Functional Differences of Grapevine Circular RNA Vv-circPTCD1 in Arabidopsis and Grapevine Callus under Abiotic Stress
Abstract
:1. Introduction
2. Results
2.1. Identification and Verification of Vv-circPTCD1
2.2. Nonconservative Back-Splicing of Vv-circPTCD1 in Plants
2.3. The Expression Analyses of PTCD1 and Vv-circPTCD1 under Abiotic Stress
2.4. Overexpression of Vv-circPTCD1 in Grapevine Callus and Arabidopsis
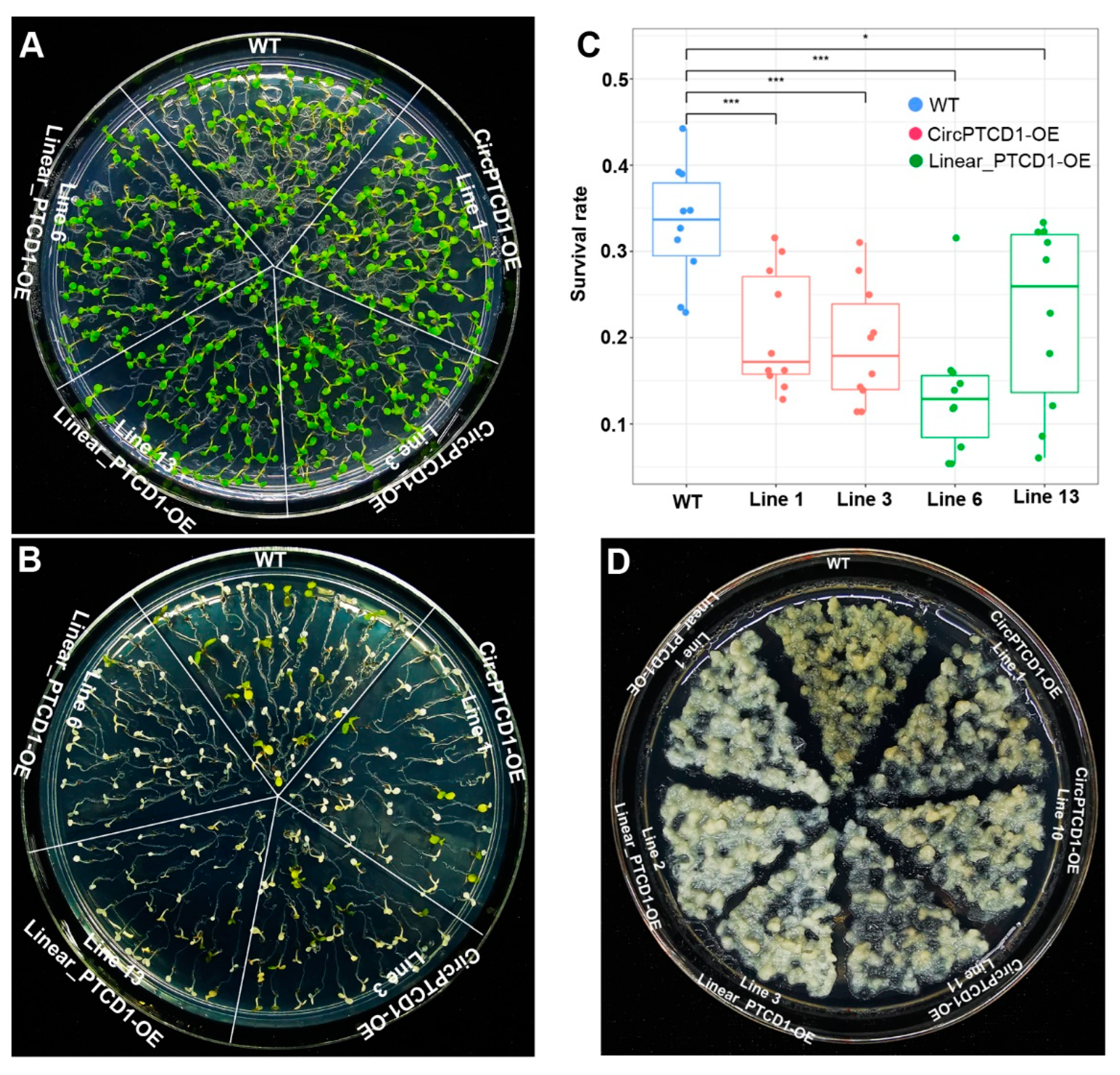
2.5. Phenotype of OE Lines under Heat Stress
2.6. Phenotype of OE Lines under Salt Stress
2.7. Phenotype of OE Lines under Drought Stress
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Treatments
4.2. Validation of circRNA and RT-qPCR
4.3. Vector Construction
4.4. Arabidopsis Transformation and Verification
4.5. Callus Transformation of ‘Thompson Seedless’ and Treatment
4.6. Statistical Analysis and Gene Structure
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Wang, Y.; Yang, M.; Wei, S.; Qin, F.; Zhao, H.; Suo, B. Identification of Circular RNAs and Their Targets in Leaves of Triticum aestivum L. under Dehydration Stress. Front. Plant Sci. 2016, 7, 2024. [Google Scholar] [CrossRef]
- Zhou, Y.; Wang, X.; Qi, K.; Bao, J.; Zhang, S.; Gu, C. Involvement of long non-coding RNAs in pear fruit senescence under high- and low-temperature conditions. Hortic. Plant J. 2022, 198, 112251. [Google Scholar] [CrossRef]
- Chen, L.L. The expanding regulatory mechanisms and cellular functions of circular RNAs. Nat. Rev. Mol. Cell Biol. 2020, 21, 475–490. [Google Scholar] [CrossRef] [PubMed]
- Chu, Q.; Zhang, X.; Zhu, X.; Liu, C.; Mao, L.; Ye, C.; Zhu, Q.H.; Fan, L. PlantcircBase: A Database for Plant Circular RNAs. Mol. Plant 2017, 10, 1126–1128. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Meng, X.; Chen, H.; Liu, Y.; Xue, J.; Zhou, Y.; Chen, M. PlantCircNet: A database for plant circRNA-miRNA-mRNA regulatory networks. Database 2017, 2017, bax089. [Google Scholar] [CrossRef] [PubMed]
- Chu, Q.; Bai, P.; Zhu, X.; Zhang, X.; Mao, L.; Zhu, Q.H.; Fan, L.; Ye, C.Y. Characteristics of plant circular RNAs. Brief. Bioinform. 2020, 21, 135–143. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef]
- Legnini, I.; Di Timoteo, G.; Rossi, F.; Morlando, M.; Briganti, F.; Sthandier, O.; Fatica, A.; Santini, T.; Andronache, A.; Wade, M.; et al. Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis. Mol. Cell 2017, 66, 22–37. [Google Scholar] [CrossRef]
- Yang, Y.; Gao, X.; Zhang, M.; Yan, S.; Sun, C.; Xiao, F.; Huang, N.; Yang, X.; Zhao, K.; Zhou, H.; et al. Novel Role of FBXW7 Circular RNA in Repressing Glioma Tumorigenesis. J. Natl. Cancer Inst. 2018, 110, 304–315. [Google Scholar] [CrossRef]
- Li, Q.; Wang, Y.; Wu, S.; Zhou, Z.; Ding, X.; Shi, R.; Thorne, R.F.; Zhang, X.D.; Hu, W.; Wu, M. CircACC1 Regulates Assembly and Activation of AMPK Complex under Metabolic Stress. Cell Metab. 2019, 30, 157–173. [Google Scholar] [CrossRef]
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264. [Google Scholar] [CrossRef] [PubMed]
- Tan, J.; Zhou, Z.; Niu, Y.; Sun, X.; Deng, Z. Identification and Functional Characterization of Tomato CircRNAs Derived from Genes Involved in Fruit Pigment Accumulation. Sci. Rep. 2017, 7, 8594. [Google Scholar] [CrossRef] [PubMed]
- Conn, V.M.; Hugouvieux, V.; Nayak, A.; Conos, S.A.; Capovilla, G.; Cildir, G.; Jourdain, A.; Tergaonkar, V.; Schmid, M.; Zubieta, C.; et al. A circRNA from SEPALLATA3 regulates splicing of its cognate mRNA through R-loop formation. Nat. Plants 2017, 3, 17053. [Google Scholar] [CrossRef]
- Zhou, J.; Yuan, M.; Zhao, Y.; Quan, Q.; Yu, D.; Yang, H.; Tang, X.; Xin, X.; Cai, G.; Qian, Q.; et al. Efficient deletion of multiple circle RNA loci by CRISPR-Cas9 reveals Os06circ02797 as a putative sponge for OsMIR408 in rice. Plant Biotechnol. J. 2021, 19, 1240–1252. [Google Scholar] [CrossRef]
- Song, Y.; Bu, C.; Chen, P.; Liu, P.; Zhang, D. Miniature inverted repeat transposable elements cis-regulate circular RNA expression and promote ethylene biosynthesis, reducing heat tolerance in Populus tomentosa. J. Exp. Bot. 2021, 72, 1978–1994. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Liu, S.; Li, X.; Yao, L.; Wu, H.; Baluška, F.; Wan, Y. An antisense circular RNA regulates expression of RuBisCO small subunit genes in Arabidopsis. Front. Plant Sci. 2021, 12, 665014. [Google Scholar] [CrossRef] [PubMed]
- Liao, X.; Li, X.J.; Zheng, G.T.; Chang, F.R.; Fang, L.; Yu, H.; Huang, J.; Zhang, Y.F. Mitochondrion-encoded circular RNAs are widespread and translatable in plants. Plant Physiol. 2022, 189, 1482–1500. [Google Scholar] [CrossRef]
- Baroi, A.M.; Popitiu, M.; Fierascu, I.; Sardarescu, I.D.; Fierascu, R.C. Grapevine Wastes: A Rich Source of Antioxidants and Other Biologically Active Compounds. Antioxidants 2022, 11, 393. [Google Scholar] [CrossRef]
- Venios, X.; Korkas, E.; Nisiotou, A.; Banilas, G. Grapevine Responses to Heat Stress and Global Warming. Plants 2020, 9, 1754. [Google Scholar] [CrossRef]
- Zhu, Z.; Quan, R.; Chen, G.; Yu, G.; Li, X.; Han, Z.; Xu, W.; Li, G.; Shi, J.; Li, B. An R2R3-MYB transcription factor VyMYB24, isolated from wild grape Vitis yanshanesis J. X. Chen., regulates the plant development and confers the tolerance to drought. Front. Plant Sci. 2022, 13, 966641. [Google Scholar] [CrossRef]
- Ju, Y.L.; Yue, X.F.; Min, Z.; Wang, X.H.; Fang, Y.L.; Zhang, J.X. VvNAC17, a novel stress-responsive grapevine (Vitis vinifera L.) NAC transcription factor, increases sensitivity to abscisic acid and enhances salinity, freezing, and drought tolerance in transgenic Arabidopsis. Plant Physiol. Biochem. 2020, 146, 98–111. [Google Scholar] [CrossRef] [PubMed]
- Zha, Q.; Xi, X.J.; He, Y.N.; Jiang, A.L. Transcriptomic analysis of the leaves of two grapevine cultivars under high-temperature stress. Sci. Hortic. 2020, 265, 109265. [Google Scholar] [CrossRef]
- Merrill, N.K.; García de Cortázar-Atauri, I.; Parker, A.K.; Walker, M.A.; Wolkovich, E.M. Exploring grapevine phenology and high temperatures response under controlled conditions. Front. Environ. Sci. 2020, 8, 516527. [Google Scholar] [CrossRef]
- Ryu, S.; Han, J.H.; Cho, J.G.; Jeong, J.H.; Lee, S.K.; Lee, H.J. High temperature at veraison inhibits anthocyanin biosynthesis in berry skins during ripening in ‘Kyoho’ grapevines. Plant Physiol. Biochem. 2020, 157, 219–228. [Google Scholar] [CrossRef] [PubMed]
- Jiang, J.; Liu, X.; Liu, C.; Liu, G.; Li, S.; Wang, L. Integrating omics and alternative splicing reveals insights into grape response to high temperature. Plant Physiol. 2017, 173, 1502–1518. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.T.; Jiang, J.F.; Liu, X.N.; Jiang, J.Z.; Sun, L.; Duan, W.; Li, R.M.; Wang, Y.; Lecourieux, D.; Liu, C.H.; et al. New insights into the heat responses of grape leaves via combined phosphoproteomic and acetylproteomic analyses. Hortic. Res. 2019, 6, 100. [Google Scholar] [CrossRef]
- Haider, S.; Iqbal, J.; Naseer, S.; Yaseen, T.; Shaukat, M.; Bibi, H.; Ahmad, Y.; Daud, H.; Abbasi, N.L.; Mahmood, T. Molecular mechanisms of plant tolerance to heat stress: Current landscape and future perspectives. Plant Cell Rep. 2021, 40, 2247–2271. [Google Scholar] [CrossRef]
- Pillet, J.; Egert, A.; Pieri, P.; Lecourieux, F.; Kappel, C.; Charon, J.; Gomes, E.; Keller, F.; Delrot, S.; Lecourieux, D. VvGOLS1 and VvHsfA2 are involved in the heat stress responses in grapevine berries. Plant Cell Physiol. 2012, 53, 1776–1792. [Google Scholar] [CrossRef] [Green Version]
- Liu, G.; Chai, F.; Wang, Y.; Jiang, J.; Duan, W.; Wang, Y.; Wang, F.; Li, S.; Wang, L. Genome-wide identification and classification of HSF family in grape, and their transcriptional analysis under heat acclimation and heat stress. Hortic. Plant J. 2018, 4, 133–143. [Google Scholar] [CrossRef]
- Gambetta, G.A.; Herrera, J.C.; Dayer, S.; Feng, Q.; Hochberg, U.; Castellarin, S.D. The physiology of drought stress in grapevine: Towards an integrative definition of drought tolerance. J. Exp. Bot. 2020, 71, 4658–4676. [Google Scholar] [CrossRef]
- Mao, H.; Jian, C.; Cheng, X.; Chen, B.; Mei, F.; Li, F.; Zhang, Y.; Li, S.; Du, L.; Li, T.; et al. The wheat ABA receptor gene TaPYL1-1B contributes to drought tolerance and grain yield by increasing water-use efficiency. Plant Biotechnol. J. 2022, 20, 846–861. [Google Scholar] [CrossRef]
- Qin, T.; Zhao, H.; Cui, P.; Albesher, N.; Xiong, L. A nucleus-localized long non-coding RNA enhances drought and salt stress tolerance. Plant Physiol. 2017, 175, 1321–1336. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Duan, Y.; Sun, N.; Wang, L.; Feng, S.; Fang, Y.; Wang, Y. The miR169n-NF-YA8 regulation module involved in drought resistance in Brassica napus L. Plant Sci. 2021, 313, 111062. [Google Scholar] [CrossRef] [PubMed]
- Pagliarani, C.; Vitali, M.; Ferrero, M.; Vitulo, N.; Incarbone, M.; Lovisolo, C.; Valle, G.; Schubert, A. The accumulation of miRNAs differentially modulated by drought stress is affected by grafting in grapevine. Plant Physiol. 2017, 173, 2180–2195. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Guo, Y. Unraveling salt stress signaling in plants. J. Integr. Plant Biol. 2018, 60, 796–804. [Google Scholar] [CrossRef]
- Yu, Y.; Ni, Y.; Qiao, T.; Ji, X.; Xu, J.; Li, B.; Sun, Q. Overexpression of VvASMT1 from grapevine enhanced salt and osmotic stress tolerance in Nicotiana benthamiana. PLoS ONE 2022, 17, e0269028. [Google Scholar] [CrossRef]
- Mzid, R.; Zorrig, W.; Ben Ayed, R.; Ben Hamed, K.; Ayadi, M.; Damak, Y.; Lauvergeat, V.; Hanana, M. The grapevine VvWRKY2 gene enhances salt and osmotic stress tolerance in transgenic Nicotiana tabacum. 3 Biotech 2018, 8, 277. [Google Scholar] [CrossRef]
- Gao, Z.; Li, J.; Luo, M.; Li, H.; Chen, Q.; Wang, L.; Song, S.; Zhao, L.; Xu, W.; Zhang, C.; et al. Characterization and cloning of Grape Circular RNAs Identified the Cold Resistance-Related Vv-circATS1. Plant Physiol. 2019, 180, 966–985. [Google Scholar] [CrossRef]
- Xing, H.; Fu, X.; Yang, C.; Tang, X.; Guo, L.; Li, C.; Xu, C.; Luo, K. Genome-wide investigation of pentatricopeptide repeat gene family in poplar and their expression analysis in response to biotic and abiotic stresses. Sci. Rep. 2018, 8, 2817. [Google Scholar] [CrossRef]
- Fujita, Y.; Nakashima, K.; Yoshida, T.; Katagiri, T.; Kidokoro, S.; Kanamori, N.; Umezawa, T.; Fujita, M.; Maruyama, K.; Ishiyama, K.; et al. Three SnRK2 protein kinases are the main positive regulators of abscisic acid signaling in response to water stress in Arabidopsis. Plant Cell Physiol. 2009, 50, 2123–2132. [Google Scholar] [CrossRef]
- Chu, Q.; Ding, Y.; Xu, X.; Ye, C.Y.; Zhu, Q.H.; Guo, L.; Fan, L. Recent origination of circular RNAs in plants. New Phytol. 2022, 233, 515–525. [Google Scholar] [CrossRef] [PubMed]
- Lurin, C.; Andres, C.; Aubourg, S.; Bellaoui, M.; Bitton, F.; Bruyere, C.; Caboche, M.; Debast, C.; Gualberto, J.; Hoffmann, B.; et al. Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell 2004, 16, 2089–2103. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Dong, F.; Wang, X.; Wang, T.; Su, R.; Hong, D.; Yang, G. A pentatricopeptide repeat protein restores nap cytoplasmic male sterility in Brassica napus. J. Exp. Bot. 2017, 68, 4115–4123. [Google Scholar] [CrossRef] [PubMed]
- Sosso, D.; Canut, M.; Gendrot, G.; Dedieu, A.; Chambrier, P.; Barkan, A.; Consonni, G.; Rogowsky, P.M. PPR8522 encodes a chloroplast-targeted pentatricopeptide repeat protein necessary for maize embryogenesis and vegetative development. J. Exp. Bot. 2012, 63, 5843–5857. [Google Scholar] [CrossRef]
- Li, X.; Gu, W.; Sun, S.; Chen, Z.; Chen, J.; Song, W.; Zhao, H.; Lai, J. Defective Kernel 39 encodes a PPR protein required for seed development in maize. J. Integr. Plant Biol. 2018, 60, 45–64. [Google Scholar] [CrossRef]
- Laluk, K.; Abuqamar, S.; Mengiste, T. The Arabidopsis mitochondria-localized pentatricopeptide repeat protein PGN functions in defense against necrotrophic fungi and abiotic stress tolerance. Plant Physiol. 2011, 156, 2053–2068. [Google Scholar] [CrossRef]
- Xu, C.; Zhang, J. Mammalian circular RNAs result largely from splicing errors. Cell Rep. 2021, 36, 109439. [Google Scholar] [CrossRef]
- Cocquerelle, C.; Mascrez, B.; Hétuin, D.; Bailleul, B. Mis-splicing yields circular RNA molecules. FASEB J. 1993, 7, 6. [Google Scholar] [CrossRef]
- Zhang, X.O.; Wang, H.B.; Zhang, Y.; Lu, X.; Chen, L.L.; Yang, L. Complementary sequence-mediated exon circularization. Cell 2014, 159, 134–147. [Google Scholar] [CrossRef]
- Liu, C.X.; Chen, L.L. Circular RNAs: Characterization, cellular roles, and applications. Cell 2022, 185, 2016–2034. [Google Scholar] [CrossRef]
- Zhang, P.; Fan, Y.; Sun, X.; Chen, L.; Terzaghi, W.; Bucher, E.; Li, L.; Dai, M. A large-scale circular RNA profiling reveals universal molecular mechanisms responsive to drought stress in maize and Arabidopsis. Plant J. 2019, 98, 697–713. [Google Scholar] [CrossRef]
- Liang, D.; Wilusz, J.E. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 2014, 28, 2233–2247. [Google Scholar] [CrossRef]
- Liu, Y.; Su, H.; Zhang, J.; Liu, Y.; Feng, C.; Han, F. Back-spliced RNA from retrotransposon binds to centromere and regulates centromeric chromatin loops in maize. PLoS Biol. 2020, 18, e3000582. [Google Scholar] [CrossRef]
- Fan, J.; Quan, W.; Li, G.B.; Hu, X.H.; Wang, Q.; Wang, H.; Li, X.P.; Luo, X.; Feng, Q.; Hu, Z.J.; et al. CircRNAs are involved in the rice-magnaporthe oryzae interaction. Plant Physiol. 2020, 182, 272–286. [Google Scholar] [CrossRef]
- Liu, R.; Ma, Y.; Guo, T.; Li, G. Identification, biogenesis, function, and mechanism of action of circular RNAs in plants. Plant Commun. 2022, 4, 100430. [Google Scholar] [CrossRef]
- Verslues, P.E.; Agarwal, M.; Katiyar-Agarwal, S.; Zhu, J.; Zhu, J.K. Methods and concepts in quantifying resistance to drought, salt and freezing, abiotic stresses that affect plant water status. Plant J. 2006, 45, 523–539. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−∆∆CT) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip a simplified method for Agrobacterium -mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 9. [Google Scholar] [CrossRef] [PubMed]
- Gambino, G.; Ruffa, P.; Vallania, R.; Gribaudo, I. Somatic embryogenesis from whole flowers, anthers and ovaries of grapevine (Vitis spp.). Plant Cell Tissue Organ Cult. 2007, 90, 79–83. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed]





Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ren, Y.; Li, J.; Liu, J.; Zhang, Z.; Song, Y.; Fan, D.; Liu, M.; Zhang, L.; Xu, Y.; Guo, D.; et al. Functional Differences of Grapevine Circular RNA Vv-circPTCD1 in Arabidopsis and Grapevine Callus under Abiotic Stress. Plants 2023, 12, 2332. https://doi.org/10.3390/plants12122332
Ren Y, Li J, Liu J, Zhang Z, Song Y, Fan D, Liu M, Zhang L, Xu Y, Guo D, et al. Functional Differences of Grapevine Circular RNA Vv-circPTCD1 in Arabidopsis and Grapevine Callus under Abiotic Stress. Plants. 2023; 12(12):2332. https://doi.org/10.3390/plants12122332
Chicago/Turabian StyleRen, Yi, Junpeng Li, Jingjing Liu, Zhen Zhang, Yue Song, Dongying Fan, Minying Liu, Lipeng Zhang, Yuanyuan Xu, Dinghan Guo, and et al. 2023. "Functional Differences of Grapevine Circular RNA Vv-circPTCD1 in Arabidopsis and Grapevine Callus under Abiotic Stress" Plants 12, no. 12: 2332. https://doi.org/10.3390/plants12122332
APA StyleRen, Y., Li, J., Liu, J., Zhang, Z., Song, Y., Fan, D., Liu, M., Zhang, L., Xu, Y., Guo, D., He, J., Song, S., Gao, Z., & Ma, C. (2023). Functional Differences of Grapevine Circular RNA Vv-circPTCD1 in Arabidopsis and Grapevine Callus under Abiotic Stress. Plants, 12(12), 2332. https://doi.org/10.3390/plants12122332





