RdDM-Associated Chromatin Remodelers in Soybean: Evolution and Stress-Induced Expression of CLASSY Genes
Abstract
1. Introduction
2. Results and Discussions
2.1. Construction of Specific Profile HMMs of CLSY Proteins
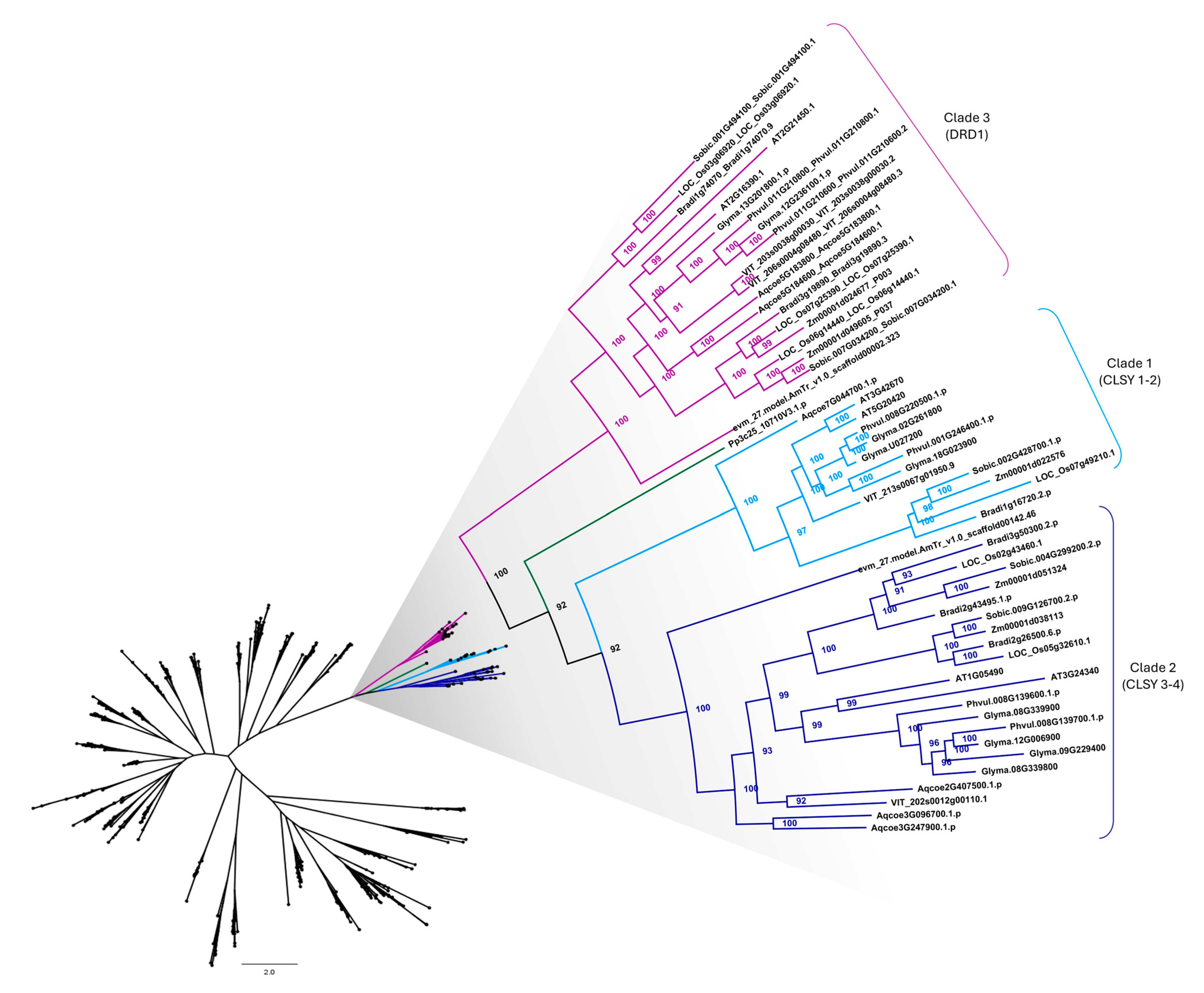
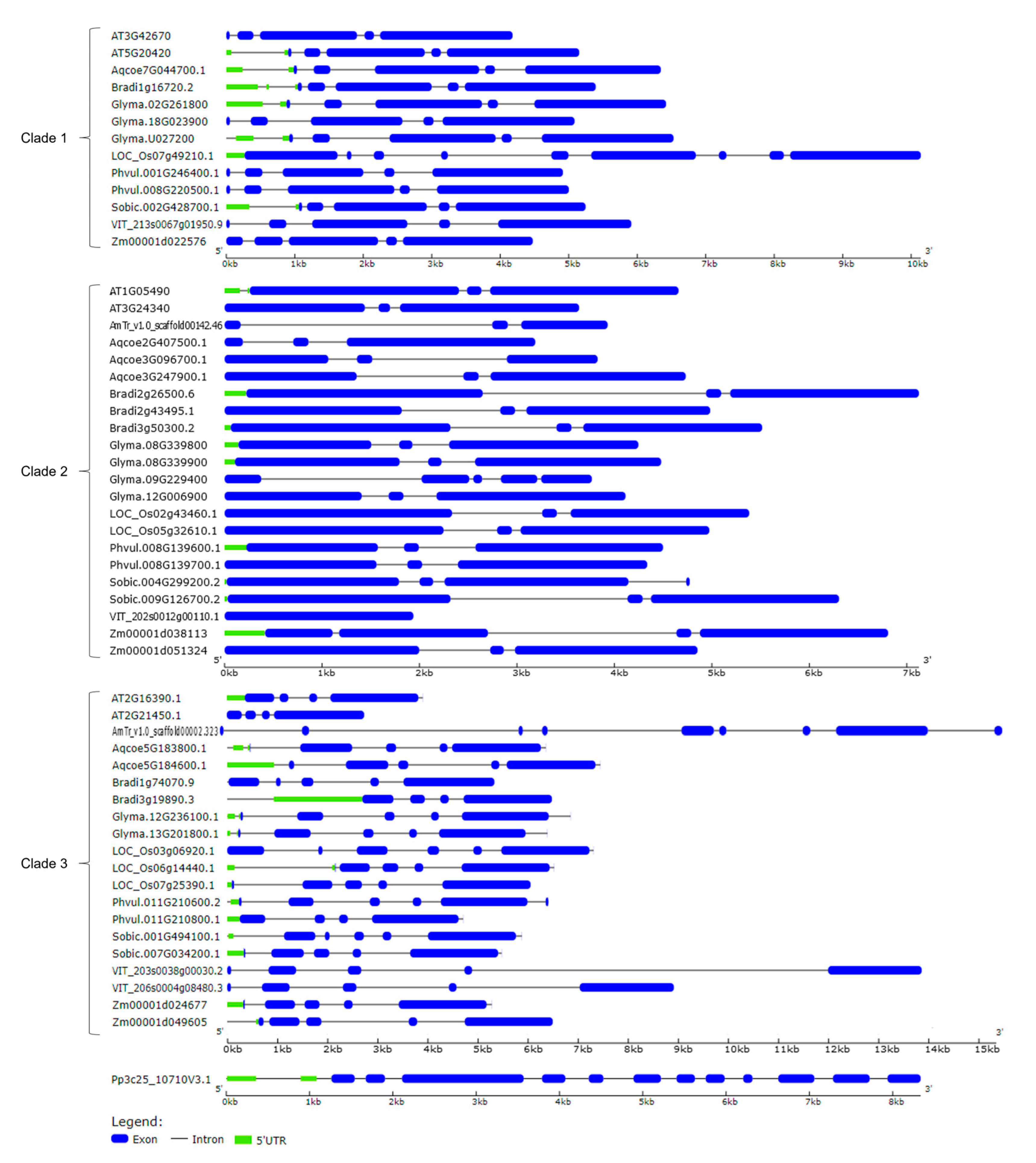
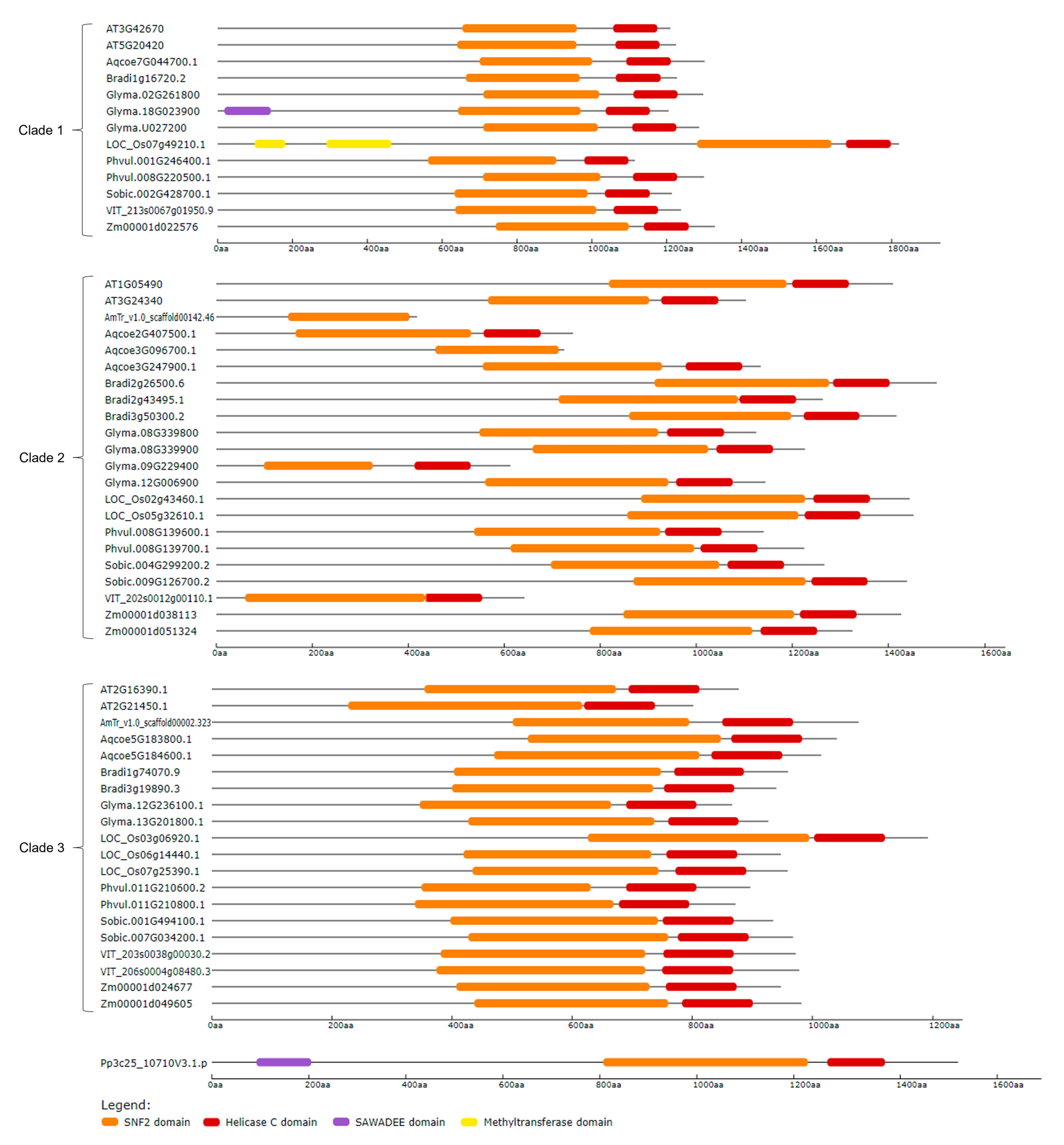
2.2. Identification, Phylogenetic Relationship, and Structural Analysis of CLSY Family in Plants
2.3. Duplication Events of CLSY Genes in Soybean
2.4. Tissue Expression Profile of CLSY Genes in Arabidopsis and Soybean
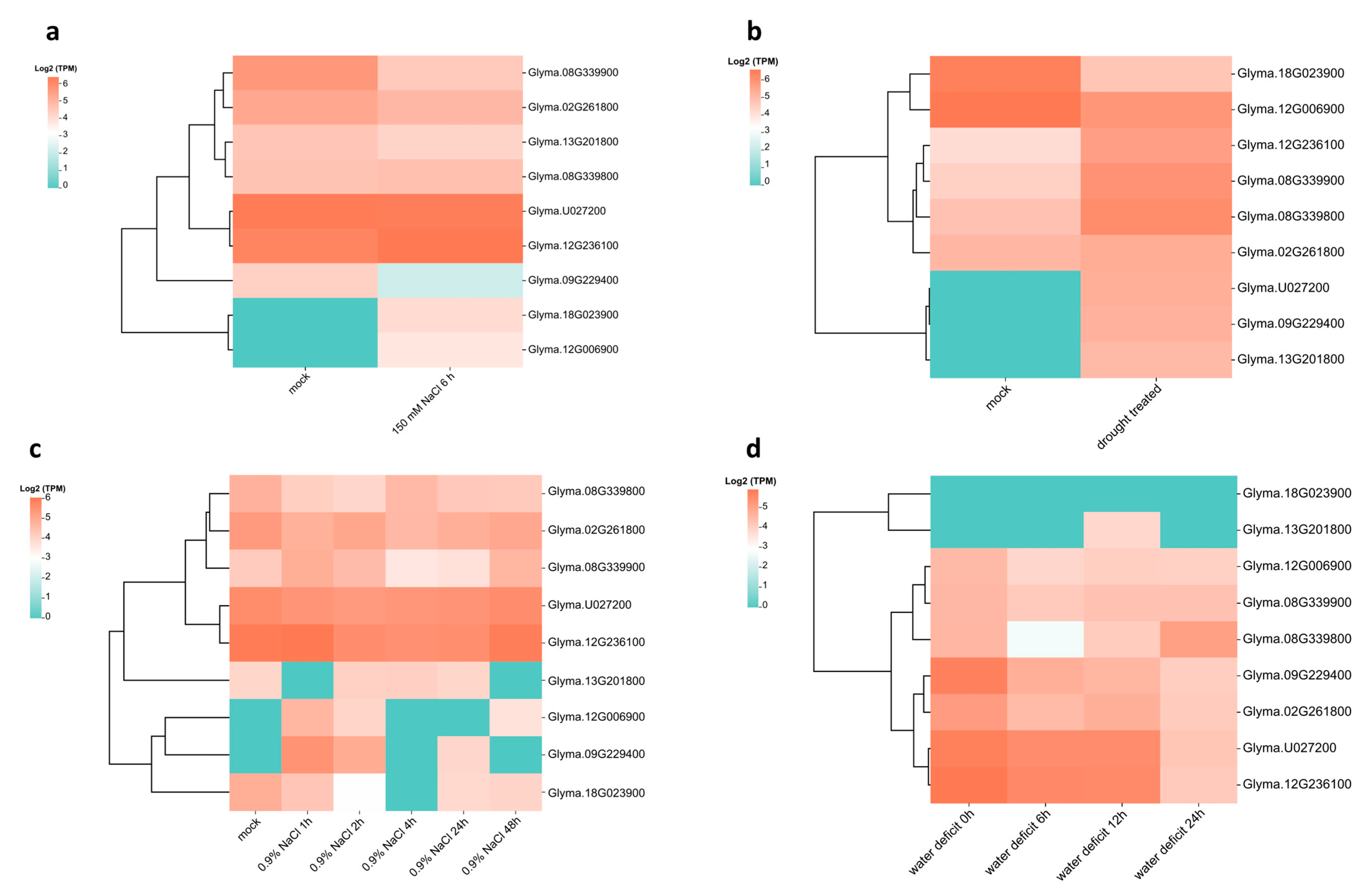
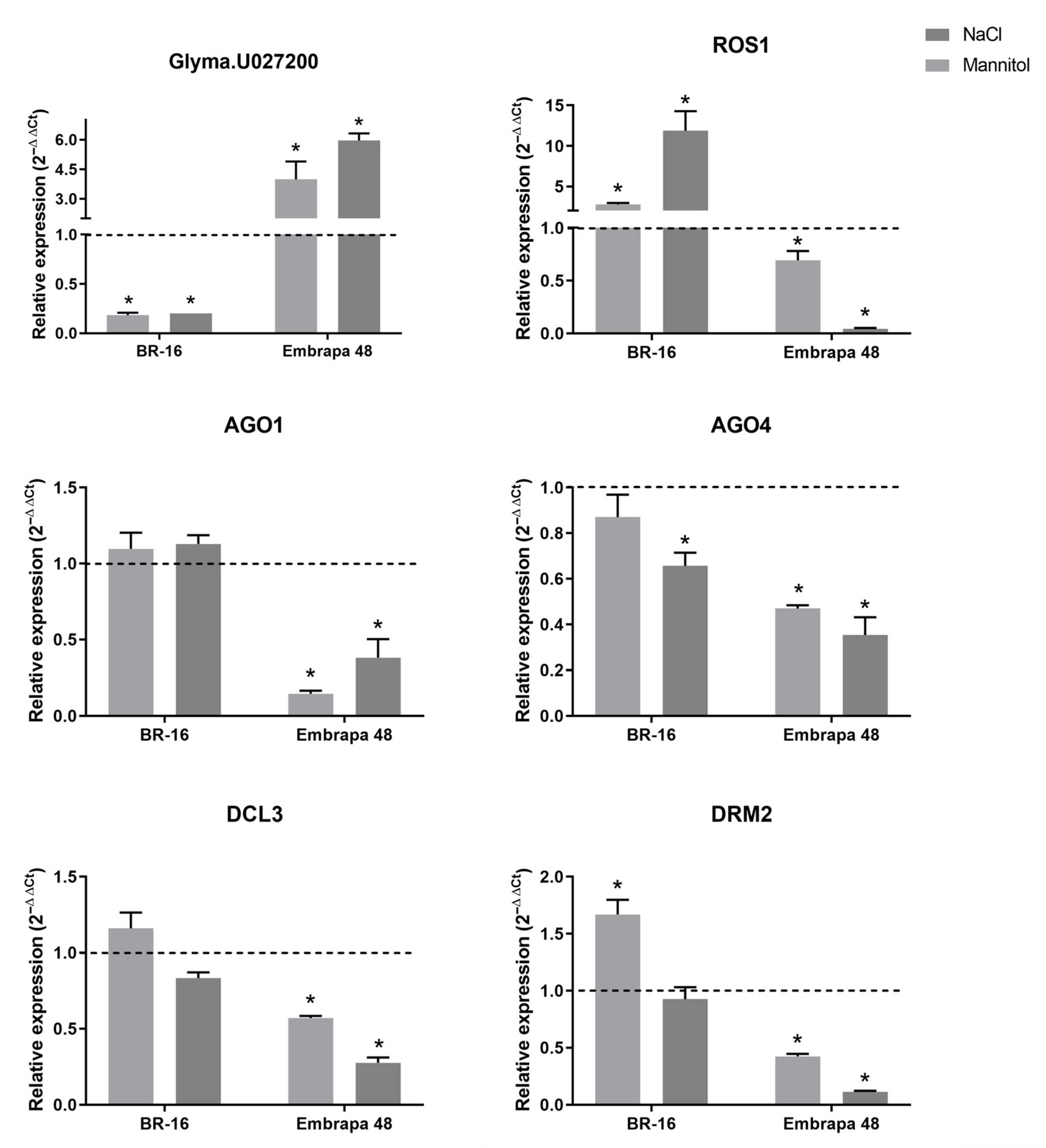
2.5. The Expression Profile of CLSY and Five Other Genes Involved in Epigenetic Regulation Can Be Modulated Under Abiotic Stresses During Soybean Germination
3. Materials and Methods
3.1. Identification of CLSY Gene Family Members
3.2. Phylogenetic Analysis
3.3. Gene Structure and Domain Prediction
3.4. Non-Synonymous and Synonymous (Ka/Ks) Analysis for Duplicated Pairs of CLASSY Genes in Soybean
3.5. Expression Analysis of CLSY Genes in Arabidopis and Soybean Tissues
3.6. Expression Analysis of a CLSY Gene During Soybean Germination Under Stress Conditions
3.7. Promoter Sequence Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AGO | ARGONAUTE |
| CLSY | CLASSY |
| DCL3 | DICER-LIKE 3 |
| DRD1 | DEFECTIVE IN RNA DIRECTED DNA METHYLATION 1 |
| DRM2 | DOMAINS REARRANGED METHYLTRANSFERASE 2 |
| dsRNA | double-stranded RNA |
| HMM | Hidden Markov models |
| Pol | RNA Polymerase |
| RdDM | RNA-directed DNA methylation |
| RDR2 | RNA-DEPENDENT RNA POLYMERASE 2 |
| ROS1 | REPRESSOR OF SILENCING 1 |
| RT-qPCR | Reverse transcription-quantitative polymerase chain reaction |
| SHH1 | SAWADEE HOMEODOMAIN HOMOLOG 1 |
| siRNA | small interfering RNA |
| sRNA | small RNA |
| ssRNA | single-stranded RNA |
| TE | Transposable element |
| WGD | Whole-genome duplication |
References
- Henderson, I.R.; Jacobsen, S.E. Epigenetic inheritance in plants. Nature 2007, 447, 418–424. [Google Scholar] [CrossRef]
- Pikaard, C.S.; Scheid, O.M. Epigenetic regulation in plants. Cold Spring Harb. Perspect. Biol. 2014, 6, a019315. [Google Scholar] [CrossRef]
- Grativol, C.; Hemerly, A.S.; Ferreira, P.C.G. Genetic and epigenetic regulation of stress responses in natural plant populations. Biochim. Biophys. Acta-Gene Regul. Mech. 2012, 1819, 176–185. [Google Scholar] [CrossRef] [PubMed]
- Brukhin, V.; Albertini, E. Epigenetic modifications in plant development and reproduction. Epigenomes 2021, 5, 25. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Mohapatra, T. Dynamics of DNA Methylation and Its Functions in Plant Growth and Development. Front. Plant Sci. 2021, 12, 596236. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Lang, Z.; Zhu, J.K. Dynamics and function of DNA methylation in plants. Nat. Rev. Mol. Cell Biol. 2018, 19, 489–506. [Google Scholar] [CrossRef]
- Hemenway, E.A.; Gehring, M. Epigenetic Regulation During Plant Development and the Capacity for Epigenetic Memory. Annu. Rev. Plant Biol. 2023, 74, 87–109. [Google Scholar] [CrossRef]
- Erdmann, R.M.; Picard, C.L. RNA-directed DNA Methylation. PLoS Genet. 2020, 16, e1009034. [Google Scholar] [CrossRef]
- Wassenegger, M.; Heimes, S.; Riedel, L.; Sänger, H.L. RNA-directed de novo methylation of genomic sequences in plants. Cell 1994, 76, 567–576. [Google Scholar] [CrossRef]
- Matzke, M.A.; Mosher, R.A. RNA-directed DNA methylation: An epigenetic pathway of increasing complexity. Nat. Rev. Genet. 2014, 15, 394–408. [Google Scholar] [CrossRef]
- Xu, R.; Wang, Y.; Zheng, H.; Lu, W.; Wu, C.; Huang, J.; Yan, K.; Yang, G.; Zheng, C. Salt-induced transcription factor MYB74 is regulated by the RNA-directed DNA methylation pathway in Arabidopsis. J. Exp. Bot. 2015, 66, 5997–6008. [Google Scholar] [CrossRef]
- Popova, O.V.; Dinh, H.Q.; Aufsatz, W.; Jonak, C. The RdDM pathway is required for basal heat tolerance in Arabidopsis. Mol. Plant 2013, 6, 396–410. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Zhao, C.; Zhang, Q.; Zinta, G.; Wang, D.; Lozano-Durán, R.; Zhu, J.K. Pathway conversion enables a double-lock mechanism to maintain DNA methylation and genome stability. Proc. Natl. Acad. Sci. USA 2021, 118, e2107320118. [Google Scholar] [CrossRef] [PubMed]
- Schnable, P.S.; Ware, D.; Fulton, R.S.; Stein, J.C.; Wei, F.; Pasternak, S.; Liang, C.; Zhang, J.; Fulton, L.; Graves, T.A.; et al. The B73 maize genome: Complexity, diversity, and dynamics. Science 2009, 326, 1112–1115. [Google Scholar] [CrossRef] [PubMed]
- Ito, H.; Gaubert, H.; Bucher, E.; Mirouze, M.; Vaillant, I.; Paszkowski, J. An siRNA pathway prevents transgenerational retrotransposition in plants subjected to stress. Nature 2011, 472, 115–120. [Google Scholar] [CrossRef]
- Lucibelli, F.; Valoroso, M.C.; Aceto, S. Plant DNA Methylation: An Epigenetic Mark in Development, Environmental Interactions, and Evolution. Int. J. Mol. Sci. 2022, 23, 8299. [Google Scholar] [CrossRef]
- Smith, L.M.; Pontes, O.; Searle, I.; Yelina, N.; Yousafzai, F.K.; Herr, A.J.; Pikaard, C.S.; Baulcombe, D.C. An SNF2 protein associated with nuclear RNA silencing and the spread of a silencing signal between cells in Arabidopsis. Plant Cell 2007, 19, 1507–1521. [Google Scholar] [CrossRef]
- Law, J.A.; Vashisht, A.A.; Wohlschlegel, J.A.; Jacobsen, S.E. SHH1, a Homeodomain protein required for DNA Methylation, as well as RDR2, RDM4, and Chromatin remodeling factors, associate with RNA Polymerase IV. PLoS Genet. 2011, 7, e1002195. [Google Scholar] [CrossRef]
- Zhou, M.; Palanca, A.M.S.; Law, J.A. Locus-specific control of the de novo DNA methylation pathway in Arabidopsis by the CLASSY family. Nat. Genet. 2018, 50, 865–873. [Google Scholar] [CrossRef]
- Yang, D.L.; Zhang, G.; Wang, L.; Li, J.; Xu, D.; Di, C.; Tang, K.; Yang, L.; Zeng, L.; Miki, D.; et al. Four putative SWI2/SNF2 chromatin remodelers have dual roles in regulating DNA methylation in Arabidopsis. Cell Discov. 2018, 4, 55. [Google Scholar] [CrossRef]
- Zhou, M.; Coruh, C.; Xu, G.; Martins, L.M.; Bourbousse, C.; Lambolez, A.; Law, J.A. The CLASSY family controls tissue-specific DNA methylation patterns in Arabidopsis. Nat. Commun. 2022, 13, 244. [Google Scholar] [CrossRef]
- Schmutz, J.; Cannon, S.B.; Schlueter, J.; Ma, J.; Mitros, T.; Nelson, W.; Hyten, D.L.; Song, Q.; Thelen, J.J.; Cheng, J.; et al. Genome sequence of the palaeopolyploid soybean. Nature 2010, 463, 178–183. [Google Scholar] [CrossRef] [PubMed]
- Dhaubhadel, S.; Marsolais, F. Transcriptomics of legume seed: Soybean a model grain legume. In Seed Development: OMICS Technologies Toward Improvement of Seed Quality and Crop Yield: OMICS in Seed Biology; Agrawal, G.K., Rakwal, R., Eds.; Springer: Berlin/Heidelberg, Germany, 2012; pp. 129–142. ISBN 9789400747494. [Google Scholar]
- Xu, D.; Zeng, L.; Wang, L.; Yang, D.L. Rice requires a chromatin remodeler for Polymerase IV-small interfering RNA production and genomic immunity. Plant Physiol. 2024, 194, 2149–2164. [Google Scholar] [CrossRef]
- Xu, Q.; Kan, Q.; Luo, Z.; Zhang, Q.; Dong, L.; Springer, N.M.; Li, Q. DNA demethylase augments RNA-directed DNA methylation by enhancing CLSY gene expression in maize and Arabidopsis. Mol. Plant 2025, 18, 1158–1170. [Google Scholar] [CrossRef] [PubMed]
- Yadav, S.; Yadava, Y.K.; Kohli, D.; Meena, S.; Kalwan, G.; Bharadwaj, C.; Gaikwad, K.; Arora, A.; Jain, P.K. Genome-wide identification, in silico characterization and expression analysis of the RNA helicase gene family in chickpea (C. arietinum L.). Sci. Rep. 2022, 12, 9778. [Google Scholar] [CrossRef] [PubMed]
- Xiang, R.; Ahmad, B.; Liang, C.; Shi, X.; Yang, L.; Du, G.; Wang, L. Systematic genome-wide and expression analysis of RNA-directed DNA methylation pathway genes in grapes predicts their involvement in multiple biological processes. Front. Plant Sci. 2022, 13, 1089392. [Google Scholar] [CrossRef] [PubMed]
- Trujillo, J.T.; Seetharam, A.S.; Hufford, M.B.; Beilstein, M.A.; Mosher, R.A. Evidence for a unique DNA-dependent RNA polymerase in cereal crops. Mol. Biol. Evol. 2018, 35, 2454–2462. [Google Scholar] [CrossRef]
- Chakraborty, T.; Trujillo, J.T.; Kendall, T.; Mosher, R.A. Charophytic Green Algae Encode Ancestral Polymerase IV/Polymerase V Subunits and a CLSY/DRD1 Homolog. Genome Biol. Evol. 2024, 16, evae119. [Google Scholar] [CrossRef]
- DiBiase, C.N.; Cheng, X.; Lee, G.; Moore, R.C.; McCoy, A.G.; Chilvers, M.I.; Sun, L.; Wang, D.; Lin, F.; Zhao, M. DNA methylation analysis reveals local changes in resistant and susceptible soybean lines in response to Phytophthora sansomeana. G3 Genes Genomes Genet. 2024, 14, jkae191. [Google Scholar] [CrossRef]
- Park, J.; Karplus, K.; Barrett, C.; Hughey, R.; Haussler, D.; Hubbard, T.; Chothia, C. Sequence comparisons using multiple sequences detect three times as many remote homologues as pairwise methods. J. Mol. Biol. 1998, 284, 1201–1210. [Google Scholar] [CrossRef]
- Oliveira, L.S.; Gruber, A. Rational Design of Profile Hidden Markov Models for Viral Classification and Discovery. In Bioinformatics; Nakaya, H.I., Ed.; Exon Publications: Brisbane, Australia, 2021; pp. 151–170. [Google Scholar]
- Matzke, M.A.; Kanno, T.; Matzke, A.J.M. RNA-directed DNA methylation: The evolution of a complex epigenetic pathway in flowering plants. Annu. Rev. Plant Biol. 2015, 66, 243–267. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.; Cokus, S.J.; Zhang, X.; Chen, P.Y.; Bostick, M.; Goll, M.G.; Hetzel, J.; Jain, J.; Strauss, S.H.; Halpern, M.E.; et al. Conservation and divergence of methylation patterning in plants and animals. Proc. Natl. Acad. Sci. USA 2010, 107, 8689–8694. [Google Scholar] [CrossRef] [PubMed]
- Bargsten, J.W.; Folta, A.; Mlynárová, L.; Nap, J.P. Snf2 family gene distribution in higher plant genomes reveals DRD1 expansion and diversification in the tomato genome. PLoS ONE 2013, 8, e81147. [Google Scholar] [CrossRef]
- Lallemand, T.; Leduc, M.; Landès, C.; Rizzon, C.; Lerat, E. An overview of duplicated gene detection methods: Why the duplication mechanism has to be accounted for in their choice. Genes 2020, 11, 1046. [Google Scholar] [CrossRef]
- Bolon, Y.T.; Stec, A.O.; Michno, J.M.; Roessler, J.; Bhaskar, P.B.; Ries, L.; Dobbels, A.A.; Campbell, B.W.; Young, N.P.; Anderson, J.E.; et al. Genome resilience and prevalence of segmental duplications following fast neutron irradiation of soybean. Genetics 2014, 198, 967–981. [Google Scholar] [CrossRef]
- Pagel, J.; Walling, J.G.; Young, N.D.; Shoemaker, R.C.; Jackson, S.A. Segmental duplications within the Glycine max genome revealed by fluorescence in situ hybridization of bacterial artificial chromosomes. Genome 2004, 47, 764–768. [Google Scholar] [CrossRef]
- Hurst, L.D. The Ka/Ks ratio: Diagnosing the form of sequence evolution. Trends Genet. 2002, 18, 486–487. [Google Scholar] [CrossRef]
- Reis, R.R.; Mertz-Henning, L.M.; Marcolino-Gomes, J.; Rodrigues, F.A.; Rockenbach-Marin, S.; Fuganti-Pagliarini, R.; Koltun, A.; Gonçalves, L.S.A.; Nepomuceno, A.L. Differential gene expression in response to water deficit in leaf and root tissues of soybean genotypes with contrasting tolerance profiles. Genet. Mol. Biol. 2020, 43, e20180290. [Google Scholar] [CrossRef]
- Neves-Borges, A.C.; Guimarães-Dias, F.; Cruz, F.; Mesquita, R.O.; Nepomuceno, A.L.; Romano, E.; Loureiro, M.E.; de Fátima Grossi-de-Sá, M.; Alves-Ferreira, M. Expression pattern of drought stress marker genes in soybean roots under two water deficit systems. Genet. Mol. Biol. 2012, 35, 212–221. [Google Scholar] [CrossRef]
- Liu, Y.; Cao, Y. GmWRKY17-mediated transcriptional regulation of GmDREB1D and GmABA2 controls drought tolerance in soybean. Plant Mol. Biol. 2023, 113, 157–170. [Google Scholar] [CrossRef]
- Wang, H.; Ni, D.; Shen, J.; Deng, S.; Xuan, H.; Wang, C.; Xu, J.; Zhou, L.; Guo, N.; Zhao, J.; et al. Genome-Wide Identification of the AP2/ERF Gene Family and Functional Analysis of GmAP2/ERF144 for Drought Tolerance in Soybean. Front. Plant Sci. 2022, 13, 848766. [Google Scholar] [CrossRef]
- Belamkar, V.; Weeks, N.T.; Bharti, A.K.; Farmer, A.D.; Graham, M.A.; Cannon, S.B. Comprehensive characterization and RNA-Seq profiling of the HD-Zip transcription factor family in soybean (Glycine max) during dehydration and salt stress. BMC Genom. 2014, 15, 950. [Google Scholar] [CrossRef]
- Bian, S.; Jin, D.; Sun, G.; Shan, B.; Zhou, H.; Wang, J.; Zhai, L.; Li, X. Characterization of the soybean R2R3-MYB transcription factor GmMYB81 and its functional roles under abiotic stresses. Gene 2020, 753, 144803. [Google Scholar] [CrossRef]
- Li, M.; Chen, R.; Jiang, Q.; Sun, X.; Zhang, H.; Hu, Z. GmNAC06, a NAC domain transcription factor enhances salt stress tolerance in soybean. Plant Mol. Biol. 2021, 105, 333–345. [Google Scholar] [CrossRef]
- Castelán-Muñoz, N.; Herrera, J.; Cajero-Sánchez, W.; Arrizubieta, M.; Trejo, C.; García-Ponce, B.; Sánchez, M.d.l.P.; Álvarez-Buylla, E.R.; Garay-Arroyo, A. MADS-box genes are key components of genetic regulatory networks involved in abiotic stress and plastic developmental responses in plants. Front. Plant Sci. 2019, 10, 853. [Google Scholar] [CrossRef] [PubMed]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 2012, 40, D1178–D1186. [Google Scholar] [CrossRef] [PubMed]
- Mistry, J.; Chuguransky, S.; Williams, L.; Qureshi, M.; Salazar, G.A.; Sonnhammer, E.L.L.; Tosatto, S.C.E.; Paladin, L.; Raj, S.; Richardson, L.J.; et al. Pfam: The protein families database in 2021. Nucleic Acids Res. 2021, 49, D412–D419. [Google Scholar] [CrossRef]
- Oliveira, L.S.; Reyes, A.; Dutilh, B.E.; Gruber, A. Rational Design of Profile HMMs for Sensitive and Specific Sequence Detection with Case Studies Applied to Viruses, Bacteriophages, and Casposons. Viruses 2023, 15, 519. [Google Scholar] [CrossRef]
- Eddy, S.R. Accelerated profile HMM searches. PLoS Comput. Biol. 2011, 7, e1002195. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; Von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; Von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.A.; Clamp, M.; Barton, G.J. Jalview Version 2-A multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; Mcgettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Zhang, Z. KaKs_Calculator 3.0: Calculating Selective Pressure on Coding and Non-coding Sequences. Genom. Proteom. Bioinforma. 2022, 20, 536–540. [Google Scholar] [CrossRef]
- Nei, M.; Gojobori, T. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol. Biol. Evol. 1986, 3, 418–426. [Google Scholar] [CrossRef]
- Lynch, M.; Conery, J.S. The evolutionary fate and consequences of duplicate genes. Science 2000, 290, 1151–1155. [Google Scholar] [CrossRef]
- Fucile, G.; Di Biase, D.; Nahal, H.; La, G.; Khodabandeh, S.; Chen, Y.; Easley, K.; Christendat, D.; Kelley, L.; Provart, N.J. Eplant and the 3D data display initiative: Integrative systems biology on the world wide web. PLoS ONE 2011, 6, e15237. [Google Scholar] [CrossRef]
- Almeida-Silva, F.; Pedrosa-Silva, F.; Venancio, T.M. The Soybean Expression Atlas v2: A comprehensive database of over 5000 RNA-seq samples. Plant J. 2023, 116, 1041–1051. [Google Scholar] [CrossRef]
- Keen, K.J. Graphics for Statistics and Data Analysis with R, 2nd ed.; Chapman and Hall/CRC: New York, NY, USA, 2018; ISBN 9780429031069. [Google Scholar]
- Sangi, S.; Olimpio, G.V.; Coelho, F.S.; Alexandrino, C.R.; Da Cunha, M.; Grativol, C. Flagellin and mannitol modulate callose biosynthesis and deposition in soybean seedlings. Physiol. Plant. 2023, 175, e13877. [Google Scholar] [CrossRef] [PubMed]
- Sangi, S.; Santos, M.L.C.; Alexandrino, C.R.; Da Cunha, M.; Coelho, F.S.; Ribeiro, G.P.; Lenz, D.; Ballesteros, H.; Hemerly, A.S.; Venâncio, T.M.; et al. Cell wall dynamics and gene expression on soybean embryonic axes during germination. Planta 2019, 250, 1325–1337. [Google Scholar] [CrossRef] [PubMed]
- Coelho, F.S.; Sangi, S.; Moraes, J.L.; da Silva Santos, W.; Gamosa, E.A.; Fernandes, K.V.S.; Grativol, C. Methyl-CpG binding proteins (MBD) family evolution and conservation in plants. Gene 2022, 824, 146404. [Google Scholar] [CrossRef]
- Rao, X.; Huang, X.; Zhou, Z.; Lin, X. An improvement of the 2ˆ(-delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostat. Bioinforma. Biomath. 2013, 3, 71–85. [Google Scholar]
- Chow, C.N.; Yang, C.W.; Wu, N.Y.; Wang, H.T.; Tseng, K.C.; Chiu, Y.H.; Lee, T.Y.; Chang, W.C. PlantPAN 4.0: Updated database for identifying conserved non-coding sequences and exploring dynamic transcriptional regulation in plant promoters. Nucleic Acids Res. 2024, 52, D1569–D1578. [Google Scholar] [CrossRef] [PubMed]


| Gene ID | Gene Length (bp) | Exons Number | Protein Length (aa) | Other Domains | Chromosome | Phylogenetic Tree Clade |
|---|---|---|---|---|---|---|
| Aqcoe7G044700.1 | 6524 | 5 | 1312 | 0 | 7 | Clade 1 |
| AT3G42670 | 4288 | 5 | 1257 | 0 | 3 | |
| AT5G20420 | 5153 | 5 | 1262 | 0 | 5 | |
| Bradi1g16720.2 | 5594 | 5 | 1261 | 0 | 1 | |
| Glyma.02G261800 | 7254 | 5 | 1311 | 0 | 2 | |
| Glyma.18G023900 | 5085 | 5 | 1236 | SAWADEE | 18 | |
| Glyma.U027200 | 7454 | 5 | 1308 | 0 | scaffold_265 | |
| LOC_Os07g49210.1 | 10,327 | 9 | 1875 | Methyltransferase | 7 | |
| Phvul.001G246400.1 | 4915 | 5 | 1179 | 0 | 1 | |
| Phvul.008G220500.1 | 5002 | 5 | 1311 | 0 | 8 | |
| Sobic.002G428700.1 | 5539 | 5 | 1231 | 0 | 2 | |
| VIT_213s0067g01950.9 | 5914 | 5 | 1264 | 0 | 13 | |
| Zm00001d022576 | 4684 | 5 | 1335 | 0 | 7 | |
| AmTr_v1.0_scaffold00142.46 | 3931 | 3 | 405 | 0 | scaffold00142 | Clade 2 |
| Aqcoe2G407500.1 | 3357 | 3 | 761 | 0 | 2 | |
| Aqcoe3G096700.1 | 3828 | 3 | 719 | 0 | 3 | |
| Aqcoe3G247900.1 | 5106 | 3 | 1173 | 0 | 3 | |
| AT1G05490 | 4851 | 3 | 1411 | 0 | 1 | |
| AT3G24340 | 3638 | 3 | 1133 | 0 | 3 | |
| Bradi2g26500.6 | 8975 | 3 | 1507 | 0 | 2 | |
| Bradi2g43495.1 | 6754 | 3 | 1286 | 0 | 2 | |
| Bradi3g50300.2 | 6593 | 3 | 1416 | 0 | 3 | |
| Glyma.08G339800 | 4517 | 3 | 1149 | 0 | 8 | |
| Glyma.08G339900 | 7386 | 3 | 1247 | 0 | 8 | |
| Glyma.09G229400 | 3769 | 5 | 618 | 0 | 9 | |
| Glyma.12G006900 | 4114 | 3 | 1167 | 0 | 12 | |
| LOC_Os02g43460.1 | 5383 | 3 | 1440 | 0 | 2 | |
| LOC_Os05g32610.1 | 5705 | 3 | 1446 | 0 | 5 | |
| Phvul.008G139600.1 | 4598 | 3 | 1143 | 0 | 8 | |
| Phvul.008G139700.1 | 4337 | 3 | 1219 | 0 | 8 | |
| Sobic.004G299200.2 | 5147 | 4 | 1279 | 0 | 4 | |
| Sobic.009G126700.2 | 7022 | 3 | 1459 | 0 | 9 | |
| VIT_202s0012g00110.1 | 1938 | 1 | 646 | 0 | 2 | |
| Zm00001d038113 | 7648 | 4 | 1436 | 0 | 6 | |
| Zm00001d051324 | 4853 | 3 | 1338 | 0 | 4 | |
| AmTr_v1.0_scaffold00002.323 | 15,347 | 9 | 1095 | 0 | scaffold00002 | Clade 3 |
| Aqcoe5G183800.1 | 8214 | 6 | 1060 | 0 | 5 | |
| Aqcoe5G184600.1 | 8072 | 6 | 1027 | 0 | 5 | |
| AT2G16390.1 | 3982 | 5 | 889 | 0 | 2 | |
| AT2G21450.1 | 2947 | 4 | 817 | 0 | 2 | |
| Bradi1g74070.9 | 5856 | 5 | 974 | 0 | 1 | |
| Bradi3g19890.3 | 7475 | 4 | 948 | 0 | 3 | |
| Glyma.12G236100.1 | 7244 | 6 | 884 | 0 | 12 | |
| Glyma.13G201800.1 | 6578 | 6 | 954 | 0 | 13 | |
| LOC_Os03g06920.1 | 7317 | 7 | 1198 | 0 | 3 | |
| LOC_Os06g14440.1 | 7192 | 6 | 952 | 0 | 6 | |
| LOC_Os07g25390.1 | 6671 | 5 | 967 | 0 | 7 | |
| Phvul.011G210600.2 | 6631 | 6 | 901 | 0 | 11 | |
| Phvul.011G210800.1 | 4894 | 5 | 872 | 0 | 11 | |
| Sobic.001G494100.1 | 6270 | 6 | 946 | 0 | 1 | |
| Sobic.007G034200.1 | 6488 | 6 | 971 | 0 | 7 | |
| VIT_203s0038g00030.2 | 13,882 | 5 | 973 | 0 | 3 | |
| VIT_206s0004g08480.3 | 8920 | 5 | 976 | 0 | 6 | |
| Zm00001d024677 | 5604 | 6 | 951 | 0 | 10 | |
| Zm00001d049605 | 7240 | 5 | 978 | 0 | 4 | |
| Pp3c25_10710V3.1 | 8807 | 12 | 1534 | SAWADEE | 25 |
| Gene ID | Gene ID | Ka | Ks | Ka/Ks | Duplication Date (Mya) | Selection Pressure | Duplication Type |
|---|---|---|---|---|---|---|---|
| Glyma.02G261800 | Glyma.18G023900 | 0.223722 | 0.618747 | 0.361573 | 47.60 | Purification or Stabilization selection | WGD or Segmental |
| Glyma.02G261800 | Glyma.U027200 | 0.0281611 | 0.117351 | 0.239974 | 9.03 | Purification or Stabilization selection | WGD or Segmental |
| Glyma.U027200 | Glyma.18G023900 | 0.223407 | 0.614387 | 0.363626 | 47.26 | Purification or Stabilization selection | WGD or Segmental |
| Glyma.08G339900 | Glyma.08G339800 | 0.219666 | 0.373098 | 0.58876 | 28.70 | Purification or Stabilization selection | Tandem |
| Glyma.08G339900 | Glyma.09G229400 | 0.166574 | 0.407391 | 0.40888 | 31.34 | Purification or Stabilization selection | WGD or Segmental |
| Glyma.08G339900 | Glyma.12G006900 | 0.225172 | 0.396627 | 0.567 | 30.51 | Purification or Stabilization selection | WGD or Segmental |
| Glyma.12G006900 | Glyma.09G229400 | 0.131003 | 0.173655 | 0.754383 | 13.36 | Purification or Stabilization selection | WGD or Segmental |
| Glyma.12G006900 | Glyma.08G339800 | 0.14434 | 0.301214 | 0.47919 | 23.17 | Purification or Stabilization selection | WGD or Segmental |
| Glyma.09G229400 | Glyma.08G339800 | 0.14488 | 0.309337 | 0.468356 | 23.80 | Purification or Stabilization selection | WGD or Segmental |
| Glyma.12G236100 | Glyma.13G201800 | 0.09762 | 0.31009 | 0.3148 | 23.85 | Purification or Stabilization selection | WGD or Segmental |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Araújo, P.M.d.; Gruber, A.; Oliveira, L.S.; Sangi, S.; Olimpio, G.V.; Paula, F.C.; Grativol, C. RdDM-Associated Chromatin Remodelers in Soybean: Evolution and Stress-Induced Expression of CLASSY Genes. Plants 2025, 14, 2543. https://doi.org/10.3390/plants14162543
Araújo PMd, Gruber A, Oliveira LS, Sangi S, Olimpio GV, Paula FC, Grativol C. RdDM-Associated Chromatin Remodelers in Soybean: Evolution and Stress-Induced Expression of CLASSY Genes. Plants. 2025; 14(16):2543. https://doi.org/10.3390/plants14162543
Chicago/Turabian StyleAraújo, Paula Machado de, Arthur Gruber, Liliane Santana Oliveira, Sara Sangi, Geovanna Vitória Olimpio, Felipe Cruz Paula, and Clícia Grativol. 2025. "RdDM-Associated Chromatin Remodelers in Soybean: Evolution and Stress-Induced Expression of CLASSY Genes" Plants 14, no. 16: 2543. https://doi.org/10.3390/plants14162543
APA StyleAraújo, P. M. d., Gruber, A., Oliveira, L. S., Sangi, S., Olimpio, G. V., Paula, F. C., & Grativol, C. (2025). RdDM-Associated Chromatin Remodelers in Soybean: Evolution and Stress-Induced Expression of CLASSY Genes. Plants, 14(16), 2543. https://doi.org/10.3390/plants14162543











