PYL Family Genes from Liriodendron chinense Positively Respond to Multiple Stresses
Abstract
1. Introduction
2. Results
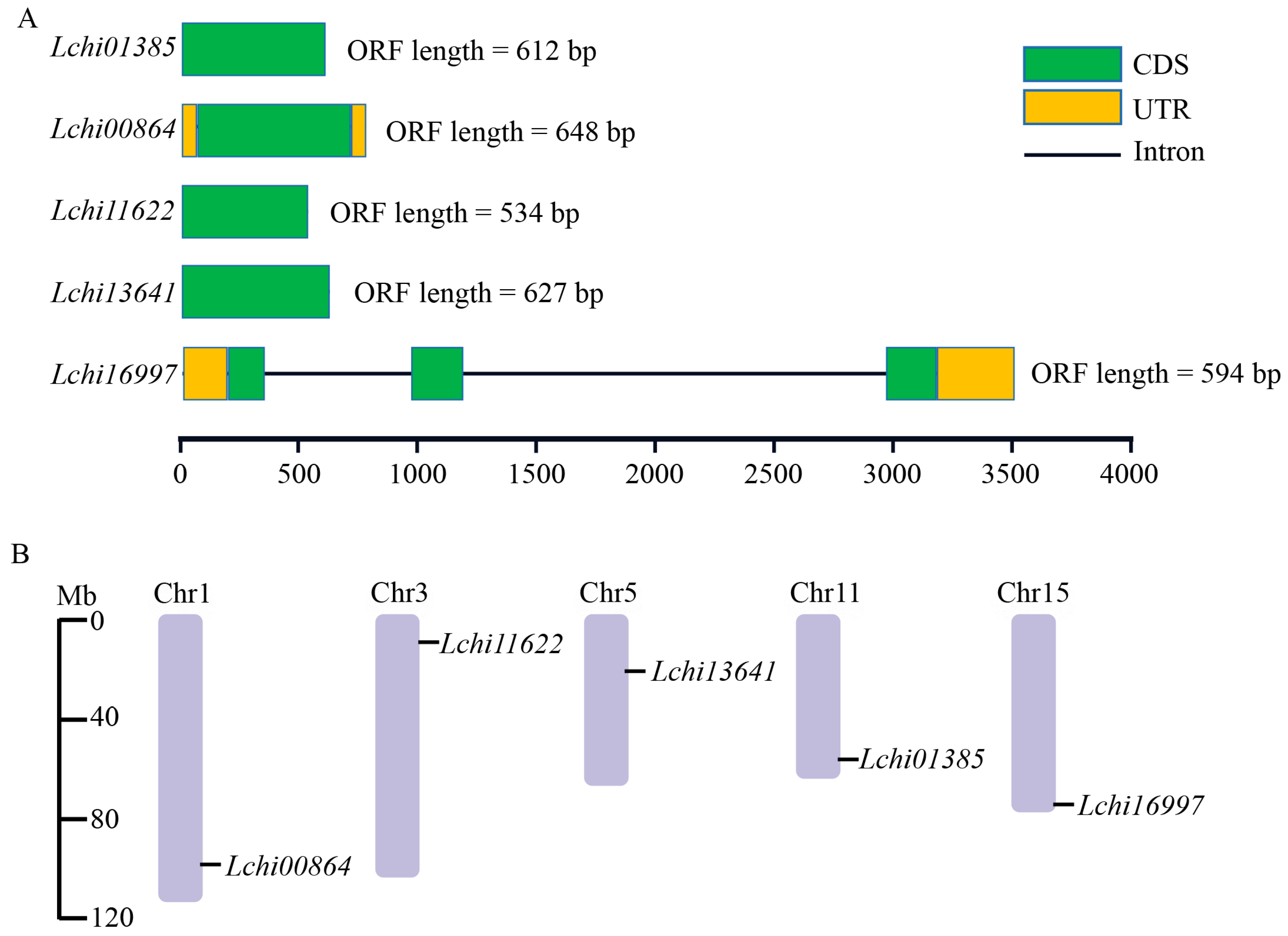
2.1. Genome-Wide Analysis and Chromosome Distribution of LcPYL Family Genes
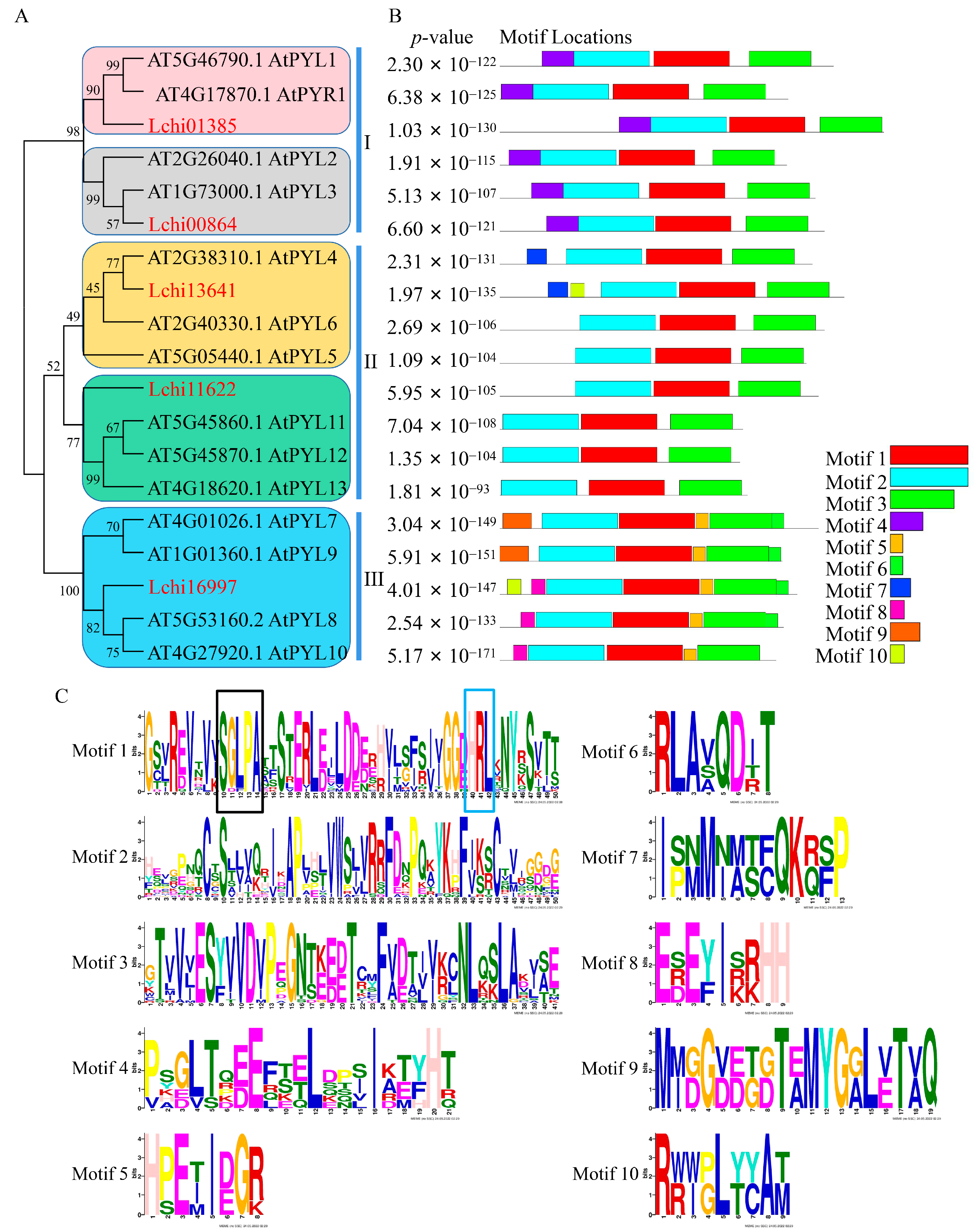
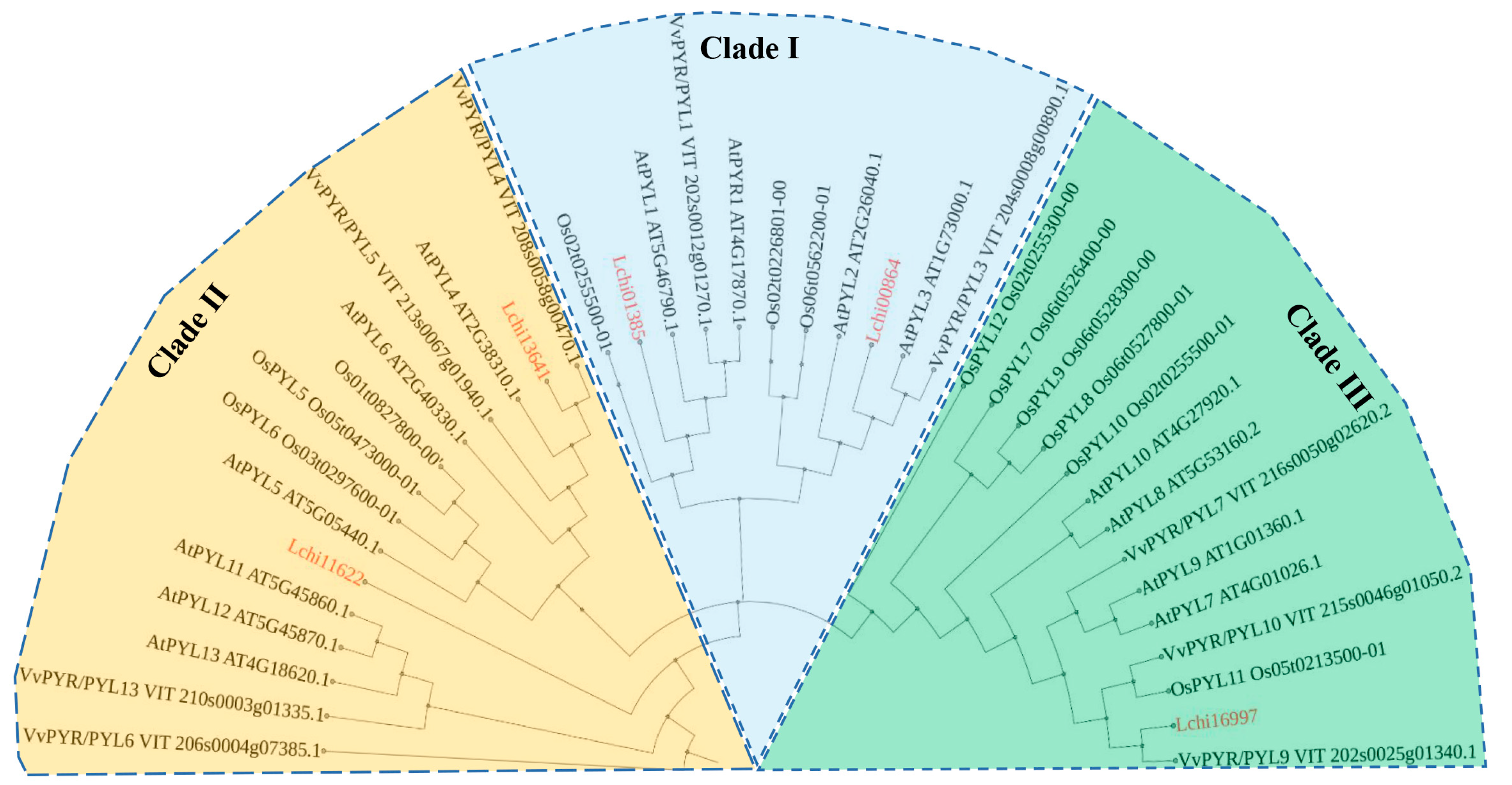
2.2. Conserved Domains and Phylogenetic Study of LcPYL Proteins
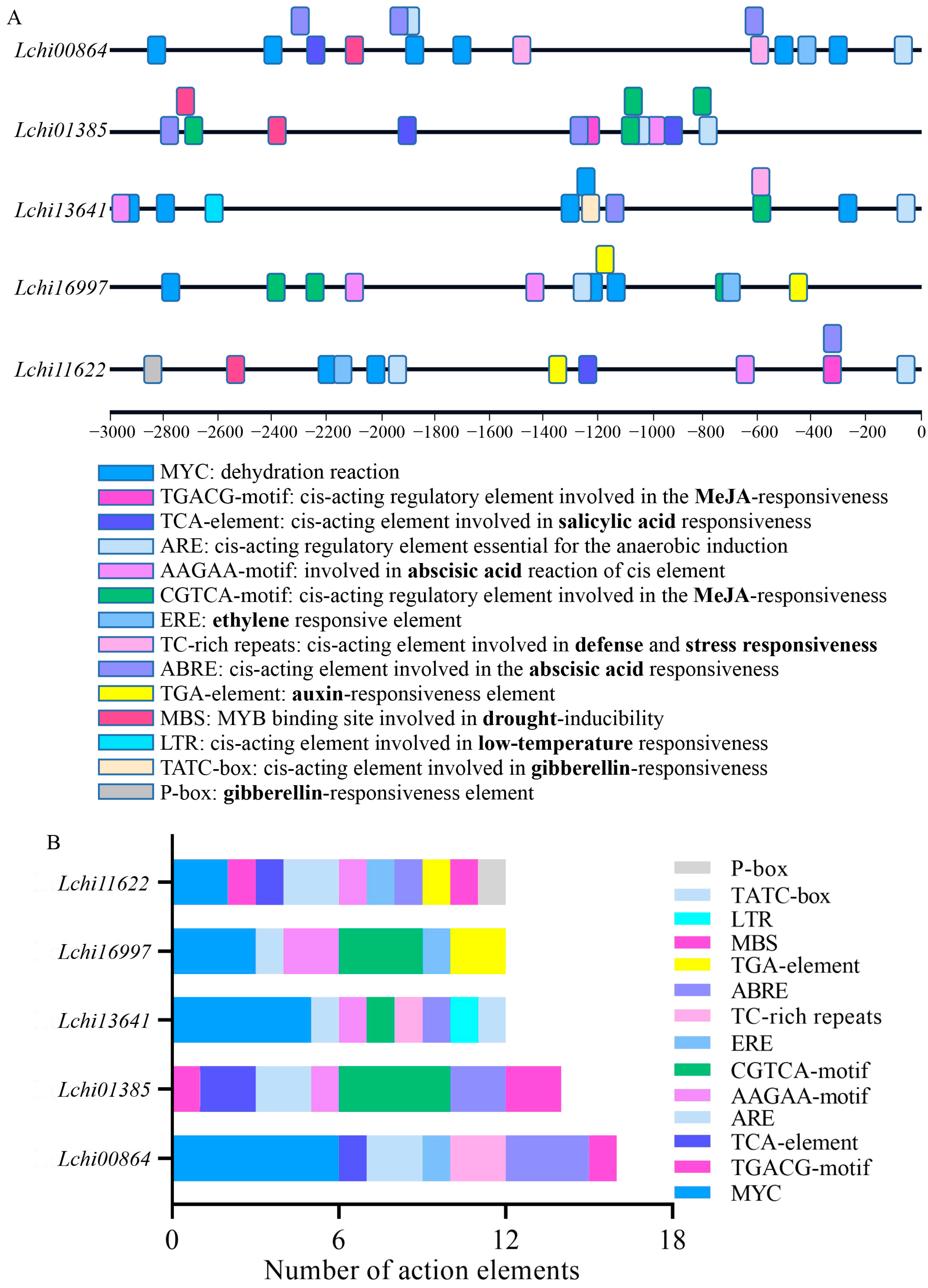
2.3. Cis-Acting Element Exploration of LcPYL Promoters
2.4. Expression Pattern of LcPYLs in Different Tissues of L. hybrid
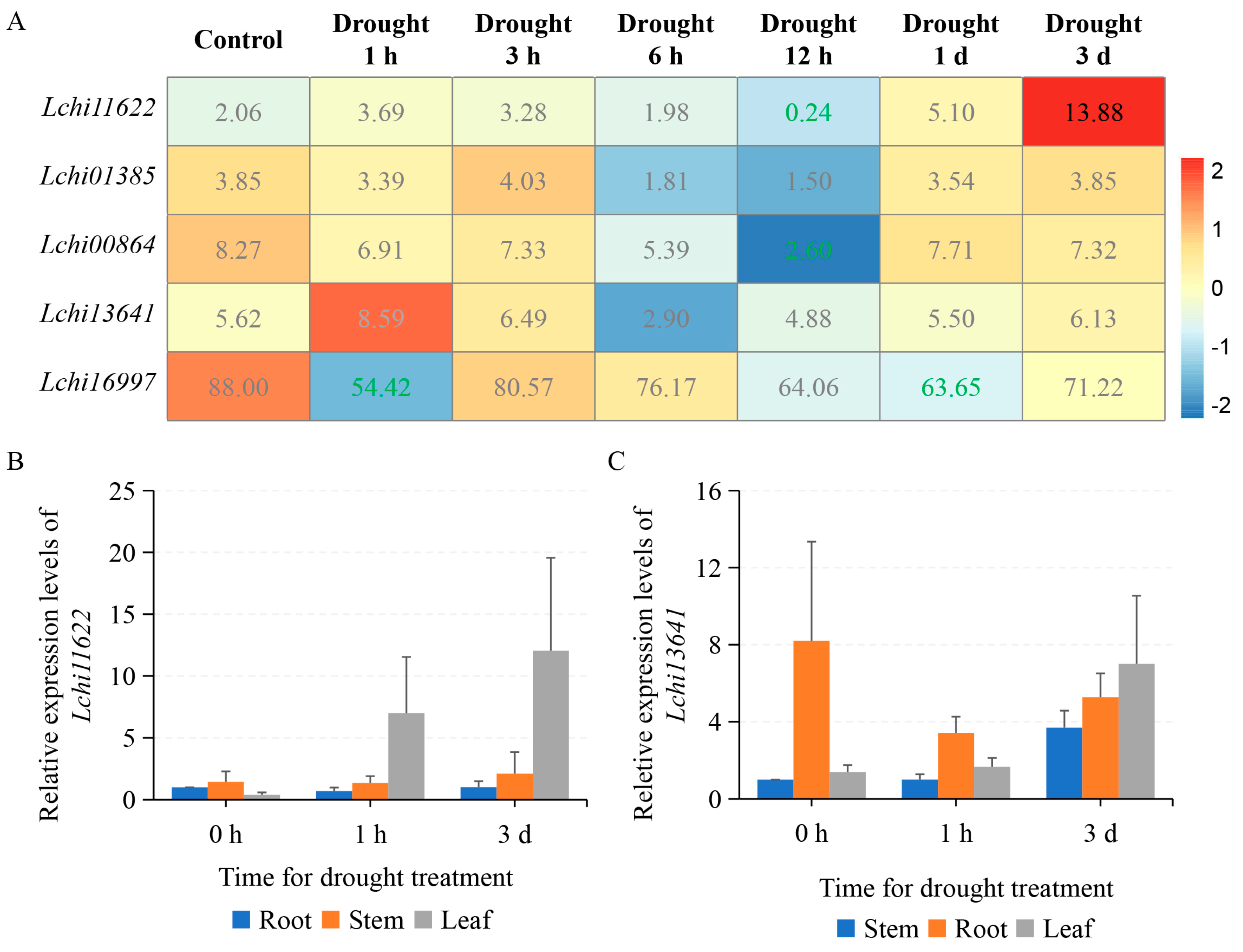
2.5. LcPYL Genes Specifically Respond to Abiotic Stresses in L. hybrid
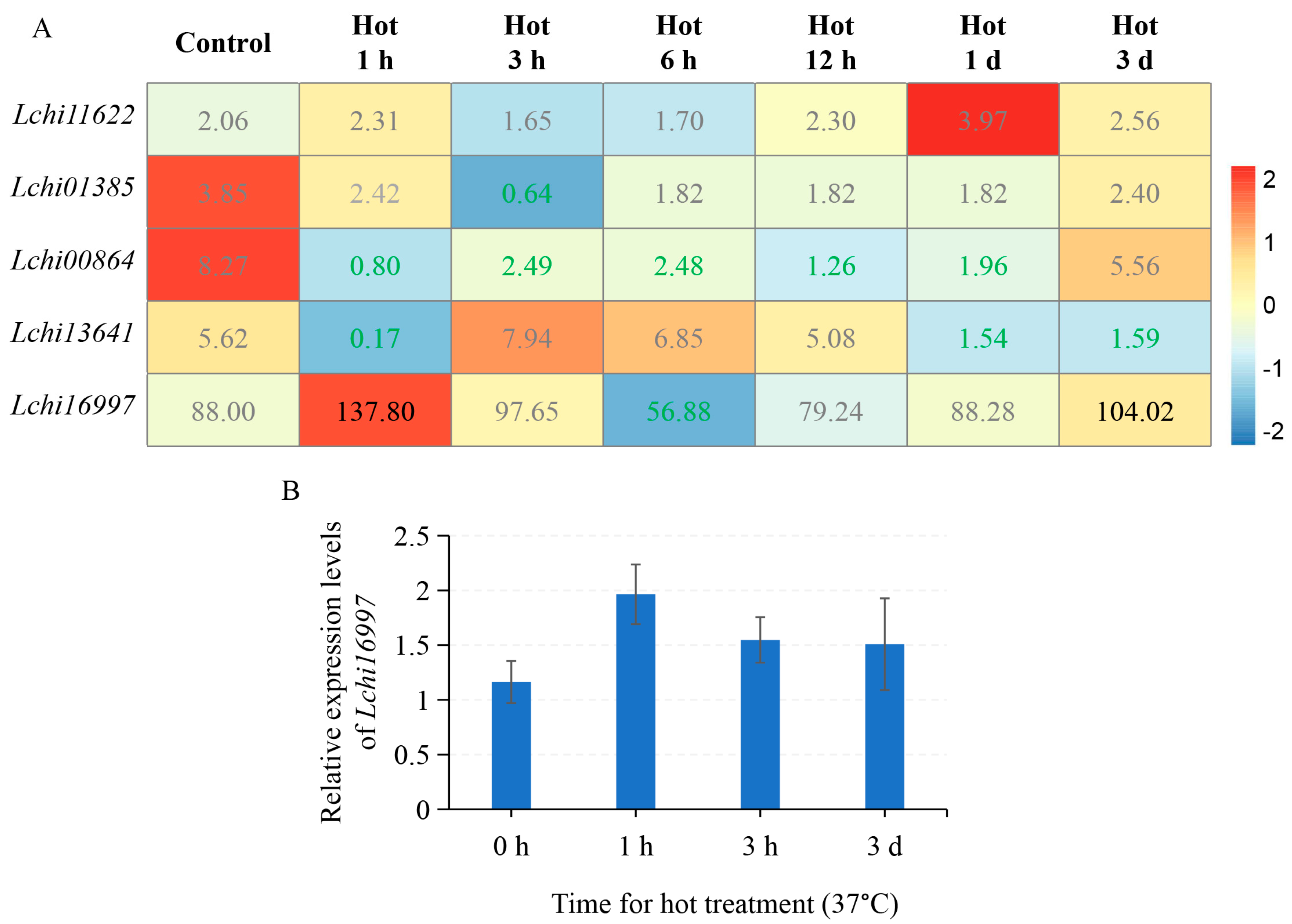
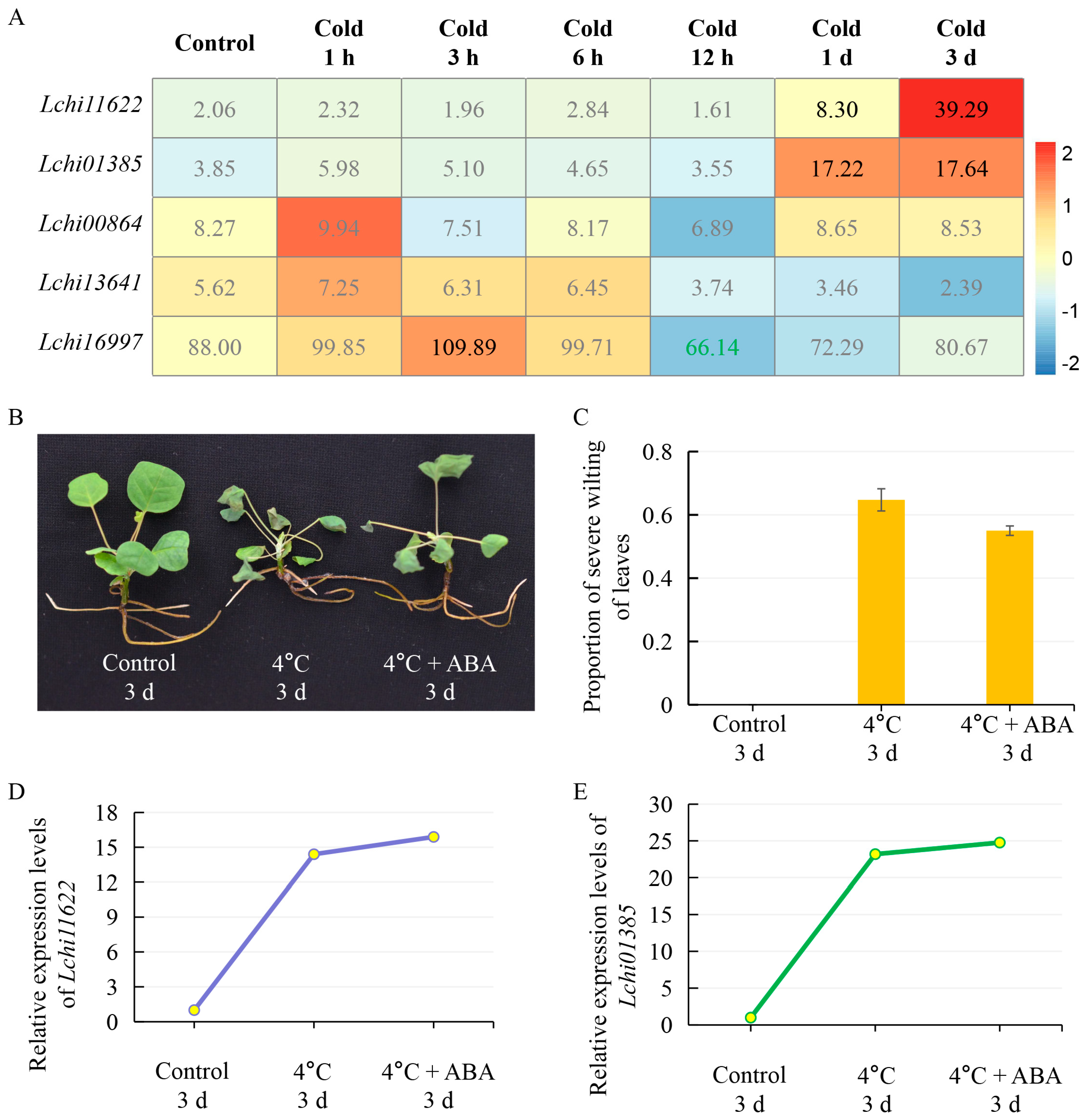
2.6. LcPYL Genes Specifically Respond to Biotic Stresses in L. hybrid
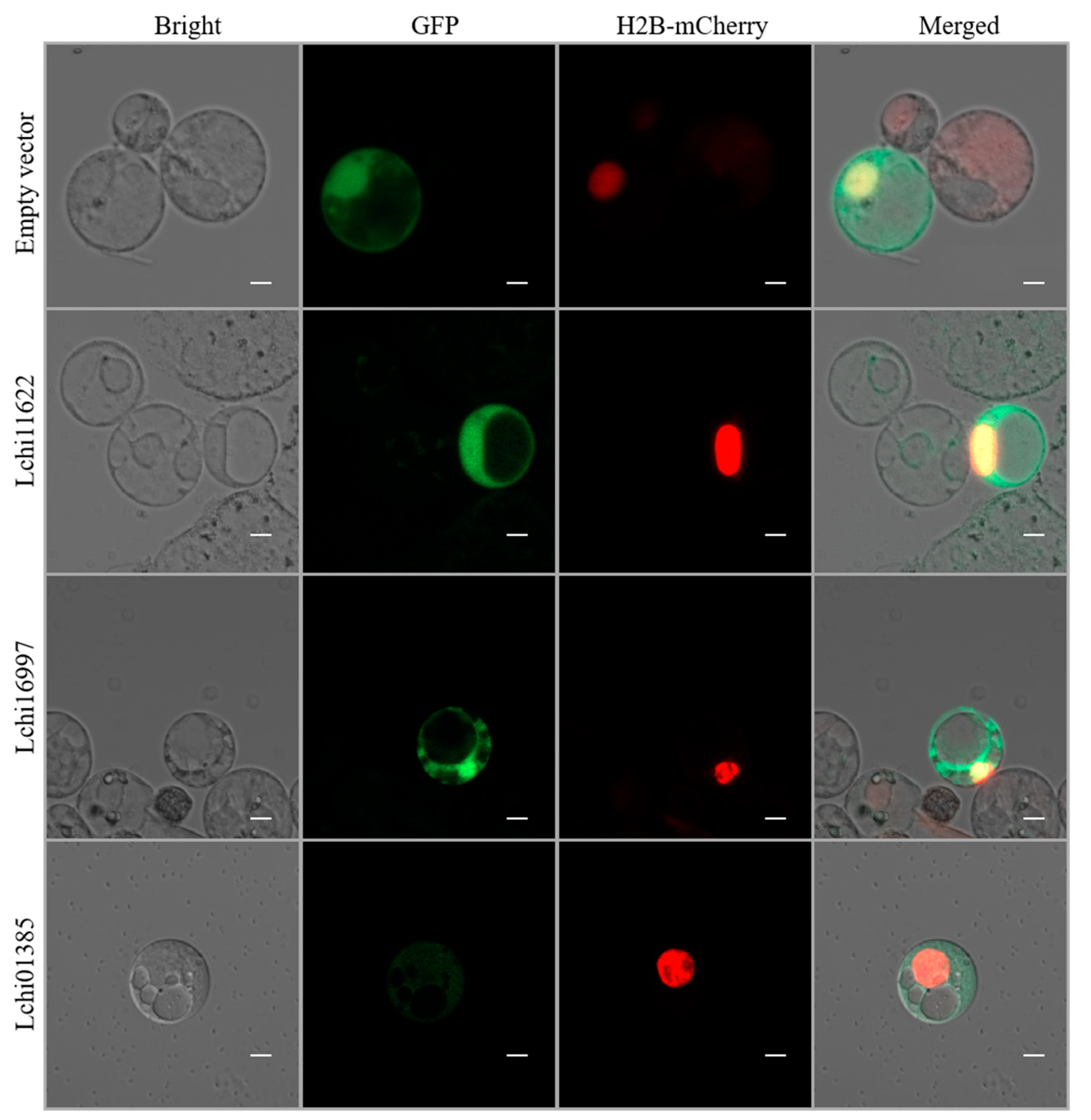
2.7. Subcellular Localization of LcPYL Genes
3. Discussion
3.1. A Limit Number of LcPYL Proteins
3.2. LcPYLs Specifically Respond to Various Abiotic Stresses
4. Materials and Methods
4.1. Plant Materials and Stress Treatment
4.2. Genome-Wide Identification of LcPYL Family Genes
4.3. Analysis of Gene Exon–Intron Structures and Protein Conserved Motifs
4.4. Basic Information of LcPYL Proteins
4.5. Cis-Acting Elemental Analysis of LcPYL Gene Promoters
4.6. Phylogenetic Analysis
4.7. Quantitative Real-Time PCR Analyses
4.8. Gene Clone and Subcellular Localization Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ali, M.; Wang, X.; Haroon, U.; Chaudhary, H.J.; Kamal, A.; Ali, Q.; Saleem, M.H.; Usman, K.; Alatawi, A.; Ali, S.; et al. Antifungal activity of Zinc nitrate derived nano Zno fungicide synthesized from Trachyspermum ammi to control fruit rot disease of grapefruit. Ecotoxicol. Environ. Saf. 2022, 233, 113311. [Google Scholar] [CrossRef]
- Gill, R.A.; Ahmar, S.; Ali, B.; Saleem, M.H.; Khan, M.U.; Zhou, W.; Liu, S. The Role of Membrane Transporters in Plant Growth and Development, and Abiotic Stress Tolerance. Int. J. Mol. Sci. 2021, 22, 12792. [Google Scholar] [CrossRef] [PubMed]
- Bashir, K.; Matsui, A.; Rasheed, S.; Seki, M. Recent advances in the characterization of plant transcriptomes in response to drought, salinity, heat, and cold stress. F1000Research 2019, 8, Rev-658. [Google Scholar] [CrossRef] [PubMed]
- Duressa, D.; Soliman, K.; Taylor, R.; Senwo, Z. Proteomic Analysis of Soybean Roots under Aluminum Stress. Int. J. Plant. Genom. 2011, 2011, 282531. [Google Scholar] [CrossRef] [PubMed]
- Weng, J.K.; Ye, M.L.; Li, B.; Noel, J.P. Co-evolution of Hormone Metabolism and Signaling Networks Expands Plant Adaptive Plasticity. Cell 2016, 166, 881–893. [Google Scholar] [CrossRef]
- Zhu, J.K. Abiotic Stress Signaling and Responses in Plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef]
- Danquah, A.; de Zelicourt, A.; Colcombet, J.; Hirt, H. The role of ABA and MAPK signaling pathways in plant abiotic stress responses. Biotechnol. Adv. 2014, 32, 40–52. [Google Scholar] [CrossRef]
- Hussain, Q.; Asim, M.; Zhang, R.; Khan, R.; Farooq, S.; Wu, J.S. Transcription Factors Interact with ABA through Gene Expression and Signaling Pathways to Mitigate Drought and Salinity Stress. Biomolecules 2021, 11, 1159. [Google Scholar] [CrossRef]
- Zhu, L.M.; Fang, H.; Lian, Z.M.; Zhang, J.B.; Li, X.L.; Shi, J.; Lu, L.; Lu, Y.; Chen, J.H.; Cheng, T.L. Genome-Wide Investigation and Expression Analysis of the Nitraria sibirica Pall. CIPK Gene Family. Int. J. Mol. Sci. 2022, 23, 11599. [Google Scholar] [CrossRef]
- Schroeder, J.I.; Kwak, J.M.; Allen, G.J. Guard cell abscisic acid signalling and engineering drought hardiness in plants. Nature 2001, 410, 327–330. [Google Scholar] [CrossRef]
- Raghavendra, A.S.; Gonugunta, V.K.; Christmann, A.; Grill, E. ABA perception and signalling. Trends Plant Sci. 2010, 15, 395–401. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Fung, P.; Nishimura, N.; Jensen, D.R.; Fujii, H.; Zhao, Y.; Lumba, S.; Santiago, J.; Rodrigues, A.; Chow, T.F.; et al. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 2009, 324, 1068–1071. [Google Scholar] [CrossRef] [PubMed]
- Fujii, H.; Chinnusamy, V.; Rodrigues, A.; Rubio, S.; Antoni, R.; Park, S.Y.; Cutler, S.R.; Sheen, J.; Rodriguez, P.L.; Zhu, J.K. In vitro reconstitution of an abscisic acid signalling pathway. Nature 2009, 462, 660–664. [Google Scholar] [CrossRef]
- Santiago, J.; Rodrigues, A.; Saez, A.; Rubio, S.; Antoni, R.; Dupeux, F.; Park, S.Y.; Marquez, J.A.; Cutler, S.R.; Rodriguez, P.L. Modulation of drought resistance by the abscisic acid receptor PYL5 through inhibition of clade A PP2Cs. Plant J. 2009, 60, 575–588. [Google Scholar] [CrossRef]
- Lumba, S.; Cutler, S.; McCourt, P. Plant nuclear hormone receptors: A role for small molecules in protein-protein interactions. Annu. Rev. Cell Dev. Biol. 2010, 26, 445–469. [Google Scholar] [CrossRef] [PubMed]
- Melcher, K.; Ng, L.M.; Zhou, X.E.; Soon, F.F.; Xu, Y.; Suino-Powell, K.M.; Park, S.Y.; Weiner, J.J.; Fujii, H.; Chinnusamy, V.; et al. A gate-latch-lock mechanism for hormone signalling by abscisic acid receptors. Nature 2009, 462, 602–608. [Google Scholar] [CrossRef] [PubMed]
- Furihata, T.; Maruyama, K.; Fujita, Y.; Umezawa, T.; Yoshida, R.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Abscisic acid-dependent multisite phosphorylation regulates the activity of a transcription activator AREB1. Proc. Natl. Acad. Sci. USA 2006, 103, 1988–1993. [Google Scholar] [CrossRef]
- Fujii, H.; Verslues, P.E.; Zhu, J.K. Identification of two protein kinases required for abscisic acid regulation of seed germination, root growth, and gene expression in Arabidopsis. Plant Cell 2007, 19, 485–494. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Zhang, Z.; Gao, J.; Wang, P.; Hu, T.; Wang, Z.; Hou, Y.J.; Wan, Y.; Liu, W.; Xie, S.; et al. Arabidopsis Duodecuple Mutant of PYL ABA Receptors Reveals PYL Repression of ABA-Independent SnRK2 Activity. Cell Rep. 2018, 23, 3340–3351.e3345. [Google Scholar] [CrossRef]
- Boneh, U.; Biton, I.; Zheng, C.; Schwartz, A.; Ben-Ari, G. Characterization of potential ABA receptors in Vitis vinifera. Plant Cell Rep. 2012, 31, 311–321. [Google Scholar] [CrossRef]
- Zhang, G.F.; Lu, T.T.; Mia, W.W.; Sun, L.R.; Tian, M.; Wang, J.; Hao, F.S. Genome-wide identification of ABA receptor PYL family and expression analysis of PYLs in response to ABA and osmotic stress in Gossypium. Peerj 2017, 5, e4126. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Guzman, M.; Rodriguez, L.; Lorenzo-Orts, L.; Pons, C.; Sarrion-Perdigones, A.; Fernandez, M.A.; Peirats-Llobet, M.; Forment, J.; Moreno-Alvero, M.; Cutler, S.R.; et al. Tomato PYR/PYL/RCAR abscisic acid receptors show high expression in root, differential sensitivity to the abscisic acid agonist quinabactin, and the capability to enhance plant drought resistance. J. Exp. Bot. 2014, 65, 4451–4464. [Google Scholar] [CrossRef] [PubMed]
- Guo, D.; Zhou, Y.; Li, H.L.; Zhu, J.H.; Wang, Y.; Chen, X.T.; Peng, S.Q. Identification and characterization of the abscisic acid (ABA) receptor gene family and its expression in response to hormones in the rubber tree. Sci. Rep. 2017, 7, 45157. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Zhang, X.; Su, H.; Liu, X.; Wang, Y.; Hong, G. Genome-wide analysis of PYL-PP2C-SnRK2s family in Camellia sinensis. Bioengineered 2020, 11, 103–115. [Google Scholar] [CrossRef]
- Bai, G.; Yang, D.H.; Zhao, Y.; Ha, S.; Yang, F.; Ma, J.; Gao, X.S.; Wang, Z.M.; Zhu, J.K. Interactions between soybean ABA receptors and type 2C protein phosphatases. Plant Mol. Biol. 2013, 83, 651–664. [Google Scholar] [CrossRef]
- Chen, Y.; Feng, L.; Wei, N.; Liu, Z.H.; Hu, S.; Li, X.B. Overexpression of cotton PYL genes in Arabidopsis enhances the transgenic plant tolerance to drought stress. Plant Physiol. Biochem. 2017, 115, 229–238. [Google Scholar] [CrossRef]
- Wilde, H.D.; Merkle, S.A. Genetic Transformation in Liriodendron tulipifera L. (Yellow Poplar). In Plant Protoplasts and Genetic Engineering V; Bajaj, Y.P.S., Ed.; Springer: Berlin/Heidelberg, Germany, 1994; pp. 337–348. [Google Scholar]
- Chen, J.; Hao, Z.; Guang, X.; Zhao, C.; Wang, P.; Xue, L.; Zhu, Q.; Yang, L.; Sheng, Y.; Zhou, Y.; et al. Liriodendron genome sheds light on angiosperm phylogeny and species-pair differentiation. Nat. Plants 2019, 5, 18–25. [Google Scholar] [CrossRef]
- Fujita, M.; Fujita, Y.; Noutoshi, Y.; Takahashi, F.; Narusaka, Y.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Crosstalk between abiotic and biotic stress responses: A current view from the points of convergence in the stress signaling networks. Curr. Opin. Plant. Biol. 2006, 9, 436–442. [Google Scholar] [CrossRef]
- Cutler, S.R.; Rodriguez, P.L.; Finkelstein, R.R.; Abrams, S.R. Abscisic acid: Emergence of a core signaling network. Annu. Rev. Plant. Biol. 2010, 61, 651–679. [Google Scholar] [CrossRef]
- Ye, Y.; Zhou, L.; Liu, X.; Liu, H.; Li, D.; Cao, M.; Chen, H.; Xu, L.; Zhu, J.-K.; Zhao, Y. A Novel Chemical Inhibitor of ABA Signaling Targets All ABA Receptors. Plant Physiol. 2017, 173, 2356–2369. [Google Scholar] [CrossRef]
- Santiago, J.; Dupeux, F.; Round, A.; Antoni, R.; Park, S.Y.; Jamin, M.; Cutler, S.R.; Rodriguez, P.L.; Marquez, J.A. The abscisic acid receptor PYR1 in complex with abscisic acid. Nature 2009, 462, 665–668. [Google Scholar] [CrossRef] [PubMed]
- Saavedra, X.; Modrego, A.; Rodriguez, D.; Gonzalez-Garcia, M.P.; Sanz, L.; Nicolas, G.; Lorenzo, O. The nuclear interactor PYL8/RCAR3 of Fagus sylvatica FsPP2C1 is a positive regulator of abscisic acid signaling in seeds and stress. Plant Physiol. 2010, 152, 133–150. [Google Scholar] [CrossRef] [PubMed]
- Panchy, N.; Lehti-Shiu, M.; Shiu, S.H. Evolution of Gene Duplication in Plants. Plant Physiol. 2016, 171, 2294–2316. [Google Scholar] [CrossRef] [PubMed]
- Yadav, S.K.; Santosh, K.V.V.; Verma, R.K.; Yadav, P.; Saroha, A.; Wankhede, D.P.; Chaudhary, B.; Chinnusamy, V. Genome-wide identification and characterization of ABA receptor PYL gene family in rice. BMC Genom. 2020, 21, 676. [Google Scholar] [CrossRef] [PubMed]
- Lei, P.; Wei, X.; Gao, R.; Huo, F.; Nie, X.; Tong, W.; Song, W. Genome-wide identification of PYL gene family in wheat: Evolution, expression and 3D structure analysis. Genomics 2021, 113, 854–866. [Google Scholar] [CrossRef] [PubMed]
- Cao, Q.; Lv, W.; Jiang, H.; Chen, X.; Wang, X.; Wang, Y. Genome-wide identification of glutathione S-transferase gene family members in tea plant (Camellia sinensis) and their response to environmental stress. Int. J. Biol. Macromol. 2022, 205, 749–760. [Google Scholar] [CrossRef]
- El-Gebali, S.; Mistry, J.; Bateman, A.; Eddy, S.R.; Luciani, A.; Potter, S.C.; Qureshi, M.; Richardson, L.J.; Salazar, G.A.; Smart, A.; et al. The Pfam protein families database in 2019. Nucleic Acids Res. 2019, 47, D427–D432. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I.; et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, D222–D226. [Google Scholar] [CrossRef]
- Zhang, Z.; Luo, S.; Liu, Z.; Wan, Z.; Gao, X.; Qiao, Y.; Yu, J.; Zhang, G. Genome-wide identification and expression analysis of the cucumber PYL gene family. PeerJ 2022, 10, e12786. [Google Scholar] [CrossRef]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Sievers, F.; Higgins, D.G. Clustal Omega for making accurate alignments of many protein sequences. Protein Sci. 2018, 27, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Biasini, M.; Bienert, S.; Waterhouse, A.; Arnold, K.; Studer, G.; Schmidt, T.; Kiefer, F.; Gallo, C.T.; Bertoni, M.; Bordoli, L.; et al. SWISS-MODEL: Modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 2014, 42, W252–W258. [Google Scholar] [CrossRef]
- Gordon, C.S.; Rajagopalan, N.; Risseeuw, E.P.; Surpin, M.; Ball, F.J.; Barber, C.J.; Buhrow, L.M.; Clark, S.M.; Page, J.E.; Todd, C.D.; et al. Characterization of Triticum aestivum Abscisic Acid Receptors and a Possible Role for These in Mediating Fusairum Head Blight Susceptibility in Wheat. PLoS ONE 2016, 11, e0164996. [Google Scholar] [CrossRef]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef]
- Lu, L.; Wu, X.R.; Wang, P.K.; Zhu, L.M.; Liu, Y.X.; Tang, Y.; Hao, Z.D.; Lu, Y.; Zhang, J.B.; Shi, J.S.; et al. Halophyte Nitraria billardieri CIPK25 mitigates salinity-induced cell damage by alleviating H2O2 accumulation. Front. Plant Sci. 2022, 13, 961651. [Google Scholar] [CrossRef]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; de Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef]
- Xiong, E.; Zheng, C.; Wu, X.; Wang, W. Protein Subcellular Location: The Gap Between Prediction and Experimentation. Plant Mol. Biol. Report. 2016, 34, 52–61. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Li, T.; Yuan, W.; Qiu, S.; Shi, J. Selection of reference genes for gene expression analysis in Liriodendron hybrids’ somatic embryogenesis and germinative tissues. Sci. Rep. 2021, 11, 4957. [Google Scholar] [CrossRef] [PubMed]
- Huo, A.L.; Chen, Z.Y.; Wang, P.K.; Yang, L.M.; Wang, G.P.; Wang, D.D.; Liao, S.C.; Cheng, T.L.; Chen, J.H.; Shi, J.S. Establishment of transient gene expression systems in protoplasts from Liriodendron hybrid mesophyll cells. PLoS ONE 2017, 12, e0172475. [Google Scholar] [CrossRef] [PubMed]



| Gene ID | Lchi01385 | Lchi00864 | Lchi11622 | Lchi13641 | Lchi16997 |
|---|---|---|---|---|---|
| aa 1 | 204 | 215 | 178 | 209 | 197 |
| MW (kd) 2 | 22.66 | 23.75 | 19.75 | 22.55 | 22.53 |
| pI 3 | 5 | 5.91 | 5.74 | 7.65 | 5.75 |
| Instability index | 38.31 | 33.28 | 60.21 | 48.32 | 47.14 |
| Aliphatic index | 78.28 | 86 | 85.9 | 90.43 | 90.41 |
| GRAVY 4 | −0.415 | −0.226 | −0.157 | −0.056 | −0.427 |
| Subcellular location | Cytoplasm | Chloroplast | Chloroplast | Chloroplast | Cytoplasm |
| Lchi01385 | Lchi00864 | Lchi11622 | Lchi13641 | Lchi16997 | |
|---|---|---|---|---|---|
| Alpha helix % | 35.78 (73 aa 1) | 37.67 (81 aa) | 42.7 (76 aa) | 32.06 (67 aa) | 40.61 (80 aa) |
| Extended strand % | 18.63 (38 aa) | 19.07 (41 aa) | 20.22 (36 aa) | 19.14 (40 aa) | 17.26 (34 aa) |
| Beta turn % | 4.41 (9 aa) | 6.05 (13 aa) | 4.49 (8 aa) | 4.78 (10 aa) | 4.06 (8 aa) |
| Random coil % | 41.18 (84 aa) | 37.21 (80 aa) | 32.58 (58 aa) | 44.02 (92 aa) | 38.07 (75 aa) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, X.; Zhu, J.; Chen, X.; Zhang, J.; Lu, L.; Hao, Z.; Shi, J.; Chen, J. PYL Family Genes from Liriodendron chinense Positively Respond to Multiple Stresses. Plants 2023, 12, 2609. https://doi.org/10.3390/plants12142609
Wu X, Zhu J, Chen X, Zhang J, Lu L, Hao Z, Shi J, Chen J. PYL Family Genes from Liriodendron chinense Positively Respond to Multiple Stresses. Plants. 2023; 12(14):2609. https://doi.org/10.3390/plants12142609
Chicago/Turabian StyleWu, Xinru, Junjie Zhu, Xinying Chen, Jiaji Zhang, Lu Lu, Zhaodong Hao, Jisen Shi, and Jinhui Chen. 2023. "PYL Family Genes from Liriodendron chinense Positively Respond to Multiple Stresses" Plants 12, no. 14: 2609. https://doi.org/10.3390/plants12142609
APA StyleWu, X., Zhu, J., Chen, X., Zhang, J., Lu, L., Hao, Z., Shi, J., & Chen, J. (2023). PYL Family Genes from Liriodendron chinense Positively Respond to Multiple Stresses. Plants, 12(14), 2609. https://doi.org/10.3390/plants12142609






