Abscisic Acid: Hidden Architect of Root System Structure
Abstract
:1. Introduction
2. ABA Regulation of the Major Control Points of Root Growth: Cell Division and Cell Elongation
2.1. ABA Regulates Root Meristem Function
2.2. Modulation of Root Cell Length by ABA
3. Control of Root Length by ABA
3.1. The Endodermis Is the Site of ABA Control of Root Elongation
3.2. Controlling the Transition from Proliferation to Differentiation
4. Lateral Root Development
4.1. Overview of Lateral Root Development
4.2. Different Developmental Roles for ABA in Different Taxa
4.2.1. Arabidopsis thaliana
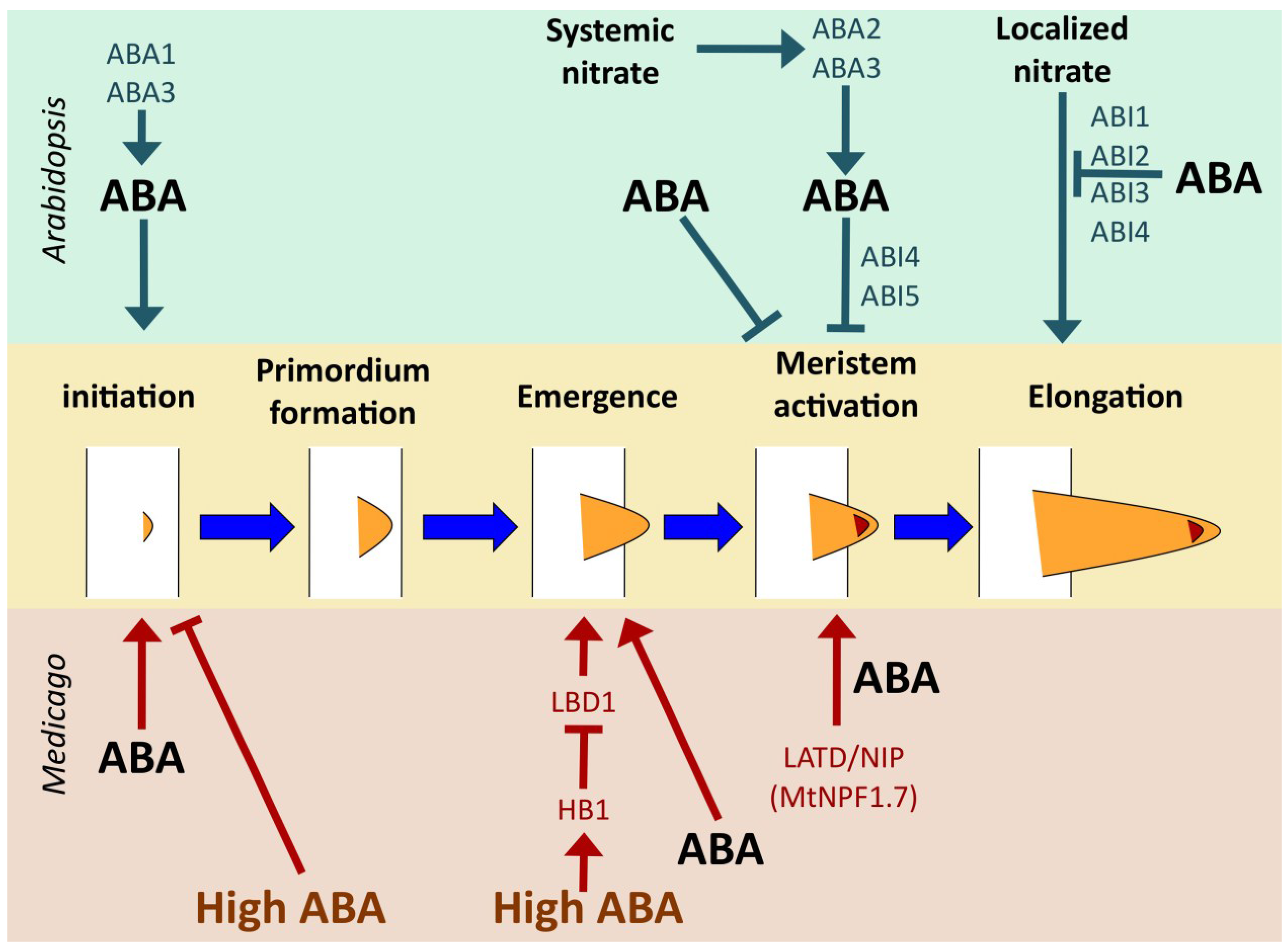
4.2.2. Medicago truncatula
4.2.3. Oryza sativa
5. ABA Shapes Root Architecture in Response to Environmental Signals
5.1. Regulation of Root Growth and Branching in Response to Abiotic Stress
5.2. Hydrotropism, Halotropism and Water Patterning
5.3. Responses to Nitrate: NPF Transporters Connect Nitrate, Hormones and Root Architecture
6. ABA Functions in a Hormone Signaling Network
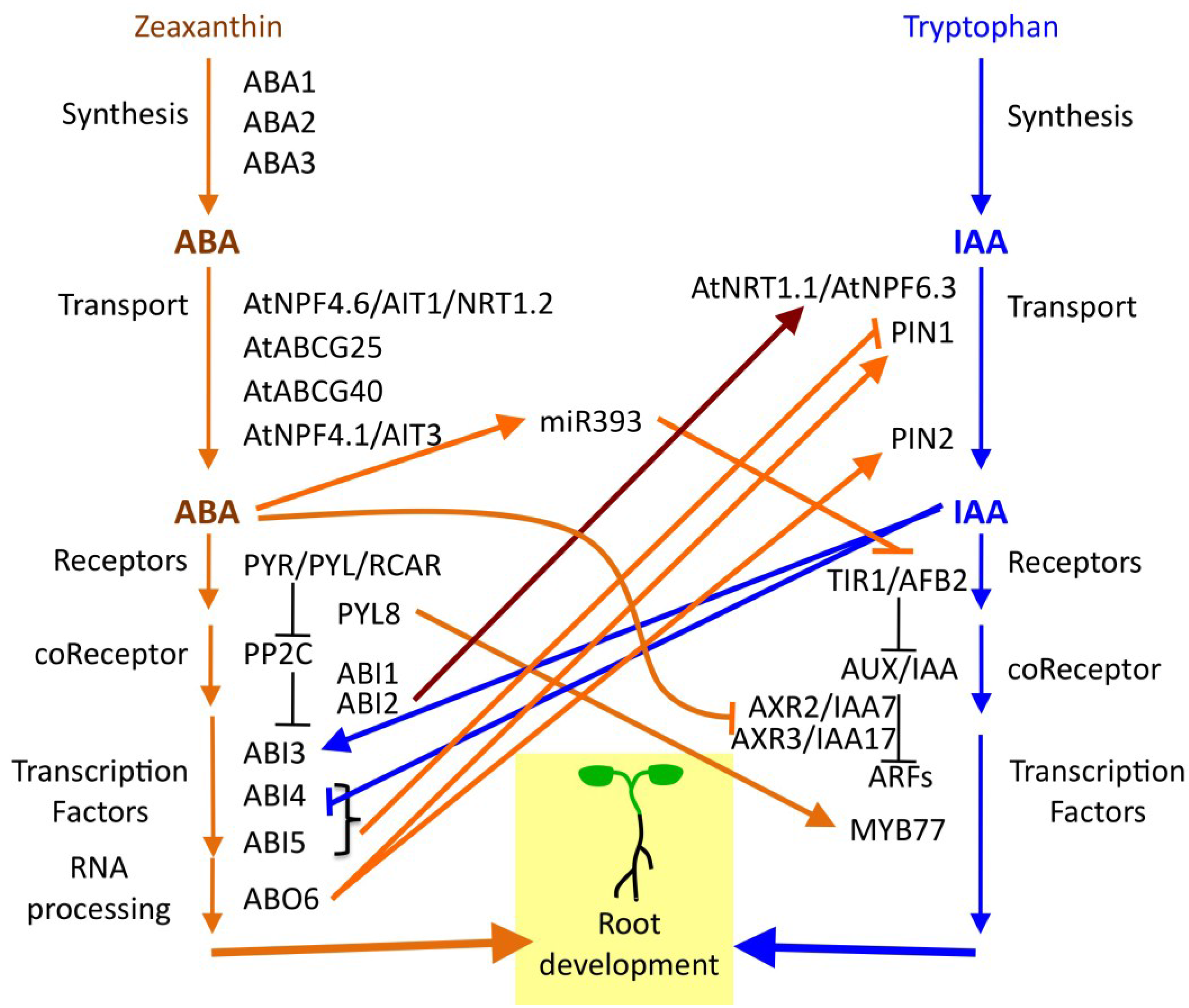
7. Diversity of Root Architecture in Different Species: ABA Signaling as a Target of Evolution?
8. Conclusions
9. Future Avenues
Acknowledgments
Conflicts of Interest
References
- Finlayson, S.A.; Krishnareddy, S.R.; Kebrom, T.H.; Casal, J.J. Phytochrome regulation of branching in Arabidopsis. Plant Physiol. 2010, 152, 1914–1927. [Google Scholar]
- Bao, Y.; Aggarwal, P.; Robbins, N.E., 2nd; Sturrock, C.J.; Thompson, M.C.; Tan, H.Q.; Tham, C.; Duan, L.; Rodriguez, P.L.; Vernoux, T.; et al. Plant roots use a patterning mechanism to position lateral root branches toward available water. Proc. Natl. Acad. Sci. USA 2014, 111, 9319–9324. [Google Scholar]
- Rostamza, M.; Richards, R.A.; Watt, M. Response of millet and sorghum to a varying water supply around the primary and nodal roots. Ann. Bot. 2013, 112, 439–446. [Google Scholar]
- Richter, G.L.; Monshausen, G.B.; Krol, A.; Gilroy, S. Mechanical stimuli modulate lateral root organogenesis. Plant Physiol. 2009, 151, 1855–1866. [Google Scholar]
- Laskowski, M.; Grieneisen, V.A.; Hofhuis, H.; Hove, C.A.; Hogeweg, P.; Maree, A.F.; Scheres, B. Root system architecture from coupling cell shape to auxin transport. PLoS Biol. 2008, 6, e307. [Google Scholar] [CrossRef] [PubMed]
- Rich, S.M.; Watt, M. Soil conditions and cereal root system architecture: Review and considerations for linking Darwin and Weaver. J. Exp. Bot. 2013, 64, 1193–1208. [Google Scholar] [CrossRef] [PubMed]
- Goh, C.H.; Veliz Vallejos, D.F.; Nicotra, A.B.; Mathesius, U. The impact of beneficial plant-associated microbes on plant phenotypic plasticity. J. Chem. Ecol. 2013, 39, 826–839. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Forde, B.G. An Arabidopsis MADS box gene that controls nutrient-induced changes in root architecture. Science 1998, 279, 407–409. [Google Scholar] [CrossRef] [PubMed]
- Zolla, G.; Heimer, Y.M.; Barak, S. Mild salinity stimulates a stress-induced morphogenic response in Arabidopsis thaliana roots. J. Exp. Bot. 2010, 61, 211–224. [Google Scholar] [CrossRef] [PubMed]
- Linkohr, B.I.; Williamson, L.C.; Fitter, A.H.; Leyser, H.M. Nitrate and phosphate availability and distribution have different effects on root system architecture of Arabidopsis. Plant J. 2002, 29, 751–760. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.P.; Brown, K.M. Topsoil foraging—An architectural adaptation of plants to low phosphorus availability. Plant Soil 2001, 237, 225–237. [Google Scholar] [CrossRef]
- Sozzani, R.; Iyer-Pascuzzi, A. Postembryonic control of root meristem growth and development. Curr. Opin. Plant Biol. 2014, 17, 7–12. [Google Scholar] [CrossRef] [PubMed]
- Vermeer, J.E.; Geldner, N. Lateral root initiation in Arabidopsis thaliana: A force awakens. F1000Prime Rep. 2015, 7. [Google Scholar] [CrossRef] [PubMed]
- Finkelstein, R. Abscisic acid synthesis and response. Arabidopsis Book 2013, 11, e0166. [Google Scholar] [CrossRef] [PubMed]
- Petricka, J.J.; Benfey, P.N. Root layers: Complex regulation of developmental patterning. Curr. Opin. Genet. Dev. 2008, 18, 354–361. [Google Scholar] [CrossRef] [PubMed]
- Heidstra, R.; Sabatini, S. Plant and animal stem cells: Similar yet different. Nat. Rev. Mol. Cell Biol. 2014, 15, 301–312. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Mitchell, D.M.; Harris, J.M. Abscisic acid rescues the root meristem defects of the Medicago truncatula latd mutant. Dev. Biol. 2007, 304, 297–307. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Han, W.; De Smet, I.; Talboys, P.; Loya, R.; Hassan, A.; Rong, H.; Jurgens, G.; Paul Knox, J.; Wang, M.H. ABA promotes quiescence of the quiescent centre and suppresses stem cell differentiation in the Arabidopsis primary root meristem. Plant J. 2010, 64, 764–774. [Google Scholar] [CrossRef] [PubMed]
- Sharp, R.E.; Poroyko, V.; Hejlek, L.G.; Spollen, W.G.; Springer, G.K.; Bohnert, H.J.; Nguyen, H.T. Root growth maintenance during water deficits: Physiology to functional genomics. J. Exp. Bot. 2004, 55, 2343–2351. [Google Scholar] [CrossRef] [PubMed]
- Sharp, R.E.; Wu, Y.J.; Voetberg, G.S.; Saab, I.N.; Lenoble, M.E. Confirmation that abscisic-acid accumulation is required for maize primary root elongation at low water potentials. J. Exp. Bot. 1994, 45, 1743–1751. [Google Scholar]
- Zhang, C.; Bousquet, A.; Harris, J.M. Abscisic acid and LATERAL ROOT ORGAN DEFECTIVE/NUMEROUS INFECTIONS AND POLYPHENOLICS modulate root elongation via reactive oxygen species in Medicago truncatula. Plant Physiol. 2014, 166, 644–658. [Google Scholar] [CrossRef] [PubMed]
- Léran, S.; Varala, K.; Boyer, J.-C.; Chiurazzi, M.; Crawford, N.M.; Daniel-Vedele, F.; David, L.; Dickstein, R.; Fernandez, E.; Forde, B.J.; et al. A unified nomenclature of nitrate transporter 1/peptide transporter family members in plants. Trends Plant Sci. 2014, 19, 5–9. [Google Scholar] [CrossRef] [PubMed]
- Yendrek, C.R.; Lee, Y.-C.; Morris, V.; Liang, Y.; Pislariu, C.I.; Burkart, G.; Meckfessel, M.H.; Salehin, M.; Kessler, H.; Lloyd, M.; et al. A putative transporter is essential for integrating nutrient and hormone signaling with lateral root growth and nodule development in Medicago truncatula. Plant J. 2010, 62, 100–112. [Google Scholar] [CrossRef] [PubMed]
- Bagchi, R.; Salehin, M.; Adeyemo, O.S.; Salazar, C.; Shulaev, V.; Sherrier, D.J.; Dickstein, R. Functional assessment of the Medicago truncatula NIP/LATD protein demonstrates that it is a high-affinity nitrate transporter. Plant Physiol. 2012, 160, 906–916. [Google Scholar] [CrossRef] [PubMed]
- Pellizzaro, A.; Clochard, T.; Cukier, C.; Bourdin, C.; Juchaux, M.; Montrichard, F.; Thany, S.; Raymond, V.; Planchet, E.; Limami, A.M.; et al. The nitrate transporter MtNPF6.8 (MtNRT1.3) transports abscisic acid and mediates nitrate regulation of primary root growth in Medicago truncatula. Plant Physiol. 2014, 166, 2152–2165. [Google Scholar] [CrossRef] [PubMed]
- Duan, L.; Dietrich, D.; Ng, C.H.; Chan, P.M.; Bhalerao, R.; Bennett, M.J.; Dinneny, J.R. Endodermal ABA signaling promotes lateral root quiescence during salt stress in Arabidopsis seedlings. Plant Cell 2013, 25, 324–341. [Google Scholar] [CrossRef] [PubMed]
- Geng, Y.; Wu, R.; Wee, C.W.; Xie, F.; Wei, X.; Chan, P.M.; Tham, C.; Duan, L.; Dinneny, J.R. A spatio-temporal understanding of growth regulation during the salt stress response in Arabidopsis. Plant Cell 2013, 25, 2132–2154. [Google Scholar] [CrossRef] [PubMed]
- Ubeda-Tomas, S.; Swarup, R.; Coates, J.; Swarup, K.; Laplaze, L.; Beemster, G.T.; Hedden, P.; Bhalerao, R.; Bennett, M.J. Root growth in Arabidopsis requires gibberellin/DELLA signalling in the endodermis. Nat. Cell Biol. 2008, 10, 625–628. [Google Scholar] [CrossRef] [PubMed]
- Kwak, J.M.; Mori, I.C.; Pei, Z.M.; Leonhardt, N.; Torres, M.A.; Dangl, J.L.; Bloom, R.E.; Bodde, S.; Jones, J.D.; Schroeder, J.I. NADPH oxidase AtRbohD and AtRbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO. J. 2003, 22, 2623–2633. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Duan, Y.; Hua, D.; Fan, G.; Wang, L.; Liu, Y.; Chen, Z.; Han, L.; Qu, L.J.; Gong, Z. DEXH box RNA helicase-mediated mitochondrial reactive oxygen species production in Arabidopsis mediates crosstalk between abscisic acid and auxin signaling. Plant Cell 2012, 24, 1815–1833. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, J.A.; Daudi, A.; Butt, V.S.; Bolwell, G.P. Reactive oxygen species and their role in plant defence and cell wall metabolism. Planta 2012, 236, 765–779. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, R.; Schippers, J.H.M. ROS-mediated redox signaling during cell differentiation in plants. Biochim. Biophys. Acta 2015, 1850, 1497–1508. [Google Scholar] [CrossRef] [PubMed]
- Tsukagoshi, H.; Busch, W.; Benfey, P.N. Transcriptional regulation of ROS controls transition from proliferation to differentiation in the root. Cell 2010, 143, 606–616. [Google Scholar] [CrossRef] [PubMed]
- Meijon, M.; Satbhai, S.B.; Tsuchimatsu, T.; Busch, W. Genome-wide association study using cellular traits identifies a new regulator of root development in Arabidopsis. Nat. Genet. 2014, 46, 77–81. [Google Scholar] [CrossRef] [PubMed]
- Wachsman, G.; Sparks, E.E.; Benfey, P.N. Genes and networks regulating root anatomy and architecture. New Phytol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Mathesius, U.; Weinman, J.J.; Rolfe, B.J.; Djordjevic, M.A. Rhizobia can induce nodules in white clover by “hijacking” mature cortical cells activated during lateral root development. Mol. Plant-Microbe Interact. 2000, 13, 170–182. [Google Scholar] [CrossRef] [PubMed]
- Mallory, T.E.; Chang, S.-H.; Cutter, E.G.; Gifford, E.M. Sequence and pattern of lateral root formation in five selected species. Am. J. Bot. 1970, 57, 800–809. [Google Scholar] [CrossRef]
- Byrne, J.M.; Pesacreta, T.C.; Fox, J.A. Development and structure of vascular connection between primary and secondary root of Glycine max (L.) Merr. Am. J. Bot. 1977, 64, 946–959. [Google Scholar] [CrossRef]
- Op den Camp, R.H.; de Mita, S.; Lillo, A.; Cao, Q.; Limpens, E.; Bisseling, T.; Geurts, R. A phylogenetic strategy based on a legume-specific whole genome duplication yields symbiotic cytokinin type-A response regulators. Plant Physiol. 2011, 157, 2013–2022. [Google Scholar] [CrossRef] [PubMed]
- Herrbach, V.; Rembliere, C.; Gough, C.; Bensmihen, S. Lateral root formation and patterning in Medicago truncatula. J. Plant Physiol. 2014, 171, 301–310. [Google Scholar] [CrossRef] [PubMed]
- Casero, P.J.; Casimiro, I.; Lloret, P.G. Pericycle proliferation pattern during the lateral root initiation in adventitious roots of Allium cepa. Protoplasma 1996, 191, 136–147. [Google Scholar] [CrossRef]
- Kawata, S.; Shibayama, H. On the lateral root primordia formation in the crown roots of rice plants. Jpn. J. Crop Sci. 1965, 33, 423–431. [Google Scholar] [CrossRef]
- Ni, J.; Shen, Y.-X.; Zhang, Y.-Y.; Liu, Y. Histological characterization of the lateral root primordium development in rice. Bot. Stud. 2014, 55. [Google Scholar] [CrossRef]
- Bell, J.K.; McCully, M.E. A histological study of lateral root initiation and development in Zea mays. Protoplasma 1970, 70, 179–205. [Google Scholar] [CrossRef]
- Moreno-Risueno, M.A.; Van Norman, J.M.; Moreno, A.; Zhang, J.; Ahnert, S.E.; Benfey, P.N. Oscillating gene expression determines competence for periodic Arabidopsis root branching. Science 2010, 329, 1306–1311. [Google Scholar] [CrossRef] [PubMed]
- Xuan, W.; Audenaert, D.; Parizot, B.; Moller, B.K.; Njo, M.F.; de Rybel, B.; de Rop, G.; van Isterdael, G.; Mahonen, A.P.; Vanneste, S.; et al. Root cap-derived auxin pre-patterns the longitudinal axis of the Arabidopsis root. Curr. Biol. 2015, 25, 1381–1388. [Google Scholar] [CrossRef] [PubMed]
- Peret, B.; de Rybel, B.; Casimiro, I.; Benkova, E.; Swarup, R.; Laplaze, L.; Beeckman, T.; Bennett, M.J. Arabidopsis lateral root development: An emerging story. Trends Plant Sci. 2009, 14, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Malamy, J.E.; Benfey, P.N. Organization and cell differentiation in lateral roots of Arabidopsis thaliana. Dev. 1997, 124, 33–44. [Google Scholar]
- Cheng, J.C.; Seeley, K.A.; Sung, Z.R. RML1 and RML2, Arabidopsis genes required for cell proliferation at the root tip. Plant Physiol. 1995, 107, 365–376. [Google Scholar] [CrossRef] [PubMed]
- Tian, H.; de Smet, I.; Ding, Z. Shaping a root system: Regulating lateral versus primary root growth. Trends Plant Sci. 2014, 19, 426–431. [Google Scholar] [CrossRef] [PubMed]
- Atkinson, J.A.; Rasmussen, A.; Traini, R.; Voss, U.; Sturrock, C.; Mooney, S.J.; Wells, D.M.; Bennett, M.J. Branching out in roots: Uncovering form, function, and regulation. Plant Physiol. 2014, 166, 538–550. [Google Scholar] [CrossRef] [PubMed]
- Vilches-Barro, A.; Maizel, A. Talking through walls: Mechanisms of lateral root emergence in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2015, 23, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Jennings, A.; Barlow, P.W.; Forde, B.G. Dual pathways for regulation of root branching by nitrate. Proc. Natl. Acad. Sci. USA 1999, 96, 6529–6534. [Google Scholar] [CrossRef] [PubMed]
- Signora, L.; de Smet, I.; Foyer, C.H.; Zhang, H. ABA plays a central role in mediating the regulatory effects of nitrate on root branching in Arabidopsis. Plant J. 2001, 28, 655–662. [Google Scholar] [CrossRef] [PubMed]
- Van Norman, J.M.; Zhang, J.; Cazzonelli, C.I.; Pogson, B.J.; Harrison, P.J.; Bugg, T.D.; Chan, K.X.; Thompson, A.J.; Benfey, P.N. Periodic root branching in Arabidopsis requires synthesis of an uncharacterized carotenoid derivative. Proc. Natl. Acad. Sci. USA 2014, 111, E1300–E1309. [Google Scholar] [CrossRef] [PubMed]
- Ariel, F.; Diet, A.; Verdenaud, M.; Gruber, V.; Frugier, F.; Chan, R.; Crespi, M. Environmental regulation of lateral root emergence in Medicago truncatula requires the HD-ZIP I transcription factor HB1. Plant Cell 2010, 22, 2171–2183. [Google Scholar] [CrossRef] [PubMed]
- De Smet, I.; Signora, L.; Beeckman, T.; Inze, D.; Foyer, C.H.; Zhang, H. An abscisic acid-sensitive checkpoint in lateral root development of Arabidopsis. Plant J. 2003, 33, 543–555. [Google Scholar] [CrossRef] [PubMed]
- Bright, L.J.; Liang, Y.; Mitchell, D.M.; Harris, J.M. The LATD gene of Medicago truncatula is required both for nodule and root development. Mol. Plant-Microbe Interact. 2005, 18, 521–532. [Google Scholar] [CrossRef] [PubMed]
- Brocard-Gifford, I.; Lynch, T.J.; Garcia, M.E.; Malhotra, B.; Finkelstein, R.R. The Arabidopsis thaliana ABSCISIC ACID-INSENSITIVE8 locus encodes a novel protein mediating abscisic acid and sugar responses essential for growth. Plant Cell 2004, 16, 406–421. [Google Scholar] [CrossRef] [PubMed]
- Lertpiriyapong, K.; Sung, Z.R. The elongation defective1 mutant of Arabidopsis is impaired in the gene encoding a serine-rich secreted protein. Plant Mol. Biol. 2003, 53, 581–595. [Google Scholar] [CrossRef] [PubMed]
- Pagant, S.; Bichet, A.; Sugimoto, K.; Lerouxel, O.; Desprez, T.; McCann, M.; Lerouge, P.; Vernhettes, S.; Hofte, H. KOBITO1 encodes a novel plasma membrane protein necessary for normal synthesis of cellulose during cell expansion in Arabidopsis. Plant Cell 2002, 14, 2001–2013. [Google Scholar] [CrossRef] [PubMed]
- Kong, D.; Karve, R.; Willet, A.; Chen, M.K.; Oden, J.; Shpak, E.D. Regulation of plasmodesmatal permeability and stomatal patterning by the glycosyltransferase-like protein KOBITO1. Plant Physiol. 2012, 159, 156–168. [Google Scholar] [CrossRef] [PubMed]
- Benitez-Alfonso, Y.; Faulkner, C.; Pendle, A.; Miyashima, S.; Helariutta, Y.; Maule, A. Symplastic intercellular connectivity regulates lateral root patterning. Dev. Cell 2013, 26, 136–147. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.; Hwang, J.U.; Lee, M.; Kim, Y.Y.; Assmann, S.M.; Martinoia, E.; Lee, Y. PDR-type ABC transporter mediates cellular uptake of the phytohormone abscisic acid. Proc. Natl. Acad. Sci. USA 2010, 107, 2355–2360. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Xing, L.; Wang, X.; Hou, Y.J.; Gao, J.; Wang, P.; Duan, C.G.; Zhu, X.; Zhu, J.K. The ABA receptor PYL8 promotes lateral root growth by enhancing MYB77-dependent transcription of auxin-responsive genes. Sci. Signal 2014, 7, ra53. [Google Scholar] [CrossRef] [PubMed]
- Deak, K.I.; Malamy, J. Osmotic regulation of root system architecture. Plant J. 2005, 43, 17–28. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.-H.; Endo, A.; Zhou, L.; Penney, J.; Chen, H.-C.; Arroyo, A.; Leon, P.; Nambara, E.; Asami, T.; Seo, M.; et al. A unique short-chain dehydrogenase/reductase in Arabidopsis glucose signaling and abscisic acid biosynthesis and functions. Plant Cell 2002, 14, 2723–2743. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, A.A.; Agbevenou, K.; Herrbach, V.; Gough, C.; Bensmihen, S. Abscisic acid promotes pre-emergence stages of lateral root development in Medicago truncatula. Plant Signal Behav. 2015, 10, e977741. [Google Scholar] [CrossRef] [PubMed]
- Pierik, R.; Tholen, D.; Poorter, H.; Visser, E.J.; Voesenek, L.A. The Janus face of ethylene: Growth inhibition and stimulation. Trends Plant Sci. 2006, 11, 176–183. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Harris, J.M. Response of root branching to abscisic acid is correlated with nodule formation both in legumes and non-legumes. Am. J. Bot. 2005, 92, 1675–1683. [Google Scholar] [CrossRef] [PubMed]
- Biswas, B.; Chan, P.K.; Gresshoff, P.M. A novel ABA insensitive mutant of Lotus japonicus with a wilty phenotype displays unaltered nodulation regulation. Mol. Plant 2009, 2, 487–499. [Google Scholar] [CrossRef] [PubMed]
- Tominaga, A.; Nagata, M.; Futsuki, K.; Abe, H.; Uchiumi, T.; Abe, M.; Kucho, K.; Hashiguchi, M.; Akashi, R.; Hirsch, A.M.; et al. Enhanced nodulation and nitrogen fixation in the abscisic acid low-sensitive mutant enhanced nitrogen fixation1 of Lotus japonicus. Plant Physiol. 2009, 151, 1965–1976. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.M.; Clairmont, L.; Macdonald, E.S.; Weiner, C.A.; Emery, R.J.; Guinel, F.C. E151 (sym15), a pleiotropic mutant of pea (Pisum sativum L.), displays low nodule number, enhanced mycorrhizae, delayed lateral root emergence, and high root cytokinin levels. J. Exp. Bot. 2015. [Google Scholar] [CrossRef]
- Guo, D.; Liang, J.; Qiao, Y.; Yan, Y.; Li, L.; Dai, Y. Involvement of G1-to-S transition and AhAUX-dependent auxin transport in abscisic acid-induced inhibition of lateral root primodia initiation in Arachis hypogaea L. J. Plant Physiol. 2012, 169, 1102–1111. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.W.; Yang, Y.W.; Lur, H.S.; Tsai, Y.G.; Chang, M.C. A novel function of abscisic acid in the regulation of rice (Oryza sativa L.) root growth and development. Plant Cell Physiol. 2006, 47, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Coudert, Y.; Perin, C.; Courtois, B.; Khong, N.G.; Gantet, P. Genetic control of root development in rice, the model cereal. Trends Plant Sci. 2010, 15, 219–226. [Google Scholar] [CrossRef] [PubMed]
- Steffens, B.; Wang, J.; Sauter, M. Interactions between ethylene, gibberellin and abscisic acid regulate emergence and growth rate of adventitious roots in deepwater rice. Planta 2006, 223, 604–612. [Google Scholar] [CrossRef] [PubMed]
- Hochholdinger, F.; Woll, K.; Sauer, M.; Dembinsky, D. Genetic dissection of root formation in maize (Zea mays) reveals root-type specific developmental programmes. Ann. Bot. 2004, 93, 359–368. [Google Scholar] [CrossRef] [PubMed]
- Puertolas, J.; Conesa, M.R.; Ballester, C.; Dodd, I.C. Local root abscisic acid (ABA) accumulation depends on the spatial distribution of soil moisture in potato: Implications for ABA signalling under heterogeneous soil drying. J. Exp. Bot. 2015, 66, 2325–2334. [Google Scholar] [CrossRef] [PubMed]
- Dodd, I.C.; Egea, G.; Davies, W.J. Abscisic acid signalling when soil moisture is heterogeneous: Decreased photoperiod sap flow from drying roots limits abscisic acid export to the shoots. Plant Cell Environ. 2008, 31, 1263–1274. [Google Scholar] [CrossRef] [PubMed]
- Dodd, I.C.; Egea, G.; Watts, C.W.; Whalley, W.R. Root water potential integrates discrete soil physical properties to influence ABA signalling during partial rootzone drying. J. Exp. Bot. 2010, 61, 3543–3551. [Google Scholar] [CrossRef] [PubMed]
- Spollen, W.G.; LeNoble, M.E.; Samuels, T.D.; Bernstein, N.; Sharp, R.E. Abscisic acid accumulation maintains maize primary root elongation at low water potentials by restricting ethylene production. Plant Physiol. 2000, 122, 967–976. [Google Scholar] [CrossRef] [PubMed]
- Volkmar, K.M. Water stressed nodal roots of wheat: Effects on leaf growth. Aust. J. Plant Physiol. 1997, 24, 49–56. [Google Scholar] [CrossRef]
- Tuteja, N. Abscisic acid and abiotic stress signaling. Plant Signal Behav 2007, 2, 135–138. [Google Scholar] [CrossRef] [PubMed]
- Murashige, T.; Skoog, F. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiologia Plantarum 1962, 15, 473–479. [Google Scholar] [CrossRef]
- Truchet, G.; Debelle, F.; Vasse, J.; Terzaghi, B.; Garnerone, A.M.; Rosenberg, C.; Batut, J.; Maillet, F.; Denarie, J. Identification of a Rhizobium meliloti pSym2011 region controlling the host specificity of root hair curling and nodulation. J. Bacteriol. 1985, 164, 1200–1210. [Google Scholar] [PubMed]
- Zhu, J.K.; Liu, J.; Xiong, L. Genetic analysis of salt tolerance in Arabidopsis. Evidence for a critical role of potassium nutrition. Plant Cell 1998, 10, 1181–1191. [Google Scholar] [CrossRef] [PubMed]
- Dubrovsky, J.G.; Soukup, A.; Napsucialy-Mendivil, S.; Jeknic, Z.; Ivanchenko, M.G. The lateral root initiation index: An integrative measure of primordium formation. Ann. Bot. 2009, 103, 807–817. [Google Scholar] [CrossRef] [PubMed]
- Cassab, G.I.; Eapen, D.; Campos, M.E. Root hydrotropism: An update. Am. J. Bot. 2013, 100, 14–24. [Google Scholar] [CrossRef] [PubMed]
- Macgregor, D.R.; Deak, K.I.; Ingram, P.A.; Malamy, J.E. Root system architecture in Arabidopsis grown in culture is regulated by sucrose uptake in the aerial tissues. Plant Cell 2008, 20, 2643–2660. [Google Scholar] [CrossRef] [PubMed]
- Galvan-Ampudia, C.S.; Julkowska, M.M.; Darwish, E.; Gandullo, J.; Korver, R.A.; Brunoud, G.; Haring, M.A.; Munnik, T.; Vernoux, T.; Testerink, C. Halotropism is a response of plant roots to avoid a saline environment. Curr. Biol. 2013, 23, 2044–2050. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, N.; Goto, N.; Okada, K.; Takahashi, H. Hydrotropism in abscisic acid, wavy, and gravitropic mutants of Arabidopsis thaliana. Planta 2002, 216, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Eapen, D.; Barroso, M.L.; Campos, M.E.; Ponce, G.; Corkidi, G.; Dubrovsky, J.G.; Cassab, G.I. A no hydrotropic response root mutant that responds positively to gravitropism in Arabidopsis. Plant Physiol. 2003, 131, 536–546. [Google Scholar] [CrossRef] [PubMed]
- Antoni, R.; Gonzalez-Guzman, M.; Rodriguez, L.; Peirats-Llobet, M.; Pizzio, G.A.; Fernandez, M.A.; de Winne, N.; de Jaeger, G.; Dietrich, D.; Bennett, M.J.; et al. PYRABACTIN RESISTANCE1-LIKE8 plays an important role for the regulation of abscisic acid signaling in root. Plant Physiol. 2013, 161, 931–941. [Google Scholar] [CrossRef] [PubMed]
- Krouk, G.; Lacombe, B.; Bielach, A.; Perrine-Walker, F.; Malinska, K.; Mounier, E.; Hoyerova, K.; Tillard, P.; Leon, S.; Ljung, K.; et al. Nitrate-regulated auxin transport by NRT1.1 defines a mechanism for nutrient sensing in plants. Dev. Cell 2010, 18, 927–937. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.Q.; Wang, R.; Chen, M.; Crawford, N.M. The Arabidopsis dual-affinity nitrate transporter gene AtNRT1.1 (CHL1) is activated and functions in nascent organ development during vegetative and reproductive growth. Plant Cell 2001, 13, 1761–1777. [Google Scholar]
- Leran, S.; Edel, K.H.; Pervent, M.; Hashimoto, K.; Corratge-Faillie, C.; Offenborn, J.N.; Tillard, P.; Gojon, A.; Kudla, J.; Lacombe, B. Nitrate sensing and uptake in Arabidopsis are enhanced by ABI2, a phosphatase inactivated by the stress hormone abscisic acid. Sci. Signal 2015, 8, ra43. [Google Scholar]
- Park, S.Y.; Fung, P.; Nishimura, N.; Jensen, D.R.; Fujii, H.; Zhao, Y.; Lumba, S.; Santiago, J.; Rodrigues, A.; Chow, T.F.; et al. Abscisic acid inhibits type 2c protein phosphatases via the PYR/PYL family of start proteins. Science 2009, 324, 1068–1071. [Google Scholar] [PubMed]
- Ma, Y.; Szostkiewicz, I.; Korte, A.; Moes, D.; Yang, Y.; Christmann, A.; Grill, E. Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 2009, 324, 1064–1068. [Google Scholar] [CrossRef] [PubMed]
- Kanno, Y.; Hanada, A.; Chiba, Y.; Ichikawa, T.; Nakazawa, M.; Matsui, M.; Koshiba, T.; Kamiya, Y.; Seo, M. Identification of an abscisic acid transporter by functional screening using the receptor complex as a sensor. Proc. Natl. Acad. Sci. USA 2012, 109, 9653–9658. [Google Scholar] [CrossRef] [PubMed]
- Huang, N.C.; Liu, K.H.; Lo, H.J.; Tsay, Y.F. Cloning and functional characterization of an Arabidopsis nitrate transporter gene that encodes a constitutive component of low-affinity uptake. Plant Cell 1999, 11, 1381–1392. [Google Scholar] [CrossRef] [PubMed]
- Kanno, Y.; Kamiya, Y.; Seo, M. Nitrate does not compete with abscisic acid as a substrate of AtNPF4.6/NRT1.2/AIT1 in Arabidopsis. Plant Signal Behav. 2013, 8, e26624. [Google Scholar] [CrossRef] [PubMed]
- Takatsuka, H.; Umeda, M. Hormonal control of cell division and elongation along differentiation trajectories in roots. J. Exp. Bot. 2014, 65, 2633–2643. [Google Scholar] [CrossRef] [PubMed]
- Ghassemian, M.; Nambara, E.; Cutler, S.; Kawaide, H.; Kamiya, Y.; McCourt, P. Regulation of abscisic acid signaling by the ethylene response pathway in Arabidopsis. Plant Cell 2000, 12, 1117–1126. [Google Scholar] [CrossRef] [PubMed]
- Beaudoin, N.; Serizet, C.; Gosti, F.; Giraudat, J. Interactions between abscisic acid and ethylene signaling cascades. Plant Cell 2000, 12, 1103–1116. [Google Scholar] [CrossRef] [PubMed]
- Thole, J.M.; Beisner, E.R.; Liu, J.; Venkova, S.V.; Strader, L.C. Abscisic acid regulates root elongation through the activities of auxin and ethylene in Arabidopsis thaliana. G3 (Bethesda) 2014, 4, 1259–1274. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Renou, J.P.; Berthome, R.; Harberd, N.P.; Genschik, P. Plant DELLAs restrain growth and promote survival of adversity by reducing the levels of reactive oxygen species. Curr. Biol. 2008, 18, 656–660. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Cheng, H.; de Grauwe, L.; Decat, J.; Schoutteten, H.; Moritz, T.; van der Straeten, D.; Peng, J.; Harberd, N.P. Integration of plant responses to environmentally activated phytohormonal signals. Science 2006, 311, 91–94. [Google Scholar] [CrossRef] [PubMed]
- Nibau, C.; Tao, L.; Levasseur, K.; Wu, H.M.; Cheung, A.Y. The Arabidopsis small GTPase AtRAC7/ROP9 is a modulator of auxin and abscisic acid signalling. J. Exp. Bot. 2013, 64, 3425–3437. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Lee, Y.; Hwang, J.-U. Arabidopsis ROP9 and ROP10 GTPases differentially regulate auxin and ABA responses. J. Plant Biol. 2014, 57, 245–254. [Google Scholar] [CrossRef]
- Yuan, T.T.; Xu, H.H.; Zhang, K.X.; Guo, T.T.; Lu, Y.T. Glucose inhibits root meristem growth via ABA INSENSITIVE 5, which represses PIN1 accumulation and auxin activity in Arabidopsis. Plant Cell Environ. 2014, 37, 1338–1350. [Google Scholar] [CrossRef] [PubMed]
- Shkolnik-Inbar, D.; Bar-Zvi, D. ABI4 mediates abscisic acid and cytokinin inhibition of lateral root formation by reducing polar auxin transport in Arabidopsis. Plant Cell 2010, 22, 3560–3573. [Google Scholar] [CrossRef] [PubMed]
- Belin, C.; Megies, C.; Hauserova, E.; Lopez-Molina, L. Abscisic acid represses growth of the Arabidopsis embryonic axis after germination by enhancing auxin signaling. Plant Cell 2009, 21, 2253–2268. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.H.; Bao, M.L.; Sun, Y.Z.; Yang, Y.J.; Xu, X.H.; Wang, J.H.; Han, N.; Bian, H.W.; Zhu, M.Y. Regulation of auxin response by miR393-targeted TRANSPORT INHIBITOR RESPONSE PROTEIN 1 is involved in normal development in Arabidopsis. Plant Mol. Biol. 2011, 77, 619–629. [Google Scholar] [CrossRef] [PubMed]
- Brady, S.M.; Sarkar, S.F.; Bonetta, D.; McCourt, P. The ABSCISIC ACID INSENSITIVE 3 (ABI3) gene is modulated by farnesylation and is involved in auxin signaling and lateral root development in Arabidopsis. Plant J. 2003, 34, 67–75. [Google Scholar] [CrossRef] [PubMed]
- Ulmasov, T.; Murfett, J.; Hagen, G.; Guilfoyle, T.J. AUX/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 1997, 9, 1963–1971. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Yang, Y.N.; Xue, L.J.; Zou, M.J.; Liu, J.Y.; Chen, F.; Xue, H.W. Rice ABI5-LIKE1 regulates abscisic acid and auxin responses by affecting the expression of ABRE-containing genes. Plant Physiol. 2011, 156, 1397–1409. [Google Scholar] [CrossRef] [PubMed]
- Okushima, Y.; Fukaki, H.; Onoda, M.; Theologis, A.; Tasaka, M. ARF7 and ARF19 regulate lateral root formation via direct activation of LBD/ASL genes in Arabidopsis. Plant Cell 2007, 19, 118–130. [Google Scholar] [CrossRef] [PubMed]
- Hauser, F.; Waadt, R.; Schroeder, J.I. Evolution of abscisic acid synthesis and signaling mechanisms. Curr. Biol. 2011, 21, R346–355. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Liu, Y.; Li, S.S.; Han, G.Z. Insights into the origin and evolution of the plant hormone signaling machinery. Plant Physiol. 2015, 167, 872–886. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Chen, Z.; Gao, J.; Gong, Z. Abscisic acid inhibits root growth in Arabidopsis through ethylene biosynthesis. Plant J. 2014, 79, 44–55. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.; Yin, C.C.; He, S.J.; Lu, X.; Zhang, W.K.; Lu, T.G.; Chen, S.Y.; Zhang, J.S. Ethylene-induced inhibition of root growth requires abscisic acid function in rice (Oryza sativa L.) seedlings. PLoS Genet. 2014, 10, e1004701. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Guo, M.; Ye, N.; Liu, Y.; Liu, R.; Xia, Y.; Cui, S.; Zhang, J. Reduced ABA accumulation in the root system is caused by ABA exudation in upland rice (Oryza sativa L. var. Gaoshan1) and this enhanced drought adaptation. Plant Cell Physiol. 2015, 56, 951–964. [Google Scholar] [CrossRef] [PubMed]
- Hartung, W.; Sauter, A.; Hose, E. Abscisic acid in the xylem: Where does it come from, where does it go to? J. Exp. Bot. 2002, 53, 27–32. [Google Scholar] [CrossRef] [PubMed]
- Belimov, A.A.; Dodd, I.C.; Safronova, V.I.; Dumova, V.A.; Shaposhnikov, A.I.; Ladatko, A.G.; Davies, W.J. Abscisic acid metabolizing rhizobacteria decrease ABA concentrations in planta and alter plant growth. Plant Physiol. Biochem. 2014, 74, 84–91. [Google Scholar] [CrossRef] [PubMed]
- Hartung, W.; Sauter, A.; Turner, N.C.; Fillery, I.; Heilmeier, H. Abscisic acid in soils: What is its function and which factors and mechanisms influence its concentration? Plant Soil 1996, 184, 105–110. [Google Scholar] [CrossRef]
- Boiero, L.; Perrig, D.; Masciarelli, O.; Penna, C.; Cassan, F.; Luna, V. Phytohormone production by three strains of Bradyrhizobium japonicum and possible physiological and technological implications. Appl. Microbiol. Biotechnol. 2007, 74, 874–880. [Google Scholar] [CrossRef] [PubMed]
- Forchetti, G.; Masciarelli, O.; Alemano, S.; Alvarez, D.; Abdala, G. Endophytic bacteria in sunflower (Helianthus annuus L.): Isolation, characterization, and production of jasmonates and abscisic acid in culture medium. Appl. Microbiol. Biotechnol. 2007, 76, 1145–1152. [Google Scholar] [CrossRef] [PubMed]
- Dangar, T.K.; Basu, P.S. Abscisic acid production in culture by some Rhizobium spp. of leguminous trees and pulses. Folia Microbiol. 1991, 36, 527–532. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Harris, J.M. Abscisic Acid: Hidden Architect of Root System Structure. Plants 2015, 4, 548-572. https://doi.org/10.3390/plants4030548
Harris JM. Abscisic Acid: Hidden Architect of Root System Structure. Plants. 2015; 4(3):548-572. https://doi.org/10.3390/plants4030548
Chicago/Turabian StyleHarris, Jeanne M. 2015. "Abscisic Acid: Hidden Architect of Root System Structure" Plants 4, no. 3: 548-572. https://doi.org/10.3390/plants4030548
APA StyleHarris, J. M. (2015). Abscisic Acid: Hidden Architect of Root System Structure. Plants, 4(3), 548-572. https://doi.org/10.3390/plants4030548




