Metagenomic Sequencing to Analyze Composition and Function of Top-Gray Chalky Grain Microorganisms from Hybrid Rice Seeds
Abstract
1. Introduction
2. Results
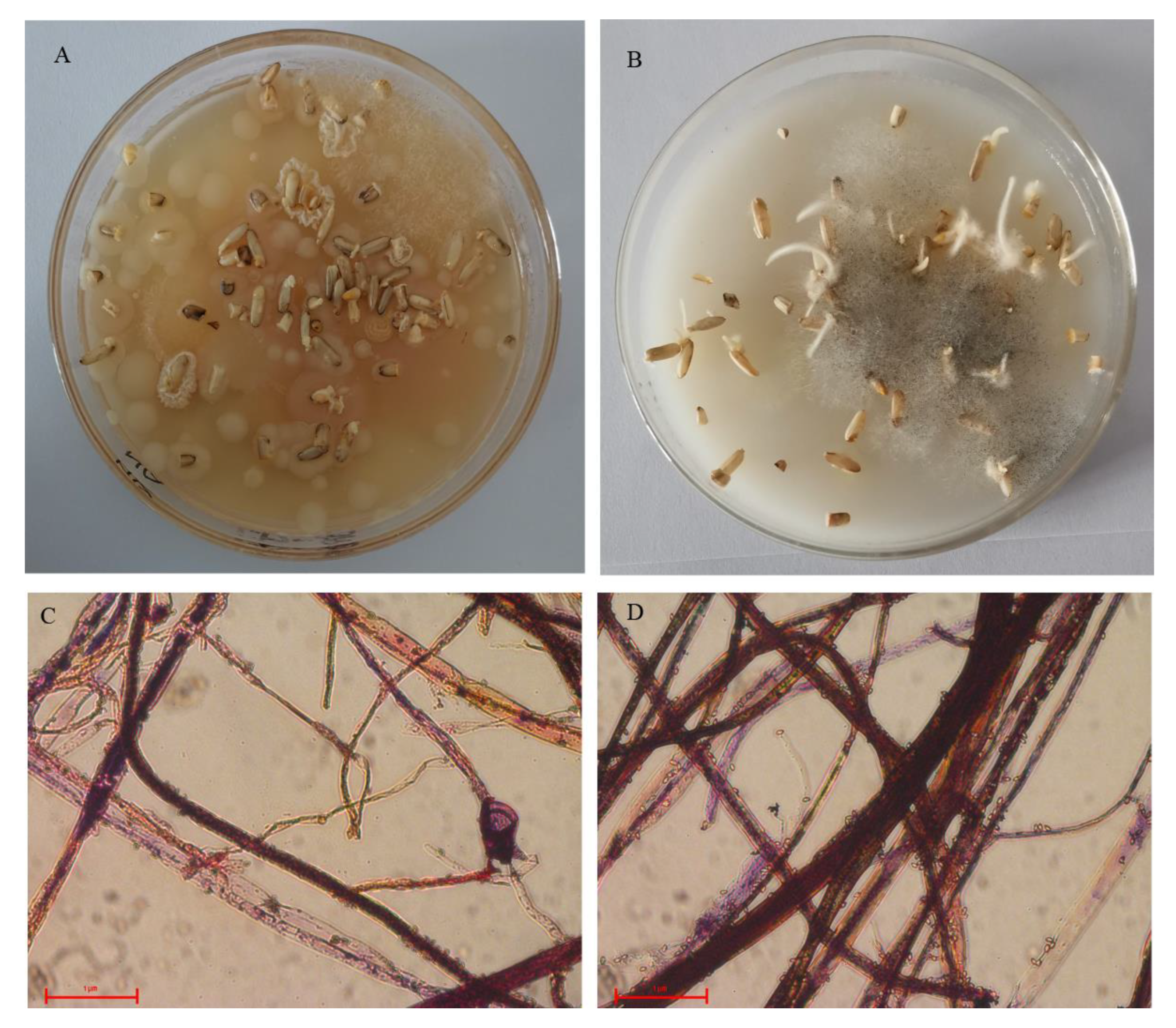
2.1. Cultivated Microorganisms from Chalky Rice Seeds
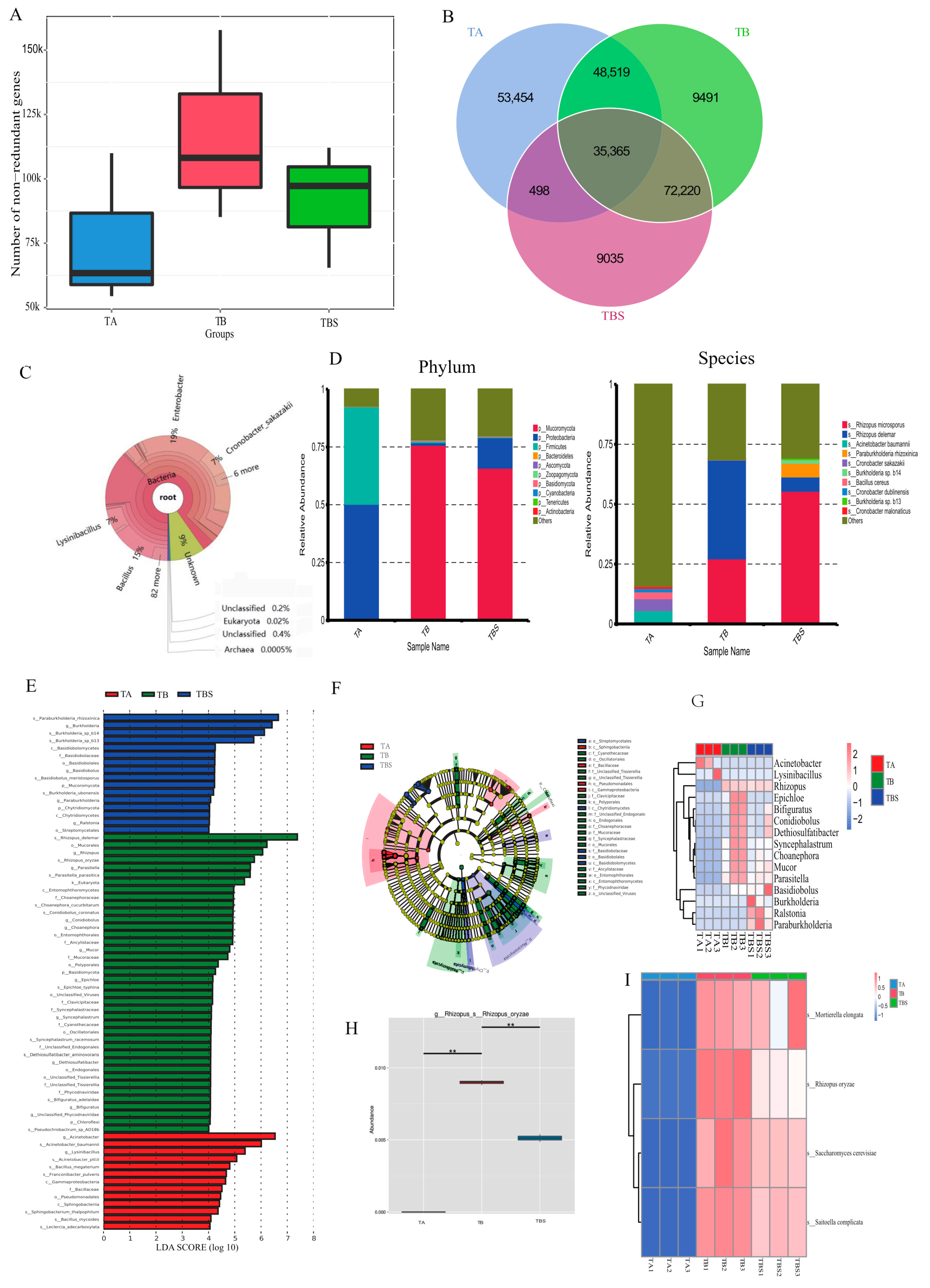
2.2. Microbiome Diversities of Chalky Rice Seeds Cultivated on Different Media
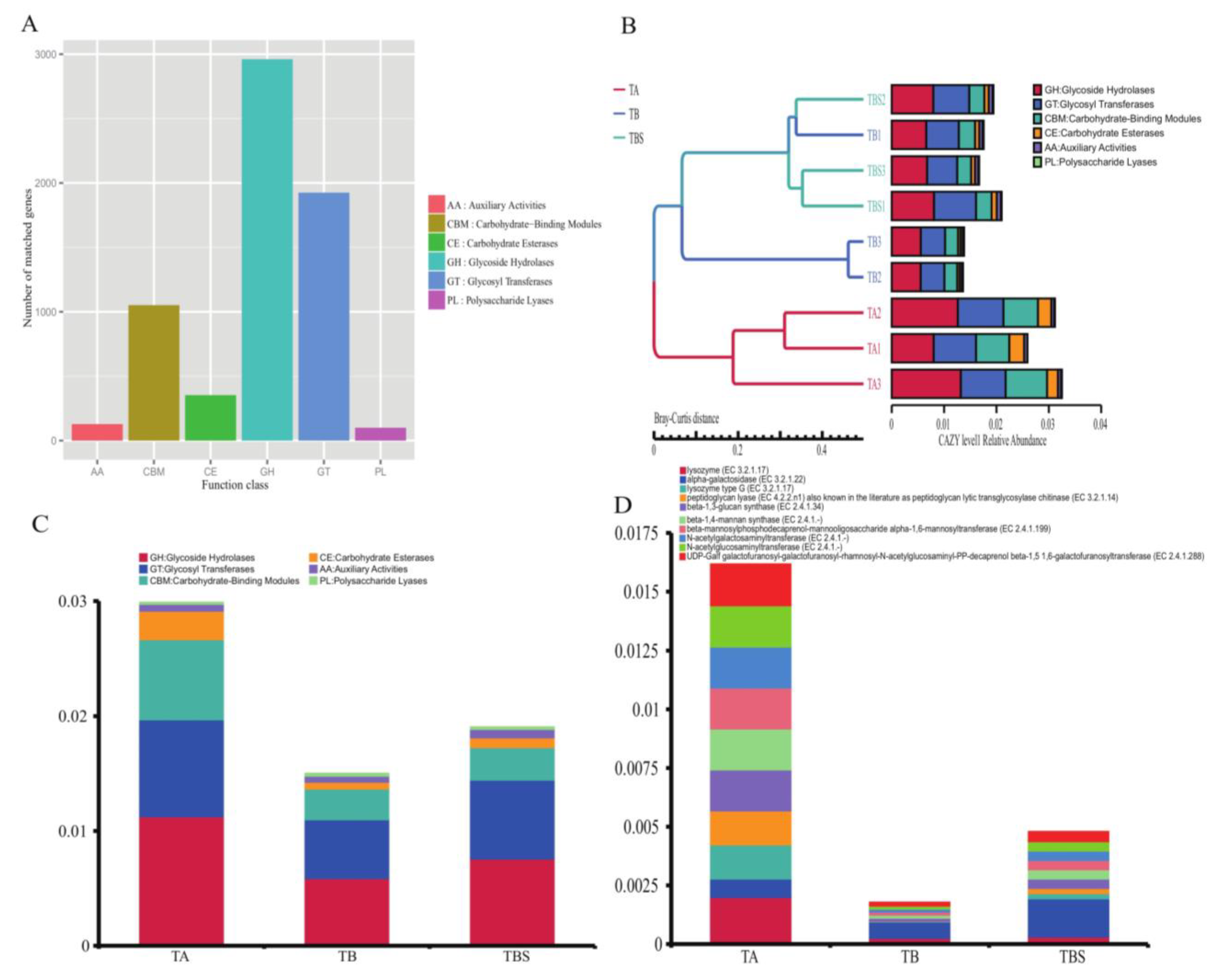
2.3. Functional Analysis of Microorganisms from Chalky Rice Seeds Cultivated on Different Media
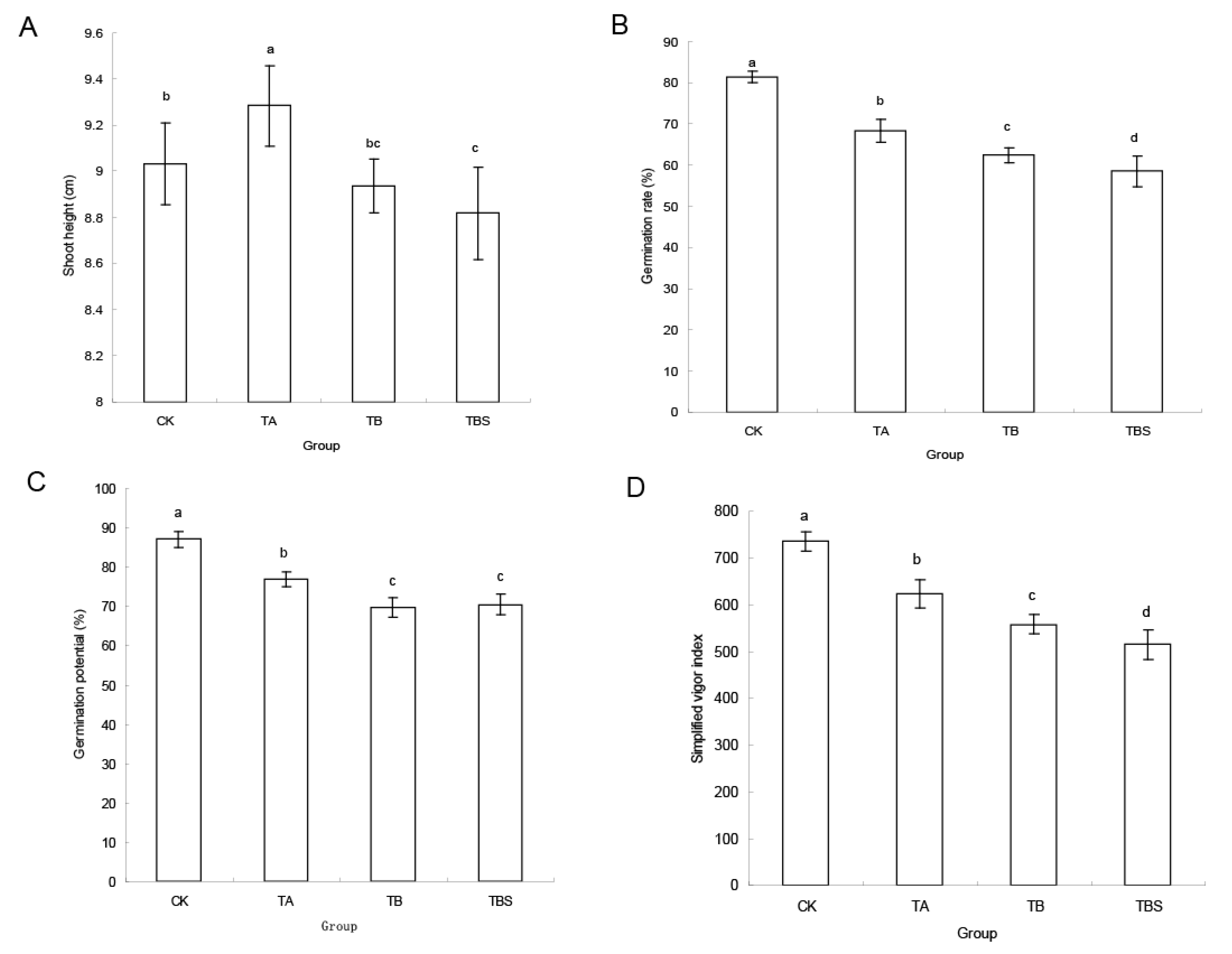
2.4. Hybrid Rice Seed Germination after Inoculated with Microorganisms from TA Group, TB Group, and TBS Group
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Microbial Cultivation
4.3. Metagenomic Shotgun Sequencing and Analysis
4.4. Infection on Seed Germination
4.5. Statistics
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Singh, R. Hybrid seed production mechanisms in rice (Oryza sativa L.). J. Biotechnol. Crop Sci. 2012, 1, 4–32. [Google Scholar]
- Li, Y.; Sun, J.; Ding, Q.; Liu, Y.; Ding, W. Effects of glume-gapping and kernel-cracking of hybrid rice seed on seed germination and seedling growth. Hybrid Rice 2015, 30, 71–74. [Google Scholar]
- Fu, C.; Wang, F.; Liu, W.; Liu, D.; Li, J.; Zhu, M.; Liao, Y.; Liu, Z.; Huang, H.; Zeng, X.; et al. Transcriptomic analysis reveals new insights into high-temperature-dependent glume-unclosing in an elite rice male sterile line. Front. Plant Sci. 2017, 8, 112. [Google Scholar] [CrossRef] [PubMed]
- Yang, J. Glume dehiscence of hybrid rice seeds and its harmfulness. Hybrid Rice 2006, 21, 57–60. [Google Scholar]
- Thu, B.V.; Chakrabarty, S.K.; Sharma, S.P.; Dadlani, M. Studies on environmental conditions and pollination management in hybrid rice seed production. Indian J. Genet. 2008, 68, 426–434. [Google Scholar]
- Zhang, H.; Tang, J.; Kuan, X.; Chen, Y.; Lu, C.; Liu, S.; Jiang, L.; Zheng, W.; Yuan, H.; Hu, Y. Germinating characteristics of “grey-matter” seeds of hybrid rice Longliangyou Huazhan. Hybrid Rice 2020, 35, 88–93. [Google Scholar]
- He, Q.; Deng, H.; Sun, P.; Zhang, W.; Shu, F.; Xing, J.; Peng, Z. Hybrid Rice. Engineering 2002, 6, 967–973. [Google Scholar] [CrossRef]
- Chen, Y.; Yuan, H.; Zhang, H.; Tang, J.; Liu, A.; Jin, C.; Hu, Y. Morphological and physiological properties of hybrid rice seeds with top-gray chalkiness. Bragantia 2021, 80, e0121. [Google Scholar] [CrossRef]
- Yoshioka, Y.; Iwata, H.; Tabata, M.; Ninomiya, S.; Ohsawa, R. Chalkiness in rice: Potential for evaluation with image analysis. Crop Sci. 2007, 47, 2113–2120. [Google Scholar] [CrossRef]
- Lurstwut, D.; Pornpanomchai, C. Image analysis based on color, shape and texture for rice seed (Oryza sativa L.) germination evaluation. Agric. Nat. Reso. 2017, 51, 383–389. [Google Scholar] [CrossRef]
- Khoenkaw, P. An image-processing based algorithm for rice seed germination rate evaluation. In Proceedings of the Computer Science & Engineering Conference, Chiang Mai, Thailand, 14–17 December 2016. [Google Scholar]
- Yin, L.; Tang, J.; Jin, C.; Hu, Y. Effect of seed-coating agents on rice seeds with dehiscent glumes. Agric. Sci. Technol. 2016, 17, 1383–1386. [Google Scholar]
- Nallathambi, P.; Umamaheswari, C.; Lal, S.K.; Manjunatha, C.; Berliner, J. Mechanism of seed transmission and seed infection in major agricultural crops in India. In Seed-Borne Diseases of Agricultural Crops: Detection, Diagnosis & Management; Kumar, R., Gupta, A., Eds.; Springer Nature Singapore Pte Ltd.: Singapore, 2020; Volume 26, pp. 749–791. [Google Scholar]
- Links, M.G.; Demeke, T.; Gräfenhan, T.; Hill, J.E.; Hemmingsen, S.M.; Dumonceaux, T.J. Simultaneous profiling of seed-associated bacteria and fungi reveals antagonistic interactions between microorganisms within a shared epiphytic microbiome on Triticum and Brassica seeds. New Phytol. 2014, 202, 542–553. [Google Scholar] [CrossRef] [PubMed]
- McGee, D.C. Seed pathology: Its place in modern seed production. Plant Dis. 1981, 65, 638–642. [Google Scholar] [CrossRef]
- Xu, F.; Zheng, J.; Zhu, Y.; Wang, G.; Yang, D.; Liu, K. Effect of super sparse cultivation on head milled rice percentage and chalkiness in hybrid rice varieties in the eastern and southern districts of Sichuan Province. Acta Phytoecol. Sin. 2004, 28, 686–691. [Google Scholar]
- Ackaah, F.M.; Nyaku, S.T.; Darkwa, E. Seed-borne fungi associated with diverse rice varieties cultivated in the western north region of Ghana. Int. J. Microbiol. 2023, 2023, 8690464. [Google Scholar] [CrossRef] [PubMed]
- Kato, H.; Ohata, K.; Kauraw, L.P.; Lee, Y.H. Fungal diseases of rice seed. In Proceedings of the International Workshop on Rice Seed Health, IRRI, Manila, Philippines, 16–19 March 1988. [Google Scholar]
- Nothling, M.D.; Xiao, Z.; Bhaskaran, A.; Blyth, M.T.; Bennett, C.; Coote, M.L.; Connal, L.A. Synthetic catalysts inspired by hydrolytic enzymes. ACS Catal. 2019, 9, 168–187. [Google Scholar] [CrossRef]
- Henrissat, B.; Davies, G. Structural and sequence-based classification of glycoside hydrolases. Curr. Opin. Struct. Biol. 1997, 7, 637–644. [Google Scholar] [CrossRef]
- Bhatia, S.; Singh, A.; Batra, N.; Singh, J. Microbial production and biotechnological applications of α-galactosidase. Int. J. Biol. Macromol. 2020, 150, 1294–1313. [Google Scholar] [CrossRef]
- Zhao, H.; Zhang, W.; Cheng, D.; You, L.; Huang, Y.; Lu, Y. Investigating dysbiosis and microbial treatment strategies in inflammatory bowel disease based on two modified Koch’s postulates. Front. Med. 2022, 9, 1023896. [Google Scholar] [CrossRef]
- Ranke, F.F.B.; Shinya, T.Y.; de Figueiredo, F.C.; Núňez, E.G.F.; Cabral, H.; Neto, P.O. Ethanol from rice byproduct using amylases secreted by Rhizopus microsporus var. oligosporus. enzyme partial purification and characterization. J. Environ. Manag. 2020, 266, 110591. [Google Scholar] [CrossRef]
- Ohnishi, M.; Higuchi, A.; Todoriki, S.; Hiromi, K.; Ohgushi, M.; Wada, A. Characterization on the conformation of Glucoamylase from Rhizopus niveus and Rhizopus delemar. Starch/Stärke 1990, 42, 273–276. [Google Scholar] [CrossRef]
- Battaglia, E.; Benoit, I.; van den Brink, J.; Wiebenga, A.; Coutinho, P.M.; Henrissat, B.; de Vries, R.P. Carbohydrate-active enzymes from the zygomycete fungus Rhizopus oryzae: A highly specialized approach to carbohydrate degradation depicted at genome level. BMC Genom. 2011, 12, 38. [Google Scholar] [CrossRef]
- Khalaf, E.M.; Raizada, M.N. Bacterial seed endophytes of domesticated cucurbits antagonize fungal and oomycete pathogens including powdery mildew. Front. Microbiol. 2018, 9, 42. [Google Scholar] [CrossRef]
- Premachandra, D.; Hudek, L.; Brau, L. Bacterial modes of action for enhancing of plant growth. J. Biotechnol. Biomater. 2016, 6, 3. [Google Scholar]
- Sanders, E.R. Aseptic laboratory techniques: Plating methods. J. Vis. Exp. 2012, 63, e3064. [Google Scholar]
- Mohan, S.K. Beyond bacteria: Interpreting fungal elements in the Gram stain. Clin. Microbiol. Newsl. 2004, 26, 108–112. [Google Scholar] [CrossRef]
- Liu, H.; Pei, H.; Han, Z.; Feng, G.; Li, D. The antimicrobial effects and synergistic antibacterial mechanism of the combination of ε-polylysine and nisin against Bacillus subtilis. Food Control 2015, 47, 444–450. [Google Scholar] [CrossRef]
- Li, J.; Jia, H.; Cai, X.; Zhong, H.; Feng, Q.; Sunagawa, S.; Arumugam, M.; Kultima, J.R.; Prifti, E.; Nielsen, T.; et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 2014, 32, 834–841. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, F.H.; Tremaroli, V.; Nookaew, I.; Bergström, G.; Behre, C.J.; Fagerberg, B.; Nielsen, J.; Bäckhed, F. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 2013, 498, 99–103. [Google Scholar] [CrossRef]
- Li, D.; Liu, C.; Luo, R.; Sadakane, K.; Lam, T. MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 2015, 31, 1674–1676. [Google Scholar] [CrossRef]
- Zhu, W.; Lomsadze, A.; Borodovsky, M. Ab initio gene identification in metagenomics sequences. Nucleic Acids Res. 2010, 38, e132. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 2006, 22, 1658–1659. [Google Scholar] [CrossRef] [PubMed]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef] [PubMed]
- Luo, B.; Pan, D.; Gao, Q.; Chen, Q.; Hou, P.; Song, P.; Wang, C. Design and experiment of rice program control germination system in cold region based on internet of things. Trans. Chin. Soc. Agricult. Eng. 2018, 34, 181–185. [Google Scholar]
- Zhao, H.; Liu, Q.; Fu, H.; Xu, X.; Wu, D.; Shu, Q. Effect of non-lethal low phytic acid mutations on grain yield and seed viability in rice. Field Crops Res. 2008, 108, 206–211. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Yuan, Y.; Yuan, H.; Wang, Y.; Jin, C.; Zhang, H.; Tang, J.; Hu, Y. Metagenomic Sequencing to Analyze Composition and Function of Top-Gray Chalky Grain Microorganisms from Hybrid Rice Seeds. Plants 2023, 12, 2358. https://doi.org/10.3390/plants12122358
Liu Y, Yuan Y, Yuan H, Wang Y, Jin C, Zhang H, Tang J, Hu Y. Metagenomic Sequencing to Analyze Composition and Function of Top-Gray Chalky Grain Microorganisms from Hybrid Rice Seeds. Plants. 2023; 12(12):2358. https://doi.org/10.3390/plants12122358
Chicago/Turabian StyleLiu, You, Yuan Yuan, Hui Yuan, Yan Wang, Chenzhong Jin, Hao Zhang, Jianliang Tang, and Yihong Hu. 2023. "Metagenomic Sequencing to Analyze Composition and Function of Top-Gray Chalky Grain Microorganisms from Hybrid Rice Seeds" Plants 12, no. 12: 2358. https://doi.org/10.3390/plants12122358
APA StyleLiu, Y., Yuan, Y., Yuan, H., Wang, Y., Jin, C., Zhang, H., Tang, J., & Hu, Y. (2023). Metagenomic Sequencing to Analyze Composition and Function of Top-Gray Chalky Grain Microorganisms from Hybrid Rice Seeds. Plants, 12(12), 2358. https://doi.org/10.3390/plants12122358




