Bioinformatic Analysis of the BTB Gene Family in Paulownia fortunei and Functional Characterization in Response to Abiotic and Biotic Stresses
Abstract
:1. Introduction
2. Results
2.1. Understanding the PfBTB Gene Family and Examining the Properties of Proteins
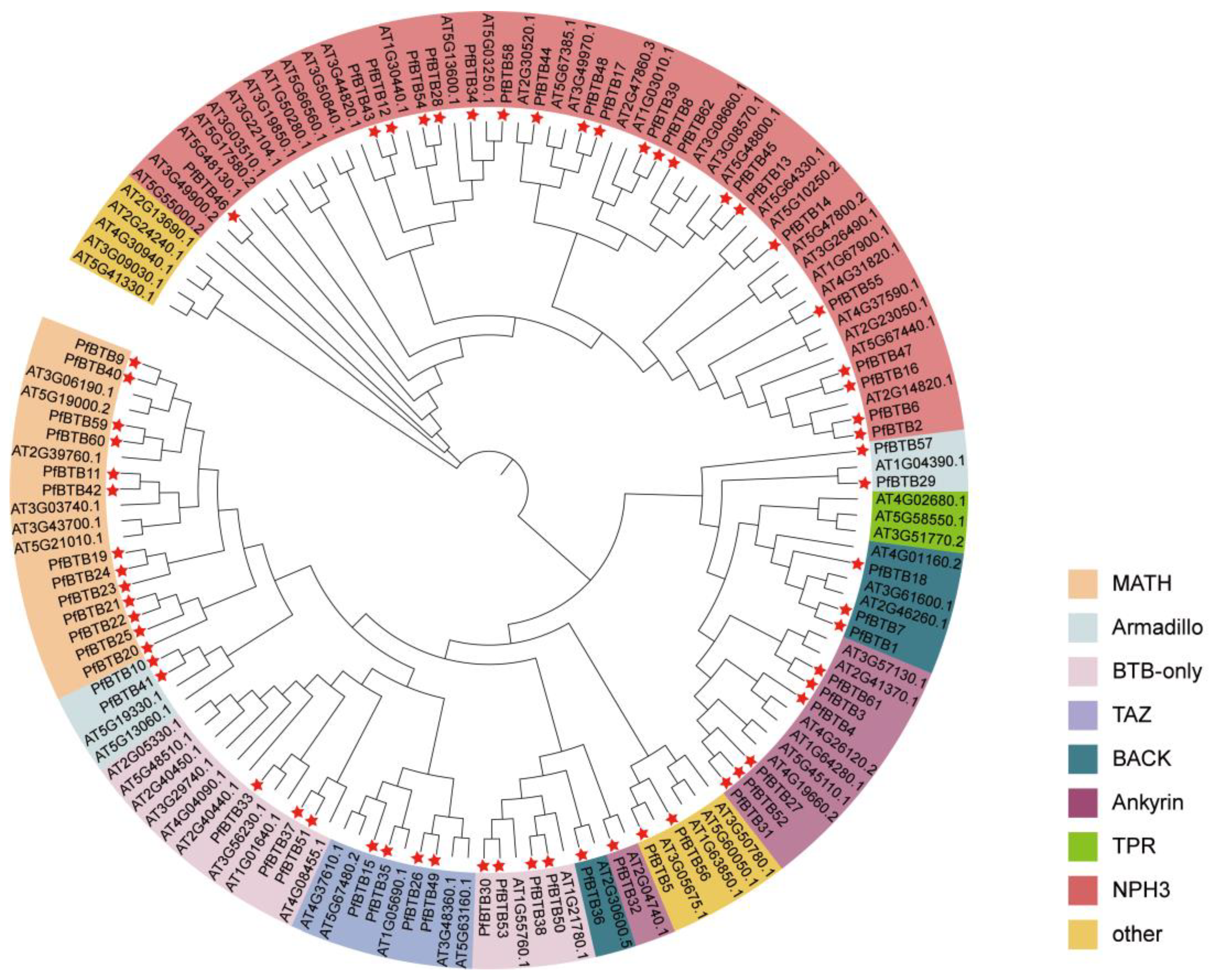
2.2. Analysis of the Phylogenetic Tree of Members of the PfBTB Gene Family
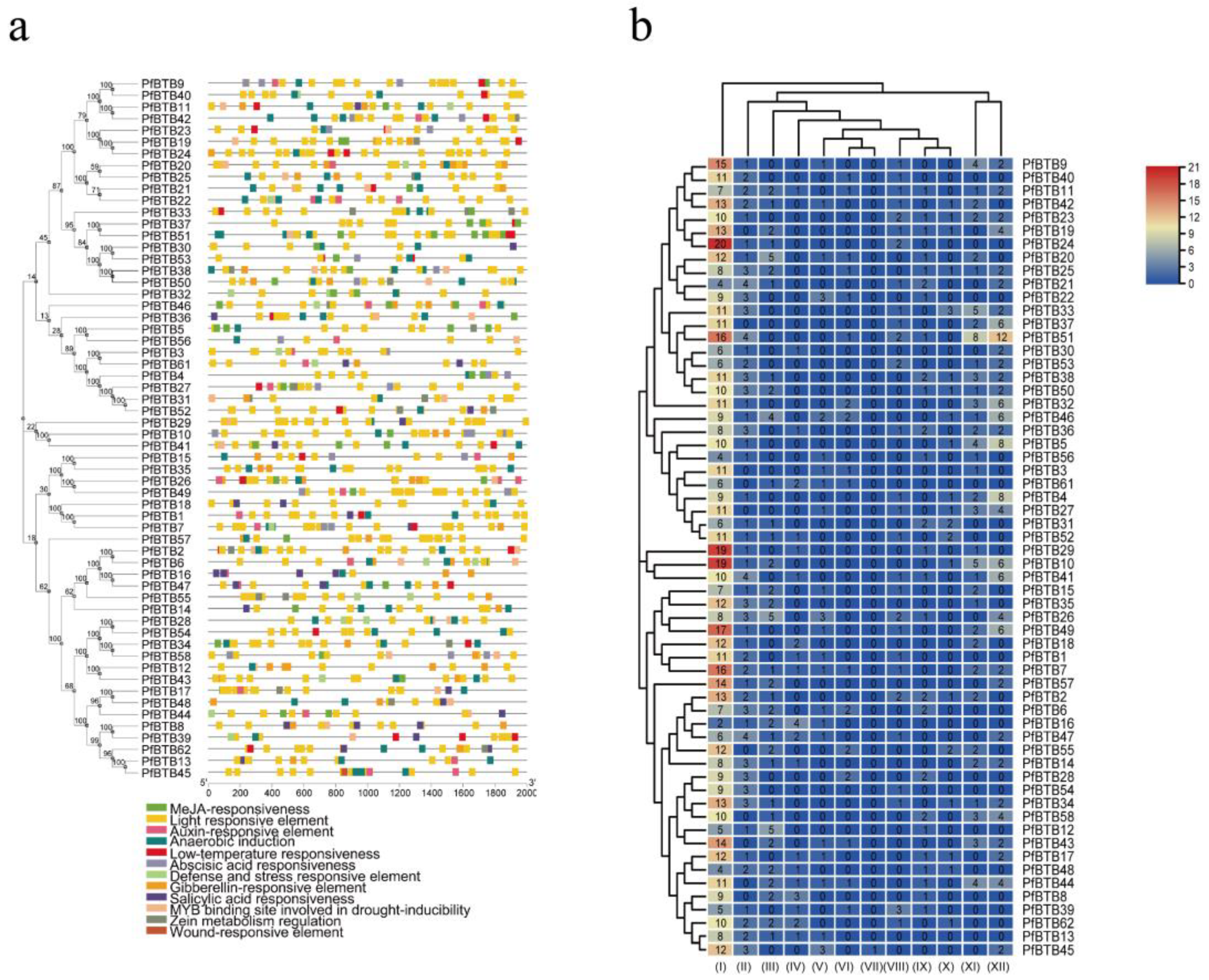
2.3. Gene Structure and Conserved Motif Analysis
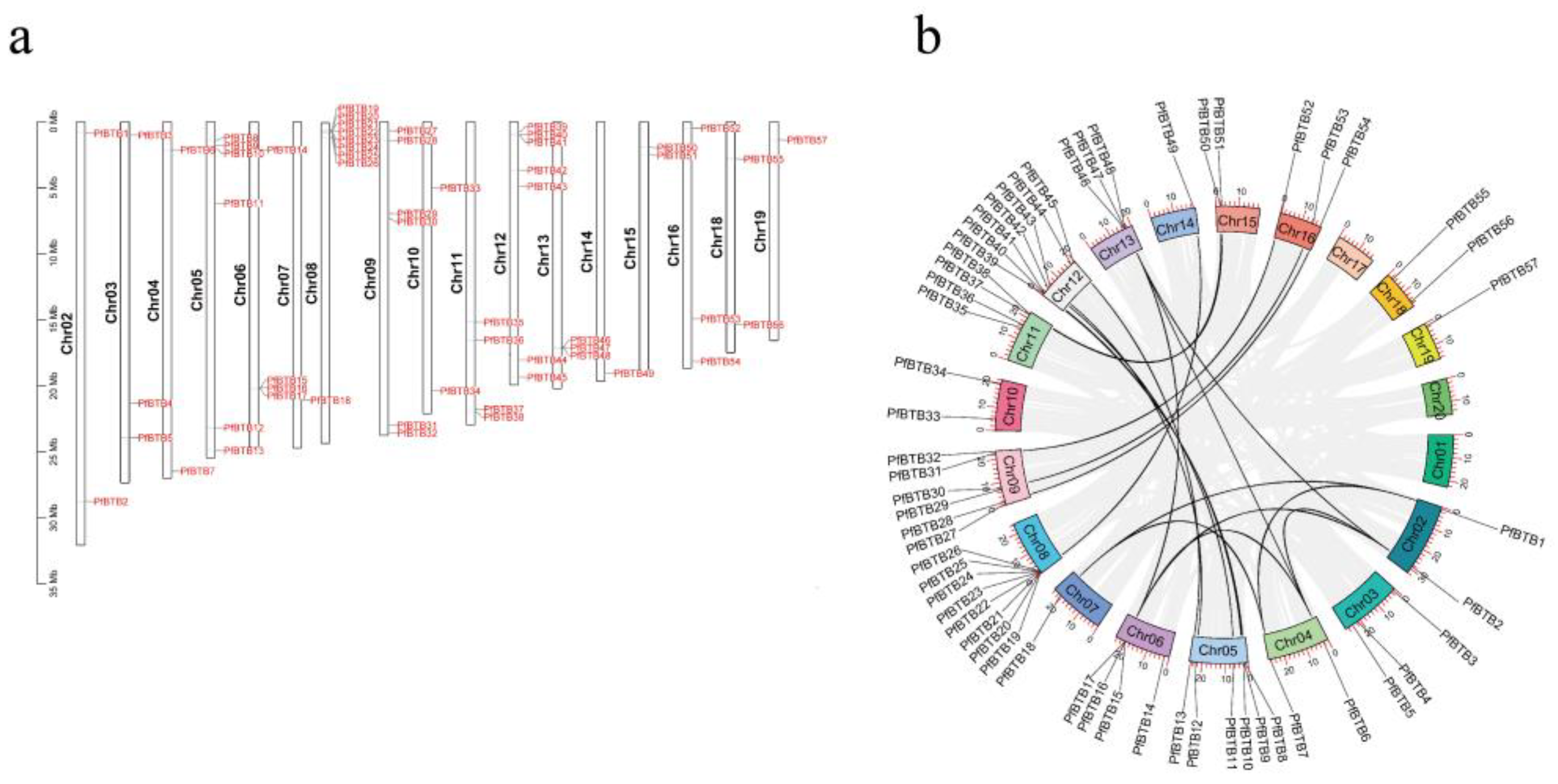
2.4. PfBTB Gene Locations on the Chromosome and Homologous Gene Analysis
2.5. Cis-Acting Element Prediction of PfBTB Genes
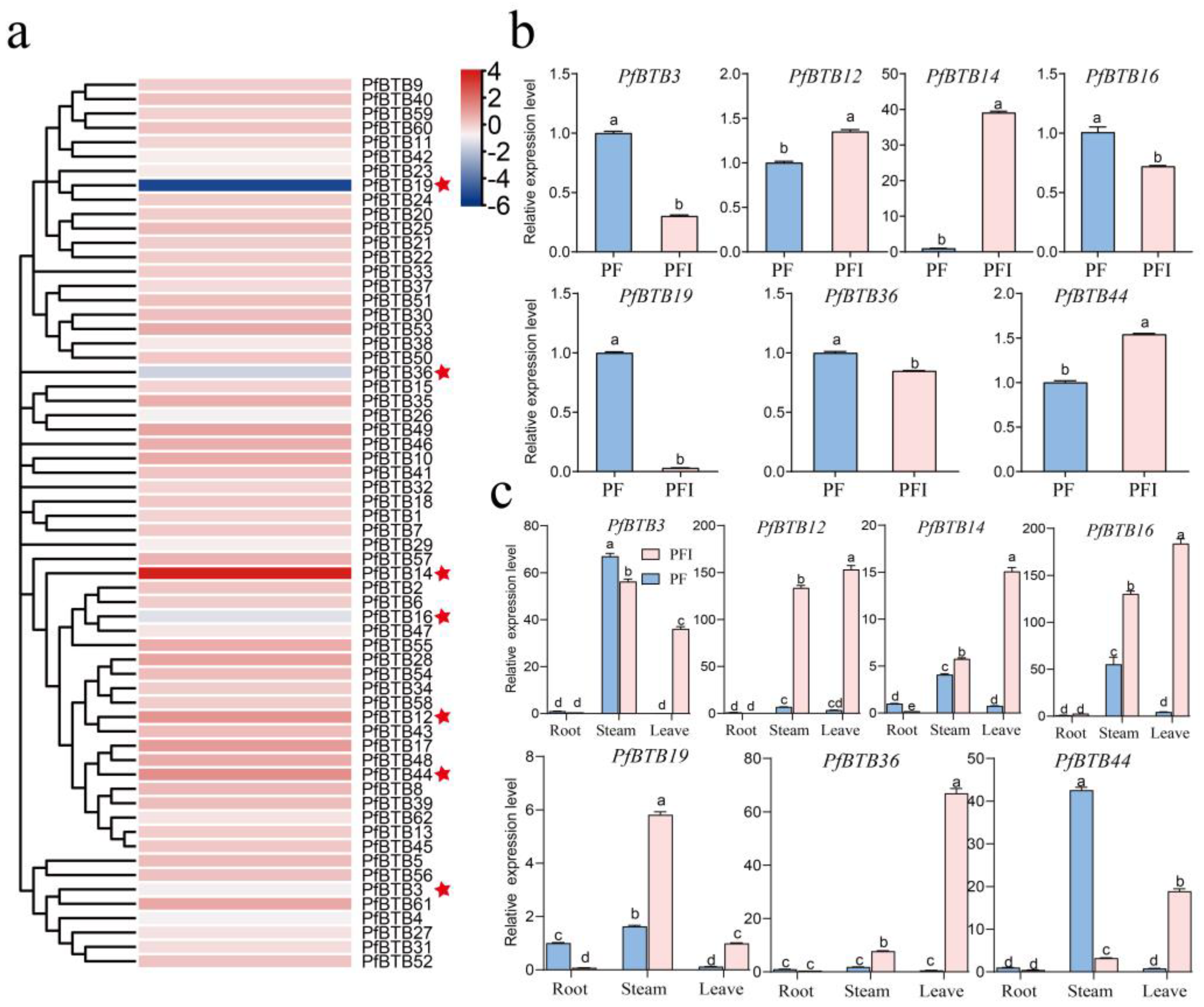
2.6. PfBTB Genes Response to Phytoplasma and Expression Patterns in Different Tissues/Organs
2.7. Expression Patterns with Different Stress Treatments
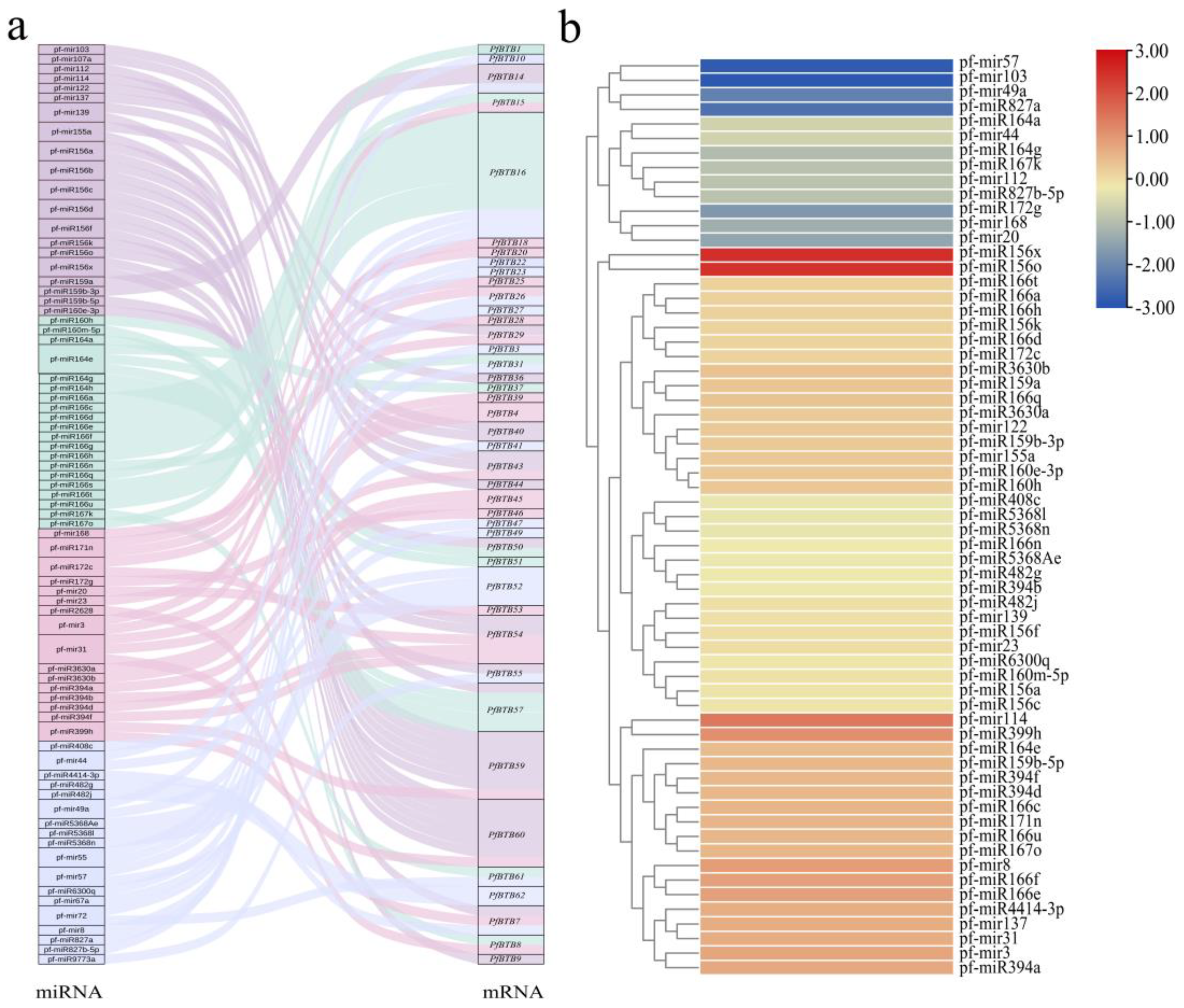
2.8. Association Analysis of PfBTB and miRNA
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Treatment
4.2. Identification of BTB Gene Family Members in P. fortunei’s Genome
4.3. PfBTB Proteins Underwent Phylogenetic Analysis
4.4. Analysis of Gene Structures, Motifs, and Cis-Acting Elements
4.5. Chromosomal Localization and Synteny Analysis
4.6. Prediction of the miRNA and PfBTB Targeting Relationship
4.7. Investigation of PfBTB Expression Patterns
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Figueroa, P.; Gusmaroli, G.; Serino, G.; Habashi, J.; Ma, L.; Shen, Y.; Feng, S.; Bostick, M.; Callis, J.; Hellmann, H.; et al. Arabidopsis has two redundant Cullin3 proteins that are essential for embryo development and that interact with RBX1 and BTB proteins to form multisubunit E3 ubiquitin ligase complexes in vivo. Plant Cell 2005, 17, 1180–1195. [Google Scholar] [CrossRef] [PubMed]
- Weber, H.; Bernhardt, A.; Dieterle, M.; Hano, P.; Mutlu, A.; Estelle, M.; Genschik, P.; Hellmann, H. Arabidopsis AtCUL3a and AtCUL3b form complexes with members of the BTB/POZ-MATH protein family. Plant Physiol. 2005, 137, 83–93. [Google Scholar] [CrossRef]
- Weber, H.; Hellmann, H. Arabidopsis thaliana BTB/ POZ-MATH proteins interact with members of the ERF/AP2 transcription factor family. FEBS J. 2009, 276, 6624–6635. [Google Scholar] [CrossRef] [PubMed]
- Gingerich, D.J.; Hanada, K.; Shiu, S.H.; Vierstra, R.D. Large-scale, lineage-specific expansion of a bric-a-brac/tramtrack/broad complex ubiquitin-ligase gene family in rice. Plant Cell 2007, 19, 2329–2348. [Google Scholar] [CrossRef]
- Li, J.; Su, X.; Wang, Y.; Yang, W.; Pan, Y.; Su, C.; Zhang, X. Genome-wide identification and expression analysis of the BTB domain-containing protein gene family in tomato. Genes Genom. 2018, 40, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Juranič, M.; Srilunchang, K.O.; Krohn, N.G.; Leljak-Levanic, D.; Sprunck, S.; Dresselhaus, T. Germline-specific MATH-BTB substrate adaptor MAB1 regulates spindle length and nuclei identity in maize. Plant Cell 2012, 24, 4974–4991. [Google Scholar] [CrossRef] [PubMed]
- Tan, B.; Lian, X.; Cheng, J.; Zeng, W.; Zheng, X.; Wang, W.; Ye, X.; Li, J.; Li, Z.; Zhang, L.; et al. Genome-wide identification and transcriptome profiling reveal that E3 ubiquitin ligase genes relevant to ethylene, auxin and abscisic acid are differentially expressed in the fruits of melting flesh and stony hard peach varieties. BMC Genom. 2019, 20, 892. [Google Scholar] [CrossRef]
- Adams, J.; Kelso, R.; Cooley, L. The kelch repeat superfamily of proteins: Propellers of cell function. Trends Cell Biol. 2000, 10, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Puccetti, E.; Sennewald, B.; Fosca-Ferrara, F.; Boehrer, S.; Bianchini, A.; Hoelzer, D.; Ottmann, O.G.; Nervi, C.; Ruthardt, M. Down-stream regions of the POZ-domain influence the interaction of the t(11;17)-associated PLZF/RARalpha fusion protein with the histone-deacetylase recruiting co-repressor complex. Hematol. J. 2001, 2, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Rivero, F.; Dislich, H.; Glockner, G.; Noegel, A.A. The Dictyostelium discoideum family of Rho-related proteins. Nucleic Acids Res. 2001, 29, 1068–1079. [Google Scholar] [CrossRef]
- Stogios, P.J.; Prive, G.G. The BACK domain in BTB-kelch proteins. Trends Biochem. Sci. 2004, 29, 634–637. [Google Scholar] [CrossRef] [PubMed]
- Salas-Vidal, E.; Meijer, A.H.; Cheng, X.; Spaink, H.P. Genomic annotation and expression analysis of the zebrafish Rho small GTPase family during development and bacterial infection. Genomics 2005, 86, 25–37. [Google Scholar] [CrossRef] [PubMed]
- Stogios, P.J.; Downs, G.S.; Jauhal, J.J.S.; Nandra, S.K.; Prive, G.G. Sequence and structural analysis of BTB domain proteins. Genome Biol. 2005, 6, R82. [Google Scholar] [CrossRef] [PubMed]
- DiBello, P.R.; Withers, D.A.; Bayer, C.A.; Fristrom, J.W.; Guild, G.M. The Drosophila Broad-Complex encodes a family of related proteins containing zinc fingers. Genetics 1991, 129, 385–397. [Google Scholar] [CrossRef] [PubMed]
- Zollman, S.; Godt, D.; Privé, G.G.; Couderc, J.L.; Laski, F.A. The BTB domain, found primarily in zinc finger proteins, defines an evolutionarily conserved family that includes several developmentally regulated genes in Drosophila. Proc. Natl. Acad. Sci. USA 1994, 91, 10717–10721. [Google Scholar] [CrossRef]
- Bomont, P.; Cavalier, L.; Blondeau, F.; Ben Hamida, C.; Belal, S.; Tazir, M.; Demir, E.; Topaloglu, H.; Korinthenberg, R.; Tüysüz, B.; et al. The gene encoding gigaxonin, a new member of the cytoskeletal BTB/kelch repeat family, is mutated in giant axonal neuropathy. Nat. Genet. 2000, 26, 370–374. [Google Scholar] [CrossRef] [PubMed]
- Collins, T.; Stone, J.R.; Williams, A.J. All in the family: The BTB/POZ, KRAB, and SCAN domains. Mol. Cell. Biol. 2001, 21, 3609–3615. [Google Scholar] [CrossRef]
- Ji, X.L.; Li, H.L.; Qiao, Z.W.; Zhang, J.C.; Sun, W.J.; You, C.X.; Hao, Y.J.; Wang, X.F. The BTB protein MdBT2 recruits auxin signaling components to regulate adventitious root formation in apple. Plant Physiol. 2022, 189, 1005–1020. [Google Scholar] [CrossRef]
- Christians, M.J.; Gingerich, D.J.; Hansen, M.; Binder, B.M.; Kieber, J.J.; Vierstra, R.D. The BTB ubiquitin ligases ETO1, EOL1 and EOL2 act collectively to regulate ethylene biosynthesis in Arabidopsis by controlling type-2 ACC synthase levels. Plant J. 2009, 57, 332–345. [Google Scholar] [CrossRef]
- Lin, W.C.; Lu, C.F.; Wu, J.W.; Cheng, M.L.; Lin, Y.M.; Yang, N.S.; Black, L.; Green, S.K.; Wang, J.F.; Cheng, C.P. Transgenic tomato plants expressing the Arabidopsis NPR1 gene display enhanced resistance to a spectrum of fungal and bacterial diseases. Transgenic Res. 2004, 13, 567–581. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.X.; Cao, L.; Li, J.; Duan, C.J.; Luo, X.M.; Le, N.; Wei, H.; Liang, S.; Chu, C.; Pan, Q. Involvement of OsNPR1/NH1 in rice basal resistance to blast fungus Magnaporthe oryzae. Eur. J. Plant Pathol. 2011, 131, 221–235. [Google Scholar] [CrossRef]
- Yuan, Y.; Zhong, S.; Li, Q.; Zhu, Z.; Lou, Y.; Wang, L.; Wang, J.; Wang, M.; Li, Q.; Yang, D.; et al. Functional analysis of rice NPR1-like genes reveals that OsNPR1/NH1 is the rice orthologue conferring disease resistance with enhanced herbivore susceptibility. Plant Biotechnol. J. 2007, 5, 313–324. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Sun, T.; Ao, K.; Peng, Y.; Zhang, Y.; Li, X.; Zhang, Y. Opposite Roles of Salicylic Acid Receptors NPR1 and NPR3/NPR4 in Transcriptional Regulation of Plant Immunity. Cell 2018, 173, 1454–1467.e1415. [Google Scholar] [CrossRef] [PubMed]
- Makandar, R.; Essig, J.S.; Schapaugh, M.A.; Trick, H.N.; Shah, J. Genetically engineered resistance to Fusarium head blight in wheat by expression of Arabidopsis NPR1. Mol. Plant-Microbe Interact. MPMI 2006, 19, 123–129. [Google Scholar] [CrossRef]
- Zhang, C.; Cheng, Q.; Wang, H.; Gao, H.; Fang, X.; Chen, X.; Zhao, M.; Wei, W.; Song, B.; Liu, S.; et al. GmBTB/POZ promotes the ubiquitination and degradation of LHP1 to regulate the response of soybean to Phytophthora sojae. Commun. Biol. 2021, 4, 372. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Li, G.; Zhang, L.; Xu, J.; Liu, S. Comprehensive genomic analysis and expression profiling of the BTB and TAZ (BT) genes in cucumber (Cucumis sativus L.). Czech J. Genet. Plant Breed. 2019, 56, 15–23. [Google Scholar] [CrossRef]
- Cao, Y.B.; Sun, G.L.; Zhai, X.Q.; Xu, P.L.; Ma, L.M.; Deng, M.J.; Zhao, Z.L.; Yang, H.B.; Dong, Y.P.; Shang, Z.H.; et al. Genomic insights into the fast growth of paulownias and the formation of Paulownia witches’ broom. Mol. Plant 2021, 14, 1668–1682. [Google Scholar] [CrossRef] [PubMed]
- Yue, H.N.; Wu, Y.F.; Shi, Y.Z.; Wu, K.K.; Li, Y.R. First Report of Paulownia Witches’-Broom Phytoplasma in China. Plant Dis. 2008, 92, 1134. [Google Scholar] [CrossRef]
- Yang, H.; Zhai, X.; Zhao, Z.; Fan, G. Comprehensive analyses of the SPL transcription factor family in Paulownia fortunei and their responses to biotic and abiotic stresses. Int. J. Biol. Macromol. 2023, 226, 1261–1272. [Google Scholar] [CrossRef]
- Zhao, X.; Li, B.; Zhai, X.; Liu, H.; Deng, M.; Fan, G. Genome-Wide Analysis of Specific PfR2R3-MYB Genes Related to Paulownia Witches’ Broom. Genes 2022, 14, 7. [Google Scholar] [CrossRef]
- Deng, M.; Dong, Y.; Xu, S.; Huang, S.; Zhai, X.; Fan, G. Genome-Wide Identification and Expression of the Paulownia fortunei MADS-Box Gene Family in Response to Phytoplasma Infection. Genes 2023, 14, 696. [Google Scholar] [CrossRef] [PubMed]
- Sharma, B.; Taganna, J. Genome-wide analysis of the U-box E3 ubiquitin ligase enzyme gene family in tomato. Sci. Rep. 2020, 10, 9581. [Google Scholar] [CrossRef] [PubMed]
- Grill, E.; Himmelbach, A. ABA signal transduction. Curr. Opin. Plant Biol. 1998, 1, 412–418. [Google Scholar] [CrossRef] [PubMed]
- Goyal, N.; Bhuria, M.; Verma, D.; Garewal, N.; Singh, K. Genome-Wide Identification of BTB Domain-Containing Gene Family in Grapevine (Vitis vinifera L.). Agriculture 2023, 13, 252. [Google Scholar] [CrossRef]
- Yang, Q.; Liu, Y.; Zhao, C.L.; Wang, X.; Ding, G.Z.; Li, Y.L.; Chen, L. Genome-Wide Identification and Expression Analysis of the BTB Domain-Containing Protein Gene Family in Sugar Beet. Agronomy 2022, 12, 253. [Google Scholar] [CrossRef]
- YuMei, H.; KeKe, L.; HuaiXia, Z.; GuoXin, C.; Muhammad, A.; Saeed Ul, H.; AiMin, W.; ZhenHui, G. Contribution of CaBPM4, a BTB Domain–Containing Gene, to the Response of Pepper to Phytophthora capsici Infection and Abiotic Stresses. Agronomy 2019, 9, 417. [Google Scholar] [CrossRef]
- Santner, A.; Estelle, M. Recent advances and emerging trends in plant hormone signalling. Nature 2009, 459, 1071–1078. [Google Scholar] [CrossRef]
- Johnson, C.; Boden, E.; Arias, J. Salicylic acid and NPR1 induce the recruitment of trans-activating TGA factors to a defense gene promoter in Arabidopsis. Plant Cell 2003, 15, 1846–1858. [Google Scholar] [CrossRef]
- Chico, J.M.; Lechner, E.; Fernandez-Barbero, G.; Canibano, E.; García-Casado, G.; Franco-Zorrilla, J.M.; Hammann, P.; Zamarreño, A.M.; García-Mina, J.M.; Rubio, V.; et al. CUL3(BPM) E3 ubiquitin ligases regulate MYC2, MYC3, and MYC4 stability and JA responses. Proc. Natl. Acad. Sci. USA 2020, 117, 6205–6215. [Google Scholar] [CrossRef]
- Silva, K.J.P.; Brunings, A.; Peres, N.A.; Mou, Z.L.; Folta, K.M. The Arabidopsis NPR1 gene confers broad-spectrum disease resistance in strawberry. Transgenic Res. 2015, 24, 693–704. [Google Scholar] [CrossRef]
- Kim, H.; Kim, S.H.; Seo, D.H.; Chung, S.; Kim, S.W.; Lee, J.S.; Kim, W.T.; Lee, J.H. ABA-HYPERSENSITIVE BTB/POZ PROTEIN 1 functions as a negative regulator in ABA-mediated inhibition of germination in Arabidopsis. Plant Mol. Biol. 2016, 90, 303–315. [Google Scholar] [CrossRef] [PubMed]
- Cai, G.; Wang, Y.; Tu, G.; Chen, P.; Luan, S.; Lan, W. Type A2 BTB Members Decrease the ABA Response during Seed Germination by Affecting the Stability of SnRK2.3 in Arabidopsis. Int. J. Mol. Sci. 2020, 21, 3153. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Zhao, Q.; Yang, Y.; Zhang, R.; Wang, X.; Zhang, T.; You, C.; Huo, H.; Hao, Y. Interaction of BTB-TAZ protein MdBT2 and DELLA protein MdRGL3a regulates nitrate-mediated plant growth. Plant Physiol. 2021, 186, 750–766. [Google Scholar] [CrossRef]
- Woodger, F.J.; Jacobsen, J.V.; Gubler, F. GMPOZ, a BTB/POZ domain nuclear protein, is a regulator of hormone responsive gene expression in barley aleurone. Plant Cell Physiol. 2004, 45, 945–950. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Zhai, H.; He, S.; Zhu, H.; Gao, S.; Xing, S.; Wei, Z.; Zhao, N.; Liu, Q. The Sweetpotato BTB-TAZ Protein Gene, IbBT4, Enhances Drought Tolerance in Transgenic Arabidopsis. Front. Plant Sci. 2020, 11, 877. [Google Scholar] [CrossRef] [PubMed]
- Gingerich, D.J.; Gagne, J.M.; Salter, D.W.; Hellmann, H.; Estelle, M.; Ma, L.G.; Vierstra, R.D. Cullins 3a and 3b assemble with members of the broad complex/tramtrack/bric-a-brac (BTB) protein family to form essential ubiquitin-protein ligases (E3s) in Arabidopsis. J. Biol. Chem. 2005, 280, 18810–18821. [Google Scholar] [CrossRef]
- Song, G.; Zhang, R.; Zhang, S.; Li, Y.; Gao, J.; Han, X.; Chen, M.; Wang, J.; Li, W.; Li, G. Response of microRNAs to cold treatment in the young spikes of common wheat. BMC Genom. 2017, 18, 212. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zhai, X.; Cao, Y.; Zhao, H.; Wang, Z.; Liu, H.; Fan, G. Transcriptome and Small RNA Sequencing Analysis Revealed Roles of PaWB-Related miRNAs and Genes in Paulownia fortunei. Forests 2018, 9, 397. [Google Scholar] [CrossRef]
- Fan, G.; Cao, X.; Zhao, Z.; Deng, M. Transcriptome analysis of the genes related to the morphological changes of Paulownia tomentosa plantlets infected with phytoplasma. Acta Physiol. Plant. 2015, 37, 202. [Google Scholar] [CrossRef]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Liu, Y.; Feng, J.; Chen, H.; He, Y.; et al. TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Mol. Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef] [PubMed]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Fan, G.; Wang, Z.; Zhai, X.; Cao, Y. ceRNA Cross-Talk in Paulownia Witches’ Broom Disease. Int. J. Mol. Sci. 2018, 19, 2463. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Niu, S.; Fan, G.; Deng, M.; Wang, Y. Genome-Wide Analysis of Gene and microRNA Expression in Diploid and Autotetraploid Paulownia fortunei (Seem) Hemsl. under Drought Stress by Transcriptome, microRNA, and Degradome Sequencing. Forests 2018, 9, 88. [Google Scholar] [CrossRef]


| Gene Name | Gene ID | Chromosome | Protein Length (aa) | Molecular Weight (Da) | Theoretical pI | Putative Subcellular Localization |
|---|---|---|---|---|---|---|
| PfBTB1 | Pfo02g001220 | Chr2 | 564 | 64,356.53 | 5.17 | nucl: 11, chlo: 1, plas: 1, golg: 1 |
| PfBTB2 | Pfo02g015880 | Chr2 | 625 | 69,264.02 | 6.7 | nucl: 7, cyto: 6, mito: 1 |
| PfBTB3 | Pfo03g001400 | Chr3 | 486 | 52,888.35 | 6.14 | chlo: 5, cyto: 5, nucl: 3, plas: 1 |
| PfBTB4 | Pfo03g008270 | Chr3 | 587 | 65,515.92 | 5.78 | cyto: 4, nucl: 3, E.R.: 3, vacu: 2, chlo: 1, |
| PfBTB5 | Pfo03g010830 | Chr3 | 496 | 55,425.03 | 4.64 | nucl: 12, cyto: 1, pero: 1 |
| PfBTB6 | Pfo04g002970 | Chr4 | 626 | 69,187.25 | 7.53 | nucl: 4, cyto: 4, chlo: 3, mito: 2, cysk: 1 |
| PfBTB7 | Pfo04g013910 | Chr4 | 572 | 64,898.01 | 5.07 | nucl: 10, chlo: 2, cyto: 1, plas: 1 |
| PfBTB8 | Pfo05g002190 | Chr5 | 623 | 69,677.81 | 8.84 | chlo: 6, cyto: 4, nucl: 3, mito: 1 |
| PfBTB9 | Pfo05g002850 | Chr5 | 410 | 45,607.9 | 6.17 | chlo: 6, mito: 3, nucl: 2, cyto: 2, plas: 1 |
| PfBTB10 | Pfo05g003240 | Chr5 | 666 | 73,530.9 | 5.62 | cyto: 11, nucl: 2, cysk: 1 |
| PfBTB11 | Pfo05g007860 | Chr5 | 403 | 44,661.49 | 5.68 | cyto: 9, nucl: 3, extr: 1, vacu: 1 |
| PfBTB12 | Pfo05g013850 | Chr5 | 632 | 71,008.93 | 7.25 | nucl: 4, cyto: 4, chlo: 3, mito: 1, plas: 1, |
| PfBTB13 | Pfo05g016000 | Chr5 | 607 | 68,367.01 | 6.25 | vacu: 7, chlo: 2, nucl: 2, mito: 1, plas: 1, |
| PfBTB14 | Pfo06g002150 | Chr6 | 639 | 72,922.16 | 7.54 | nucl: 9, chlo: 3, mito: 1, pero: 1 |
| PfBTB15 | Pfo06g010190 | Chr6 | 379 | 43,425.7 | 9.09 | cyto: 6, nucl: 5, chlo: 1, plas: 1 |
| PfBTB16 | Pfo06g010240 | Chr6 | 605 | 67,284.45 | 5.58 | cyto: 8, nucl: 3, chlo: 2, cysk: 1 |
| PfBTB17 | Pfo06g010330 | Chr6 | 619 | 68,873.48 | 6 | cyto: 5.5, cyto_nucl: 5, nucl: 3.5, chlo: 2 |
| PfBTB18 | Pfo07g012470 | Chr7 | 544 | 61,639.22 | 5.23 | nucl: 9, cyto: 2, plas: 1, cysk: 1, golg: 1 |
| PfBTB19 | Pfo08g000530 | Chr8 | 393 | 44,326.69 | 5.94 | nucl: 6, cyto: 6, chlo: 1, vacu: 1 |
| PfBTB20 | Pfo08g000540 | Chr8 | 365 | 41,226.8 | 7.54 | chlo: 8, mito: 5.5, cyto_mito: 3.5 |
| PfBTB21 | Pfo08g000630 | Chr8 | 227 | 24,879.48 | 6.35 | mito: 6.5, cyto_mito: 4, nucl: 3, chlo: 2 |
| PfBTB22 | Pfo08g000660 | Chr8 | 396 | 44,387.72 | 5.65 | chlo: 8.5, chlo_mito: 5.5, nucl: 3, mito: 1.5 |
| PfBTB23 | Pfo08g000690 | Chr8 | 411 | 46,884.72 | 7.21 | nucl: 7, mito: 6, chlo: 1 |
| PfBTB24 | Pfo08g000720 | Chr8 | 394 | 44,389.13 | 6.48 | cyto: 9, nucl: 3, mito: 2 |
| PfBTB25 | Pfo08g000730 | Chr8 | 382 | 42,939.41 | 7.21 | chlo: 12, mito: 1.5, cyto_mito: 1.5 |
| PfBTB26 | Pfo08g001180 | Chr8 | 358 | 41,041.9 | 9.52 | nucl: 8, mito: 3, plas: 1.5, golg_plas: 1.5 |
| PfBTB27 | Pfo09g000940 | Chr9 | 586 | 65,718.53 | 6.57 | cyto: 4, E.R.: 4, golg: 2, chlo: 1, nucl: 1 |
| PfBTB28 | Pfo09g001930 | Chr9 | 602 | 66,580.42 | 5.78 | nucl: 5, cyto: 4, chlo: 3, plas: 1, cysk: 1 |
| PfBTB29 | Pfo09g009710 | Chr9 | 1016 | 115,074.68 | 6.61 | plas: 6, E.R.: 5, nucl: 1, mito: 1, pero: 1 |
| PfBTB30 | Pfo09g010190 | Chr9 | 328 | 37,502.9 | 5.63 | cyto: 8, nucl: 4, mito: 1, plas: 1 |
| PfBTB31 | Pfo09g019460 | Chr9 | 603 | 67,411.24 | 6.07 | chlo: 8, cyto: 3, nucl: 2, extr: 1 |
| PfBTB32 | Pfo09g020220 | Chr9 | 596 | 67,308.42 | 4.92 | nucl: 10.5, cyto_nucl: 6.5, cyto: 1.5, chlo: 1 |
| PfBTB33 | Pfo10g006800 | Chr10 | 270 | 30,234.37 | 5.59 | nucl: 7, chlo: 4, mito: 2, cyto: 1 |
| PfBTB34 | Pfo10g012330 | Chr10 | 616 | 69,548.02 | 5.6 | chlo: 8, nucl: 2, cyto: 2, plas: 1, extr: 1 |
| PfBTB35 | Pfo11g007120 | Chr11 | 401 | 45,685.27 | 9.1 | nucl: 6, cysk: 3, chlo: 2, cyto: 2, plas: 1 |
| PfBTB36 | Pfo11g008220 | Chr11 | 804 | 92,144.78 | 5.44 | E.R.: 4, nucl: 3, cyto: 3, plas: 3, chlo: 1 |
| PfBTB37 | Pfo11g014130 | Chr11 | 256 | 28,694.63 | 4.9 | chlo: 6, cyto: 4, mito: 2, nucl: 1, plas: 1 |
| PfBTB38 | Pfo11g014430 | Chr11 | 327 | 37,633.11 | 5.62 | cyto: 9, chlo: 1, nucl: 1, mito: 1, cysk: 1 |
| PfBTB39 | Pfo12g001080 | Chr12 | 633 | 70,819.11 | 9.13 | cyto: 6, chlo: 5, nucl: 2, mito: 1 |
| PfBTB40 | Pfo12g001360 | Chr12 | 410 | 45,439.7 | 6.07 | chlo: 3, nucl: 3, cyto: 3, mito: 3, plas: 1 |
| PfBTB41 | Pfo12g001600 | Chr12 | 709 | 78,578.93 | 6.22 | cyto: 7, nucl: 4, mito: 1, vacu: 1, cysk: 1 |
| PfBTB42 | Pfo12g004960 | Chr12 | 404 | 44,534.33 | 5.88 | cyto: 7, nucl: 4, chlo: 1, mito: 1, cysk: 1 |
| PfBTB43 | Pfo12g006350 | Chr12 | 641 | 72,314.29 | 8.01 | cyto: 3, chlo: 2, nucl: 2, mito: 2, vacu: 2 |
| PfBTB44 | Pfo12g010520 | Chr12 | 563 | 63,201.63 | 7.9 | plas: 9, vacu: 2, nucl: 1, extr: 1, E.R.: 1 |
| PfBTB45 | Pfo12g011820 | Chr12 | 607 | 68,499.04 | 6.03 | nucl: 6, cysk: 3, chlo: 2, cyto: 1, mito: 1 |
| PfBTB46 | Pfo13g007700 | Chr13 | 555 | 62,398.01 | 8.3 | chlo: 7, nucl: 5, cyto: 1, golg_plas: 1 |
| PfBTB47 | Pfo13g007770 | Chr13 | 545 | 60,899.48 | 5.5 | cyto: 7, nucl: 3, chlo: 2, mito: 1, vacu: 1 |
| PfBTB48 | Pfo13g007850 | Chr13 | 606 | 68,444.12 | 7.91 | nucl: 8, cyto: 4, chlo: 1, cysk: 1 |
| PfBTB49 | Pfo14g010540 | Chr14 | 399 | 45,657.17 | 9 | nucl: 13, cyto: 1 |
| PfBTB50 | Pfo15g002860 | Chr15 | 345 | 39,347.06 | 5.25 | cyto: 6, nucl_plas: 3, nucl: 2.5, plas: 2.5 |
| PfBTB51 | Pfo15g003440 | Chr15 | 292 | 33,412.69 | 5.08 | nucl: 6, mito: 4, chlo: 3, cysk: 1 |
| PfBTB52 | Pfo16g000510 | Chr16 | 588 | 65,792.84 | 5.99 | cyto: 6, nucl: 4, chlo: 1, mito: 1, plas: 1 |
| PfBTB53 | Pfo16g008780 | Chr16 | 328 | 37,300.61 | 5.39 | cyto: 7, nucl: 2, mito: 2, plas: 2, pero: 1 |
| PfBTB54 | Pfo16g013570 | Chr16 | 633 | 70,396.02 | 5.94 | nucl: 6, chlo: 4, cyto: 2, plas: 1, cysk: 1 |
| PfBTB55 | Pfo18g003810 | Chr18 | 617 | 68,606.8 | 8.82 | cyto: 6, nucl: 4, vacu: 2, chlo: 1, E.R.: 1 |
| PfBTB56 | Pfo18g011610 | Chr18 | 428 | 48,041.23 | 4.82 | chlo: 4, nucl: 3.5, cysk_nucl: 2.5, plas: 2 |
| PfBTB57 | Pfo19g001900 | Chr19 | 857 | 94,729.13 | 5.59 | nucl: 12, cyto: 1, vacu: 1 |
| PfBTB58 | Pfoxxg008290 | Contig01580 | 616 | 68,975.67 | 5.67 | cyto: 8, nucl: 5, vacu: 1 |
| PfBTB59 | Pfoxxg012560 | Contig01619 | 383 | 42,468.21 | 5.71 | nucl: 8, chlo: 4, cyto: 1, cysk: 1 |
| PfBTB60 | Pfoxxg012630 | Contig01622 | 312 | 34,410.13 | 6.19 | nucl: 8.5, cyto_nucl: 6, chlo: 3, cyto: 2.5 |
| PfBTB61 | Pfoxxg018320 | tig00016041 | 452 | 49,057.37 | 6.05 | cyto: 6, chlo: 5, nucl: 1, plas: 1 |
| PfBTB62 | Pfoxxg021290 | tig00016941 | 549 | 61,153.44 | 6.11 | chlo: 10.5, chlo_mito: 6, nucl: 3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, P.; Fan, Y.; Xu, P.; Fan, G. Bioinformatic Analysis of the BTB Gene Family in Paulownia fortunei and Functional Characterization in Response to Abiotic and Biotic Stresses. Plants 2023, 12, 4144. https://doi.org/10.3390/plants12244144
Zhu P, Fan Y, Xu P, Fan G. Bioinformatic Analysis of the BTB Gene Family in Paulownia fortunei and Functional Characterization in Response to Abiotic and Biotic Stresses. Plants. 2023; 12(24):4144. https://doi.org/10.3390/plants12244144
Chicago/Turabian StyleZhu, Peipei, Yujie Fan, Pingluo Xu, and Guoqiang Fan. 2023. "Bioinformatic Analysis of the BTB Gene Family in Paulownia fortunei and Functional Characterization in Response to Abiotic and Biotic Stresses" Plants 12, no. 24: 4144. https://doi.org/10.3390/plants12244144
APA StyleZhu, P., Fan, Y., Xu, P., & Fan, G. (2023). Bioinformatic Analysis of the BTB Gene Family in Paulownia fortunei and Functional Characterization in Response to Abiotic and Biotic Stresses. Plants, 12(24), 4144. https://doi.org/10.3390/plants12244144





