Transposon Dynamics Drive Genome Evolution and Regulate Genetic Mechanisms of Agronomic Traits in Cotton
Abstract
1. Introduction
2. Results
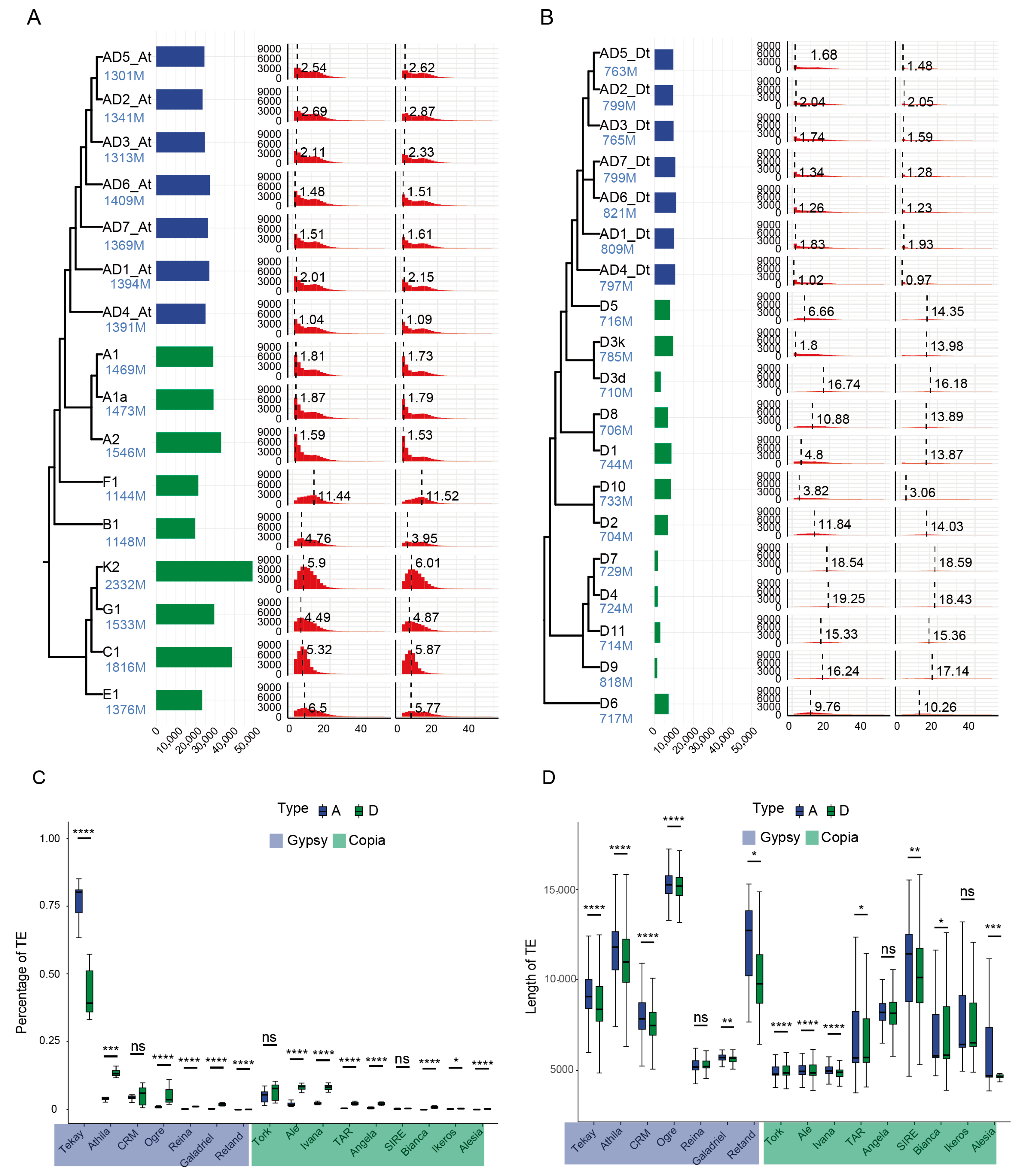
2.1. Tekay Shapes the Genomes of Diploid Cotton Species
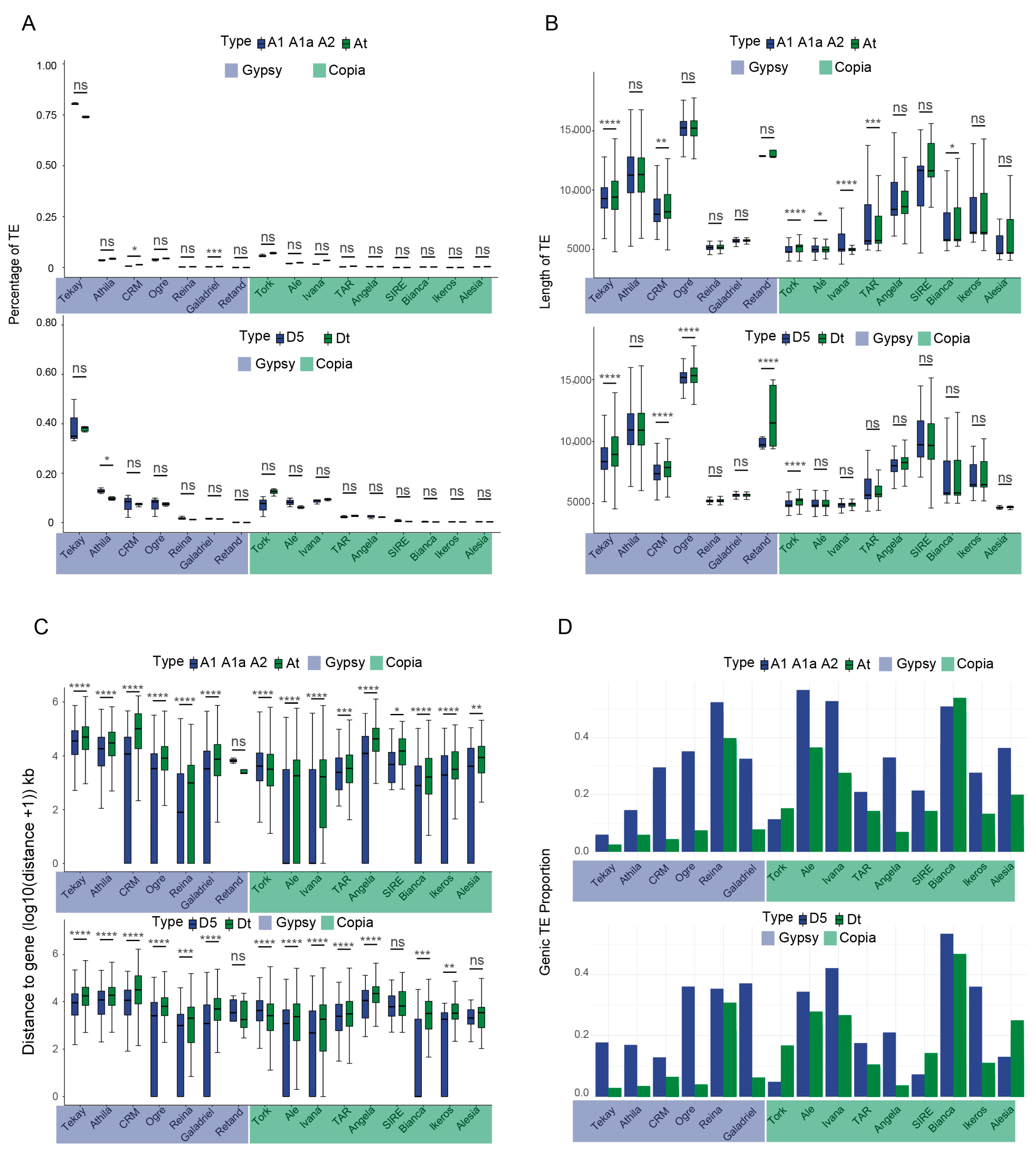
2.2. TE Dynamics Following Polyploidization in Cotton Species
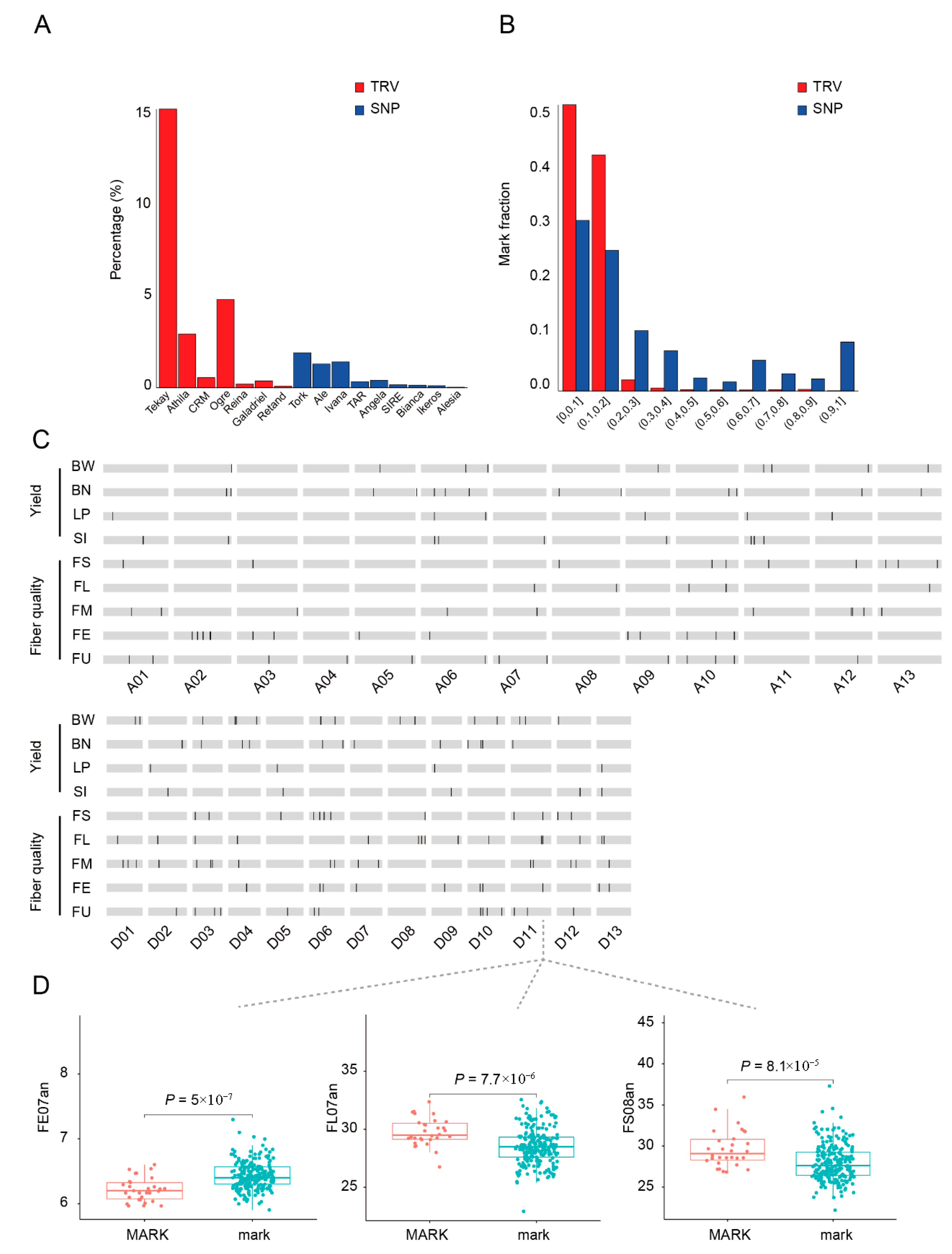
2.3. Construction of the TRV Genetic Map and Its Transcriptional Effects
2.4. Identification of TRV-Associated Loci Contributing to Agronomic Traits in Cultivated Cotton
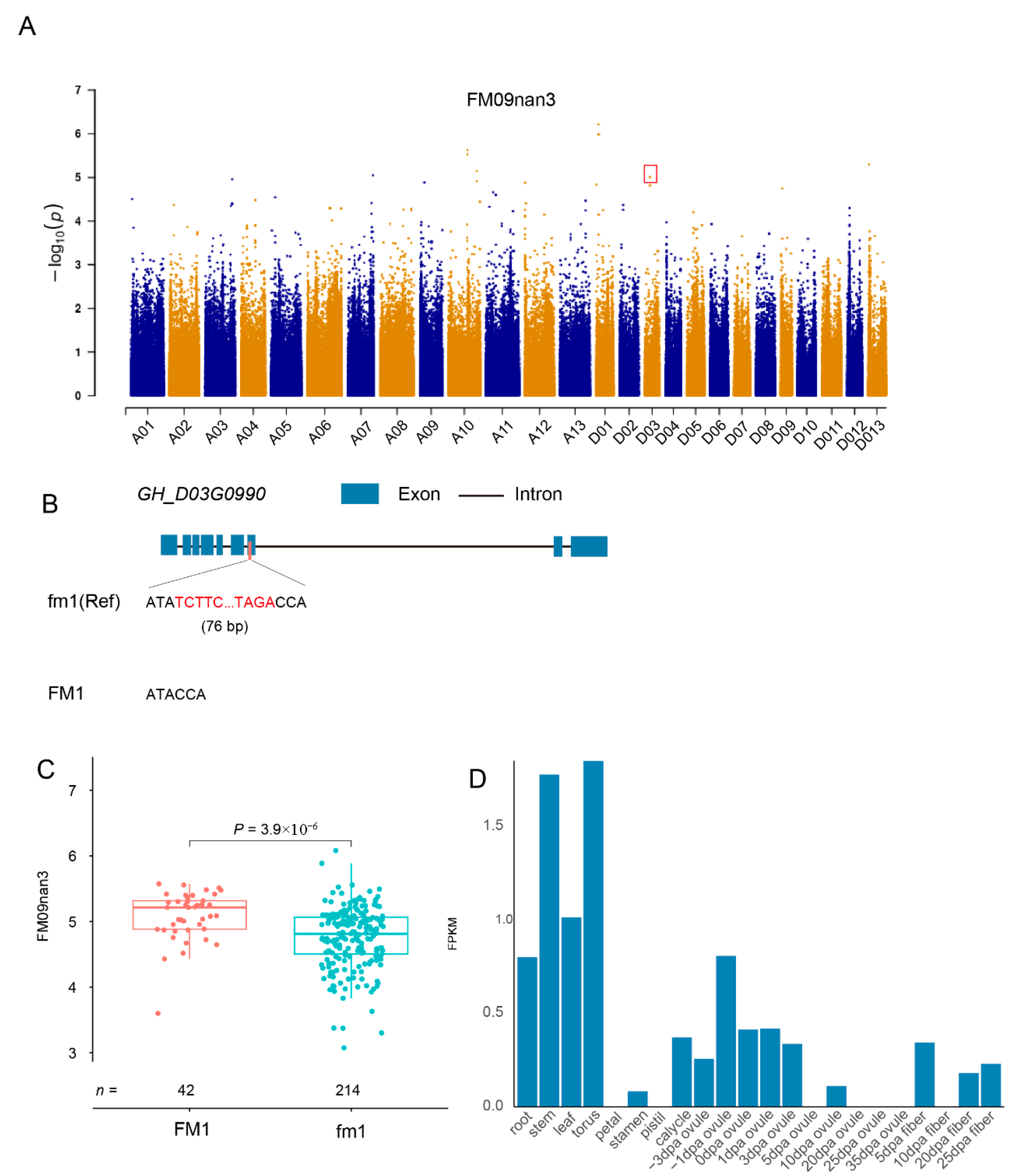
2.5. Key TRV Influencing Agronomic Traits in Cotton
3. Discussion
3.1. Dynamics of Lineage-Specific TEs During Polyploidization in Gossypium
3.2. TEs Are a Major Source of Phenotypic Variation in Cultivated Gossypium Species
4. Materials and Methods
4.1. Published Genome and Transcriptome Data Collection
4.2. Identification of Intact LTR
4.3. Classification of Intact LTR Retrotransposons
4.4. Estimation of Insertion Time of the LTR-RTs
4.5. Identification of Interspecific Transposon-Related Variations Among the Gossypium Genus
4.6. Pan-TRV Construction
4.7. TRV Genotyping and Annotation
4.8. Transcriptional Impact of TRV and eQTL Analysis
4.9. Genome-Wide Association Analysis Based on TRV
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Mita, P.; Boeke, J.D. How retrotransposons shape genome regulation. Curr. Opin. Genet. Dev. 2016, 37, 90–100. [Google Scholar] [CrossRef]
- Said, I.; McGurk, M.P.; Clark, A.G.; Barbash, D.A. Patterns of piRNA Regulation in Drosophila Revealed through Transposable Element Clade Inference. Mol. Biol. Evol. 2022, 39, msab336. [Google Scholar] [CrossRef] [PubMed]
- Seidl, M.F.; Thomma, B. Transposable Elements Direct The Coevolution between Plants and Microbes. Trends Genet. 2017, 33, 842–851. [Google Scholar] [CrossRef] [PubMed]
- Wicker, T.; Sabot, F.; Hua-Van, A.; Bennetzen, J.L.; Capy, P.; Chalhoub, B.; Flavell, A.; Leroy, P.; Morgante, M.; Panaud, O. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 2007, 8, 973–982. [Google Scholar] [CrossRef] [PubMed]
- Gozashti, L.; Hoekstra, H.E. Accounting for diverse transposable element landscapes is key to developing and evaluating accurate de novo annotation strategies. Genome Biol. 2024, 25, 4. [Google Scholar] [CrossRef]
- Riehl, K.; Riccio, C.; Miska, E.A.; Hemberg, M. TransposonUltimate: Software for transposon classification, annotation and detection. Nucleic Acids Res. 2022, 50, e64. [Google Scholar] [CrossRef]
- Qin, P.; Lu, H.; Du, H.; Wang, H.; Chen, W.; Chen, Z.; He, Q.; Ou, S.; Zhang, H.; Li, X. Pan-genome analysis of 33 genetically diverse rice accessions reveals hidden genomic variations. Cell 2021, 184, 3542–3558.e3516. [Google Scholar] [CrossRef]
- Gao, L.; Gonda, I.; Sun, H.; Ma, Q.; Bao, K.; Tieman, D.M.; Burzynski-Chang, E.A.; Fish, T.L.; Stromberg, K.A.; Sacks, G.L. The tomato pan-genome uncovers new genes and a rare allele regulating fruit flavor. Nat. Genet. 2019, 51, 1044–1051. [Google Scholar] [CrossRef]
- Liu, Y.; Du, H.; Li, P.; Shen, Y.; Peng, H.; Liu, S.; Zhou, G.-A.; Zhang, H.; Liu, Z.; Shi, M. Pan-genome of wild and cultivated soybeans. Cell 2020, 182, 162–176.e13. [Google Scholar] [CrossRef]
- Klumpe, S.; Senti, K.A.; Beck, F.; Sachweh, J.; Hampoelz, B.; Ronchi, P.; Oorschot, V.; Brandstetter, M.; Yeroslaviz, A.; Briggs, J.A.G.; et al. In-cell structure and snapshots of copia retrotransposons in intact tissue by cryo-ET. Cell 2025, 188, 2094–2110.e18. [Google Scholar] [CrossRef]
- Sanchez, D.H.; Gaubert, H.; Drost, H.G.; Zabet, N.R.; Paszkowski, J. High-frequency recombination between members of an LTR retrotransposon family during transposition bursts. Nat. Commun. 2017, 8, 1283. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Bennetzen, J.L. Plant retrotransposons. Annu. Rev. Genet. 1999, 33, 479–532. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Eickbush, T.H. Origin and evolution of retroelements based upon their reverse transcriptase sequences. EMBO J. 1990, 9, 3353–3362. [Google Scholar] [CrossRef] [PubMed]
- Llorens, C.; Futami, R.; Covelli, L.; Domínguez-Escribá, L.; Viu, J.M.; Tamarit, D.; Aguilar-Rodríguez, J.; Vicente-Ripolles, M.; Fuster, G.; Bernet, G.P. The Gypsy Database (GyDB) of mobile genetic elements: Release 2.0. Nucleic Acids Res. 2010, 39, D70–D74. [Google Scholar] [CrossRef]
- Baduel, P.; Quadrana, L.; Hunter, B.; Bomblies, K.; Colot, V. Relaxed purifying selection in autopolyploids drives transposable element over-accumulation which provides variants for local adaptation. Nat. Commun. 2019, 10, 5818. [Google Scholar] [CrossRef]
- Eriksson, M.C.; Mandakova, T.; McCann, J.; Temsch, E.M.; Chase, M.W.; Hedren, M.; Weiss-Schneeweiss, H.; Paun, O. Repeat Dynamics across Timescales: A Perspective from Sibling Allotetraploid Marsh Orchids (Dactylorhiza majalis s.l.). Mol. Biol. Evol. 2022, 39, msac167. [Google Scholar] [CrossRef]
- Choi, J.Y.; Purugganan, M.D. Evolutionary Epigenomics of Retrotransposon-Mediated Methylation Spreading in Rice. Mol. Biol. Evol. 2018, 35, 365–382. [Google Scholar] [CrossRef]
- Gordon, S.P.; Contreras-Moreira, B.; Levy, J.J.; Djamei, A.; Czedik-Eysenberg, A.; Tartaglio, V.S.; Session, A.; Martin, J.; Cartwright, A.; Katz, A.; et al. Gradual polyploid genome evolution revealed by pan-genomic analysis of Brachypodium hybridum and its diploid progenitors. Nat. Commun. 2020, 11, 3670. [Google Scholar] [CrossRef]
- Almeida, R.; Allshire, R.C. RNA silencing and genome regulation. Trends Cell Biol. 2005, 15, 251–258. [Google Scholar] [CrossRef]
- Huettel, B.; Kanno, T.; Daxinger, L.; Bucher, E.; van der Winden, J.; Matzke, A.J.; Matzke, M. RNA-directed DNA methylation mediated by DRD1 and Pol IVb: A versatile pathway for transcriptional gene silencing in plants. Biochim. Biophys. Acta 2007, 1769, 358–374. [Google Scholar] [CrossRef]
- Matzke, M.A.; Birchler, J.A. RNAi-mediated pathways in the nucleus. Nat. Rev. Genet. 2005, 6, 24–35. [Google Scholar] [CrossRef]
- Langmuller, A.M.; Nolte, V.; Dolezal, M.; Schlotterer, C. The genomic distribution of transposable elements is driven by spatially variable purifying selection. Nucleic Acids Res. 2023, 51, 9203–9213. [Google Scholar] [CrossRef] [PubMed]
- Peng, Y.; Zhang, Y.; Gui, Y.; An, D.; Liu, J.; Xu, X.; Li, Q.; Wang, J.; Wang, W.; Shi, C.; et al. Elimination of a Retrotransposon for Quenching Genome Instability in Modern Rice. Mol. Plant 2019, 12, 1395–1407. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.; Huang, J.-Q.; Chen, X.-Y.; Zhu, Y.-X. Recent advances and future perspectives in cotton research. Annu. Rev. Plant Biol. 2021, 72, 437–462. [Google Scholar] [CrossRef] [PubMed]
- Wen, X.; Chen, Z.; Yang, Z.; Wang, M.; Jin, S.; Wang, G.; Zhang, L.; Wang, L.; Li, J.; Saeed, S. A comprehensive overview of cotton genomics, biotechnology and molecular biological studies. Sci. China-Life Sci. 2023, 66, 2214–2256. [Google Scholar] [CrossRef]
- Huang, G.; Wu, Z.; Percy, R.G.; Bai, M.; Li, Y.; Frelichowski, J.E.; Hu, J.; Wang, K.; Yu, J.Z.; Zhu, Y. Genome sequence of Gossypium herbaceum and genome updates of Gossypium arboreum and Gossypium hirsutum provide insights into cotton A-genome evolution. Nat. Genet. 2020, 52, 516–524. [Google Scholar] [CrossRef]
- Chen, Z.J.; Sreedasyam, A.; Ando, A.; Song, Q.; De Santiago, L.M.; Hulse-Kemp, A.M.; Ding, M.; Ye, W.; Kirkbride, R.C.; Jenkins, J. Genomic diversifications of five Gossypium allopolyploid species and their impact on cotton improvement. Nat. Genet. 2020, 52, 525–533. [Google Scholar] [CrossRef]
- Stephens, S. Phenogenetic Evidence for the Amphidiploid Origin of New World Cottons. Nature 1944, 153, 53–54. [Google Scholar] [CrossRef]
- Gerstel, D. Chromosomal translocations in interspecific hybrids of the genus Gossypium. Evolution 1953, 7, 234–244. [Google Scholar] [CrossRef]
- Abdul Kadir, Z.B. DNA evolution in the genus Gossypium. Chromosoma 1976, 56, 85–94. [Google Scholar] [CrossRef]
- Johnson, B.L.; Thein, M.M. Assessment of evolutionary affinities in Gossypium by protein electrophoresis. Am. J. Bot. 1970, 57, 1081–1092. [Google Scholar] [CrossRef]
- Mei, H.; Zhao, T.; Dong, Z.; Han, J.; Xu, B.; Chen, R.; Zhang, J.; Zhang, J.; Hu, Y.; Zhang, T. Population-scale polymorphic short tandem repeat provides an alternative strategy for allele mining in cotton. Front. Plant Sci. 2022, 13, 916830. [Google Scholar] [CrossRef]
- Fang, L.; Wang, Q.; Hu, Y.; Jia, Y.; Chen, J.; Liu, B.; Zhang, Z.; Guan, X.; Chen, S.; Zhou, B. Genomic analyses in cotton identify signatures of selection and loci associated with fiber quality and yield traits. Nat. Genet. 2017, 49, 1089–1098. [Google Scholar] [CrossRef] [PubMed]
- Merenciano, M.; Gonzalez, J. The Interplay Between Developmental Stage and Environment Underlies the Adaptive Effect of a Natural Transposable Element Insertion. Mol. Biol. Evol. 2023, 40, msad044. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Gao, X.; Mao, J.; Liu, Y.; Tong, L.; Chen, X.; Liu, Y.; Kou, W.; Chang, C.; Foster, T.; et al. Genome sequencing of ‘Fuji’ apple clonal varieties reveals genetic mechanism of the spur-type morphology. Nat. Commun. 2024, 15, 10082. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.; Han, Z.; Hu, Y.; Si, Z.; Dai, F.; He, L.; Cheng, Y.; Li, Y.; Zhao, T.; Fang, L. Structural variation (SV)-based pan-genome and GWAS reveal the impacts of SVs on the speciation and diversification of allotetraploid cottons. Mol. Plant 2023, 16, 678–693. [Google Scholar] [CrossRef]
- Cai, X.; Wu, J.; Liang, J.; Lin, R.; Zhang, K.; Cheng, F.; Wang, X. Improved Brassica oleracea JZS assembly reveals significant changing of LTR-RT dynamics in different morphotypes. Theor. Appl. Genet. 2020, 133, 3187–3199. [Google Scholar] [CrossRef]
- Li, X.; Dai, X.; He, H.; Lv, Y.; Yang, L.; He, W.; Liu, C.; Wei, H.; Liu, X.; Yuan, Q.; et al. A pan-TE map highlights transposable elements underlying domestication and agronomic traits in Asian rice. Natl. Sci. Rev. 2024, 11, nwae188. [Google Scholar] [CrossRef]
- Kou, Y.; Liao, Y.; Toivainen, T.; Lv, Y.; Tian, X.; Emerson, J.J.; Gaut, B.S.; Zhou, Y. Evolutionary Genomics of Structural Variation in Asian Rice (Oryza sativa) Domestication. Mol. Biol. Evol. 2020, 37, 3507–3524. [Google Scholar] [CrossRef]
- Liu, S.; Cheng, H.; Zhang, Y.; He, M.; Zuo, D.; Wang, Q.; Lv, L.; Lin, Z.; Liu, J.; Song, G. Cotton transposon-related variome reveals roles of transposon-related variations in modern cotton cultivation. J. Adv. Res. 2024, 71, 17–28. [Google Scholar] [CrossRef]
- Dong, Z.; Jin, S.; Fan, R.; Sun, P.; Shao, L.; Zhao, T.; Jiang, H.; Zhang, Z.; Shang, H.; Guan, X.; et al. High-quality genome of Firmiana hainanensis provides insights into the evolution of Malvaceae subfamilies and the mechanism of their wood density formation. J. Genet. Genom. 2024, 52, 812–825. [Google Scholar] [CrossRef]
- Ranawaka, B.; An, J.; Lorenc, M.T.; Jung, H.; Sulli, M.; Aprea, G.; Roden, S.; Llaca, V.; Hayashi, S.; Asadyar, L.; et al. A multi-omic Nicotiana benthamiana resource for fundamental research and biotechnology. Nat. Plants 2023, 9, 1558–1571. [Google Scholar] [CrossRef]
- Feng, X.; Chen, Q.; Wu, W.; Wang, J.; Li, G.; Xu, S.; Shao, S.; Liu, M.; Zhong, C.; Wu, C.I.; et al. Genomic evidence for rediploidization and adaptive evolution following the whole-genome triplication. Nat. Commun. 2024, 15, 1635. [Google Scholar] [CrossRef] [PubMed]
- Domínguez, M.; Dugas, E.; Benchouaia, M.; Leduque, B.; Jiménez-Gómez, J.M.; Colot, V.; Quadrana, L. The impact of transposable elements on tomato diversity. Nat. Commun. 2020, 11, 4058. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Dong, G.; Noinaj, N.; Huang, R. Discovery of Bisubstrate Inhibitors for Protein N-Terminal Methyltransferase 1. J. Med. Chem. 2019, 62, 3773–3779. [Google Scholar] [CrossRef]
- Jia, K.; Huang, G.; Wu, W.; Shrestha, R.; Wu, B.; Xiong, Y.; Li, P. In vivo methylation of OLA1 revealed by activity-based target profiling of NTMT1. Chem. Sci. 2019, 10, 8094–8099. [Google Scholar] [CrossRef]
- Si, Z.; Liu, H.; Zhu, J.; Chen, J.; Wang, Q.; Fang, L.; Gao, F.; Tian, Y.; Chen, Y.; Chang, L.; et al. Mutation of SELF-PRUNING homologs in cotton promotes short-branching plant architecture. J. Exp. Bot. 2018, 69, 2543–2553. [Google Scholar] [CrossRef]
- Chen, D.; Dong, G.; Deng, Y.; Noinaj, N.; Huang, R. Structure-based Discovery of Cell-Potent Peptidomimetic Inhibitors for Protein N-Terminal Methyltransferase 1. ACS Med. Chem. Lett. 2021, 12, 485–493. [Google Scholar] [CrossRef]
- Thyssen, G.N.; Fang, D.D.; Zeng, L.; Song, X.; Delhom, C.D.; Condon, T.L.; Li, P.; Kim, H.J. The Immature Fiber Mutant Phenotype of Cotton (Gossypium hirsutum) Is Linked to a 22-bp Frame-Shift Deletion in a Mitochondria Targeted Pentatricopeptide Repeat Gene. G3-Genes Genomes Genet. 2016, 6, 1627–1633. [Google Scholar] [CrossRef]
- Wang, Y.; Zou, D.; Cheng, C.H.; Zhang, J.; Zhang, J.B.; Zheng, Y.; Li, Y.; Li, X.B. GhTBL3 is required for fiber secondary cell wall (SCW) formation via maintaining acetylation of xylan in cotton. Plant J. 2025, 121, e17167. [Google Scholar] [CrossRef]
- Krzyszton, M.; Zakrzewska-Placzek, M.; Kwasnik, A.; Dojer, N.; Karlowski, W.; Kufel, J. Defective XRN3-mediated transcription termination in Arabidopsis affects the expression of protein-coding genes. Plant J. 2018, 93, 1017–1031. [Google Scholar] [CrossRef]
- Liu, L.; Chen, G.; Li, S.; Gu, Y.; Lu, L.; Qanmber, G.; Mendu, V.; Liu, Z.; Li, F.; Yang, Z. A brassinosteroid transcriptional regulatory network participates in regulating fiber elongation in cotton. Plant Physiol. 2023, 191, 1985–2000. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Qi, Z.; Liu, Z.; Chang, X.; Zhang, X.; Li, J.; Wang, M. Pangenome analysis reveals transposon-driven genome evolution in cotton. BMC Biol. 2024, 22, 92. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, J.S.; Kim, H.; Nason, J.D.; Wing, R.A.; Wendel, J.F. Differential lineage-specific amplification of transposable elements is responsible for genome size variation in Gossypium. Genome Res. 2006, 16, 1252–1261. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.E.; Cui, G.; Wang, X.; Liew, Y.J.; Aranda, M. Recent expansion of heat-activated retrotransposons in the coral symbiont Symbiodinium microadriaticum. ISME J. 2018, 12, 639–643. [Google Scholar] [CrossRef]
- Lu, X.; Zhang, F.; Zhang, C.; Li, G.; Du, Y.; Zhao, C.; Zhao, W.; Gao, F.; Fu, L.; Liu, X. TaTPS11 enhances wheat cold resistance by regulating source-sink factor. Plant Physiol. Biochem. 2024, 211, 108695. [Google Scholar] [CrossRef]
- Zhang, C.; Liu, Z.; Shu, S.; Li, X.; Li, Y.; Liu, L.; Liu, L.; Wang, X.; Li, F.; Qanmber, G.; et al. GhEXL3 participates in brassinosteroids regulation of fiber elongation in Gossypium hirsutum. Plant J. 2024, 120, 491–504. [Google Scholar] [CrossRef]
- Dai, F.; Chen, J.; Zhang, Z.; Liu, F.; Li, J.; Zhao, T.; Hu, Y.; Zhang, T.; Fang, L. COTTONOMICS: A comprehensive cotton multi-omics database. Database 2022, 2022, baac080. [Google Scholar] [CrossRef]
- Yu, J.; Jung, S.; Cheng, C.-H.; Lee, T.; Zheng, P.; Buble, K.; Crabb, J.; Humann, J.; Hough, H.; Jones, D. CottonGen: The community database for cotton genomics, genetics, and breeding research. Plants 2021, 10, 2805. [Google Scholar] [CrossRef]
- Zhao, T.; Guan, X.; Hu, Y.; Zhang, Z.; Yang, H.; Shi, X.; Han, J.; Mei, H.; Wang, L.; Shao, L. Population-wide DNA methylation polymorphisms at single-nucleotide resolution in 207 cotton accessions reveal epigenomic contributions to complex traits. Cell Res. 2024, 34, 859–872. [Google Scholar] [CrossRef]
- Xu, Z.; Wang, H. LTR_FINDER: An efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 2007, 35, W265–W268. [Google Scholar] [CrossRef] [PubMed]
- Ellinghaus, D.; Kurtz, S.; Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinform. 2008, 9, 18. [Google Scholar] [CrossRef] [PubMed]
- Neumann, P.; Novak, P.; Hostakova, N.; Macas, J. Systematic survey of plant LTR-retrotransposons elucidates phylogenetic relationships of their polyprotein domains and provides a reference for element classification. Mob. DNA 2019, 10, 1. [Google Scholar] [CrossRef] [PubMed]
- Kielbasa, S.M.; Wan, R.; Sato, K.; Horton, P.; Frith, M.C. Adaptive seeds tame genomic sequence comparison. Genome Res. 2011, 21, 487–493. [Google Scholar] [CrossRef]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef]
- Hu, Y.; Chen, J.; Fang, L.; Zhang, Z.; Ma, W.; Niu, Y.; Ju, L.; Deng, J.; Zhao, T.; Lian, J.; et al. Gossypium barbadense and Gossypium hirsutum genomes provide insights into the origin and evolution of allotetraploid cotton. Nat. Genet. 2019, 51, 739–748. [Google Scholar] [CrossRef]
- Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef]
- Goel, M.; Sun, H.; Jiao, W.-B.; Schneeberger, K. SyRI: Finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 2019, 20, 277. [Google Scholar] [CrossRef]
- Jeffares, D.C.; Jolly, C.; Hoti, M.; Speed, D.; Shaw, L.; Rallis, C.; Balloux, F.; Dessimoz, C.; Bähler, J.; Sedlazeck, F.J. Transient structural variations have strong effects on quantitative traits and reproductive isolation in fission yeast. Nat. Commun. 2017, 8, 14061. [Google Scholar] [CrossRef]
- Sirén, J.; Monlong, J.; Chang, X.; Novak, A.M.; Eizenga, J.M.; Markello, C.; Sibbesen, J.A.; Hickey, G.; Chang, P.-C.; Carroll, A. Pangenomics enables genotyping of known structural variants in 5202 diverse genomes. Science 2021, 374, abg8871. [Google Scholar] [CrossRef]
- Narasimhan, V.; Danecek, P.; Scally, A.; Xue, Y.; Tyler-Smith, C.; Durbin, R. BCFtools/RoH: A hidden Markov model approach for detecting autozygosity from next-generation sequencing data. Bioinformatics 2016, 32, 1749–1751. [Google Scholar] [CrossRef]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010, 38, e164. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.-C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Kang, H.M.; Sul, J.H.; Service, S.K.; Zaitlen, N.A.; Kong, S.-y.; Freimer, N.B.; Sabatti, C.; Eskin, E. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 2010, 42, 348–354. [Google Scholar] [CrossRef]
- Yin, Y.; Fan, H.; Zhou, B.; Hu, Y.; Fan, G.; Wang, J.; Zhou, F.; Nie, W.; Zhang, C.; Liu, L.; et al. Molecular mechanisms and topological consequences of drastic chromosomal rearrangements of muntjac deer. Nat. Commun. 2021, 12, 6858. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dong, Z.; Jin, S.; Hao, Y.; Zhao, T.; Shang, H.; Zhang, Z.; Fang, L.; Zheng, Z.; Li, J. Transposon Dynamics Drive Genome Evolution and Regulate Genetic Mechanisms of Agronomic Traits in Cotton. Plants 2025, 14, 2509. https://doi.org/10.3390/plants14162509
Dong Z, Jin S, Hao Y, Zhao T, Shang H, Zhang Z, Fang L, Zheng Z, Li J. Transposon Dynamics Drive Genome Evolution and Regulate Genetic Mechanisms of Agronomic Traits in Cotton. Plants. 2025; 14(16):2509. https://doi.org/10.3390/plants14162509
Chicago/Turabian StyleDong, Zeyu, Shangkun Jin, Yupeng Hao, Ting Zhao, Haihong Shang, Zhiyuan Zhang, Lei Fang, Zhihong Zheng, and Jun Li. 2025. "Transposon Dynamics Drive Genome Evolution and Regulate Genetic Mechanisms of Agronomic Traits in Cotton" Plants 14, no. 16: 2509. https://doi.org/10.3390/plants14162509
APA StyleDong, Z., Jin, S., Hao, Y., Zhao, T., Shang, H., Zhang, Z., Fang, L., Zheng, Z., & Li, J. (2025). Transposon Dynamics Drive Genome Evolution and Regulate Genetic Mechanisms of Agronomic Traits in Cotton. Plants, 14(16), 2509. https://doi.org/10.3390/plants14162509





