Screening of Alfalfa Varieties Resistant to Phytophthora cactorum and Related Resistance Mechanism
Abstract
1. Introduction
2. Results
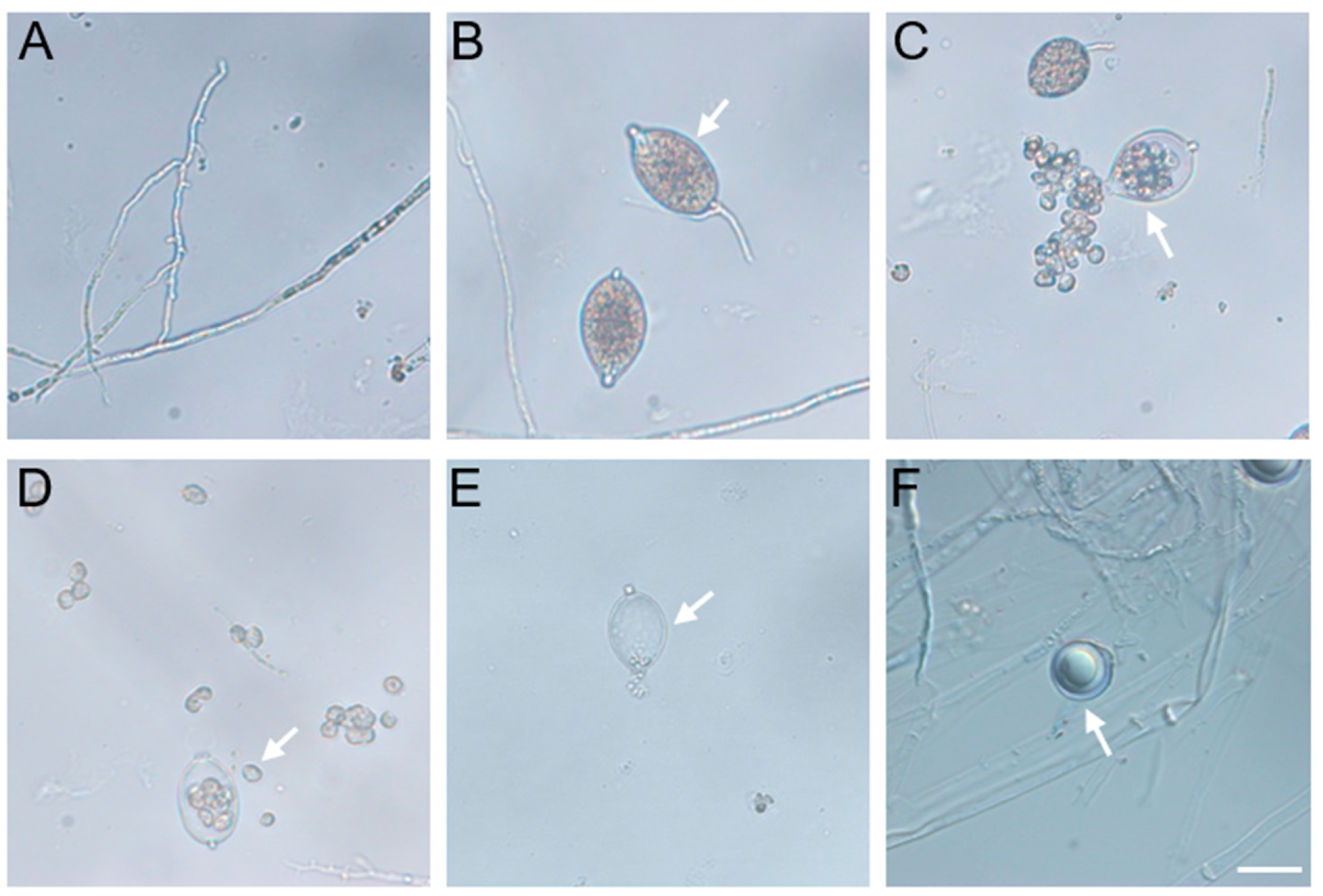
2.1. Isolation and Identification of Alfalfa Root Rot Pathogen P. cactorum
2.2. Pathogenicity Test Confirmed That P. cactorum Caused Severe Seed and Root Rot of Alfalfa
2.3. Staining Observation on the Infection of Alfalfa by P. cactorum
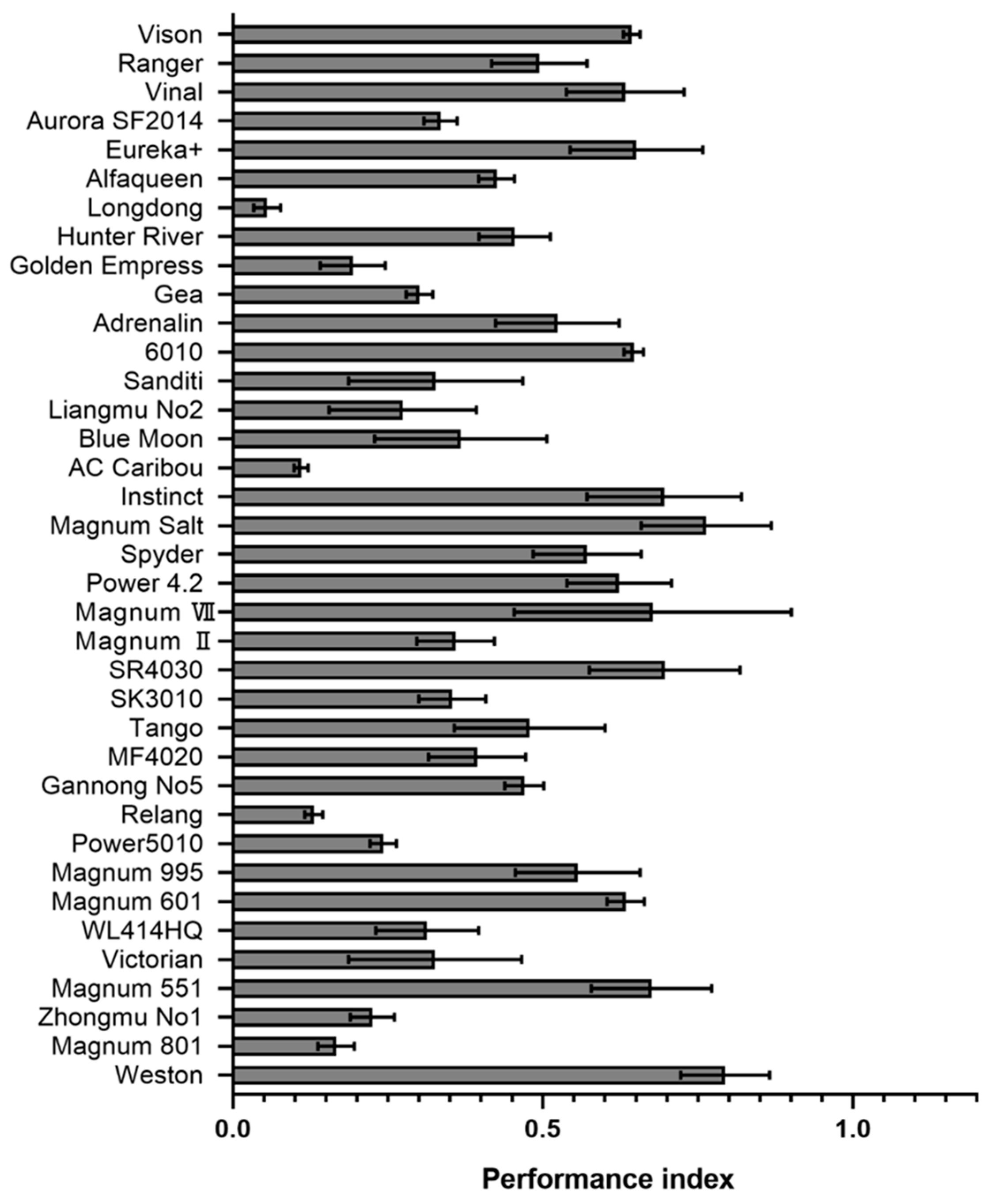
2.4. Evaluation of the Resistance Level of Different Alfalfa Varieties to P. cactorum
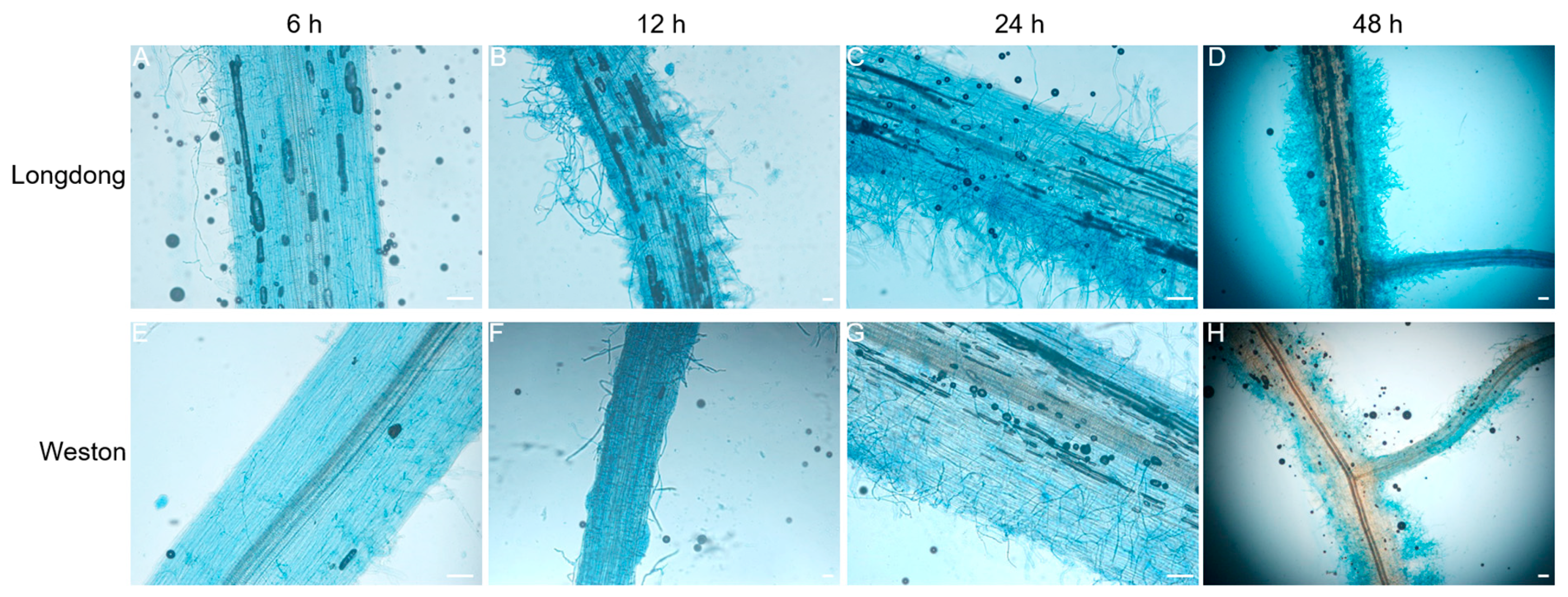
2.5. Microscopic Observation of Resistant or Susceptible Alfalfa Varieties Infected by P. cactorum at Different Time Points
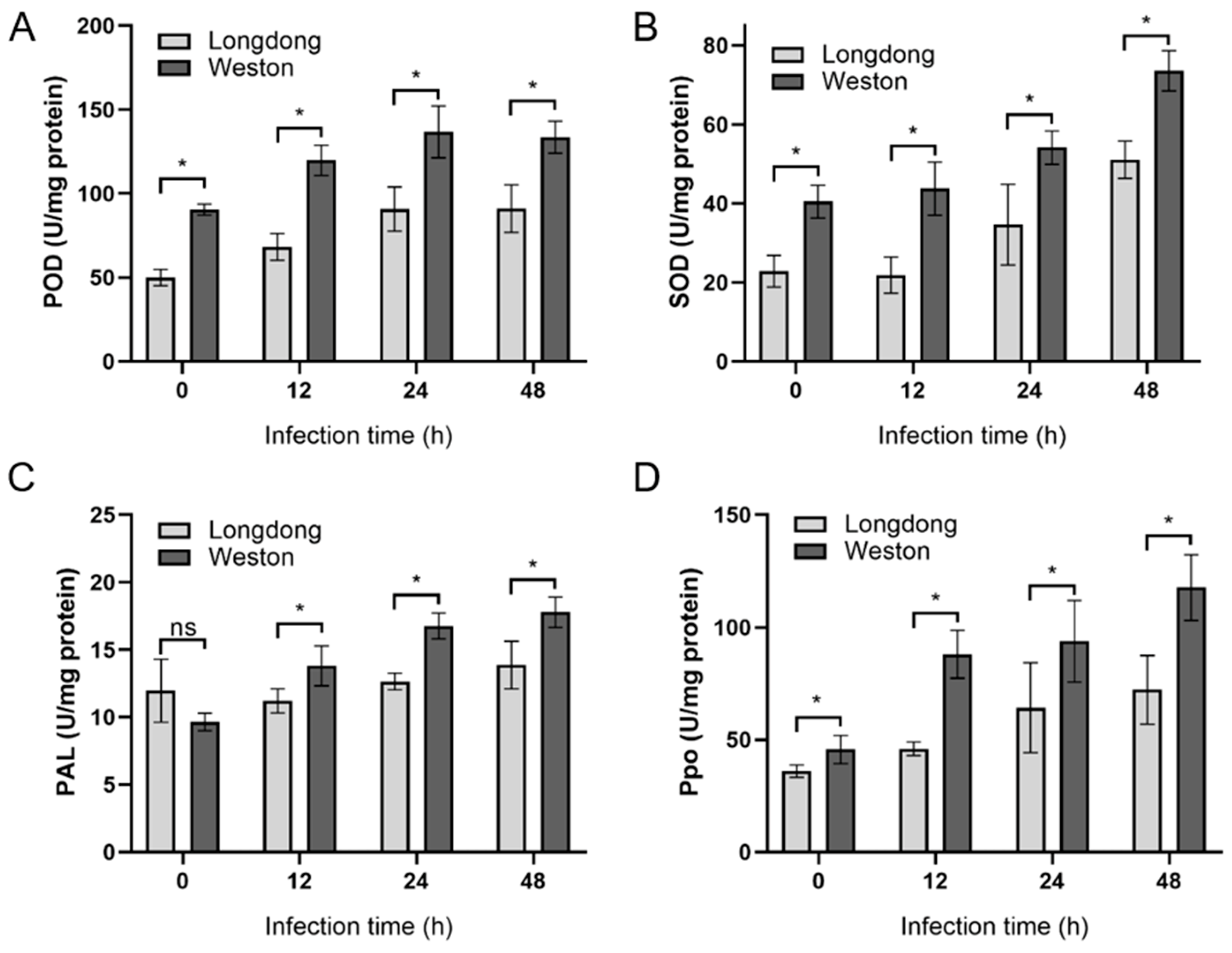
2.6. Antioxidant Enzyme Activities of Resistant or Susceptible Alfalfa Varieties Infected by P. cactorum at Different Time Points
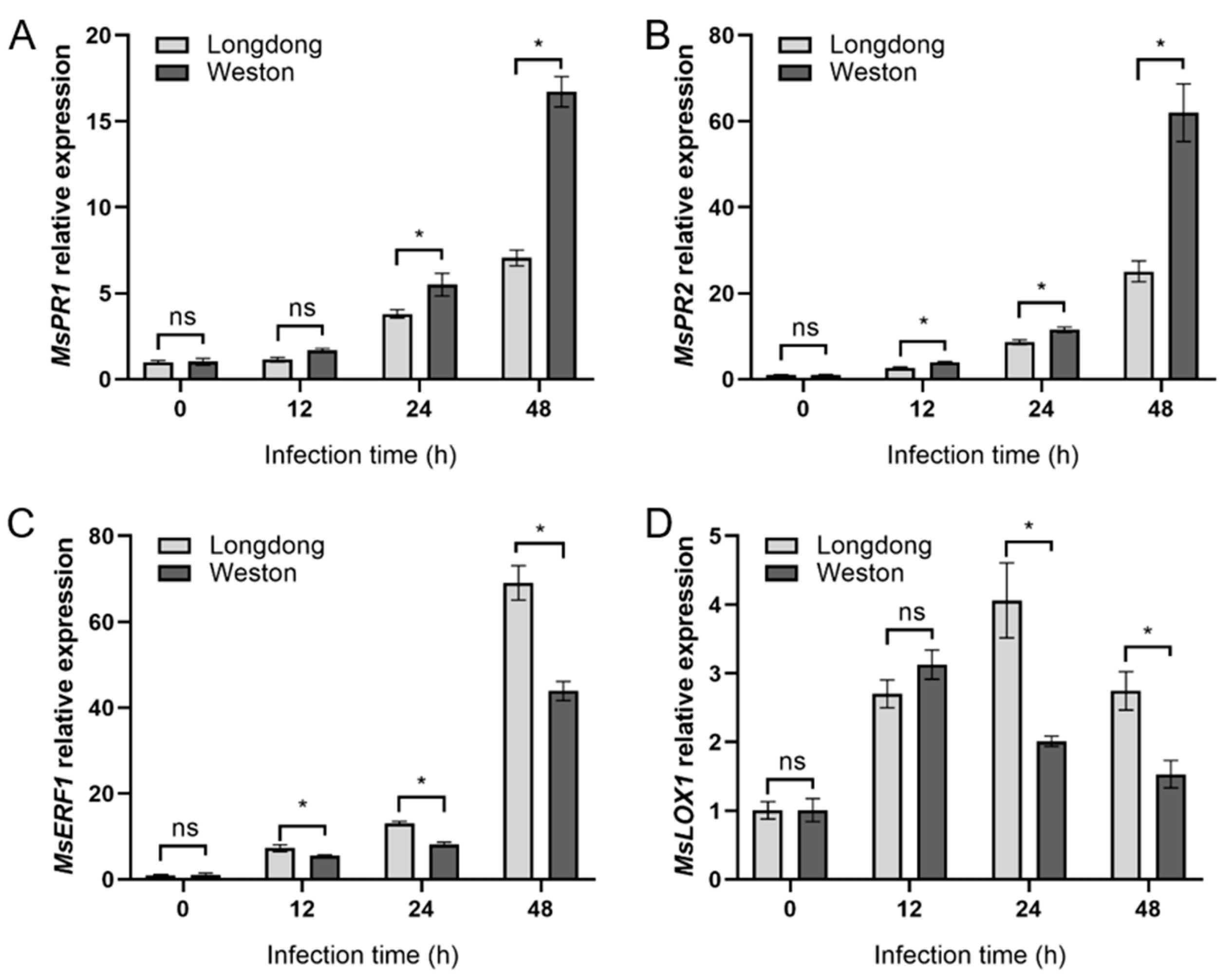
2.7. Relative Expression of Phytohormone-Related Genes in Resistant or Susceptible Alfalfa Varieties Infected by P. cactorum at Different Time Points
3. Discussion
4. Materials and Methods
4.1. Sample Collection
4.2. Isolation and Identification
4.3. Pathogenicity Test
4.4. Source of Tested Alfalfa Cultivars
4.5. qRT-PCR Analysis
4.6. Microscopic Examination
4.7. Phylogenetic Analysis of P. cactorum
4.8. Measurements of Plant Antioxidant Enzyme Activities
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Shi, S.; Nan, L.; Smith, K. The current status, problems, and prospects of Alfalfa (Medicago sativa L.) breeding in China. Agronomy 2017, 7, 1. [Google Scholar] [CrossRef]
- Fang, X.; Zhang, C.; Wang, Z.; Duan, T.; Yu, B.; Jia, X.; Pang, J.; Ma, L.; Wang, Y.; Nan, Z. Co-infection by soil-borne fungal pathogens alters disease responses among diverse alfalfa varieties. Front. Microbiol. 2021, 12, 664385. [Google Scholar] [CrossRef]
- Yang, B.; Zhao, Y.; Guo, Z. Research Progress and Prospect of Alfalfa Resistance to Pathogens and Pests. Plants 2022, 11, 2008. [Google Scholar] [CrossRef] [PubMed]
- Berg, L.E.; Miller, S.S.; Dornbusch, M.R.; Samac, D.A. Seed rot and damping-off of Alfalfa in minnesota caused by Pythium and Fusarium species. Plant Dis. 2017, 101, 1860–1867. [Google Scholar] [CrossRef] [PubMed]
- Hancock, J. Seedling diseases of alfalfa in California. Plant Dis. 1983, 67, 1203–1208. [Google Scholar] [CrossRef]
- Yang, B.; Zhao, Y.; Lu, Y.; Tao, M.; Wang, Y.; Guo, Z. First report of Alfalfa root rot caused by Fusarium commune in China. Plant Dis. 2022, PDIS-06. [Google Scholar] [CrossRef]
- Whitham, S.A.; Qi, M.; Innes, R.W.; Ma, W.; Lopes-Caitar, V.; Hewezi, T. Molecular soybean-pathogen interactions. Rev. Phytopathol. 2016, 54, 443–468. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Asano, T.; Suga, H.; Kageyama, K. A multiplex PCR for the detection of Phytophthora nicotianae and P. cactorum, and a survey of their occurrence in strawberry production areas of Japan. Plant Dis. 2011, 95, 1270–1278. [Google Scholar] [CrossRef]
- Chen, X.R.; Huang, S.X.; Zhang, Y.; Sheng, G.L.; Zhang, B.Y.; Li, Q.Y.; Zhu, F.; Xu, J.Y. Transcription profiling and identification of infection-related genes in Phytophthora cactorum. Mol. Genet. Genom. 2018, 293, 541–555. [Google Scholar] [CrossRef] [PubMed]
- Cai, W.; Tian, H.; Liu, J.; Fang, X.; Nan, Z. Phytophthora cactorum as a pathogen associated with root rot on Alfalfa (Medicago sativa) in China. Plant Dis. 2021, 105, 231. [Google Scholar] [CrossRef]
- Chen, X.R.; Brurberg, M.B.; Elameen, A.; Klemsdal, S.S.; Martinussen, I. Expression of resistance gene analogs in woodland strawberry (Fragaria vesca) during infection with Phytophthora cactorum. Mol. Genet. Genom. 2016, 291, 1967–1978. [Google Scholar] [CrossRef]
- Jones, J.D.; Dangl, J.L. The plant immune system. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tyler, B.M.; Wang, Y. Defense and counterdefense during plant-pathogenic oomycete infection. Annu. Rev. Microbiol. 2019, 73, 667–696. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Mir, Z.A.; Tyagi, A.; Mehari, H.; Meena, R.P.; Bhat, J.A.; Yadav, P.; Papalou, P.; Rawat, S.; Grover, A. Overexpression of NPR1 in Brassica juncea confers broad spectrum resistance to fungal pathogens. Front. Plant Sci. 2017, 8, 1693. [Google Scholar] [CrossRef]
- Bari, R.; Jones, J.D. Role of plant hormones in plant defence responses. Plant Mol. Biol. 2009, 69, 473–488. [Google Scholar] [CrossRef]
- Alizadeh-Moghaddam, G.; Rezayatmand, Z.; Esfahani, M.N.; Khozaei, M. Bio-genetic analysis of resistance in tomato to early blight disease, Alternaria alternata. Phytochemistry 2020, 179, 112486. [Google Scholar] [CrossRef]
- Khatediya, N.K.; Parmar, D.V.; Mahatma, M.K.; Pareek, M. Increased accumulation of phenolic metabolites in groundnut (Arachis hypogaea L.) genotypes contribute to defense against Sclerotium rolfsii infection. Arch. Phytopathol. Plant Prot. 2018, 51, 530–549. [Google Scholar] [CrossRef]
- Wildermuth, M.; Dewdney, J.; Wu, G.; Ausubel, F. Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 2001, 414, 562–565. [Google Scholar] [CrossRef]
- Adam, A.L.; Nagy, Z.A.; Katay, G.; Mergenthaler, E.; Viczian, O. Signals of systemic immunity in plants: Progress and open questions. Int. J. Mol. Sci. 2018, 19, 1146. [Google Scholar] [CrossRef]
- Spoel, S.H.; Dong, X. Making sense of hormone crosstalk during plant immune responses. Cell Host Microbe 2008, 3, 348–351. [Google Scholar] [CrossRef] [PubMed]
- Ma, K.W.; Ma, W. Phytohormone pathways as targets of pathogens to facilitate infection. Plant Mol. Biol. 2016, 91, 713–725. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Han, X.; Feng, D.; Yuan, D.; Huang, L.J. Signaling crosstalk between salicylic acid and ethylene/jasmonate in plant defense: Do we understand what they are whispering? Int. J. Mol. Sci. 2019, 20, 671. [Google Scholar] [CrossRef] [PubMed]
- Thomma, B.P.; Nelissen, I.; Eggermont, K.; Broekaert, W.F. Deficiency in phytoalexin production causes enhanced susceptibility of Arabidopsis thaliana to the fungus Alternaria brassicicola. Plant J. 1999, 19, 163–171. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Wang, T.; Cheng, Y.; Wang, L.; Zhang, J.; JIing, H.; Chong, K. Current situation and prospect of forage breeding in China. Bull. Chin. Acad. Sci. 2021, 36, 660–665. [Google Scholar] [CrossRef]
- Liu, Z.; Zhou, Q.; Liu, W.; Zhang, J.; Xie, W.; Fang, L.; Wang, Y.; Nan, Z. Some scientific issues of forage breeding in China. Acta Prataculturae Sin. 2021, 30, 184–193. [Google Scholar]
- Abbas, A.; Mubeen, M.; Sohail, M.A.; Solanki, M.K.; Hussain, B.; Nosheen, S.; Kashyap, B.K.; Zhou, L.; Fang, X. Root rot a silent alfalfa killer in China: Distribution, fungal, and oomycete pathogens, impact of climatic factors and its management. Front Microbiol. 2022, 13, 961794. [Google Scholar] [CrossRef]
- Yang, B.; Wang, Y.; Guo, B.; Jing, M.; Zhou, H.; Li, Y.; Wang, H.; Huang, J.; Wang, Y.; Ye, W.; et al. The Phytophthora sojae RXLR effector Avh238 destabilizes soybean Type2 GmACSs to suppress ethylene biosynthesis and promote infection. New Phytol. 2019, 222, 425–437. [Google Scholar] [CrossRef]
- Qiu, X.; Kong, L.; Chen, H.; Lin, Y.; Tu, S.; Wang, L.; Chen, Z.; Zeng, M.; Xiao, J.; Yuan, P.; et al. The Phytophthora sojae nuclear effector PsAvh110 targets a host transcriptional complex to modulate plant immunity. Plant Cell 2023, 35, 574–597. [Google Scholar] [CrossRef]
- Lin, X.; Olave-Achury, A.; Heal, R.; Pais, M.; Witek, K.; Ahn, H.K.; Zhao, H.; Bhanvadia, S.; Karki, H.S.; Song, T.; et al. A potato late blight resistance gene protects against multiple Phytophthora species by recognizing a broadly conserved RXLR-WY effector. Mol. Plant 2022, 15, 1457–1469. [Google Scholar] [CrossRef]
- Vera-Estrella, R.; Barkla, B.; Higgins, V.; Blumwald, E. Plant defense response to fungal pathogens (activation of host-plasma membrane H+-ATPase by elicitor-induced enzyme dephosphorylating). Plant Physiol. 1994, 104, 209–215. [Google Scholar] [CrossRef]
- Paranidharan, V.; Palaniswami, A.; Vidhyasekaran, P.; Velazhahan, R. Induction of enzymatic scavengers of active oxygen species in rice in response to infection by Rhizoctonia solani. Acta Physiol. Plant. 2003, 25, 91–96. [Google Scholar] [CrossRef]
- Robert-Seilaniantz, A.; Grant, M.; Jones, J.D. Hormone crosstalk in plant disease and defense: More than just jasmonate-salicylate antagonism. Annu. Rev. Phytopathol. 2011, 49, 317–343. [Google Scholar] [CrossRef] [PubMed]
- Berrocal-Lobo, M.; Molina, A.; Solano, R. Constitutive expression of ETHYLENE-RESPONSE-FACTOR1 in Arabidopsis confers resistance to several necrotrophic fungi. Plant J. 2002, 29, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Leah, R.; Tommerup, H.; Svendsen, I.; Mundy, J. Biochemical and molecular characterization of three barley seed proteins with antifungal properties. J. Biol. Chem. 1991, 266, 1564–1573. [Google Scholar] [CrossRef]
- Stintzi, A.; Heitz, T.; Prasad, V.; Wiedemann-Merdinoglu, S.; Kauffmann, S.; Geoffroy, P.; Legrand, M.; Fritig, B. Plant ‘pathogenesis-related’ proteins and their role in defense against pathogens. Biochimie 1993, 75, 687–706. [Google Scholar] [CrossRef]
- Gamir, J.; Darwiche, R.; Van’t Hof, P.; Choudhary, V.; Stumpe, M.; Schneiter, R.; Mauch, F. The sterol-binding activity of PATHOGENESIS-RELATED PROTEIN 1 reveals the mode of action of an antimicrobial protein. Plant J. 2017, 89, 502–509. [Google Scholar] [CrossRef]
- Breen, S.; Williams, S.J.; Outram, M.; Kobe, B.; Solomon, P.S. Emerging insights into the functions of pathogenesis-related protein 1. Trends Plant Sci. 2017, 22, 871–879. [Google Scholar] [CrossRef]
- Murray, M.; Thompson, W. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4325. [Google Scholar] [CrossRef]
- Rahman, M.Z.; Mukobata, H.; Suga, H.; Kageyama, K. Phytophthora asiatica sp. nov., a new species causing leaf and stem blight of kudzu in Japan. Mycol. Prog. 2014, 13, 759–769. [Google Scholar] [CrossRef]
- Mesny, F.; Miyauchi, S.; Thiergart, T.; Pickel, B.; Atanasova, L.; Karlsson, M.; Huttel, B.; Barry, K.W.; Haridas, S.; Chen, C.; et al. Genetic determinants of endophytism in the Arabidopsis root mycobiome. Nat. Commun. 2021, 12, 7227. [Google Scholar] [CrossRef]
- Zhuo, C.; Liang, L.; Zhao, Y.; Guo, Z.; Lu, S. A cold responsive ethylene responsive factor from Medicago falcata confers cold tolerance by up-regulation of polyamine turnover, antioxidant protection, and proline accumulation. Plant Cell Env. 2018, 41, 2021–2032. [Google Scholar] [CrossRef] [PubMed]
- Soheili-Moghaddam, B.; Nasr-Esfahani, M.; Mousanejad, S.; Hassanzadeh-Khankahdani, H.; Karbalaie-Khiyavie, H. Biochemical defense mechanism associated with host-specific disease resistance pathways against Rhizoctonia solani AG3-PT potatoes canker disease. Planta 2022, 257, 13. [Google Scholar] [CrossRef] [PubMed]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tao, M.; Zhao, Y.; Hu, T.; Zhang, Q.; Feng, H.; Lu, Y.; Guo, Z.; Yang, B. Screening of Alfalfa Varieties Resistant to Phytophthora cactorum and Related Resistance Mechanism. Plants 2023, 12, 702. https://doi.org/10.3390/plants12040702
Tao M, Zhao Y, Hu T, Zhang Q, Feng H, Lu Y, Guo Z, Yang B. Screening of Alfalfa Varieties Resistant to Phytophthora cactorum and Related Resistance Mechanism. Plants. 2023; 12(4):702. https://doi.org/10.3390/plants12040702
Chicago/Turabian StyleTao, Menghuan, Yao Zhao, Tianxue Hu, Quan Zhang, Hui Feng, Yiwen Lu, Zhenfei Guo, and Bo Yang. 2023. "Screening of Alfalfa Varieties Resistant to Phytophthora cactorum and Related Resistance Mechanism" Plants 12, no. 4: 702. https://doi.org/10.3390/plants12040702
APA StyleTao, M., Zhao, Y., Hu, T., Zhang, Q., Feng, H., Lu, Y., Guo, Z., & Yang, B. (2023). Screening of Alfalfa Varieties Resistant to Phytophthora cactorum and Related Resistance Mechanism. Plants, 12(4), 702. https://doi.org/10.3390/plants12040702







