Identification of Daphnane Diterpenoids from Flower Buds and Blooming Flowers of Daphne odora Using UHPLC-Q-Exactive-Orbitrap MS
Abstract
1. Introduction
2. Results and Discussion
2.1. Detection of Daphnane Diterpenoids in the Flower Buds and Flowers of D. odora Using LC-MS/MS
| No. | tR, min (+/−) | Precursor Ion, m/z (Error, ppm) | Molecular Formula | Tentative Identification | Reference | |
|---|---|---|---|---|---|---|
| [M + H]+ a | [M–HCOO]− b | |||||
| 1 | 4.21/4.19 | 487.2325 (−0.33) | 485.2189 (1.74) | C27H34O8 | 3-deoxo-1,2-dihydro-3-hydroxydaphnetoxin | [23,24] |
| 2 | 6.09/6.08 | 543.2587 (−0.24) | 587.2507 (1.51) | C30H38O9 | peddiea factor A1 | [25] |
| 3 | 6.93/6.95 | 545.2746 (0.16) | 589.2662 (1.25) | C30H40O9 | 12-hydroxyexcoecariatoxin | [26] |
| 4 | 8.10/8.08 | 585.2686 (−1.38) | 629.2609 (0.91) | C32H40O10 | peddiea factor V2 | [25] |
| 5 c | 8.10/8.11 | 1028.4264 (−1.02) | 1055.3934 (1.53) | C55H62O18 | daphneodrin B | [10] |
| 6 e | 8.13/8.14 | 705.2899 (−0.90) | 703.2770 (1.39) | C39H44O12 | 12-O-(E)-caffeoyl-9,13,14-ortho-(2E,4E,6E)-decatrienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | - |
| 7 e | 8.68/8.70 | 689.2951 (−0.77) | 687.2814 (0.42) | C39H44O11 | 12-O-(Z)-coumaroyl-9,13,14-ortho-(2E,4E,6E)-decatrienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | - |
| 8 e | 8.88/8.87 | 719.3056 (−0.83) | 717.2920 (0.48) | C40H46O12 | 12-O-(Z)-feruloyl-9,13,14-ortho-(2E,4E,6E)-decatrienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | - |
| 9 c | 8.88/8.89 | 970.4219 (−0.09) | 997.3874 (1.05) | C53H60O16 | daphneodorin A | [10] |
| 10 e | 8.93/8.95 | 689.2952 (−0.60) | 687.2819 (1.13) | C39H44O11 | 12-O-(E)-coumaroyl-9,13,14-ortho-(2E,4E,6Z)-decatrienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | - |
| 11 | 8.97/8.95 | 587.2850 (−0.17) | 631.2769 (1.36) | C32H42O10 | yuanhuadin | [27] |
| 12 e | 8.97/8.95 | 707.3051 (−1.63) | 705.2924 (1.09) | C39H46O12 | 12-O-(E)-caffeoyl-9,13,14-ortho-(2E,4E)-decadienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | - |
| 13 c | 9.10/9.12 | 689.2953 (−0.42) | 687.2814 (0.42) | C39H44O11 | daphneodorin E | [11] |
| 14 e | 9.18/9.17 | 719.3061 (−0.16) | 763.2982 (1.35) | C40H46O12 | 12-O-(E)-feruloyl-9,13,14-ortho-(2E,4E,6Z)-decatrienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | - |
| 15 c | 9.35/9.34 | 719.3051 (−1.60) | 763.2970 (−0.17) | C40H46O12 | actilobin C | [28] |
| 16 e,f | 9.47/9.49 | 691.3111 (−0.28) | 689.2972 (0.65) | C39H46O11 | 12-O-(Z)-coumaroyl-9,13,14-ortho-(2E,4E)-decadienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | - |
| 17 e | 9.67/9.68 | 721.3220 (0.15) | 719.3081 (1.12) | C40H48O12 | 12-O-(Z)-feruloyl-9,13,14-ortho-(2E,4E)-decadienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | - |
| 18 c | 9.91/9.92 | 691.3109 (−0.54) | 689.2968 (0.11) | C39H46O11 | daphneodorin D | [11] |
| 19 d | 10.05/10.03 | 649.3000 (−1.04) | 693.2921 (0.58) | C37H44O10 | 1,2-dihydroyuanhuajine | [24] |
| 20 c | 10.15/10.16 | 721.3212 (−0.95) | 765.3127 (−0.05) | C40H48O12 | actilobin D | [28] |
| 21 c | 10.15/10.16 | 775.3693 (0.67) | 819.3587 (−1.25) | C44H54O12 | gnidimacrin | [29] |
| 22 c | 10.36/10.35 | 647.2845 (−0.91) | 691.2759 (−0.09) | C37H42O10 | yuanhuajine | [7] |
| 23 e | 10.93/10.91 | 673.3004 (−0.46) | 717.2928 (1.58) | C39H44O10 | 12-O-(E)-cinnamoyl-9,13,14-ortho-(2E,4E,6Z)-decatrienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | - |
| 24 c | 11.07/11.06 | 673.2999 (−1.27) | 717.2918 (0.22) | C39H44O10 | 12-O-(E)-cinnamoyl-9,13,14-ortho-(2E,4E,6E)-decatrienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | [7] |
| 25 c | 11.18/11.16 | 649.2998 (−1.42) | 693.2917 (0.14) | C37H44O10 | yuanhuacine/odoracin | [30] |
| 26 c | 11.81/11.82 | 675.3157 (−0.95) | 719.3074 (0.10) | C39H46O10 | 12-O-(E)-cinnamoyl-9,13,14-ortho-(2E,4E)-decadienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | [7] |
| 27 | 12.01/12.00 | 635.3215 (0.02) | 679.3135 (1.70) | C37H46O9 | actilobin F | [29] |
| 28 e | 12.74/12.73 | 691.3474 (−0.38) | 735.3394 (1.03) | C40H50O10 | 12-O-(2E,4E,6E)-decatrienoyl-9,13,14-ortho-(2E,4E,6E)-decatrienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | - |
| 29 | 12.77/12.76 | 637.3369 (−0.31) | 681.3286 (0.76) | C37H48O9 | wikstroemia factor M1 | [27] |
| 30 d | 13.39/13.38 | 693.3630 (−0.50) | 737.3547 (0.66) | C40H52O10 | 12-O-(2E,4E,6E)-decatrienoyl-9,13,14-ortho-(2E,4E)-decadienylidyne-5β,12β-dihydroxyresiniferonol-6α,7α-oxide | [24] |
2.2. Identification of Previously Unreported Daphnane Diterpenoids by MS/MS Fragmentation Analysis
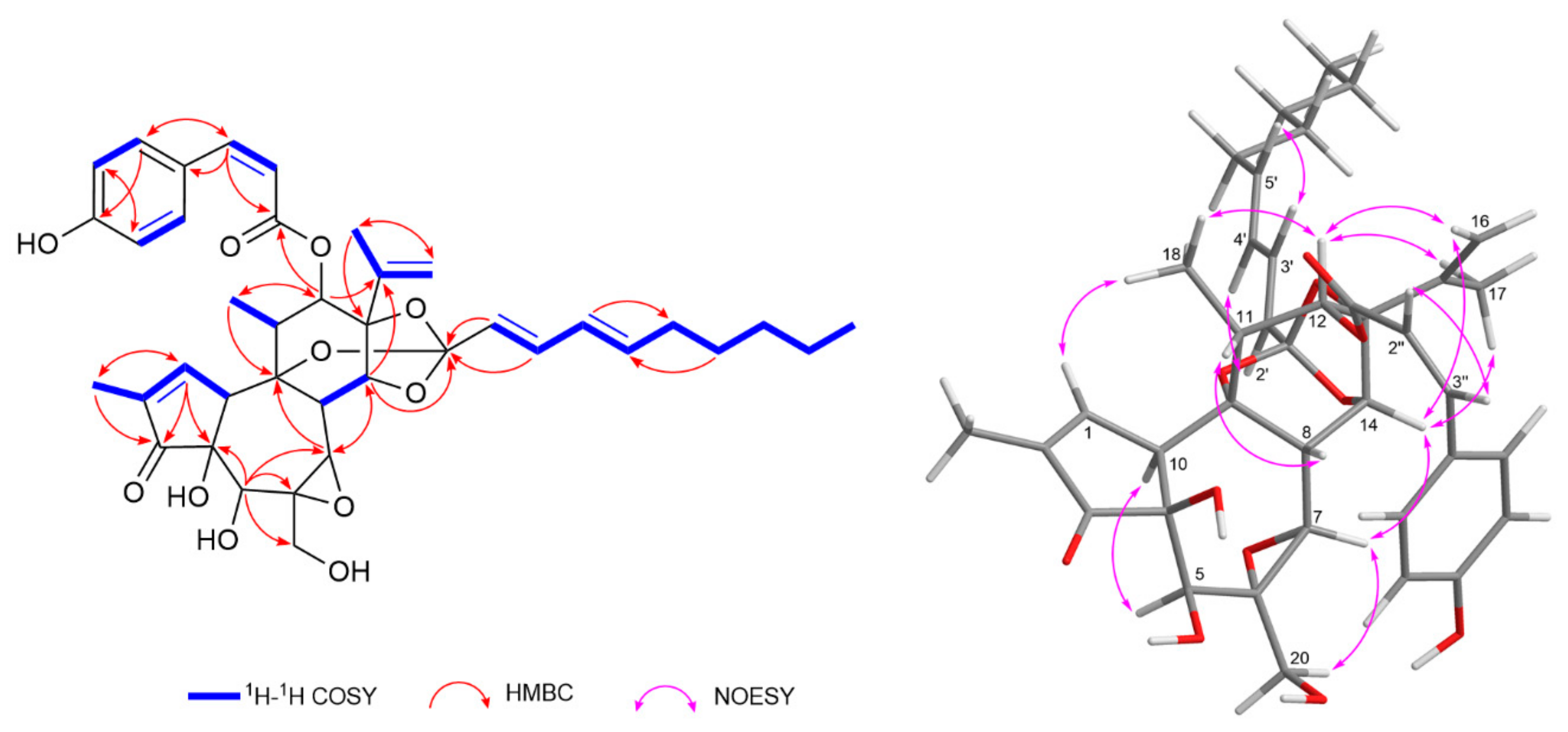
2.3. LC-MS Guided Isolation and Structural Elucidation of Compound 16
3. Materials and Methods
3.1. General Experimental Producers
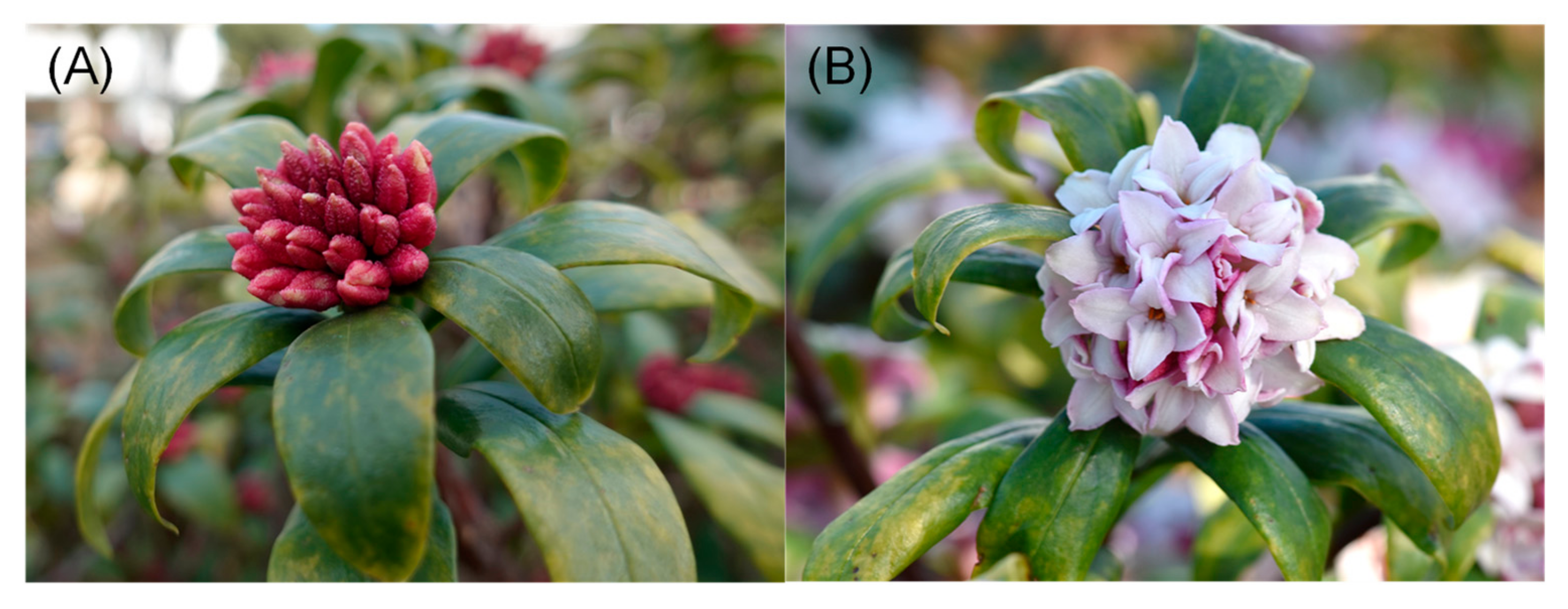
3.2. Plant Materials
3.3. Extraction and Isolation
Daphneodorin I (16)
3.4. Qualitative Analysis of Flower Buds and Flower of D. odora
3.4.1. Sample Preparation
3.4.2. LC-MS/MS Conditions
3.4.3. Data Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wu, Z.; Peter, H.R.; Hong, D. Flora of China; Science Press: Beijing, China; Missouri Botanical Garden: St. Louis, MO, USA, 1994; Volume 13, p. 238. [Google Scholar]
- Shirai, K. Tyuyakudaiziten; Shanghai Scientific & Technical Publishers: Shanghai, China; Shougakukan, Inc.: Tokyo, Japan, 1985; Volume 2. [Google Scholar]
- Kogiso, S.; Hosozawa, S.; Wada, K.; Munakata, K. Daphneolone in roots of Daphne odora. Phytochemistry 1974, 13, 2332–2334. [Google Scholar] [CrossRef]
- Satô, M.; Hasegawa, M. Conversion of daphnin to daphnetin-8-glucoside in Daphne odora. Phytochemistry 1969, 8, 1211–1214. [Google Scholar] [CrossRef]
- Baba, K.; Takeuchi, K.; Hamasaki, F.; Kozawa, M. Chemical studies on the constituents of the Thymelaeaceous plants. I.: Structures of two new flavans from Daphne odora Thunb. Chem. Pharm. Bull. 1986, 34, 595–602. [Google Scholar] [CrossRef]
- Baba, K.; Yoshikawa, M.; Taniguchi, M.; Kozawa, M. Biflavonoids from Daphne odora. Phytochemistry 1995, 38, 1021–1026. [Google Scholar] [CrossRef]
- Okunishi, T.; Umezawa, T.; Shimada, M. Isolation and enzymatic formation of lignans of Daphne genkwa and Daphne odora. J. Wood Sci. 2001, 47, 383–388. [Google Scholar] [CrossRef]
- Ohigashi, H.; Hirota, M.; Ohtsuka, T.; Koshimizu, K.; Fujiki, H.; Suganuma, M.; Tamaizumi, Z.; Sugimura, T. Resiniferonol-related diterpene esters from Daphne odora Thunb. and their ornithine decarboxylase-inducing activity in mouse skin. Agric. Biol. Chem. 1982, 46, 2605–2608. [Google Scholar] [CrossRef]
- Shigemori, H.; Nakasone, R.; Kurisu, M.; Onodera, M.; Miyamae, Y.; Matsuura, D.; Kanatani, H.; Yano, S. Promoting effects on hepatocyte growth factor production of daphnane diterpenoids from Daphne odora. Heterocycles 2013, 87, 1087. [Google Scholar] [CrossRef]
- Otsuki, K.; Li, W.; Asada, Y.; Chen, C.H.; Lee, K.H.; Koike, K. Daphneodorins A–C, anti-HIV gnidimacrin related macrocyclic daphnane orthoesters from Daphne odora. Org. Lett. 2020, 22, 11–15. [Google Scholar] [CrossRef]
- Otsuki, K.; Li, W.; Miura, K.; Asada, Y.; Huang, L.; Chen, C.H.; Lee, K.H.; Koike, K. Isolation, structural elucidation, and anti-HIV activity of daphnane diterpenoids from Daphne odora. J. Nat. Prod. 2020, 83, 3270–3277. [Google Scholar] [CrossRef]
- Otsuki, K.; Li, W. Tigliane and daphnane diterpenoids from Thymelaeaceae family: Chemistry, biological activity, and potential in drug discovery. J. Nat. Med. 2023, 77, 625–643. [Google Scholar] [CrossRef]
- Adolf, W.; Sorg, B.; Hergenhahn, M.; Hecker, E. Structure-activity relations of polyfunctional diterpenes of the daphnane type I. Revised structure for resiniferatoxin and structure-activity relations of resiniferonol and some of its esters. J. Nat. Prod. 1982, 45, 347–354. [Google Scholar] [CrossRef]
- Hou, Z.L.; Yao, G.D.; Song, S.J. Daphnane-type diterpenes from genus Daphne and their anti-tumor activity. Chin. Herb. Med. 2021, 13, 145–156. [Google Scholar] [CrossRef] [PubMed]
- El-Desoky, A.H.H.; Eguchi, K.; Kishimoto, N.; Asano, T.; Kato, H.; Hitora, Y.; Kotani, S.; Nakamura, T.; Tsuchiya, S.; Kawahara, T.; et al. Isolation, synthesis, and structure-activity relationship study on daphnane and tigliane diterpenes as HIV latency-reversing agents. J. Med. Chem. 2022, 65, 3460–3472. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, L.; Pereira, J.A.; Saraiva, J.A.; Ramalhosa, E.; Casal, S. Phytochemical characterization of Borago officinalis L. and Centaurea cyanus L. during flower development. Food Res. Int. 2019, 123, 771–778. [Google Scholar] [CrossRef] [PubMed]
- Önder, S.; Tonguç, M.; Erbaş, S.; Önder, D.; Mutlucan, M. Investigation of phenological, primary and secondary metabolites changes during flower developmental of Rosa damascena. Plant Physiol. Biochem. 2022, 192, 20–34. [Google Scholar] [CrossRef]
- Zhao, H.D.; Lu, Y.; Yan, M.; Chen, C.H.; Morris-Natschke, S.L.; Lee, K.H.; Chen, D.F. Rapid recognition and targeted isolation of anti-HIV daphnane diterpenes from Daphne genkwa guided by UPLC-MSn. J. Nat. Prod. 2020, 83, 134–141. [Google Scholar] [CrossRef]
- Pan, R.R.; Zhang, C.Y.; Li, Y.; Zhang, B.B.; Zhao, L.; Ye, Y.; Song, Y.N.; Zhang, M.; Tie, H.Y.; Zhang, H.; et al. Daphnane diterpenoids from Daphne genkwa inhibit PI3K/Akt/mTOR signaling and induce cell cycle arrest and apoptosis in human colon cancer cells. J. Nat. Prod. 2020, 83, 1238–1248. [Google Scholar] [CrossRef]
- Pluskal, T.; Castillo, S.; Villar-Briones, A.; Oresic, M. MZmine 2: Modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinform. 2010, 11, 395. [Google Scholar] [CrossRef]
- Zhang, M.; Otsuki, K.; Tan, L.J.; Kikuchi, T.; Li, N.; Li, W. Automated annotation of complex natural products using a Modular Fragmentation-based Structure Assembly (MFSA) strategy. Sci. Adv. 2025, 11, eadw4693. [Google Scholar] [CrossRef]
- Otsuki, K.; Zhang, M.; Tan, L.; Komaki, M.; Shimada, A.; Kikuchi, T.; Zhou, D.; Li, N.; Li, W. Isomer differentiation by UHPLC-Q-Exactive-Orbitrap MS led to enhanced identification of daphnane diterpenoids in Daphne tangutica. Phytochem. Anal. 2025, 36, 1053–1062. [Google Scholar] [CrossRef]
- Wu, D.; Sorg, B.; Adolf, W.; Seip, E.H.; Hecker, E. Oligo- and macrocyclic diterpenes in Thymelaeaceae and Euphorbiaceae occurring and utilized in Yunnan (Southwest China) 3. Two new daphnane type 9,13,14-orthoesters from Wikstroemia mekongenia. Phytother. Res. 1993, 7, 72–75. [Google Scholar] [CrossRef]
- Li, L.Z.; Gao, P.Y.; Wang, L.H.; Song, S.J. A novel daphnane-type diterpene from the flower bud of Daphne genkwa. Chem. Nat. Compd. 2010, 46, 380–382. [Google Scholar] [CrossRef]
- Adolf, W.; Dossaji, S.F.; Seip, E.H.; Hecker, E. Skin irritant diterpene orthoesters of the daphnane type from Peddiea africana and P. volkensii. Phytochemistry 1985, 24, 2047–2049. [Google Scholar] [CrossRef]
- Zhang, S.; Li, X.; Zhang, F.; Yang, P.; Gao, X.; Song, Q. Preparation of yuanhuacine and relative daphne diterpene esters from Daphne genkwa and structure-activity relationship of potent inhibitory activity against DNA topoisomerase I. Bioorg. Med. Chem. 2006, 14, 3888–3895. [Google Scholar] [CrossRef]
- Adolf, W.; Seip, E.H.; Hecker, E.; Dossaji, S.F. Irritant principles of the mezereon family (Thymelaeaceae), V. New skin irritants and tumor promoters of the daphnane and 1 alpha-alkyldaphnane type from Synaptolepis kirkii and Synaptolepis retusa. J. Nat. Prod. 1988, 51, 662–674. [Google Scholar] [CrossRef]
- Huang, S.Z.; Zhang, X.J.; Li, X.Y.; Kong, L.M.; Jiang, H.Z.; Ma, Q.Y.; Liu, Y.Q.; Hu, J.M.; Zheng, Y.T.; Li, Y.; et al. Daphnane-type diterpene esters with cytotoxic and anti-HIV-1 activities from Daphne acutiloba Rehd. Phytochemistry 2012, 75, 99–107. [Google Scholar] [CrossRef]
- Kupchan, S.M.; Shizuri, Y.; Murae, T.; Swenny, J.G.; Haynes, H.R.; Shen, M.S.; Barrick, J.C.; Bryan, A.F.; vander Helm, D.; Wu, K.K. Letter: Gnidimacrin and gnidimacrin 20-palmitate, novel macrocyclic antileukemic diterpenoid esters from Gnidia subcordata. J. Am. Chem. Soc. 1976, 98, 5719–5720. [Google Scholar] [CrossRef]
- Kogiso, S.; Wada, K.; Munakata, K. Odoracin, a nematicidal constituent from Daphne odora. Agric. Biol. Chem. 1976, 40, 2119–2120. [Google Scholar] [CrossRef]
- Otsuki, K.; Zhang, M.; Kikuchi, T.; Tsuji, M.; Tejima, M.; Bai, Z.S.; Zhou, D.; Huang, L.; Chen, C.H.; Lee, K.H.; et al. Identification of anti-HIV macrocyclic daphnane orthoesters from Wikstroemia ligustrina by LC-MS analysis and phytochemical investigation. J. Nat. Med. 2021, 75, 1058–1066. [Google Scholar] [CrossRef]
- Liu, F.; Yang, X.; Liang, Y.; Dong, B.; Su, G.; Tuerhong, M.; Jin, D.Q.; Xu, J.; Guo, Y. Daphnane diterpenoids with nitric oxide inhibitory activities and interactions with iNOS from the leaves of Trigonostemon thyrsoideus. Phytochemistry 2018, 147, 57–67. [Google Scholar] [CrossRef]
- Clifford, M.N.; Kirkpatrick, J.; Kuhnert, N.; Roozendaal, H.; Salgado, P.R. LC–MSn analysis of the cis isomers of chlorogenic acids. Food Chem. 2008, 106, 379–385. [Google Scholar] [CrossRef]
- Clifford, M.N.; Jaganath, I.B.; Ludwig, I.A.; Crozier, A. Chlorogenic acids and the acyl-quinic acids: Discovery, biosynthesis, bioavailability and bioactivity. Nat. Prod. Rep. 2017, 34, 1391–1421. [Google Scholar] [CrossRef]
- Bermúdez-Oria, A.; Castejón, M.L.; Rubio-Senent, F.; Fernández-Prior, Á.; Rodríguez-Gutiérrez, G.; Fernández-Bolaños, J. Isolation and structural determination of cis- and trans-p-coumaroyl-secologanoside (comselogoside) from olive oil waste (alperujo). Photoisomerization with ultraviolet irradiation and antioxidant activities. Food Chem. 2024, 432, 137233. [Google Scholar] [CrossRef]



| No. | δH (J in Hz) | C, Type | No. | δH (J in Hz) | C, Type | ||
|---|---|---|---|---|---|---|---|
| 1 | 7.57 (1H, m) | 161.3 | CH | 1′ | 116.6 | C | |
| 2 | 136.6 | C | 2′ | 5.56 (1H, d, 15.4) | 122.3 | CH | |
| 3 | 209.6 | C | 3′ | 6.57 (1H, dd, 15.4, 10.9) | 134.9 | CH | |
| 4 | 72.1 | C | 4′ | 5.99 (1H, dd, 15.2, 10.9) | 128.6 | CH | |
| 5 | 4.27 (1H, s) | 74.1 | CH | 5′ | 5.80 (1H, dt, 15.2, 7.2) | 139.2 | CH |
| 6 | 58.6 | C | 6′ | 2.06 (2H, q, 7.2) | 32.6 | CH2 | |
| 7 | 3.23 (1H, s) | 66.3 | CH | 7′ | 1.35 (2H, quin, 7.2) | 28.7 | CH2 |
| 8 | 2.88 (1H, d, 2.5) | 35.8 | CH | 8′ | 1.25 (2H, m) | 31.3 | CH2 |
| 9 | 77.9 | C | 9′ | 1.28 (2H, m) | 22.5 | CH2 | |
| 10 | 3.73 (1H, t, 2.6) | 47.8 | CH | 10′ | 0.86 (3H, t, 7.0) | 14.0 | CH3 |
| 11 | 2.31 (1H, q, 7.4) | 44.5 | CH | 1″ | 165.9 | C | |
| 12 | 4.91 (1H, s) | 79.1 | CH | 2″ | 5.78 (1H, d, 12.2) | 119.4 | CH |
| 13 | 83.7 | C | 3″ | 7.05 (1H, d, 12.2) | 144.6 | CH | |
| 14 | 4.23(1H, d, 2.5) | 79.9 | CH | 4″ | 127.7 | C | |
| 15 | - | 142.7 | C | 5″ | 7.15 (1H, d, 8.4) | 130.7 | CH |
| 16 | 4.74 (1H, br s) | 113.7 | CH2 | 6″ | 6.89 (1H, d, 8.4) | 115.0 | CH |
| 4.92 (1H, br s) | 7″ | 157.2 | C | ||||
| 17 | 1.69 (3H, br s) | 18.8 | CH3 | 8″ | 6.89 (1H, d, 8.4) | 115.0 | CH |
| 18 | 1.28 (3H, d, 7.2) | 18.4 | CH3 | 9″ | 7.15 (1H, d, 8.4) | 130.7 | CH |
| 19 | 1.78 (3H, m) | 9.8 | CH3 | ||||
| 20 | 3.36 (1H, d, 12.6) | 68.2 | CH2 | ||||
| 4.40 (1H, d, 12.6) | |||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Otsuki, K.; Miyamoto, K.; Goto, M.; Zhang, M.; Kikuchi, T.; Li, W. Identification of Daphnane Diterpenoids from Flower Buds and Blooming Flowers of Daphne odora Using UHPLC-Q-Exactive-Orbitrap MS. Plants 2025, 14, 2616. https://doi.org/10.3390/plants14172616
Otsuki K, Miyamoto K, Goto M, Zhang M, Kikuchi T, Li W. Identification of Daphnane Diterpenoids from Flower Buds and Blooming Flowers of Daphne odora Using UHPLC-Q-Exactive-Orbitrap MS. Plants. 2025; 14(17):2616. https://doi.org/10.3390/plants14172616
Chicago/Turabian StyleOtsuki, Kouharu, Kousei Miyamoto, Mami Goto, Mi Zhang, Takashi Kikuchi, and Wei Li. 2025. "Identification of Daphnane Diterpenoids from Flower Buds and Blooming Flowers of Daphne odora Using UHPLC-Q-Exactive-Orbitrap MS" Plants 14, no. 17: 2616. https://doi.org/10.3390/plants14172616
APA StyleOtsuki, K., Miyamoto, K., Goto, M., Zhang, M., Kikuchi, T., & Li, W. (2025). Identification of Daphnane Diterpenoids from Flower Buds and Blooming Flowers of Daphne odora Using UHPLC-Q-Exactive-Orbitrap MS. Plants, 14(17), 2616. https://doi.org/10.3390/plants14172616








