) | 41 | [9] |
| Acacia xanthine (white-stemmed wattle) | 36 | [9] |
| Arabidopsis thaliana (thale cress) | 75 | [6,7] |
| Artabotrys hongkongensis (talon wild vine) | 41 | [15] |
| Banksia attenuate (slender banksia) | 69 | [9] |
| Banksia chamaephyton (fishbone banksia) | 82 | [16] |
| Banksia serrata (saw banksia) | 95 | [9] |
| Calophyllum polyanthum (sirpoon tree) | 53 | [15] |
| Cladium jamaicense (Jamaica swamp grass) | 78 | [17] |
| Empertrum hermaphroditum (mountain crowberry) | 70 | [18] |
| Eriophorum vaginatum (tussock cottongrass) | 90 | [18] |
| Glyceria maxima (reed mannagrass) | 22 | [19] |
| Glycine max (soybean) | 50 | [8] |
| Hakea prostrata (harsh hakea) | 85 | [10,11] |
| Michelia floribunda | 80 | [15] |
| Phragmites australis (common reed) | 50 | [19] |
| Vaccinium uliginosum (bog blueberry) | 40 | [18] |
2. Transcriptome Changes Promote Phosphorus Remobilization during Leaf Senescence
2.1. Differential Gene Expression over the Course of Senescence
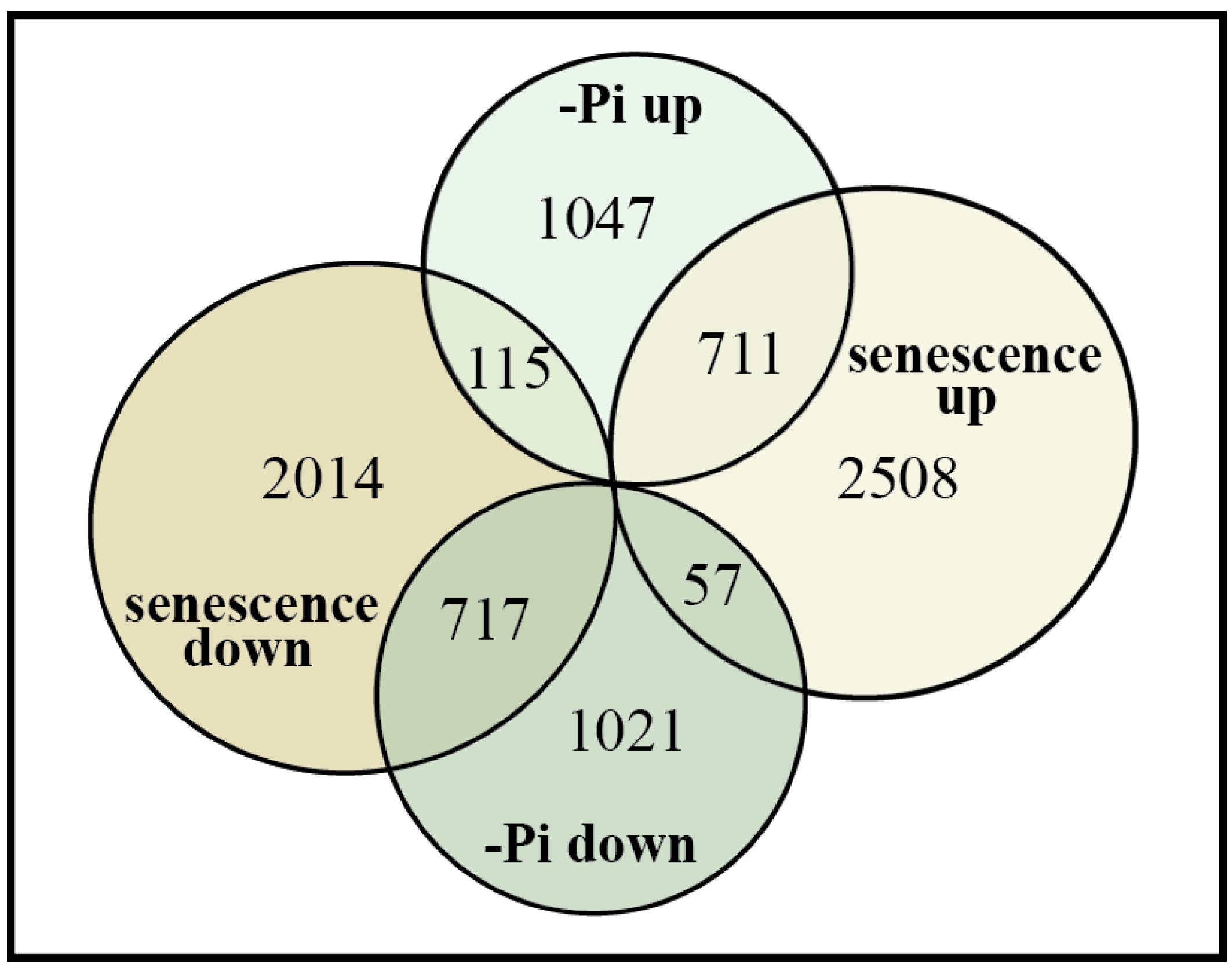
2.2. Transcriptome Changes during Leaf Senescence versus Nutritional Pi-Deficiency
| Phosphate Source | Gene | Species (Common Name) | Proposed Function (s) | Reference |
|---|---|---|---|---|
| Nucleic acids | RNS2 | Arabidopsis thaliana (thale cress) | class II RNase Housekeeping rRNA degradation | [46,47,48] |
| NGR2 | Nicotiana glutinosa (tobacco) | class II RNase Housekeeping rRNA degradation | [49] | |
| AhSL28 | Antirrhinum (snapdragon) | class II RNase Housekeeping rRNA degradation RNA degradation during Pi starvation RNA degradation during senescence response | [50] | |
| RNaseLER | Solanum lycopersicum (tomato) | class II RNase Housekeeping rRNA degradation | [51,52] | |
| RNaseLX | Solanum lycopersicum (tomato) | intracellular class I RNase RNA degradation during xylem differentiation RNA degradation during leaf abscission | [53,54,55,56] | |
| RNaseLE | Solanum lycopersicum (tomato) | extracellular class I RNase RNA degradation during sieve element development RNA degradation during mechanical wounding response | [53,54,57] | |
| ZRNase I | Zinnia elegans (common zinnia) | extracellular class I RNase RNA degradation during tracheary element differentiation | [58] | |
| RNS1 | Arabidopsis thaliana (thale cress) | extracellular class I RNase RNA degradation during Pi-starvation RNA degradation during senescence | [7,41,47] | |
| NvRN1 | Nepenthes ventricosa (tropical pitcher plant) | extracellular class I RNase RNA degradation from insect prey | [59] | |
| BFN1 | Arabidopsis thaliana (thale cress) | type I nuclease Nucleic acid degradation during PCD | [60,61,62] | |
| LeNUC1 | Solanum lycopersicum (tomato) | type I nuclease Nucleic acid degradation during senescence | [63] | |
| Phospholipids | PLA1 gene AF026480 | Dianthus caryophyllus (carnation) | Phospholipase A1 Hydrolysis of phospholipids Promote senescence progress | [64,65] |
| NPC4 | Arabidopsis thaliana (thale cress) | Phospholipase C Phospholipid hydrolysis during Pi-starvation | [66] | |
| PLDα | Arabidopsis thaliana (thale cress) | Phospholipase Hydrolysis of phospholipids Promote senescence progress | [67] | |
| SQD1 | Arabidopsis thaliana (thale cress) | Sulfoquinovosyldiacylglycerol Sulfolipid biosynthesis | [32] | |
| Other Pi-monoesters | AtPAP26 | Arabidopsis thaliana (thale cress) | Purple acid phosphatase (dual-targeted to cell vacuole and cell wall/secretome) Scavenge Pi during Pi-starvation Scavenge Pi during senescence | [7,41,42,43,44,45] |
| HpPAP1 | Hakea prostrata (harsh hakea) | Purple acid phosphatase Scavenge Pi during senescence | [41] | |
| AtPAP17 | Arabidopsis thaliana (thale cress) | Purple acid phosphatase Scavenge Pi during Pi-starvation ROS metabolism | [7,41,43,68] | |
| AtPPsPase1 | Arabidopsis thaliana (thale cress) | HAD pyrophosphatase Scavenge Pi during Pi-starvation | [69] | |
| AtPECP1 | Arabidopsis thaliana (thale cress) | HAD phosphoethanolamine/phosphocholine phosphatase Scavenge Pi during Pi-starvation Phospholipid degradation | [70] | |
| AtSgpp | Arabidopsis thaliana (thale cress) | HAD phosphosugar phosphatase Scavenge Pi during Pi-starvation | [71] | |
| LePS2;1 | Solanum lycopersicum (tomato) | HAD protein phosphatase Pi signaling during Pi-starvation | [72,73] | |
| PvPS2:1 | Phaseolus vulgaris (common bean) | HAD protein phosphatase Pi signaling during Pi-starvation | [74,75] |
3. Catabolism of Macromolecules Frees Phosphate for Remobilization
3.1. Catabolism of Nucleic Acids
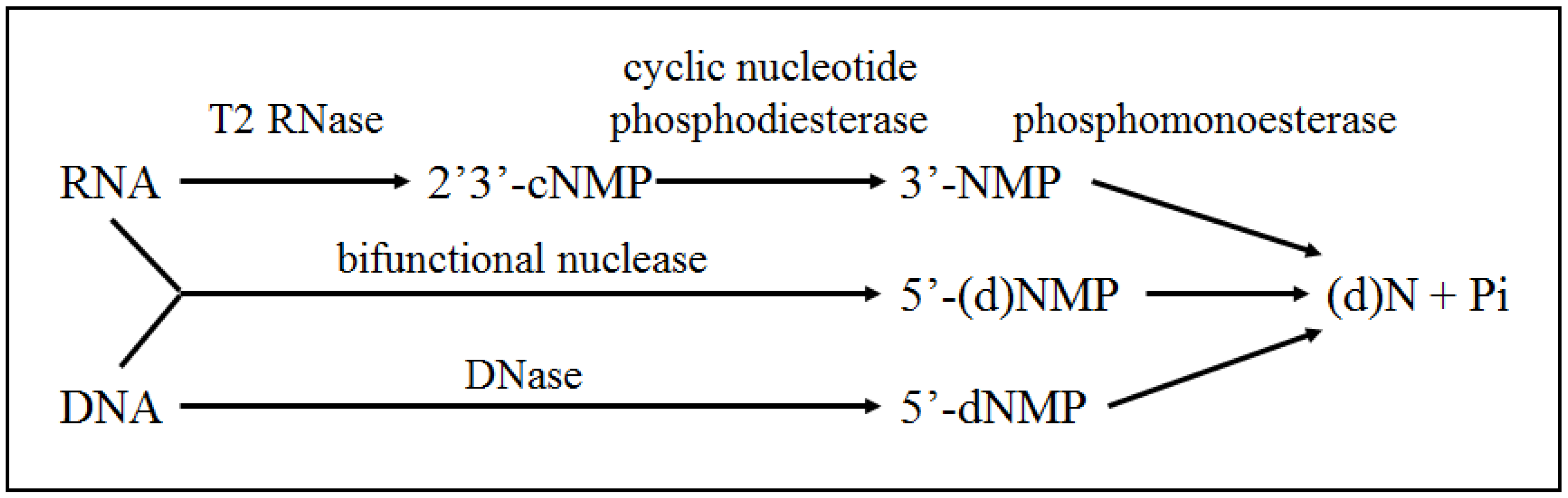
3.2. Catabolism of Lipids
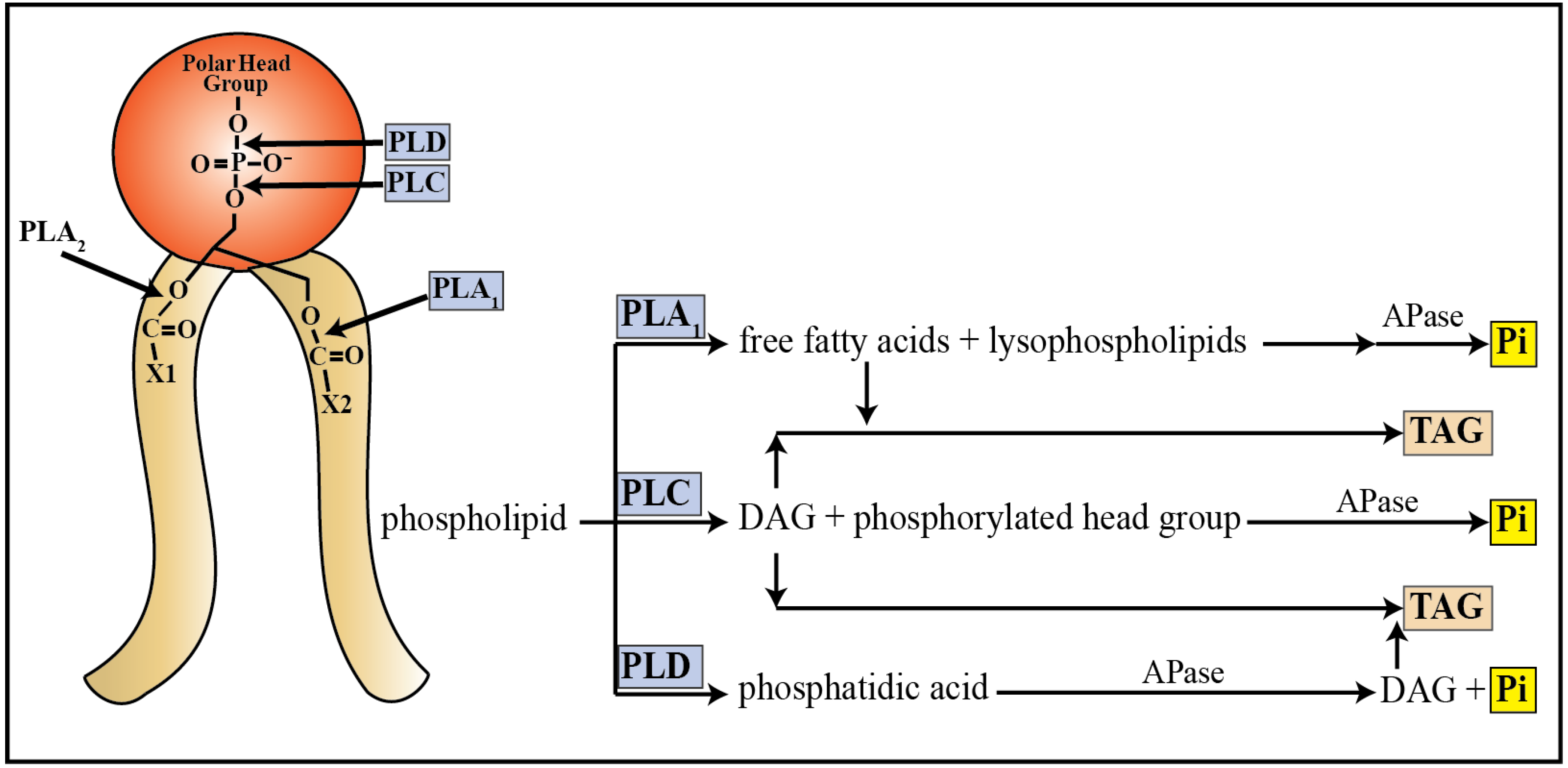
3.3. Other Phosphate Pools
4. Phosphate Transport from Senescing Leaves to Growing Tissue
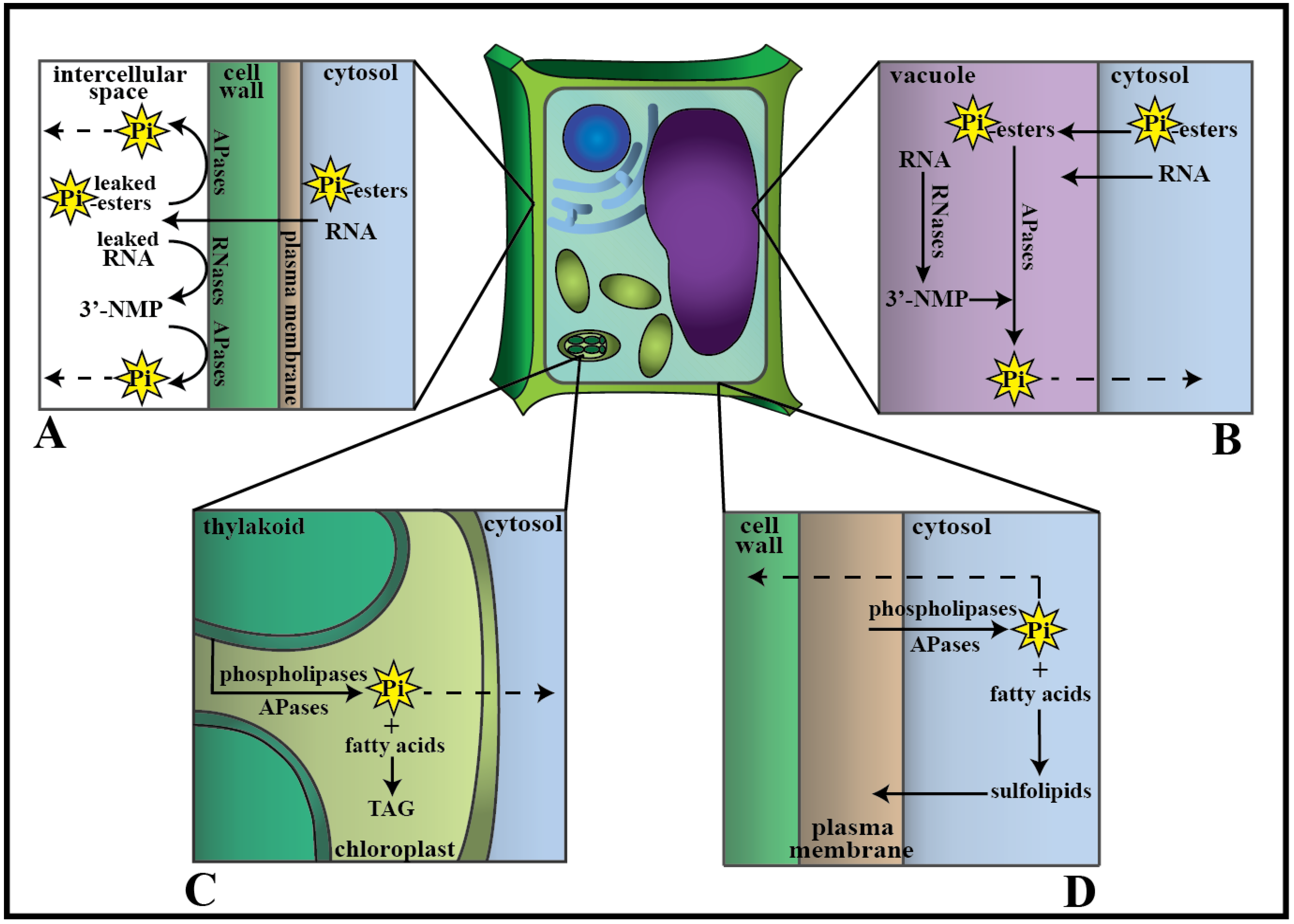
5. Concluding Remarks
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Plaxton, W.C.; Tran, H.T. Metabolic adaptations of phosphate-starved plants. Plant Physiol. 2011, 156, 1006–1015. [Google Scholar] [CrossRef] [PubMed]
- Baligar, V.C.; Fageria, N.K.; He, Z.L. Nutrient use efficiency in plants. Commun. Soil Sci. Plant Anal. 2001, 32, 921–950. [Google Scholar] [CrossRef]
- Cordell, D.; Drangert, J.-O.; White, S. The story of phosphorus: Global food security and food for thought. Glob. Environ. Change 2009, 19, 292–305. [Google Scholar] [CrossRef]
- Veneklaas, E.J.; Lambers, H.; Bragg, J.; Finnegan, P.M.; Lovelock, C.E.; Plaxton, W.C.; Price, C.A.; Scheible, W.; Shane, M.W.; White, P.J.; et al. Opportunities for improving phosphorus-use efficiency in crop plants. New Phytol. 2012, 195, 306–320. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.P.; Fontenot, E.B.; Zahraeifard, S.; Feuer DiTusa, S. Molecular components that drive phosphorus-remobilisation during leaf senescence. Annu. Plant Rev. 2015, 48, 159–186. [Google Scholar]
- Himelblau, E.; Amasino, R.M. Nutrients mobilized from leaves of Arabidopsis thaliana during leaf senescence. J. Plant Physiol. 2001, 158, 1317–1323. [Google Scholar] [CrossRef]
- Robinson, W.D.; Carson, I.; Ying, S.; Ellis, K.; Plaxton, W.C. Eliminating the purple acid phosphatase AtPAP26 in Arabidopsis thaliana delays leaf senescence and impairs phosphorus remobilization. New Phytol. 2012, 196, 1024–1029. [Google Scholar] [CrossRef] [PubMed]
- Crafts-Brandner, S.J. Phosphorus nutrition influence on leaf senescence in soybean. Plant Physiol. 1992, 98, 1128–1132. [Google Scholar] [CrossRef] [PubMed]
- De Campos, M.C.R.; Pearse, S.J.; Oliveira, R.S.; Lambers, H. Downregulation of new phosphorus-uptake capacity is inversely related to leaf phosphorus-resorption proficiency in four species from a phosphorus-impoverished environment. Ann. Bot. 2013. [Google Scholar] [CrossRef] [PubMed]
- Shane, M.W.; Cramer, M.D.; Funayama-Noguchi, S.; Cawthray, G.R.; Millar, A.H.; Day, D.A.; Lambers, H. Developmental physiology of cluster-root carboxylate synthesis and exudation in Harsh Hakea. Expression of phosphoenolpyruvate carboxylase and the alternative oxidase. Plant Physiol. 2004, 135, 549–560. [Google Scholar] [CrossRef] [PubMed]
- Lambers, H.; Finnegan, P.M.; Jost, R.; Plaxton, W.C.; Shane, M.W.; Stitt, M. Phosphorus nutrition in Proteaceae and beyond. Nat. Plants 2015, 1, 1–9. [Google Scholar] [CrossRef]
- Kobe, R.K.; Lepczyk, C.A.; Iyer, M. Resorption efficiency decreases with increasing green leaf nutrients in a global data set. Ecology 2005, 86, 2780–2792. [Google Scholar] [CrossRef]
- Vergutz, L.; Manzoni, S.; Porporato, A.; Novais, R.F.; Jackson, R.B. Global resorption efficiencies and concentrations of carbon and nutrients in leaves of terrestrial plants. Ecol. Monogr. 2012, 82, 205–220. [Google Scholar] [CrossRef]
- Aerts, R. Nutrient resorption from senescing leaves of perennials: Are there general patterns? J. Ecol. 1996, 84, 597–608. [Google Scholar] [CrossRef]
- Cai, Z.; Bongers, F. Contrasting nitrogen and phosphorus efficiencies in trees and lianas from a tropical montane rainforest in Xishuangbanna, south-west China. J. Trop. Ecol. 2007, 23, 115–118. [Google Scholar] [CrossRef]
- Denton, M.D.; Veneklaas, E.J.; Freimoser, F.M.; Lambers, H. Banksia species (Proteaceae) from severely phosphorus-impoverished soils exhibit extreme efficiency in the use and re-mobilization of phosphorus. Plant Cell Environ. 2007, 30, 1557–1565. [Google Scholar] [CrossRef] [PubMed]
- Richardson, C.J.; Ferrell, G.M.; Vaithiyanathan, P. Nutrient effects on stand structure, resorption efficiency, and secondary compounds in everglades sawgrass. Ecology 1999, 80, 2182–2192. [Google Scholar] [CrossRef]
- Van Heerwaarden, L.M.; Toet, S.; Aerts, R. Nitrogen and phosphorus resorption efficiency and proficiency in six sub-arctic bog species after 4 years of nitrogen fertilization. J. Ecol. 2003, 91, 1060–1070. [Google Scholar] [CrossRef]
- Lawniczak, A.E. Nitrogen, phosphorus, and potassium resorption efficiency and proficiency of four emergent macrophytes from nutrient-rich wetlands. Pol. J. Environ. Stud. 2011, 5, 1227–1234. [Google Scholar]
- Balazadeh, S.; Riano-Pachan, D.M.; Mueller-Rocher, B. Transcription factors regulating leaf senescence in Arabidopsis thaliana. Plant Biol. 2008, 10, 63–75. [Google Scholar] [CrossRef] [PubMed]
- Breeze, E.; Harrison, E.; McHattie, S.; Hughs, L.; Hichman, R.; Hill, C.; Kiddle, S.; Kim, Y.; Penfold, C.A.; Jenkins, D.; et al. High resolution temporal profiling of transcripts during Arabidopsis leaf senescence reveals a distinct chronology of processes and regulation. Plant Cell 2011, 23, 873–894. [Google Scholar] [CrossRef] [PubMed]
- Buchanan-Wollaston, V.; Earl, S.; Harrison, E.; Mathas, E.; Navabpour, S.; Page, T.; Pink, D. The molecular analysis of leaf senescence—A genomics approach. Plant Biotechnol. J. 2003, 1, 3–22. [Google Scholar] [CrossRef] [PubMed]
- Gepstein, S.; Sabehi, G.; Carp, M.-J.; Hajouj, T.; Falah, M.; Nesher, O.; Yariv, I.; Dor, C.; Bassani, M. Large-scale identification of leaf senescence-associated genes. Plant J. 2003, 36, 629–642. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Cai, Z.; Gan, S. Transcriptome of Arabidopsis leaf senescence. Plant Cell Environ. 2004, 27, 521–549. [Google Scholar] [CrossRef]
- Gregersen, P.L.; Holm, P.B. Transcriptome analysis of senescence in the flag leaf of wheat (Triticum aestivum L.). Plant Biotechnol. J. 2007, 5, 192–206. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Zhou, Y.; Zhou, G.; Ye, R.; Zhao, L.; Li, X.; Lin, Y. Identification of early senescence-associated genes in rice flag leaves. Plant Mol. Biol. 2008, 67, 37–55. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.Y.; Xu, Y.C.; Li, W.L.; Yang, L.; Yue, X.; Zhange, X.S.; Zhao, X.Y. Transcriptional analyses of natural leaf senescence in maize. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.; Pang, C.; Fan, S.; Song, M.; Wei, H.; Yu, S. Global analysis of the Gossypium hirsutum L. transcriptome during leaf senescence by RNA-Seq. BMC Plant Biol. 2015, 15. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.F.; Gan, S.S. AtNAP, a NAC family transcription factor, has an important role in leaf senescence. Plant. J. 2006, 46, 601–612. [Google Scholar] [CrossRef] [PubMed]
- Matallana-Ramirez, L.P.; Rauf, M.; Farage-Barrhom, S.; Dortay, H.; Xue, G.; Droge-Laser, W.; Lers, A.; Balazadeh, S.; Mueller-Roeber, B. NAC transcription factor ORE1 and senescence-induced BIFUNCTIONAL NUCLEASE1 (BFN1) constitute a regulatory cascade in Arabidopsis. Mol. Plant 2013, 6, 1438–1452. [Google Scholar] [CrossRef] [PubMed]
- Hinderhofer, K.; Zentgraf, U. Identification of a transcription factor specifically expressed at the onset of leaf senescence. Planta 2001, 213, 469–473. [Google Scholar] [CrossRef] [PubMed]
- Miao, Y.; Laun, T.; Zimmermann, P.; Zentgraf, U. Targets of the WRKY53 transcription factor and its role during leaf senescence in Arabidopsis. Plant Mol. Biol. 2004, 55, 853–867. [Google Scholar] [CrossRef] [PubMed]
- Maio, Y.; Laun, T.M.; Smykowski, A.; Zentgraf, U. Arabidopsis MEKK1 can take a short cut: It can directly interact with senescence-related WRKY53 transcription factor on the protein level and can bind to its promoter. Plant Mol. Biol. 2007, 65, 63–76. [Google Scholar] [CrossRef] [PubMed]
- Mission, J.; Raghothama, K.G.; Jain, A.; Jouhet, J.; Block, M.A.; Bligny, R.; Ortet, P.; Creff, A.; Somerville, S.; Rolland, N.; et al. A genome-wide transcriptional analysis using Arabidopsis thaliana Affymatrix gene chips determined plant responses to phosphate deprivation. Proc. Natl. Acad. Sci. USA 2005, 102, 11934–11939. [Google Scholar] [CrossRef] [PubMed]
- Robatzek, S.; Somssich, I.E. Targets of AtWRKY6 regulation during plant senescence and pathogen defense. Genes & Dev. 2002, 16, 1139–1149. [Google Scholar]
- Chen, Y.-F.; Li, L.-Q.; Xu, Q.; Kong, Y.-H.; Wong, H.; Wu, W.-H. The WRKY6 transcription factor modulates PHOSPHATE1 expression in response to low Pi stress in Arabidopsis. Plant Cell 2009, 21, 3554–3566. [Google Scholar] [CrossRef] [PubMed]
- Bustos, R.; Castrillo, G.; Linhares, F.; Puga, M.I.; Rubio, V.; Perez-Perez, J.; Solano, R.; Leyva, A.; Poz-Ares, J. A central regulatory system largely control transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PLoS Genet. 2010, 6, e1001102. [Google Scholar] [CrossRef] [PubMed]
- Tran, H.T.; Hurley, B.A.; Plaxton, W.C. Feeding hungry plants: The role of purple acid phosphatases in phosphate nutrition. Plant Sci. 2010, 179, 14–27. [Google Scholar] [CrossRef]
- Franco-Zorrilla, J.M.; Gonzalez, E.; Bustos, R.; Linhares, F.; Leyva, A.; Paz-Ares, J. The transcriptional control of plant responses to phosphate limitation. J. Exp. Bot. 2004, 55, 285–293. [Google Scholar] [CrossRef] [PubMed]
- Rubio, V.; Linahares, F.; Solano, R.; Martin, A.C.; Iglesias, J.; Leyva, P.; Paz-Ares, J. A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and unicellular algae. Genes Dev. 2001, 15, 2122–2133. [Google Scholar] [CrossRef] [PubMed]
- Shane, M.W.; Stigter, K.; Fedosejevs, E.T.; Plaxton, W.C. Senescence-inducible cell wall and intracellular purple acid phosphatases: Implications for phosphorus remobilization in Hakea prostrata (Proteaceae) and Arabidopsis thaliana (Brassicaceae). J. Exp. Bot. 2014, 65, 6097–6106. [Google Scholar] [CrossRef] [PubMed]
- Robinson, W.D.; Park, J.; Tran, H.T.; Del Vecchio, H.A.; Ying, S.; Zins, J.L.; Patel, K.; McKnight, T.D.; Plaxton, W.C. The secreted purple acid phosphatase isozymes AtPAP12 and AtPAP26 play a pivotal role in extracellular phosphate-scavenging by Arabidopsis thaliana. J. Exp. Bot. 2012, 63, 6531–6542. [Google Scholar] [CrossRef] [PubMed]
- Veljanovski, V.; Vanderbeld, B.; Knowles, V.L.; Snedden, W.A.; Plaxton, W.C. Biochemical and molecular characterization of AtPAP26, a vacuolar purple acid phosphatase up-regulated in phosphate-deprived Arabidopsis suspension cells and seedlings. Plant Physiol. 2006, 142, 1282–1293. [Google Scholar] [CrossRef] [PubMed]
- Hurley, B.A.; Tran, H.T.; Marty, N.J.; Park, J.; Snedden, W.A.; Mullen, R.T.; Plaxton, W.C. The dual-targeted purple acid phosphatase isozyme AtPAP26 is essential for efficient acclimation of Arabidopsis to phosphate deprivation. Plant Physiol. 2010, 153, 1112–1122. [Google Scholar] [CrossRef] [PubMed]
- Tran, H.T.; Qian, W.; Hurley, B.A.; She, Y.-M.; Wang, D.; Plaxton, W.C. Biochemical and molecular characterization of AtPAP12 and AtPAP26: The predominant phosphatase isozymes secreted by phosphate-starved Arabidopsis thaliana. Plant Cell Environ. 2010, 33, 1789–1803. [Google Scholar] [CrossRef] [PubMed]
- Hillwig, M.S.; Contento, A.L.; Meyer, A.; Ebany, D.; Bassham, D.C.; MacIntosh, G.C. RNS2, a conserved member of the RNase T2 family, is necessary for ribosomal RNA decay in plants. Proc. Natl. Acad. Sci. USA 2011, 108, 1093–1098. [Google Scholar] [CrossRef] [PubMed]
- Bariola, P.A.; MacIntosh, G.C.; Green, P.J. Regulation of S-like ribonuclease levels in Arabidopsis. Antisense inhibition of RNS1 or RNS2 elevates anthocyanin accumulation. Plant Physiol. 1999, 119, 331–342. [Google Scholar] [CrossRef] [PubMed]
- Taylor, C.B.; Bariola, P.A.; delCardayre, S.B.; Raines, R.T.; Green, P.J. RNS2: A senescence-associated RNase of Arabidopsis that diverged from the S-RNases before speciation. Proc. Natl. Acad. Sci. USA 1993, 90, 5118–5122. [Google Scholar] [CrossRef] [PubMed]
- Kurata, N.; Kariu, T.; Kawano, S.; Kimura, M. Molecular cloning of cDNAs encoding ribonuclease-related proteins in Nicotiana glutinosa leaves, as induced in response to wounding or to TMV-infection. Biosci. Biotechnol. Biochem. 2002, 66, 391–397. [Google Scholar] [CrossRef] [PubMed]
- Liang, L.; Lai, Z.; Ma, W.; Zhang, Y.; Xue, Y. AhSL28, a senescence- and phosphate starvation-induced S-like RNase gene in Antirrhinum. Biochim. et Biophy. Acta—Gene. Struct. Expr. 2002, 1579, 64–71. [Google Scholar] [CrossRef]
- MacIntosh, G.C.; Hillwig, M.S.; Meyer, A.; Flagel, L. RNase T2 genes from rice and the evolution of secretory ribonuclease in plants. Mol. Genet. Genom. 2010, 283, 381–396. [Google Scholar] [CrossRef] [PubMed]
- Kothke, S.; Kock, M. The Solanum lycopersicum RNaseLER is a class II enzyme of the RNase T2 family and shows preferential expression in guard cells. J. Plant Physiol. 2011, 168, 840–847. [Google Scholar] [CrossRef] [PubMed]
- Kock, M.; Loffler, A.; Abel, S.; Glund, K. cDNA structure and regulatory properties of a family of starvation-induced ribonucleases from tomato. Plant Mol. Biol. 1995, 27, 477–485. [Google Scholar] [CrossRef] [PubMed]
- Lers, A.; Khalchitski, A.; Lomaniec, E.; Burd, S.; Green, P.J. Senescence-induced RNases in tomato. Plant Mol. Biol. 1998, 36, 439–449. [Google Scholar] [CrossRef] [PubMed]
- Lers, A.; Sonego, L.; Green, P.J.; Burd, S. Suppression of LX ribonuclease in tomato results in a delay of leaf senescence and abscission. Plant Physiol. 2006, 142, 710–721. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, K.; Hause, B.; Altmann, D.; Kock, M. Tomato ribonuclease LX with the functional endoplasmic reticulum retention motif HDEF is expressed during programmed cell death processes, including xylem differentiation, germination, and senescence. Plant Physiol. 2001, 127, 436–449. [Google Scholar] [CrossRef] [PubMed]
- Kock, M.; Gross, N.; Stenzel, I.; Hause, G. Phloem-specific expression of the wound-inducible ribonuclease LE from tomato (Lycopersicon esculentum cv. Lukullus). Planta 2004, 219, 233–242. [Google Scholar] [CrossRef] [PubMed]
- Ye, Z.-H.; Droste, D.L. Isolation and characterization of cDNAs encoding xylogenesis-associated and wound-induced ribonucleases in Zinnia elegans. Plant Mol. Biol. 1996, 30, 697–709. [Google Scholar] [CrossRef] [PubMed]
- Stephenson, P.; Hogan, J. Cloning and characterization of a ribonuclease, a cysteine proteinase, and an aspartic proteinase from pitchers of the carnivorous plant Nepenthes ventricosa Blanco. Int. J. Plant Sci. 2006, 167, 239–248. [Google Scholar] [CrossRef]
- Farage-Barhom, S.; Burd, S.; Sonego, L.; Mett, A.; Belausov, E.; Gidoni, D.; Lers, A. Localization of the Arabidopsis senescence- and cell death-associated BFN1 nuclease: From the ER to fragmented nuclei. Mol. Plant 2011, 4, 1062–1073. [Google Scholar] [CrossRef] [PubMed]
- Perez-Amador, M.A.; Abler, M.L.; DeRocher, E.J.; Thompson, D.M.; vanHoof, A.; LeBrasseur, N.D.; Lers, A.; Green, P.J. Identification of BFN1, a bifunctional nuclease induced during leaf and stem senescence in Arabidopsis. Plant Physiol. 2000, 122, 169–179. [Google Scholar] [CrossRef] [PubMed]
- Wagstaff, C.; Yang, T.J.W.; Stead, A.D.; Buchanan-Wollaston, V.; Roberts, J.A. A molecular and structural characterization of senescing Arabidopsis siiliques and comparison of transcriptional profiles with senescing petals and leaves. Plant J. 2009, 57, 690–705. [Google Scholar] [CrossRef] [PubMed]
- Lers, A.; Lomaniec, E.; Burd, S.; Khalchitski, A. The characterization of LeNUC1, a nuclease associated with leaf senescence of tomato. Physiol. Plant 2001, 112, 176–182. [Google Scholar] [CrossRef] [PubMed]
- Hong, Y.W.; Wang, T.W.; Hudak, K.A.; Schade, F.; Froese, C.D.; Thompson, J. An ethylene-induced cDNA encoding a lipase expressed at the onset of senescence. Proc. Natl. Acad. Sci. USA 2000, 97, 8717–8722. [Google Scholar] [CrossRef] [PubMed]
- Tompson, J.; Taylor, C.; Wang, T.W. Altered membrane lipase expression delays leaf senescence. Biochem. Soc. Trans. 2000, 28, 775–777. [Google Scholar] [CrossRef]
- Nakamura, Y.; Awai, K.; Masuda, T.; Yoshioka, Y.; Takamiya, K.-I.; Ohta, H. A novel phosphatidylcholine-hydrolyzing phospholipase C induced by phosphate starvation in Arabidopsis. J. Biol. Chem. 2005, 280, 7469–7476. [Google Scholar] [CrossRef] [PubMed]
- Jia, Y.; Tao, F.; Li, W. Lipid profiling demonstrates that suppressing Arabidopsis phospholipase Dα retards ABA-promoted leaf senescence by attenuating lipid degradation. PLoS ONE 2013, 8. [Google Scholar] [CrossRef]
- Del Pozo, J.C.; Allona, I.; Ruio, V.; Leyva, A.; de La Pena, A.; Aragoncilla, C.; Paz-Ares, J. A type 5 acid phosphatase gene from Arabidopsis thaliana is induced by phosphate starvation and by some other types of phosphate mobilising/oxidative stress conditions. Plant J. 1999, 19, 579–589. [Google Scholar] [CrossRef] [PubMed]
- May, A.; Berger, S.; Hertel, T.; Kock, M. The Arabidopsis thaliana phosphate starvation responsive gene AtPPsPase1 encodes a novel type of inorganic pyrophosphatase. Biochim. et Biophy. Acta—Gen. Subj. 2011, 1810, 178–185. [Google Scholar] [CrossRef] [PubMed]
- May, A.; Spinka, M.; Kock, M. Arabidopsis thaliana PECP1—Enzymatic characterization and structural organization of the first plant phosphoethanolamine/phosphocholine phosphatase. Biochim. et Biophy. Acta—Proteins Proteom. 2012, 1824, 319–325. [Google Scholar] [CrossRef] [PubMed]
- Caparros-Martin, J.A.; McCarthy-Suarez, I.; Culianez-Macia, F.A. HAD hydrolase function unveiled by substrate screening: Enzymatic characterization of Arabidopsis thaliana subclass I phosphosugar phosphatase AtSgpp. Planta 2013, 237, 943–954. [Google Scholar] [CrossRef] [PubMed]
- Baldwin, J.C.; Karthikeyan, A.S.; Raghothama, K.G. LEPS2, a phosphorus starvation-induced novel acid phosphatase from tomato. Plant Physiol. 2001, 125, 728–737. [Google Scholar] [CrossRef] [PubMed]
- Baldwin, J.C.; Karthikeyan, A.S.; Cao, A.; Raghothama, K.G. Biochemical and molecular analysis of LePS2;1: A phosphate starvation induced protein phosphatase gene from tomato. Planta 2008, 228, 273–280. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.-Q.; Allan, D.L.; Vance, C.P. Systematic signaling and local sensing of phosphate in common bean: Cross-talk between photosynthate and microRNA399. Mol. Plant 2010, 3, 428–437. [Google Scholar] [CrossRef] [PubMed]
- Liang, C.Y.; Chen, Z.J.; Yao, Z.-F.; Tian, J.; Liao, H. Characterization of two putative protein phosphatase genes and their involvement in phosphorus efficiency in Phaseolus vulgaris. J. Integr. Plant Biol. 2012, 54, 400–411. [Google Scholar] [CrossRef] [PubMed]
- Hensel, L.L.; Grbic, V.; Baumgarten, D.A.; Bleecker, A.B. Developmental and age-related processes that influence the longevity and senescence of photosynthetic tissues in Arabidopsis. Plant Cell 1993, 5, 553–564. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, Y.; Kihara-Doi, T.; Kawazu, T.; Miyake, C.; Makino, A. Differences in Rubiso content and its synthesis in leaves at different positions in Eucalyptus globulus seedlings. Plant Cell Environ. 2010, 33, 1314–1323. [Google Scholar] [PubMed]
- Hua, Z.-H.; Fields, A.; Kao, T. Biochemical models for S-RNase-based self-incompatibility. Mol. Plant 2008, 1, 575–585. [Google Scholar] [CrossRef] [PubMed]
- Abel, S.; Nurnberger, T.; Ahnert, V.; Krauss, G.-J.; Glund, K. Induction of an extracellular cyclic nucleotide phosphodiesterase as an accessory ribonucleolytic activity during phosphate starvation of cultured tomato cells. Plant Physiol. 2000, 122, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Okabe, T.; Iwakiri, Y.; Mori, H.; Ogawa, T.; Ohyama, T. An S-like ribonuclease gene is used to generate a trap-leaf enzyme in the carnivorous plant Drosera adelae. FEBS Lett. 2005, 579, 5729–5733. [Google Scholar] [CrossRef] [PubMed]
- Lim, P.O.; Kim, H.J.; Nam, H.G. Leaf Senescence. Annu. Rev. Plant Biol. 2007, 58, 115–136. [Google Scholar] [CrossRef] [PubMed]
- Farage-Barhom, S.; Burd, S.; Sonego, L.; Perl-Treves, R.; Lers, A. Expression analysis of the BFN1 nuclease gene promoter during senescence, abscission, and programmed cell death-related processes. J. Exp. Bot. 2008, 59, 3247–3258. [Google Scholar] [CrossRef] [PubMed]
- Tronsco-Ponce, M.A.; Cao, X.; Yang, Z.; Ohlrogge, J.B. Lipid turnover during senescence. Plant Sci. 2013, 205, 13–19. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.E.; Froese, C.D.; Madey, E.; Smith, M.D.; Hong, Y. Lipid metabolism during plant senescence. Prog. Lipid Res. 1998, 37, 119–141. [Google Scholar] [CrossRef]
- Yang, Z.I.; Ohlrogge, J.B. Turnover of fatty acids during natural senescence of Arabidopsis, Brachypodium, and switchgrass and in Arabidopsis beta-oxidation mutants. Plant Physiol. 2009, 150, 1981–1989. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Snyder, C.L.; Greer, M.S.; Weselake, R.J. Biology and biochemistry of plant phospholipases. Crit. Rev. Plant Sci. 2011, 30, 239–258. [Google Scholar] [CrossRef]
- Fan, L.; Zheng, S.; Wang, X. Antisense suppression of phospholipase D alpha retards abscisic acid- and ethylene-promoted senescence of postharvest Arabidopsis leaves. Plant Cell 1997, 9, 2183–2196. [Google Scholar] [CrossRef] [PubMed]
- Kaup, M.T.; Froese, C.D.; Thompson, J.E. A role for diacylglycerol acyltransferase during leaf senescence. Plant Physiol. 2002, 129, 1616–1626. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Zhu, H.; Liu, K.; Liu, X.; Leggewie, G.; Udvard, M.; Wang, D. Purple acid phosphatases of Arabidopsis thaliana—Comparative analysis and differential regulation by phosphate deprivation. J. Biol. Chem. 2002, 277, 27772–27781. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Gan, S.S. Convergence and divergence in gne expression profiles induced by leaf senescence and 27 senescence-promoting hormonal, pathological and environmental stress treatments. Plant Cell Environ. 2012, 35, 644–655. [Google Scholar] [CrossRef] [PubMed]
- Burroughs, A.M.; Allen, K.N.; Dunaway-Mariano, D.; Aravind, L. Evolutionary genomics of the HAD superfamily: Understanding the structural adaptations and catalytic diversity in a superfamily of phosphoesterases and allied enzymes. J. Mol. Biol. 2006, 361, 1003–1034. [Google Scholar] [CrossRef] [PubMed]
- Van der Graaff, E.; Schwacke, R.; Schneider, A.; Desimone, M.; Flugge, U.-I.; Kunze, R. Transcription analysis of Arabidopsis membrane transporters and hormone pathways during developmental and induced leaf senescence. Plant Physiol. 2006, 141, 776–792. [Google Scholar] [CrossRef] [PubMed]
- Smith, F.W.; Mudge, S.R.; Rae, A.L.; Glassop, D. Phosphate transport in plants. Plant Soil 2003, 248, 71–83. [Google Scholar] [CrossRef]
- Nussaume, L.; Kanno, S.; Javot, H.; Marin, E.; Pochon, N.; Ayadi, A.; Nakanishi, T.M.; Thibaud, M.-C. Phosphate import in plants: Focus on the PHT1 transporters. Front. Plant Sci. 2011, 2, 83. [Google Scholar] [CrossRef] [PubMed]
- Nagarajan, V.K.; Jain, A.; Poling, M.D.; Lewis, A.J.; Raghothama, K.G.; Smith, A.P. Arabidopsis Pht1;5 mobilizes phosphate between source and sink organs and influences the interaction between phosphate homeostasis and ethylene signaling. Plant Physiol. 2011, 156, 1149–1163. [Google Scholar] [CrossRef] [PubMed]
- Rae, A.L.; Cybinski, D.H.; Jarmey, J.M.; Smith, F.W. Characterization of two phosphate transporters from barley; evidence for diverse function and kinetic properties among members of the Pht1 family. Plant Mol. Biol. 2003, 53, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Ai, P.; Sun, S.; Zhao, J.; Fan, X.; Xin, W.; Guo, Q.; Yu, L.; Shen, Q.; Wu, P.; Miller, A.J.; et al. Two rice phosphate transporters, OsPht1;2 and OsPht1;6, have different functions and kinetic properties in uptake and translocation. Plant J. 2009, 57, 798–809. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.; Gu, M.; Cao, Y.; Huang, X.; Zhang, X.; Ai, P.; Zhao, J.; Fan, X.; Xu, G. A constitutive expressed phosphate transporter, OsPht1;1, modulates phosphate uptake and translocation in phosphate-replete rice. Plant Physiol. 2012, 159, 1571–1581. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Ren, H.; Gu, M.; Zhao, J.; Sun, S.; Zhang, X.; Chen, J.; Wu, P.; Xu, G. The phosphate transporter gene OsPht1;8 is involved in phosphate homeostasis in rice. Plant Physiol. 2011, 156, 1164–1175. [Google Scholar] [CrossRef] [PubMed]
- Daram, P.; Brunner, S.; Rausch, C.; Steiner, C.; Amrhein, N.; Bucher, M. Pht2;1 encodes a low-affinity phosphate transporter from Arabidopsis. Plant Cell 1999, 11, 2153–2166. [Google Scholar] [CrossRef] [PubMed]
- Versaw, W.K.; Harrison, M.J. A chloroplast phosphate transporter, PHT2;1, influences allocation of phosphate within the plant and phosphate-starvation responses. Plant Cell 2002, 14, 1751–1766. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Zhao, X.; Liu, X.; Zhang, L.; Gu, J.; Li, X.; Lu, W.; Xiao, K. Function of wheat phosphate transporter gene TaPHT2;1 in Pi translocation and plant growth regulation under replete and limited Pi supply conditions. Planta 2013, 237, 1163–1178. [Google Scholar] [CrossRef] [PubMed]
- Aziz, T.; Finnegan, P.M.; Lambers, H.; Jost, R. Organ-specific phosphorus-allocation patterns and transcript profiles linked to phosphorus efficiency in two contrasting wheat genotypes. Plant Cell Environ. 2014, 37, 943–960. [Google Scholar] [CrossRef] [PubMed]
- Porier, Y.; Thoma, S.; Somerville, C.; Schiefelbein, J. Mutant of Arabidopsis deficient in xylem loading of phosphate. Plant Physiol. 1991, 97, 1087–1093. [Google Scholar] [CrossRef]
- Zimmerli, C.; Ribot, C.; Vavasseur, A.; Bauer, H.; Hedrich, R.; Poirer, Y. PHO1 expression in guard cells mediates the stomatal response to abscisic acid in Arabidopsis. Plant J. 2012, 72, 199–211. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Huang, W.; Ying, Y.; Li, S.; Secco, D.; Tyerman, S.; Whelan, J.; Shou, H. Function characterization of the rice SPX-MFS family reveals a key role of OsSPX-MFS1 incontrolling phosphate homeostasis in leaves. New Phytol. 2012, 196, 139–148. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.-Y.; Huang, T.-K.; Chiou, T.-J. NITROGEN LIMITATION ADAPTATION, a target of microRNA827, mediates degradation of plasma membrane-localized phosphate transporters to maintain phosphate homeostasis in Arabidopsis. Plant Cell 2013, 25, 4061–4074. [Google Scholar] [CrossRef] [PubMed]
- Peng, M.; Hannam, C.; Gu, H.; Bi, Y.-M.; Rothstein, S.J. A mutation in NLA, which encodes a RING-type ubiquitin ligase, disrupts the adaptability of Arabidopsis to nitrogen limitation. Plant. J. 2007, 50, 320–337. [Google Scholar] [CrossRef] [PubMed]
- Kant, S.; Peng, M.; Rothstein, S.J. Genetic regulation by NLA and microRNA827 for maintaining nitrate-dependent phosphate homeostasis in Arabidopsis. PLoS Genet. 2011. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Stigter, K.A.; Plaxton, W.C. Molecular Mechanisms of Phosphorus Metabolism and Transport during Leaf Senescence. Plants 2015, 4, 773-798. https://doi.org/10.3390/plants4040773
Stigter KA, Plaxton WC. Molecular Mechanisms of Phosphorus Metabolism and Transport during Leaf Senescence. Plants. 2015; 4(4):773-798. https://doi.org/10.3390/plants4040773
Chicago/Turabian StyleStigter, Kyla A., and William C. Plaxton. 2015. "Molecular Mechanisms of Phosphorus Metabolism and Transport during Leaf Senescence" Plants 4, no. 4: 773-798. https://doi.org/10.3390/plants4040773
APA StyleStigter, K. A., & Plaxton, W. C. (2015). Molecular Mechanisms of Phosphorus Metabolism and Transport during Leaf Senescence. Plants, 4(4), 773-798. https://doi.org/10.3390/plants4040773





