High-Precision Tomato Disease Detection Using NanoSegmenter Based on Transformer and Lightweighting
Abstract
:1. Introduction
- A dataset containing ten kinds of tomato diseases and healthy states is collected and annotated at the pixel level. This dataset, which consists of 15,383 images, covers various disease states from early to late stages, as well as healthy tomato leaves. This dataset is not only useful for training and validating the model proposed here but also serves as a rich resource for other researchers.
- To address the issue of class imbalance in the dataset, a diffusion model is used to generate samples of weak classes, making the number of instances for each class in the dataset balanced. The principle of the diffusion model, as well as how it is applied to the task in this work, is introduced.
- Furthermore, the NanoSegmenter model, which is based on the task of instance segmentation and employs the Transformer structure, inverse bottleneck technique, and sparse attention mechanism, is proposed. This model can achieve high-precision tomato disease detection.
- Additionally, a grading model is utilized in combination with an expert system to perform disease grading based on the diseased area, offering corresponding advice. This grading model can assist farmers in more accurately assessing the severity of diseases, thereby developing more effective control strategies.
- Lastly, the model undergoes lightweight processing and is deployed on a smartphone. This allows farmers to perform disease detection and grading in the field, greatly improving detection efficiency.
2. Results and Discussion
2.1. Segmentation Results
2.2. Visualization Analysis
2.3. Test on Other Dataset
2.4. Ablation Study of Lightweight Methods
2.4.1. Theoretical Analysis
2.4.2. Ablation Experiment Results on Different Platform
2.5. Model Deployment
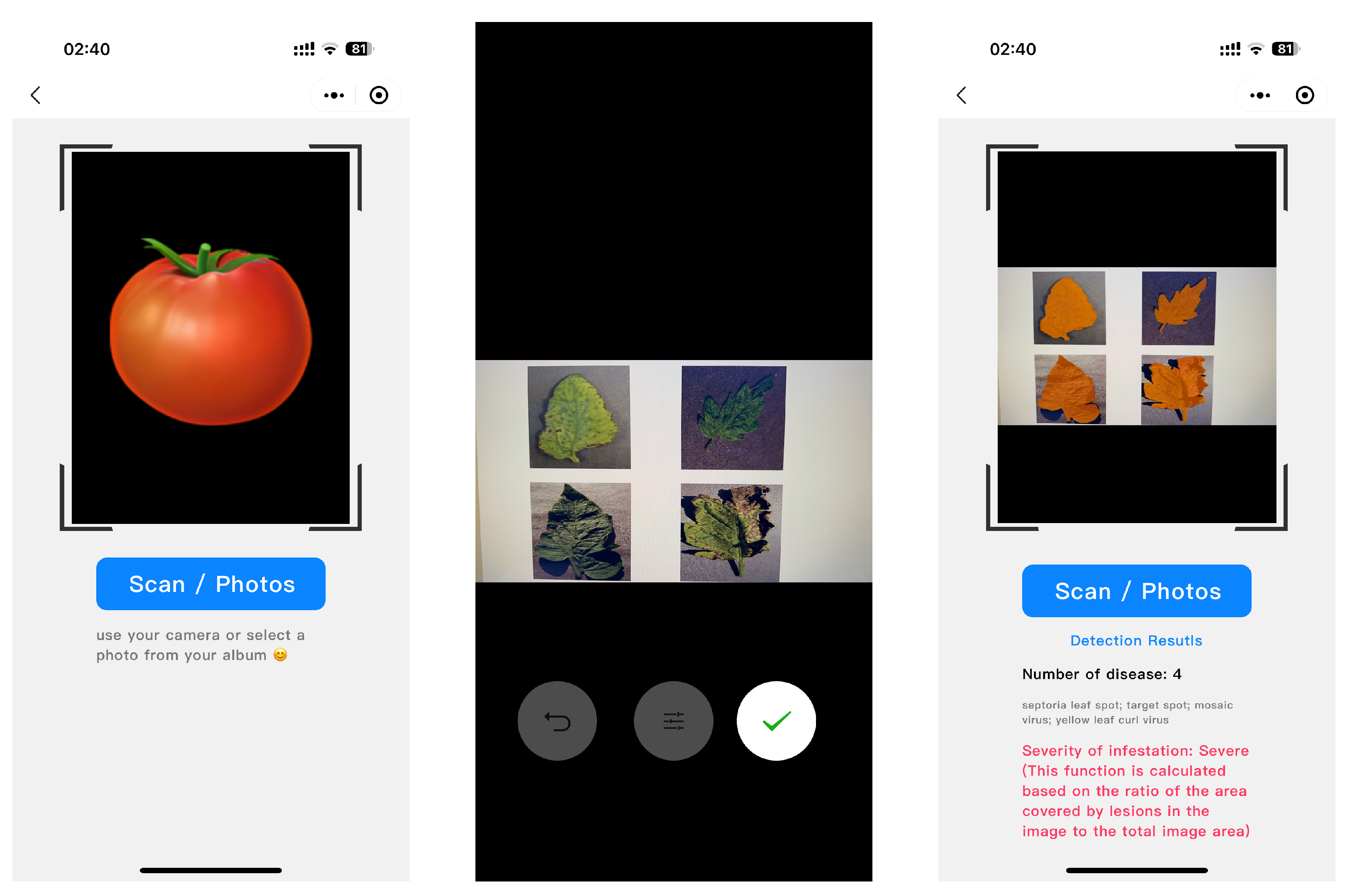
2.5.1. Deployment on Smartphones
2.5.2. Federated Learning-Based Training Framework
3. Materials
3.1. Dataset Collection and Analysis
3.2. Dataset Augmentation
4. Methods
4.1. NanoSegmenter
4.1.1. Overall
- Introduction of the Transformer structure into instance segmentation tasks: The Transformer structure was initially proposed by Vaswani et al. [29] in “Attention Is All You Need”, designed to handle sequence-to-sequence tasks. The structure centers around the self-attention mechanism, allowing the model to automatically learn the interdependencies among different parts of the input sequence. Although Transformers have achieved remarkable success in the NLP domain, their application in visual tasks is still relatively limited. This can be primarily attributed to the strong locality of dependencies between pixels in visual tasks, whereas Transformer structures often capture global dependencies. To address this issue, the Transformer structure was incorporated into instance segmentation tasks to simultaneously capture global and local dependencies. It was found that this approach significantly improves the model’s segmentation precision.
- Model lightweight processing using inverse bottleneck technology: Despite the impressive performance of the Transformer structure, its substantial parameter quantity makes it challenging to deploy on resource-constrained devices. To solve this issue, the technique of inverse bottleneck was introduced to achieve lightweight processing by reducing model complexity. The inverse bottleneck is an effective model compression technique, where the key idea is to add a lower-dimensional hidden layer between the model’s input and output, thereby substantially reducing the model’s computational cost. By applying this technique to the Transformer structure, the model’s parameter quantity was successfully reduced by an order of magnitude, while maintaining comparable performance.
- Model lightweight processing using sparse attention: In addition to the inverse bottleneck technique, a sparse attention mechanism was introduced to further decrease the model size and computational complexity. In traditional Transformer structures, the output at each position is the weighted sum of the inputs from all positions, resulting in a computational complexity of O(), where n is the length of the input. To address this issue, a sparse attention mechanism was introduced so that the output at each position depends only on a small subset of the input. By doing so, the model’s computational complexity was reduced to O(n log n), enabling deployment on resource-constrained devices.
4.1.2. Segment by Transformer
- Input embedding layer: The task of this layer is to convert the original RGB image into a feature vector suitable for Transformer input. A pre-trained convolutional neural network (such as ResNet50) is used as the feature extractor, and a linear transformation is then applied to map the features to a designated dimension. Assuming the original image is , the convolutional feature extractor is , and the linear transformation is , the output of the input embedding layer iswhere represents the use of convolutional neural networks; denotes the process of flattening the output of the neural network, that is, the flatten operation; and d is the set feature dimension.
- Transformer layer: The task of this layer is to understand the interdependencies between pixels. The output E from the input embedding layer is converted into a sequence format and then input into the Transformer. Specifically, assuming the Transformer structure is , the output of the Transformer layer iswhere , is the total number of pixels and signifies the procedure of feeding inputs into the structure depicted in Figure 8. Note that the core of the Transformer is the self-attention mechanism, which can automatically learn the interdependencies between pixels.
- Output classification layer: The task of this layer is to convert the output from the Transformer into classification results for each pixel. A simple linear transformation is used as the classifier. Specifically, assuming the linear transformation is , the final output iswhere , , and c is the total number of categories.
4.1.3. Inverted Bottleneck
4.1.4. Sparse Attention
4.2. Experimental Settings
4.2.1. Hardware and Software Platform
4.2.2. Optimizer, Loss Function and Hyperparameters
4.2.3. Training Strategy
4.2.4. Experiment Metric
- mAP (mean average precision): mAP is a common measure for assessing the performance of object detection or instance segmentation tasks. It computes the average of the area under curve (AUC) of precision and recall of the predicted bounding boxes. mAP evaluates the performance of the model at all thresholds comprehensively, and its mathematical expression iswhere Q is the set of all queries, is the number of relevant documents for the q-th query, and is the precision of the k-th document. A higher mAP signifies better model performance.
- Precision: Precision is a metric used to assess the accuracy of the model prediction, and its mathematical expression iswhere represents the number of true positives, and represents the number of false positives. In this task, precision reflects the proportion of correctly predicted disease regions out of all predicted regions.
- Recall: Recall is a metric used to assess the coverage of the model prediction, and its mathematical expression iswhere represents the number of false negatives. In this task, recall reflects the proportion of correctly predicted disease regions out of all actual disease regions.
- FPS (frames per second): FPS is a metric used to evaluate the computational efficiency of the model. In practical applications, especially in scenarios requiring real-time processing, FPS is critical. A higher FPS indicates that the model can process more images in a short period, denoting higher computational efficiency.
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Zhang, Y.; Wa, S.; Liu, Y.; Zhou, X.; Sun, P.; Ma, Q. High-accuracy detection of maize leaf diseases CNN based on multi-pathway activation function module. Remote Sens. 2021, 13, 4218. [Google Scholar] [CrossRef]
- Zhang, Y.; Wa, S.; Sun, P.; Wang, Y. Pear defect detection method based on resnet and dcgan. Information 2021, 12, 397. [Google Scholar] [CrossRef]
- Vishal, M.K.; Saluja, R.; Aggrawal, D.; Banerjee, B.; Raju, D.; Kumar, S.; Chinnusamy, V.; Sahoo, R.N.; Adinarayana, J. Leaf Count Aided Novel Framework for Rice (Oryza sativa L.) Genotypes Discrimination in Phenomics: Leveraging Computer Vision and Deep Learning Applications. Plants 2022, 11, 2663. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Zhang, Y.; Yang, G. Small unopened cotton boll counting by detection with MRF-YOLO in the wild. Comput. Electron. Agric. 2023, 204, 107576. [Google Scholar] [CrossRef]
- Li, D.; Ahmed, F.; Wu, N.; Sethi, A.I. YOLO-JD: A Deep Learning Network for Jute Diseases and Pests Detection from Images. Plants 2022, 11, 937. [Google Scholar] [CrossRef]
- Abbas, I.; Liu, J.; Amin, M.; Tariq, A.; Tunio, M.H. Strawberry Fungal Leaf Scorch Disease Identification in Real-Time Strawberry Field Using Deep Learning Architectures. Plants 2021, 10, 2643. [Google Scholar] [CrossRef]
- Lin, J.; Bai, D.; Xu, R.; Lin, H. TSBA-YOLO: An Improved Tea Diseases Detection Model Based on Attention Mechanisms and Feature Fusion. Forests 2023, 14, 619. [Google Scholar] [CrossRef]
- Li, M.; Cheng, S.; Cui, J.; Li, C.; Li, Z.; Zhou, C.; Lv, C. High-Performance Plant Pest and Disease Detection Based on Model Ensemble with Inception Module and Cluster Algorithm. Plants 2023, 12, 200. [Google Scholar] [CrossRef]
- Zhang, X.; Zhu, D.; Wen, R. SwinT-YOLO: Detection of densely distributed maize tassels in remote sensing images. Comput. Electron. Agric. 2023, 210, 107905. [Google Scholar] [CrossRef]
- Dang, F.; Chen, D.; Lu, Y.; Li, Z. YOLOWeeds: A novel benchmark of YOLO object detectors for multi-class weed detection in cotton production systems. Comput. Electron. Agric. 2023, 205, 107655. [Google Scholar] [CrossRef]
- Nan, Y.; Zhang, H.; Zeng, Y.; Zheng, J.; Ge, Y. Intelligent detection of Multi-Class pitaya fruits in target picking row based on WGB-YOLO network. Comput. Electron. Agric. 2023, 208, 107780. [Google Scholar] [CrossRef]
- Rodrigues, L.; Magalhães, S.A.; da Silva, D.Q.; dos Santos, F.N.; Cunha, M. Computer Vision and Deep Learning as Tools for Leveraging Dynamic Phenological Classification in Vegetable Crops. Agronomy 2023, 13, 463. [Google Scholar] [CrossRef]
- Xu, D.; Zhao, H.; Lawal, O.M.; Lu, X.; Ren, R.; Zhang, S. An Automatic Jujube Fruit Detection and Ripeness Inspection Method in the Natural Environment. Agronomy 2023, 13, 451. [Google Scholar] [CrossRef]
- Phan, Q.H.; Nguyen, V.T.; Lien, C.H.; Duong, T.P.; Hou, M.T.K.; Le, N.B. Classification of Tomato Fruit Using Yolov5 and Convolutional Neural Network Models. Plants 2023, 12, 790. [Google Scholar] [CrossRef]
- Zhang, Y.; Wa, S.; Zhang, L.; Lv, C. Automatic plant disease detection based on tranvolution detection network with GAN modules using leaf images. Front. Plant Sci. 2022, 13, 875693. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, M.; Ma, X.; Wu, X.; Wang, Y. High-Precision Wheat Head Detection Model Based on One-Stage Network and GAN Model. Front. Plant Sci. 2022, 13, 787852. [Google Scholar] [CrossRef] [PubMed]
- Daniya, T.; Vigneshwari, S. Rider Water Wave-enabled deep learning for disease detection in rice plant. Adv. Eng. Softw. 2023, 182, 103472. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, H.; Xu, R.; Yang, X.; Wang, Y.; Liu, Y. High-Precision Seedling Detection Model Based on Multi-Activation Layer and Depth-Separable Convolution Using Images Acquired by Drones. Drones 2022, 6, 152. [Google Scholar] [CrossRef]
- Chen, L.C.; Zhu, Y.; Papandreou, G.; Schroff, F.; Adam, H. Encoder-decoder with atrous separable convolution for semantic image segmentation. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; pp. 801–818. [Google Scholar]
- Chen, L.C.; Papandreou, G.; Schroff, F.; Adam, H. Rethinking atrous convolution for semantic image segmentation. arXiv 2017, arXiv:1706.05587. [Google Scholar]
- Zhou, Z.; Siddiquee, M.M.R.; Tajbakhsh, N.; Liang, J. Unet++: A nested u-net architecture for medical image segmentation. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support; Springer: Berlin/Heidelberg, Germany, 2018; pp. 3–11. [Google Scholar]
- Zhao, H.; Shi, J.; Qi, X.; Wang, X.; Jia, J. Pyramid scene parsing network. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 2881–2890. [Google Scholar]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. Segnet: A deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015: 18th International Conference, Munich, Germany, 5–9 October 2015, Proceedings, Part III 18; Springer: Berlin/Heidelberg, Germany, 2015; pp. 234–241. [Google Scholar]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar]
- Potnis, N.; Timilsina, S.; Strayer, A.; Shantharaj, D.; Barak, J.D.; Paret, M.L.; Vallad, G.E.; Jones, J.B. Bacterial spot of tomato and pepper: Diverse X anthomonas species with a wide variety of virulence factors posing a worldwide challenge. Mol. Plant Pathol. 2015, 16, 907–920. [Google Scholar] [CrossRef] [PubMed]
- Chowdappa, P.; Kumar, S.M.; Lakshmi, M.J.; Upreti, K. Growth stimulation and induction of systemic resistance in tomato against early and late blight by Bacillus subtilis OTPB1 or Trichoderma harzianum OTPB3. Biol. Control 2013, 65, 109–117. [Google Scholar] [CrossRef]
- He, H.; Garcia, E.A. Learning from imbalanced data. IEEE Trans. Knowl. Data Eng. 2009, 21, 1263–1284. [Google Scholar]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. In Advances in Neural Information Processing Systems 30 (NIPS 2017); Curran Associates Inc.: Red Hook, NY, USA, 2017. [Google Scholar]
- Loshchilov, I.; Hutter, F. Decoupled weight decay regularization. arXiv 2017, arXiv:1711.05101. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]










| Model | Precision | Recall | mIoU | FPS |
|---|---|---|---|---|
| NanoSegmenter | 0.98 | 0.97 | 0.95 | 30 |
| DeepLabv3+ [19] | 0.97 | 0.96 | 0.93 | 27 |
| DeepLabv3 [20] | 0.96 | 0.95 | 0.91 | 26 |
| UNet++ [21] | 0.94 | 0.93 | 0.90 | 23 |
| PSPNet [22] | 0.93 | 0.91 | 0.88 | 21 |
| SegNet [23] | 0.91 | 0.90 | 0.86 | 20 |
| UNet [24] | 0.90 | 0.88 | 0.85 | 18 |
| FCN [25] | 0.88 | 0.86 | 0.83 | 15 |
| Model | Precision | Recall | mAP on Wheat Dataset |
|---|---|---|---|
| YOLOv5 | 0.61 | 0.58 | 0.59 |
| SSD | 0.58 | 0.53 | 0.55 |
| NanoTransformer + Detection Network | 0.61 | 0.57 | 0.58 |
| Accuracy on pear dataset | |||
| ResNet50 | 0.95 | 0.92 | 0.93 |
| VGG19 | 0.91 | 0.89 | 0.91 |
| AlexNet | 0.88 | 0.87 | 0.89 |
| NanoTransformer + Softmax | 0.97 | 0.92 | 0.94 |
| Method | Parameter Quantity | Computation | GPU Memory Usage |
|---|---|---|---|
| Inverted Bottleneck | Decrease by 30% | Decrease by 20% | Decrease by 20% |
| Sparse Attention | No Change | Decrease by 50% | Decrease by 30% |
| Quantization | No Change | Decrease by 90% | Decrease by 75% |
| Method | Inverted Bottleneck | Quantization | Sparse Attention | Precision | Recall | mIoU | FPS1 | FPS2 | FPS3 |
|---|---|---|---|---|---|---|---|---|---|
| ours | None | None | None | 0.98 | 0.97 | 0.96 | 14 | 11 | 27 |
| ours | √ | None | None | 0.98 | 0.97 | 0.95 | 31 | 18 | 42 |
| ours | None | √ | None | 0.97 | 0.96 | 0.94 | 32 | 18 | 44 |
| ours | None | None | √ | 0.96 | 0.95 | 0.93 | 33 | 18 | 48 |
| ours | √ | √ | None | 0.95 | 0.94 | 0.92 | 34 | 19 | 50 |
| ours | None | √ | √ | 0.94 | 0.93 | 0.91 | 35 | 22 | 49 |
| ours | √ | None | √ | 0.93 | 0.92 | 0.90 | 36 | 19 | 51 |
| ours | √ | √ | √ | 0.92 | 0.91 | 0.89 | 37 | 31 | 51 |
| ShuffleNet | None | None | None | 0.92 | 0.90 | 0.90 | 33 | 15 | 48 |
| MobileNet | None | None | None | 0.93 | 0.92 | 0.92 | 35 | 21 | 45 |
| Category | Number of Images | Proportion |
|---|---|---|
| bacterial spot | 468 | 0.030 |
| early blight | 1580 | 0.103 |
| healthy | 523 | 0.034 |
| late blight | 1478 | 0.096 |
| leaf mold | 2495 | 0.162 |
| Septoria leaf spot | 2510 | 0.163 |
| spider mites: two-spotted spider mite | 1569 | 0.102 |
| target spot | 2534 | 0.165 |
| mosaic virus | 1522 | 0.099 |
| yellow leaf curl virus | 504 | 0.033 |
| Category | Number of Images | Proportion |
|---|---|---|
| bacterial spot | 2534 | 0.104 |
| early blight | 2534 | 0.104 |
| healthy | 2534 | 0.104 |
| late blight | 2534 | 0.104 |
| leaf mold | 2534 | 0.104 |
| Septoria leaf spot | 2534 | 0.104 |
| spider mites two-spotted spider mite | 2534 | 0.104 |
| target spot | 2534 | 0.104 |
| mosaic virus | 2534 | 0.104 |
| yellow leaf curl virus | 2534 | 0.104 |
| Total | 25,340 | 1.000 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Song, Y.; Ye, R.; Zhu, S.; Huang, Y.; Chen, T.; Zhou, J.; Li, J.; Li, M.; Lv, C. High-Precision Tomato Disease Detection Using NanoSegmenter Based on Transformer and Lightweighting. Plants 2023, 12, 2559. https://doi.org/10.3390/plants12132559
Liu Y, Song Y, Ye R, Zhu S, Huang Y, Chen T, Zhou J, Li J, Li M, Lv C. High-Precision Tomato Disease Detection Using NanoSegmenter Based on Transformer and Lightweighting. Plants. 2023; 12(13):2559. https://doi.org/10.3390/plants12132559
Chicago/Turabian StyleLiu, Yufei, Yihong Song, Ran Ye, Siqi Zhu, Yiwen Huang, Tailai Chen, Junyu Zhou, Jiapeng Li, Manzhou Li, and Chunli Lv. 2023. "High-Precision Tomato Disease Detection Using NanoSegmenter Based on Transformer and Lightweighting" Plants 12, no. 13: 2559. https://doi.org/10.3390/plants12132559
APA StyleLiu, Y., Song, Y., Ye, R., Zhu, S., Huang, Y., Chen, T., Zhou, J., Li, J., Li, M., & Lv, C. (2023). High-Precision Tomato Disease Detection Using NanoSegmenter Based on Transformer and Lightweighting. Plants, 12(13), 2559. https://doi.org/10.3390/plants12132559






