Genome-Wide Identification and Expression Analysis of the Zinc Finger Protein Gene Subfamilies under Drought Stress in Triticum aestivum
Abstract
1. Introduction
2. Results
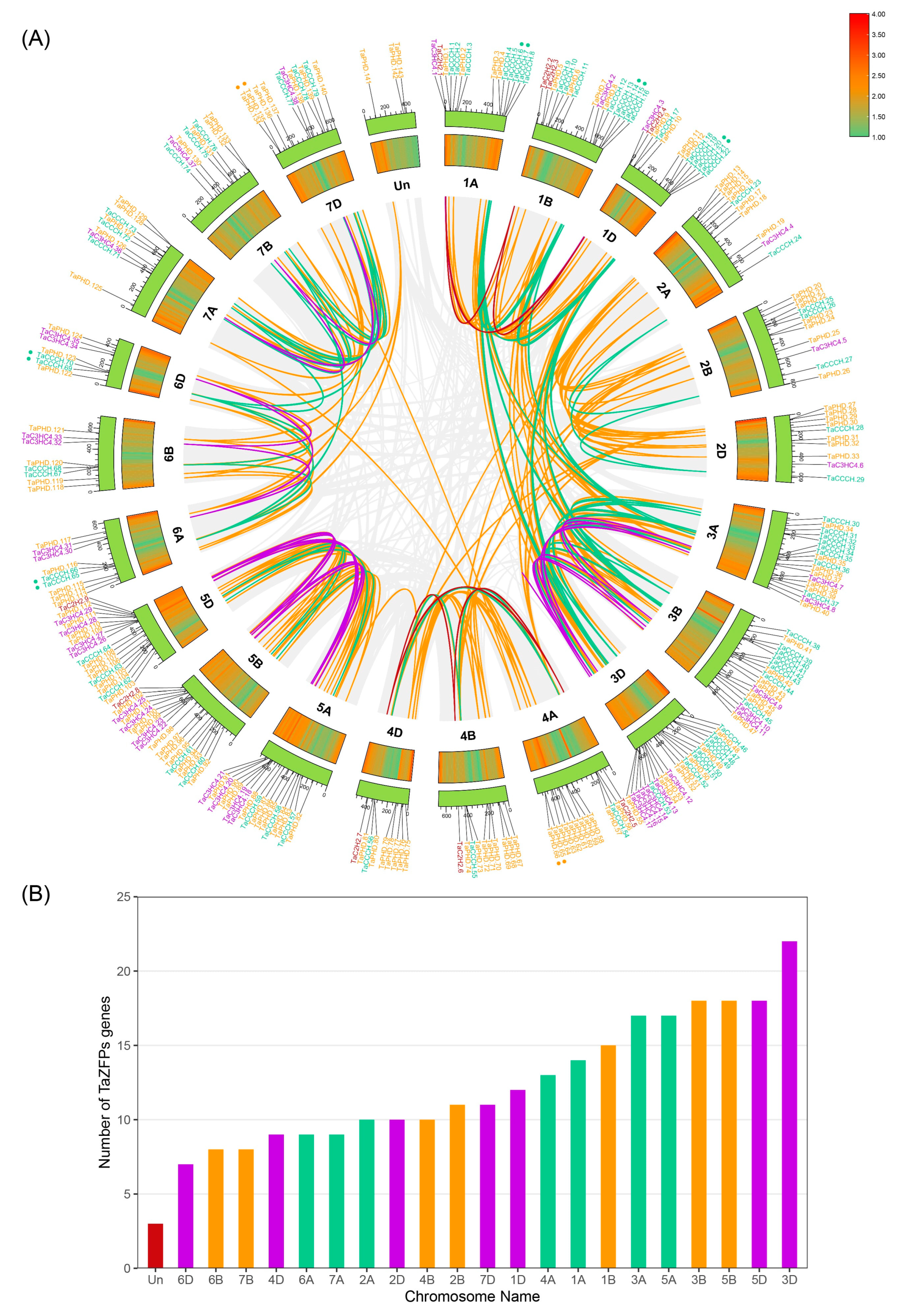
2.1. Identification and Chromosomal Distribution of the TaZFPs
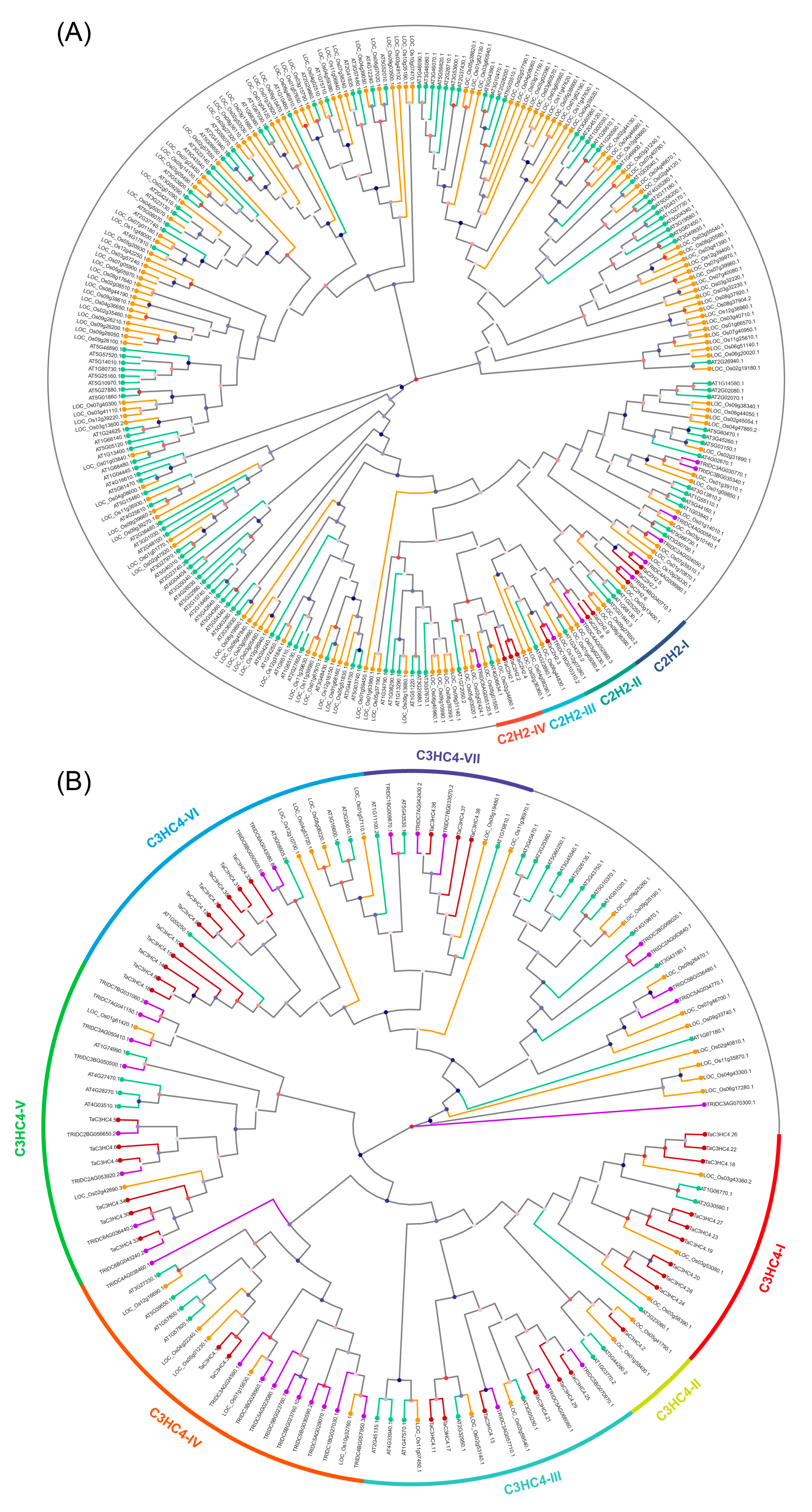
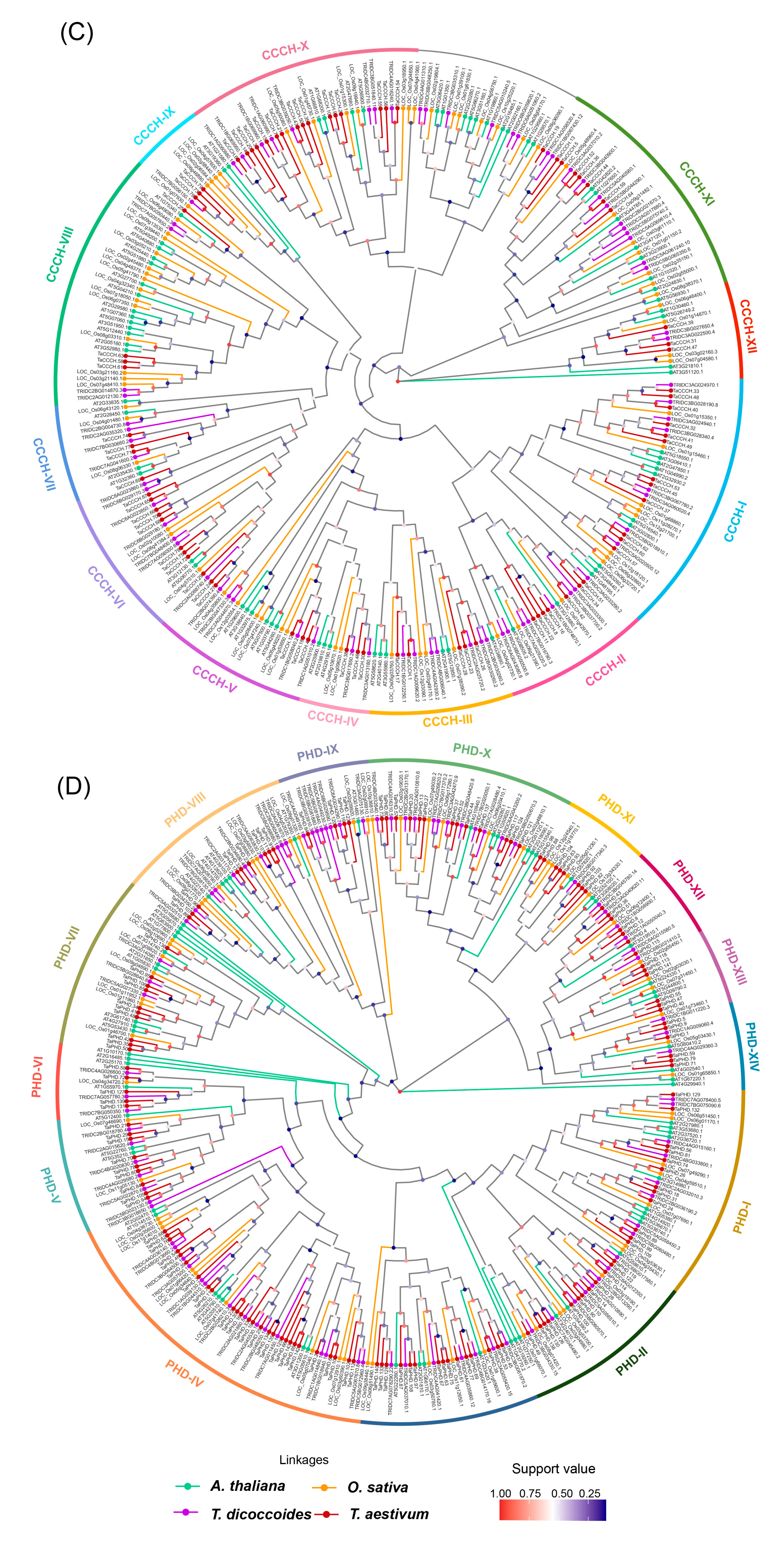
2.2. Multiple Sequence Alignment and Phylogenetic Analysis of TaZFPs
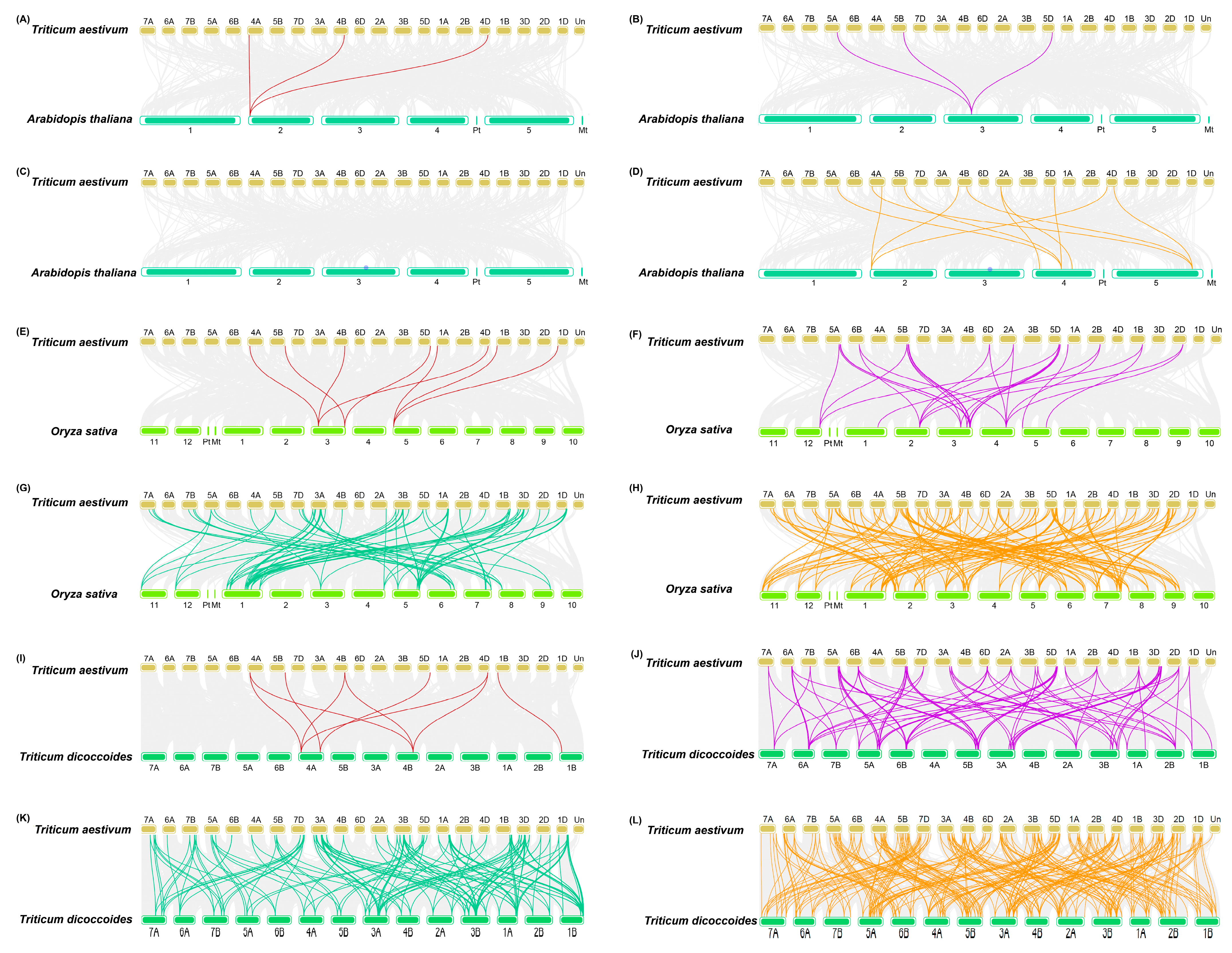
2.3. Gene Duplication, Synteny, and Ka/Ks Analysis of TaZFPs
2.4. Gene Structure and Conserved Motifs of Four TaZFP Gene Subfamilies
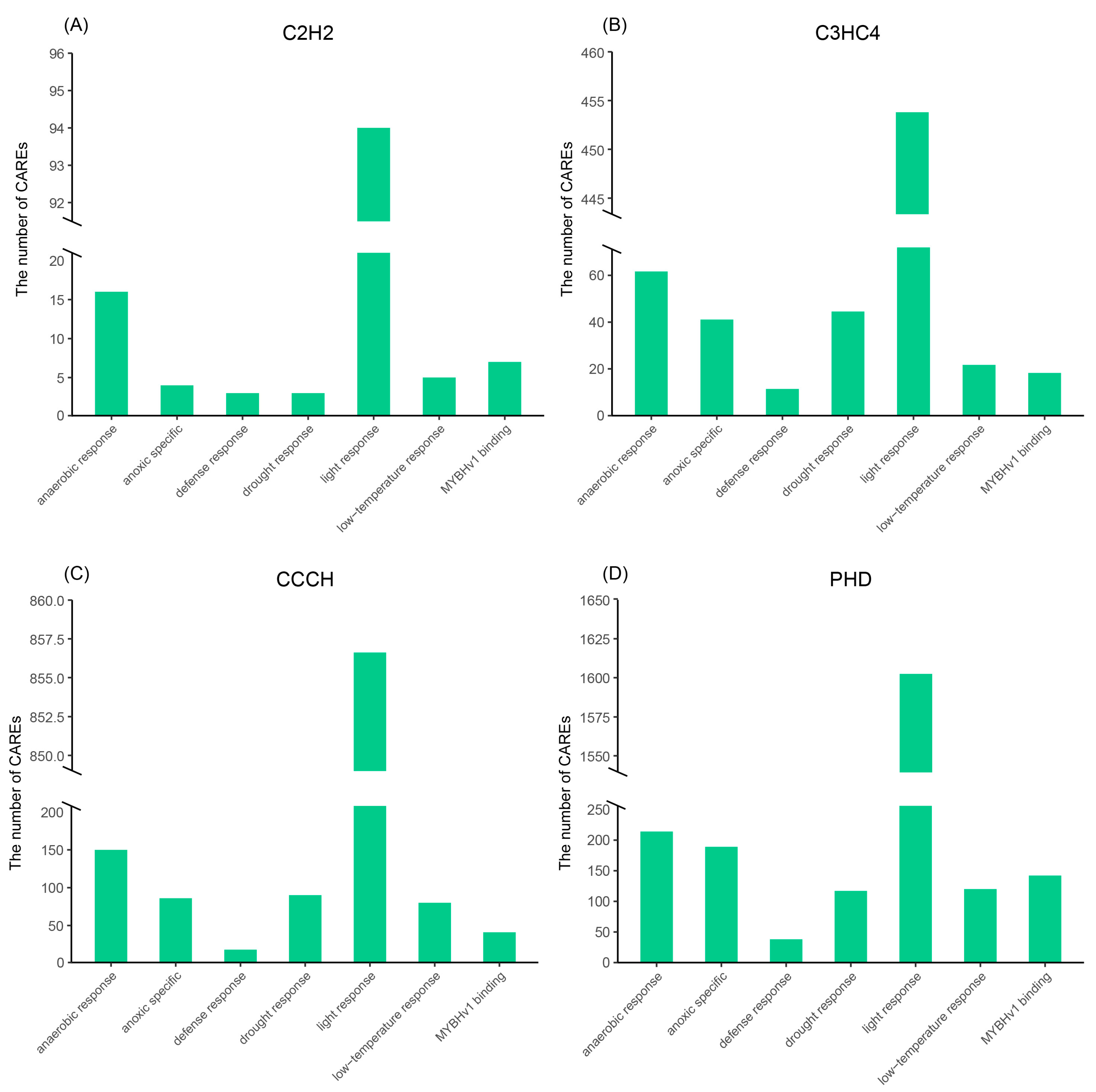
2.5. Stress-Related Cis-Acting Regulatory Elements (CAREs) of the TaZFP Gene Promoter
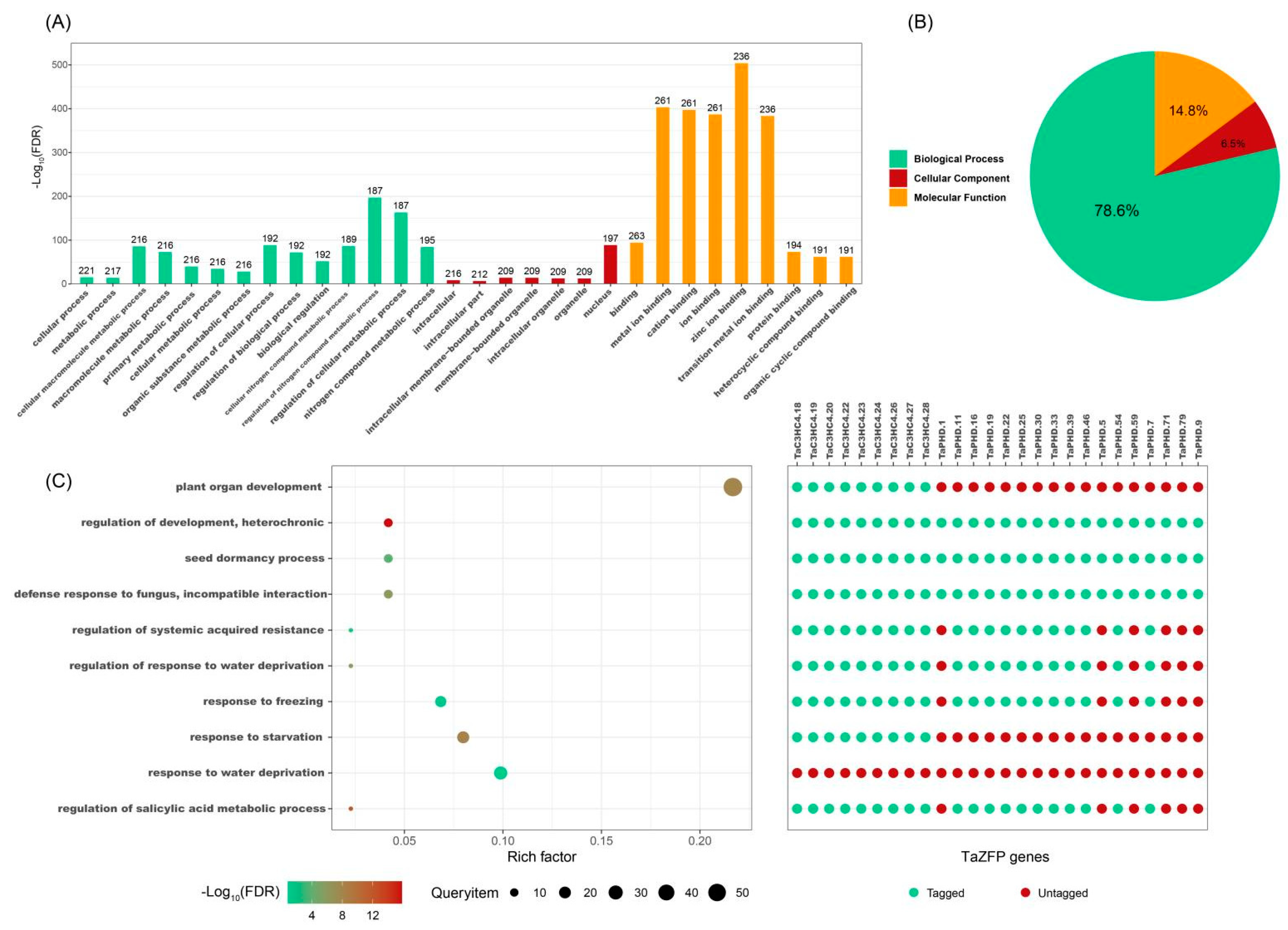
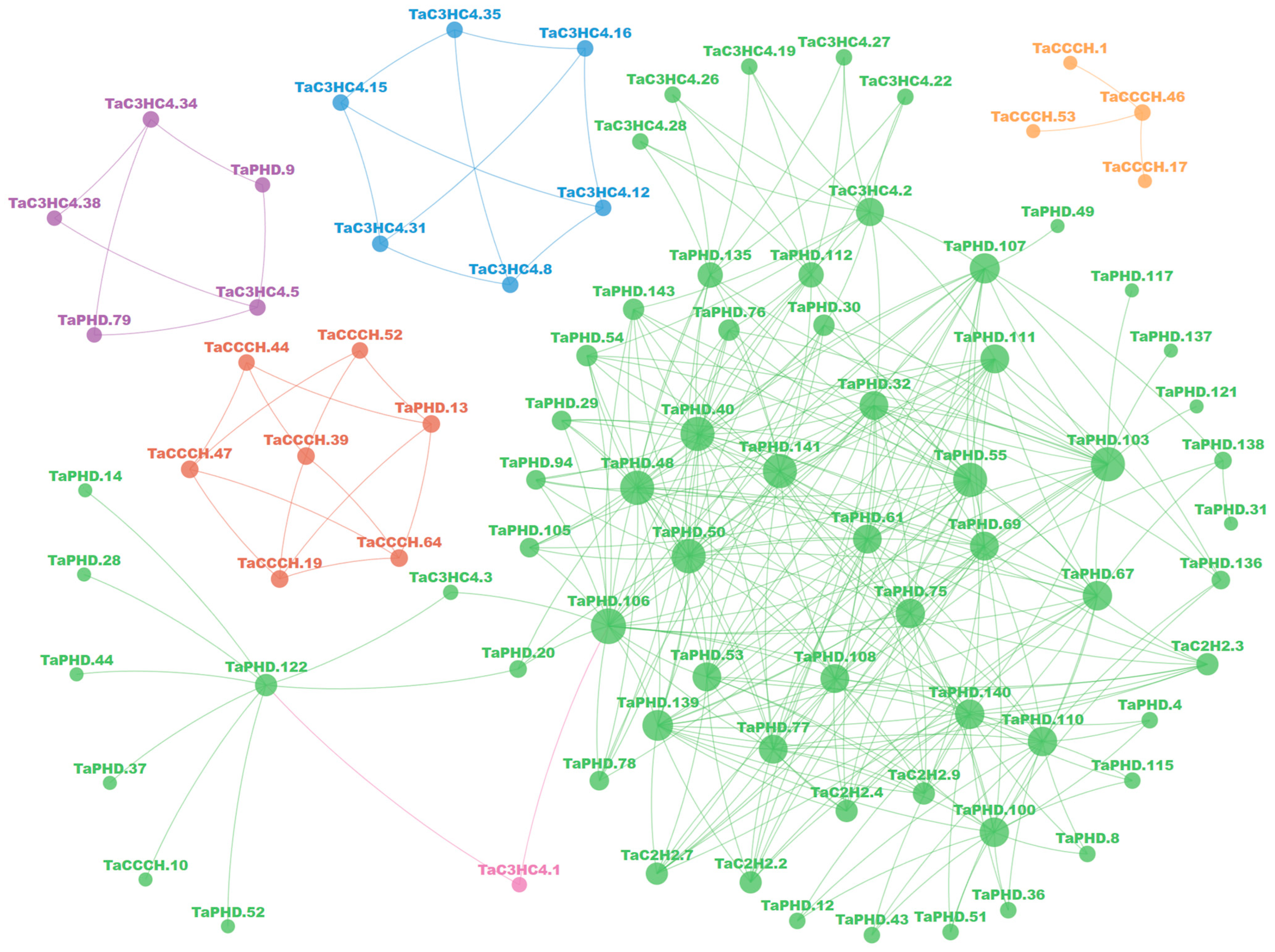
2.6. Gene Ontology (GO) Enrichment and Protein-Protein Interaction Network of TaZFPs
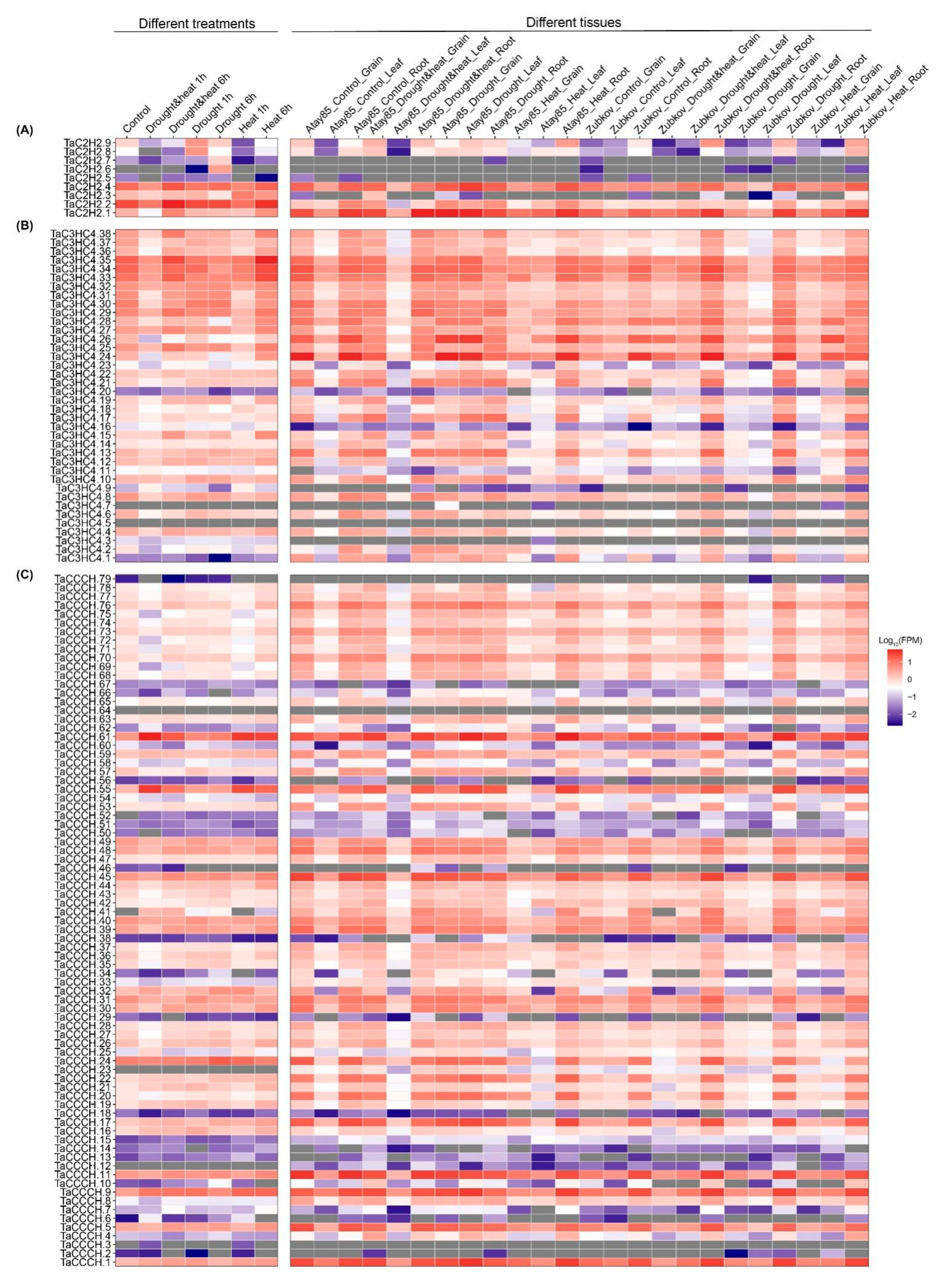
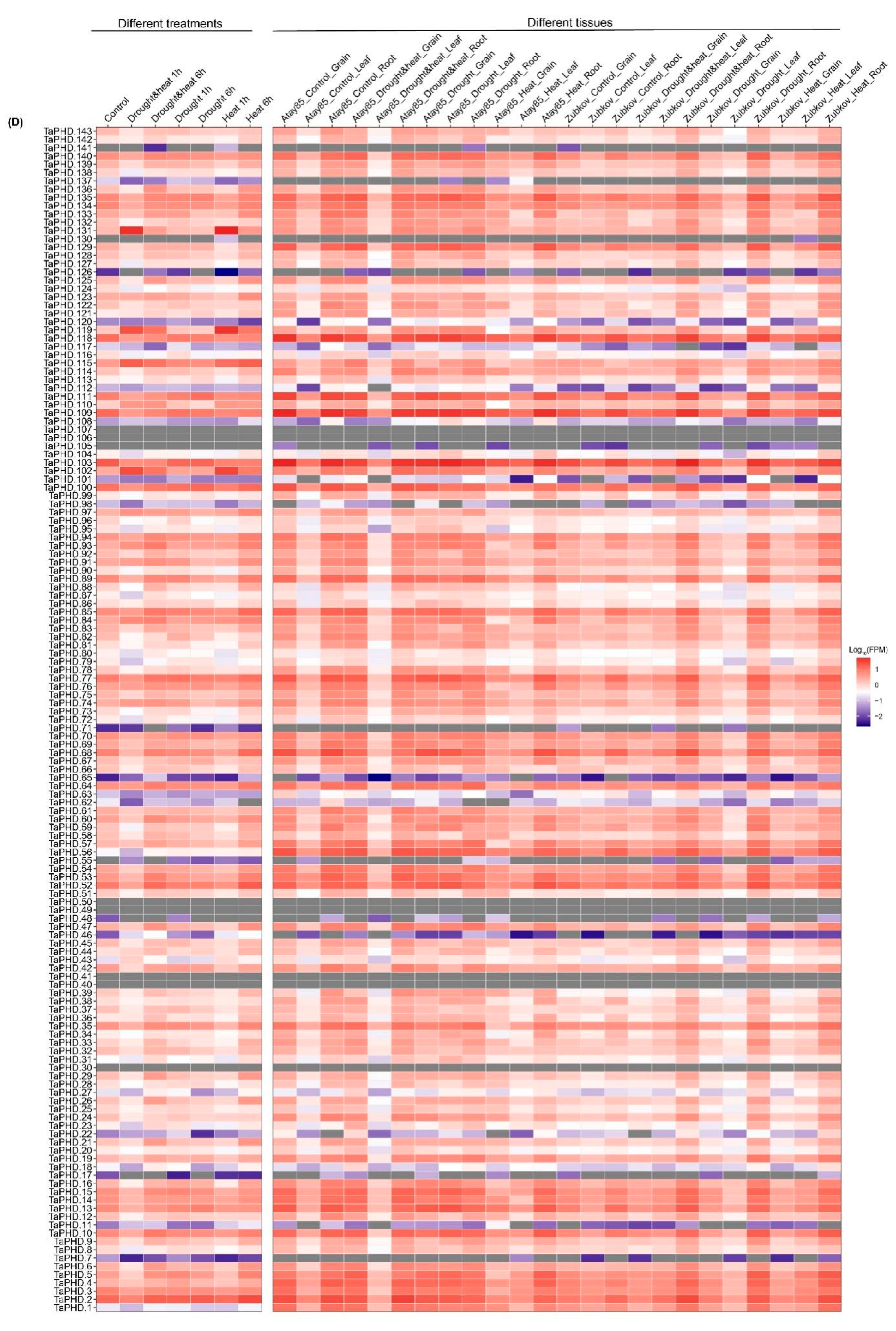
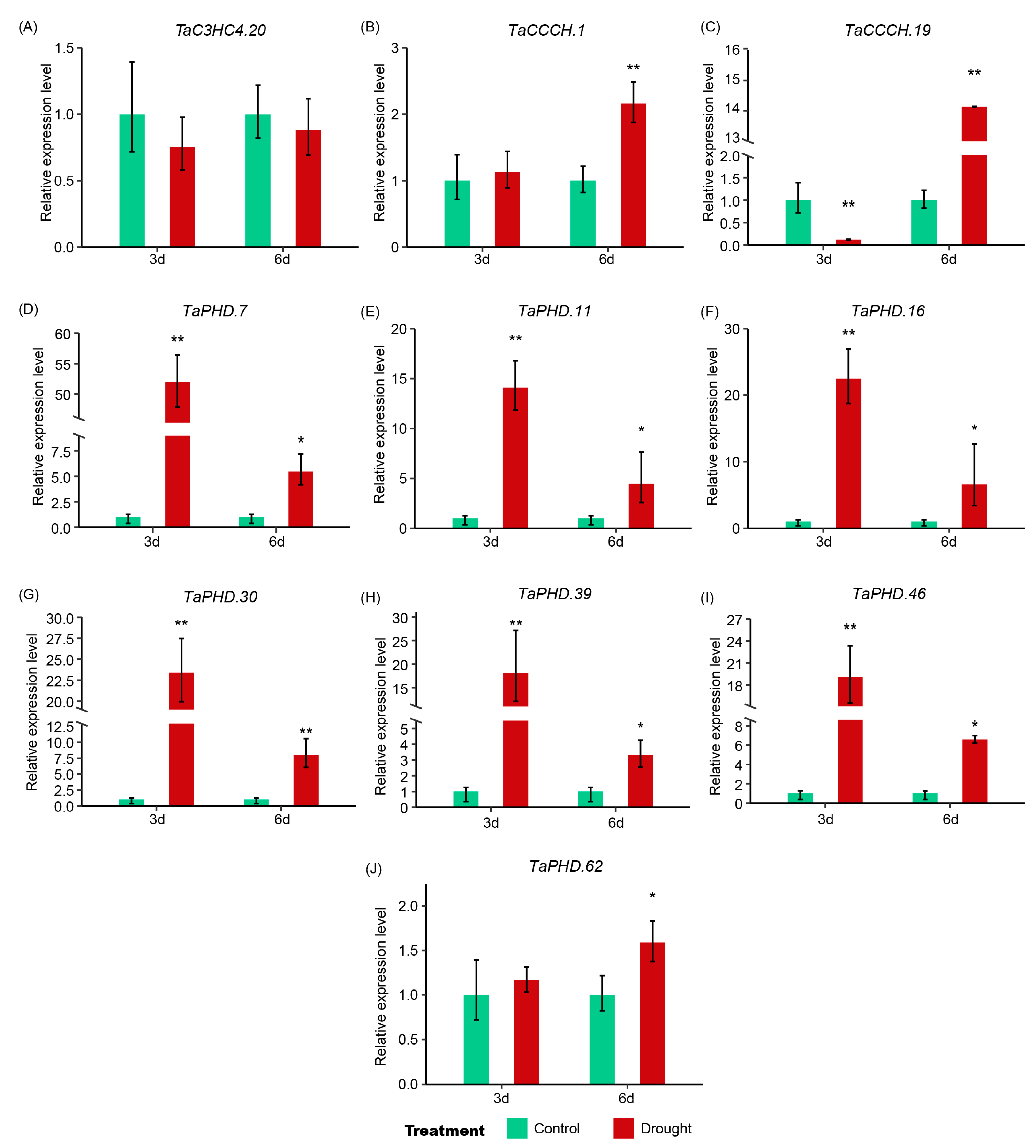
2.7. Expression Pattern Analysis of TaZFPs under Abiotic Stresses
3. Discussion
4. Materials and Methods
4.1. Identification of TaTFPs in Triticum aestivum
4.2. Multiple Sequence Alignment and Phylogenetic Analysis
4.3. Determination of Chromosome Distribution, Synteny, and Ka/Ks of 4 TaZFP Gene Subfamilies
4.4. Identification of Gene Structure, Conserved Motifs, and Cis-Acting Regulatory Elements
4.5. GO Enrichment and Protein Interaction Network Establishment
4.6. Gene Expression Analysis
4.7. Plants Material and Culture
4.8. RNA Extraction and qRT-PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ahuja, I.; de Vos, R.C.H.; Bones, A.M.; Hall, R.D. Plant Molecular Stress Responses Face Climate Change. Trends Plant Sci. 2010, 15, 664–674. [Google Scholar] [CrossRef] [PubMed]
- Miryeganeh, M. Plants’ Epigenetic Mechanisms and Abiotic Stress. Genes 2021, 12, 1106. [Google Scholar] [CrossRef] [PubMed]
- Rehaman, A.; Mishra, A.K.; Ferdose, A.; Per, T.S.; Hanief, M.; Jan, A.T.; Asgher, M. Melatonin in Plant Defense against Abiotic Stress. Forests 2021, 12, 1404. [Google Scholar] [CrossRef]
- Naeem, M.; Shahzad, K.; Saqib, S.; Shahzad, A.; Nasrullah; Younas, M.; Afridi, M.I. The Solanum Melongena COP1LIKE Manipulates Fruit Ripening and Flowering Time in Tomato (Solanum lycopersicum). Plant Growth Regul. 2022, 96, 369–382. [Google Scholar] [CrossRef]
- Han, G.; Qiao, Z.; Li, Y.; Yang, Z.; Wang, C.; Zhang, Y.; Liu, L.; Wang, B. RING Zinc Finger Proteins in Plant Abiotic Stress Tolerance. Front. Plant Sci. 2022, 13, 877011. [Google Scholar] [CrossRef]
- Ayaz, A.; Huang, H.; Zheng, M.; Zaman, W.; Li, D.; Saqib, S.; Zhao, H.; Lü, S. Molecular Cloning and Functional Analysis of GmLACS2-3 Reveals Its Involvement in Cutin and Suberin Biosynthesis along with Abiotic Stress Tolerance. Int. J. Mol. Sci. 2021, 22, 9175. [Google Scholar] [CrossRef]
- Rathour, M.; Shumayla; Alok, A.; Upadhyay, S.K. Investigation of Roles of TaTALE Genes during Development and Stress Response in Bread Wheat. Plants 2022, 11, 587. [Google Scholar] [CrossRef]
- Bollier, N.; Gonzalez, N.; Chevalier, C.; Hernould, M. Zinc Finger-Homeodomain and Mini Zinc Finger Proteins Are Key Players in Plant Growth and Responses to Environmental Stresses. J. Exp. Bot. 2022, 73, 4662–4673. [Google Scholar] [CrossRef]
- Han, G.; Li, Y.; Qiao, Z.; Wang, C.; Zhao, Y.; Guo, J.; Chen, M.; Wang, B. Advances in the Regulation of Epidermal Cell Development by C2H2 Zinc Finger Proteins in Plants. Front. Plant Sci. 2021, 12, 754512. [Google Scholar] [CrossRef]
- Wang, K.; Ding, Y.; Cai, C.; Chen, Z.; Zhu, C. The Role of C2H2 Zinc Finger Proteins in Plant Responses to Abiotic Stresses. Physiol. Plant. 2019, 165, 690–700. [Google Scholar] [CrossRef]
- Han, G.; Lu, C.; Guo, J.; Qiao, Z.; Sui, N.; Qiu, N.; Wang, B. C2H2 Zinc Finger Proteins: Master Regulators of Abiotic Stress Responses in Plants. Front. Plant Sci. 2020, 11, 115. [Google Scholar] [CrossRef]
- Jiao, Z.; Wang, L.; Du, H.; Wang, Y.; Wang, W.; Liu, J.; Huang, J.; Huang, W.; Ge, L. Genome-Wide Study of C2H2 Zinc Finger Gene Family in Medicago Truncatula. BMC Plant Biol. 2020, 20, 401. [Google Scholar] [CrossRef] [PubMed]
- Dutta, S.K.; Nimmakayala, P.; Reddy, U.K. Genome-Wide Identification, Characterisation, and Expression of C3HC4-Type RING Finger Gene Family in Capsicum annuum L. J. Hortic. Sci. Biotechnol. 2022, 97, 603–614. [Google Scholar] [CrossRef]
- Kim, D.H.; Yamaguchi, S.; Lim, S.; Oh, E.; Park, J.; Hanada, A.; Kamiya, Y.; Choi, G. SOMNUS, a CCCH-Type Zinc Finger Protein in Arabidopsis, Negatively Regulates Light-Dependent Seed Germination Downstream of PIL5. Plant Cell 2008, 20, 1260. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.-Q.; Huang, J.; Guo, S.-Q.; Yang, X.; Bao, Y.-M.; Tang, H.-J.; Zhang, H.-S. Overexpression of a TFIIIA-Type Zinc Finger Protein Gene ZFP252 Enhances Drought and Salt Tolerance in Rice (Oryza sativa L.). FEBS Lett. 2008, 582, 1037–1043. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.; Chen, J.; Liu, M.; Zhang, H.; Zhang, S.; Liu, D.; Chen, S. Genome-Wide Analysis of C2H2 Zinc Finger Gene Family and Its Response to Cold and Drought Stress in Sorghum [Sorghum bicolor (L.) Moench]. Int. J. Mol. Sci. 2022, 23, 5571. [Google Scholar] [CrossRef]
- Tang, L.; Cai, H.; Ji, W.; Luo, X.; Wang, Z.; Wu, J.; Wang, X.; Cui, L.; Wang, Y.; Zhu, Y.; et al. Overexpression of GsZFP1 Enhances Salt and Drought Tolerance in Transgenic Alfalfa (Medicago sativa L.). Plant Physiol. Biochem. 2013, 71, 22–30. [Google Scholar] [CrossRef]
- Aceituno-Valenzuela, U.; Micol-Ponce, R.; Ponce, M.R. Genome-Wide Analysis of CCHC-Type Zinc Finger (ZCCHC) Proteins in Yeast, Arabidopsis, and Humans. Cell. Mol. Life Sci. 2020, 77, 3991–4014. [Google Scholar] [CrossRef]
- Wu, S.; Tong, X.; Li, C.; Lu, K.; Tan, D.; Hu, H.; Liu, H.; Dai, F. Genome-Wide Identification and Expression Profiling of the C2H2-Typ Zinc Finger Protein Genes in the Silkworm Bombyx Mori. PeerJ 2019, 7, e7222. [Google Scholar] [CrossRef]
- Zhao, T.; Wu, T.; Zhang, J.; Wang, Z.; Pei, T.; Yang, H.; Li, J.; Xu, X. Genome-Wide Analyses of the Genetic Screening of C2H2-Type Zinc Finger Transcription Factors and Abiotic and Biotic Stress Responses in Tomato (Solanum lycopersicum) Based on RNA-Seq Data. Front. Genet. 2020, 11, 540. [Google Scholar] [CrossRef]
- Ding, Q.; Zhao, H.; Zhu, P.; Jiang, X.; Nie, F.; Li, G. Genome-Wide Identification and Expression Analyses of C2H2 Zinc Finger Transcription Factors in Pleurotus Ostreatus. PeerJ 2022, 10, e12654. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Wang, G.; Pan, J.; Wen, H.; Du, H.; Sun, J.; Zhang, K.; Lv, D.; He, H.; Cai, R.; et al. Comprehensive Genomic Analysis and Expression Profiling of the C2H2 Zinc Finger Protein Family under Abiotic Stresses in Cucumber (Cucumis sativus L.). Genes 2020, 11, 171. [Google Scholar] [CrossRef] [PubMed]
- Brouns, F.R.; van Rooy, G.; Shewry, P.; Rustgi, S.; Jonkers, D. Adverse Reactions to Wheat or Wheat Components. Compr. Rev. Food Sci. Food Saf. 2019, 18, 1437–1452. [Google Scholar] [CrossRef] [PubMed]
- Shewry, P.R. Wheat. J. Exp. Bot. 2009, 60, 1537–1553. [Google Scholar] [CrossRef] [PubMed]
- Sallam, A.; Alqudah, A.M.; Dawood, M.F.A.; Baenziger, P.S.; Börner, A. Drought Stress Tolerance in Wheat and Barley: Advances in Physiology, Breeding and Genetics Research. Int. J. Mol. Sci. 2019, 20, 3137. [Google Scholar] [CrossRef]
- Sun, A.; Li, Y.; He, Y.; Zou, X.; Chen, F.; Ji, R.; You, C.; Yu, K.; Li, Y.; Xiao, W.; et al. Comprehensive Genome-Wide Identification, Characterization, and Expression Analysis of CCHC-Type Zinc Finger Gene Family in Wheat (Triticum aestivum L.). Front. Plant Sci. 2022, 13, 892105. [Google Scholar] [CrossRef]
- Faraji, S.; Rasouli, S.H.; Kazemitabar, S.K. Genome-Wide Exploration of C2H2 Zinc Finger Family in Durum Wheat (Triticum Turgidum Ssp. Durum): Insights into the Roles in Biological Processes Especially Stress Response. Biometals 2018, 31, 1019–1042. [Google Scholar] [CrossRef]
- Ma, R.; Chen, J.; Huang, B.; Huang, Z.; Zhang, Z. The BBX Gene Family in Moso Bamboo (Phyllostachys edulis): Identification, Characterization and Expression Profiles. BMC Genom. 2021, 22, 533. [Google Scholar] [CrossRef]
- He, P.; Yang, Y.; Wang, Z.; Zhao, P.; Yuan, Y.; Zhang, L.; Ma, Y.; Pang, C.; Yu, J.; Xiao, G. Comprehensive Analyses of ZFP Gene Family and Characterization of Expression Profiles during Plant Hormone Response in Cotton. BMC Plant Biol. 2019, 19, 329. [Google Scholar] [CrossRef]
- Kesawat, M.S.; Kherawat, B.S.; Singh, A.; Dey, P.; Routray, S.; Mohapatra, C.; Saha, D.; Ram, C.; Siddique, K.H.M.; Kumar, A.; et al. Genome-Wide Analysis and Characterization of the Proline-Rich Extensin-like Receptor Kinases (PERKs) Gene Family Reveals Their Role in Different Developmental Stages and Stress Conditions in Wheat (Triticum aestivum L.). Plants 2022, 11, 496. [Google Scholar] [CrossRef]
- Iuchi, S. Three Classes of C2H2 Zinc Finger Proteins. CMLS Cell. Mol. Life Sci. 2001, 58, 625–635. [Google Scholar] [CrossRef] [PubMed]
- Winicov, I. Alfin1 Transcription Factor Overexpression Enhances Plant Root Growth under Normal and Saline Conditions and Improves Salt Tolerance in Alfalfa. Planta 2000, 210, 416–422. [Google Scholar] [CrossRef] [PubMed]
- Alam, I.; Batool, K.; Cui, D.-L.; Yang, Y.-Q.; Lu, Y.-H. Comprehensive Genomic Survey, Structural Classification and Expression Analysis of C2H2 Zinc Finger Protein Gene Family in Brassica rapa L. PLoS ONE 2019, 14, e0216071. [Google Scholar] [CrossRef]
- Smith, A.E.F.; Farzaneh, F.; Ford, K.G. Single Zinc-Finger Extension: Enhancing Transcriptional Activity and Specificity of Three-Zinc-Finger Proteins. Biol. Chem. 2005, 386, 95–99. [Google Scholar] [CrossRef] [PubMed]
- Hiratsuka, K.; Chua, N.-H. Light Regulated Transcription in Higher Plants. J. Plant Res. 1997, 110, 131–139. [Google Scholar] [CrossRef]
- Manjunath, S.; Sachs, M.M. Molecular Characterization and Promoter Analysis of the Maize Cytosolic Glyceraldehyde 3-Phosphate Dehydrogenase Gene Family and Its Expression during Anoxia. Plant Mol. Biol. 1997, 33, 97–112. [Google Scholar] [CrossRef]
- Thomashow, M.F. Plant Cold Acclimation: Freezing Tolerance Genes and Regulatory Mechanisms. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1999, 50, 571–599. [Google Scholar] [CrossRef]
- Geffers, R.; Sell, S.; Cerff, R.; Hehl, R. The TATA Box and a Myb Binding Site Are Essential for Anaerobic Expression of a Maize GapC4 Minimal Promoter in Tobacco. Biochim. Biophys. Acta (BBA)-Gene Struct. Expr. 2001, 1521, 120–125. [Google Scholar] [CrossRef]
- Xu, R. Genome-Wide Analysis and Identification of Stress-Responsive Genes of the CCCH Zinc Finger Family in Solanum lycopersicum. Mol. Genet. Genom. 2014, 289, 965–979. [Google Scholar] [CrossRef]
- Ming, N.; Ma, N.; Jiao, B.; Lv, W.; Meng, Q. Genome Wide Identification of C2H2-Type Zinc Finger Proteins of Tomato and Expression Analysis Under Different Abiotic Stresses. Plant Mol. Biol. Rep. 2020, 38, 75–94. [Google Scholar] [CrossRef]
- Guk, J.-Y.; Jang, M.-J.; Kim, S. Identification of Novel PHD-Finger Genes in Pepper by Genomic Re-Annotation and Comparative Analyses. BMC Plant Biol. 2022, 22, 206. [Google Scholar] [CrossRef] [PubMed]
- Pi, B.; He, X.; Ruan, Y.; Jang, J.-C.; Huang, Y. Genome-Wide Analysis and Stress-Responsive Expression of CCCH Zinc Finger Family Genes in Brassica Rapa. BMC Plant Biol. 2018, 18, 373. [Google Scholar] [CrossRef] [PubMed]
- Lu, P.; Magwanga, R.O.; Guo, X.; Kirungu, J.N.; Lu, H.; Cai, X.; Zhou, Z.; Wei, Y.; Wang, X.; Zhang, Z.; et al. Genome-Wide Analysis of Multidrug and Toxic Compound Extrusion (MATE) Family in Gossypium Raimondii and Gossypium Arboreum and Its Expression Analysis Under Salt, Cadmium, and Drought Stress. G3-Genes Genomes Genet. 2018, 8, 2483–2500. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Xin, M.; Qin, J.; Peng, H.; Ni, Z.; Yao, Y.; Sun, Q. Temporal Transcriptome Profiling Reveals Expression Partitioning of Homeologous Genes Contributing to Heat and Drought Acclimation in Wheat (Triticum aestivum L.). BMC Plant Biol. 2015, 15, 152. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Yang, Y.; Zhang, L. Zinc Finger-Homeodomain Transcriptional Factors (ZF-HDs) in Wheat (Triticum aestivum L.): Identification, Evolution, Expression Analysis and Response to Abiotic Stresses. Plants 2021, 10, 593. [Google Scholar] [CrossRef] [PubMed]
- Han, G.; Qiao, Z.; Li, Y.; Wang, C.; Wang, B. The Roles of CCCH Zinc-Finger Proteins in Plant Abiotic Stress Tolerance. Int. J. Mol. Sci. 2021, 22, 8327. [Google Scholar] [CrossRef]
- Gao, Y.; Liu, H.; Wang, Y.; Li, F.; Xiang, Y. Genome-Wide Identification of PHD-Finger Genes and Expression Pattern Analysis under Various Treatments in Moso Bamboo (Phyllostachys edulis). Plant Physiol. Biochem. 2018, 123, 378–391. [Google Scholar] [CrossRef]
- Peng, X.; Zhao, Y.; Cao, J.; Zhang, W.; Jiang, H.; Li, X.; Ma, Q.; Zhu, S.; Cheng, B. CCCH-Type Zinc Finger Family in Maize: Genome-Wide Identification, Classification and Expression Profiling under Abscisic Acid and Drought Treatments. PLoS ONE 2012, 7, e40120. [Google Scholar] [CrossRef]
- Liang, Y.; Xiong, Z.; Zheng, J.; Xu, D.; Zhu, Z.; Xiang, J.; Gan, J.; Raboanatahiry, N.; Yin, Y.; Li, M. Genome-Wide Identification, Structural Analysis and New Insights into Late Embryogenesis Abundant (LEA) Gene Family Formation Pattern in Brassica napus. Sci. Rep. 2016, 6, 24265. [Google Scholar] [CrossRef]
- Shumayla; Mendu, V.; Singh, K.; Upadhyay, S.K. Insight into the Roles of Proline-Rich Extensin-like Receptor Protein Kinases of Bread Wheat (Triticum aestivum L.). Life 2022, 12, 941. [Google Scholar] [CrossRef]
- Shumayla; Madhu; Singh, K.; Upadhyay, S.K. LysM Domain-Containing Proteins Modulate Stress Response and Signalling in Triticum aestivum L. Environ. Exp. Bot. 2021, 189, 104558. [Google Scholar] [CrossRef]
- Park, H.-Y.; Lee, K.C.; Jang, Y.H.; Kim, S.-K.; Thu, M.P.; Lee, J.H.; Kim, J.-K. The Arabidopsis Splicing Factors, AtU2AF65, AtU2AF35, and AtSF1 Shuttle between Nuclei and Cytoplasms. Plant Cell Rep. 2017, 36, 1113–1123. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.B.; Brendel, V. Molecular Characterization and Phylogeny of U2AF(35) Homologs in Plants. Plant Physiol. 2006, 140, 624–636. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, P.; Arora, R.; Ray, S.; Singh, A.K.; Singh, V.P.; Takatsuji, H.; Kapoor, S.; Tyagi, A.K. Genome-Wide Identification of C2H2 Zinc-Finger Gene Family in Rice and Their Phylogeny and Expression Analysis. Plant Mol. Biol. 2007, 65, 467–485. [Google Scholar] [CrossRef]
- Zhang, S.; Liu, J.; Zhong, G.; Wang, B. Genome-Wide Identification and Expression Patterns of the C2H2-Zinc Finger Gene Family Related to Stress Responses and Catechins Accumulation in Camellia sinensis [L.] O. Kuntze. Int. J. Mol. Sci. 2021, 22, 4197. [Google Scholar] [CrossRef]
- Wang, D.; Guo, Y.; Wu, C.; Yang, G.; Li, Y.; Zheng, C. Genome-Wide Analysis of CCCH Zinc Finger Family in Arabidopsis and Rice. BMC Genom. 2008, 9, 44. [Google Scholar] [CrossRef]
- Wan, Y.; Wang, Z.; Xia, J.; Shen, S.; Guan, M.; Zhu, M.; Qiao, C.; Sun, F.; Liang, Y.; Li, J.; et al. Genome-Wide Analysis of Phosphorus Transporter Genes in Brassica and Their Roles in Heavy Metal Stress Tolerance. Int. J. Mol. Sci. 2020, 21, 2209. [Google Scholar] [CrossRef]
- Taylor, J.S.; Raes, J. Duplication and Divergence: The Evolution of New Genes and Old Ideas. Annu. Rev. Genet. 2004, 38, 615–643. [Google Scholar] [CrossRef]
- Hao, Y.; Hao, M.; Cui, Y.; Kong, L.; Wang, H. Genome-Wide Survey of the Dehydrin Genes in Bread Wheat (Triticum aestivum L.) and Its Relatives: Identification, Evolution and Expression Profiling under Various Abiotic Stresses. BMC Genom. 2022, 23, 73. [Google Scholar] [CrossRef]
- Dong, L.; Lu, Y.; Liu, S. Genome-Wide Member Identification, Phylogeny and Expression Analysis of PEBP Gene Family in Wheat and Its Progenitors. PeerJ 2020, 8, e10483. [Google Scholar] [CrossRef]
- Jiang, Y.; Liu, L.; Pan, Z.; Zhao, M.; Zhu, L.; Han, Y.; Li, L.; Wang, Y.; Wang, K.; Liu, S.; et al. Genome-Wide Analysis of the C2H2 Zinc Finger Protein Gene Family and Its Response to Salt Stress in Ginseng, Panax Ginseng Meyer. Sci. Rep. 2022, 12, 10165. [Google Scholar] [CrossRef] [PubMed]
- Malik, W.A.; Wang, X.; Wang, X.; Shu, N.; Cui, R.; Chen, X.; Wang, D.; Lu, X.; Yin, Z.; Wang, J.; et al. Genome-Wide Expression Analysis Suggests Glutaredoxin Genes Response to Various Stresses in Cotton. Int. J. Biol. Macromol. 2020, 153, 470–491. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Sharma, S.; Chunduri, V.; Kaur, A.; Kaur, S.; Malhotra, N.; Kumar, A.; Kapoor, P.; Kumari, A.; Kaur, J.; et al. Genome-Wide Identification and Characterization of Heat Shock Protein Family Reveals Role in Development and Stress Conditions in Triticum aestivum L. Sci. Rep. 2020, 10, 7858. [Google Scholar] [CrossRef] [PubMed]
- Fedorova, L.; Fedorov, A. Introns in Gene Evolution. Genetica 2003, 118, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Deshmukh, R.K.; Sonah, H.; Singh, N.K. Intron Gain, a Dominant Evolutionary Process Supporting High Levels of Gene Expression in Rice. J. Plant Biochem. Biotechnol. 2016, 25, 142–146. [Google Scholar] [CrossRef]
- Yu, P.; Shinde, H.; Dudhate, A.; Tsugama, D.; Gupta, S.K.; Liu, S.; Takano, T. Genome-Wide Investigation of SQUAMOSA Promoter Binding Protein-like Transcription Factor Family in Pearl Millet (Pennisetum glaucum (L.) R. Br.). Plant Gene 2021, 27, 100313. [Google Scholar] [CrossRef]
- Singh, S.; Kudapa, H.; Garg, V.; Varshney, R.K. Comprehensive Analysis and Identification of Drought-Responsive Candidate NAC Genes in Three Semi-Arid Tropics (SAT) Legume Crops. BMC Genom. 2021, 22, 289. [Google Scholar] [CrossRef]
- Li, K.; Xing, C.; Yao, Z.; Huang, X. PbrMYB21, a Novel MYB Protein of Pyrus Betulaefolia, Functions in Drought Tolerance and Modulates Polyamine Levels by Regulating Arginine Decarboxylase Gene. Plant Biotechnol. J. 2017, 15, 1186–1203. [Google Scholar] [CrossRef]
- Tyagi, A.K.; Gaur, T. Light Regulation of Nuclear Photosynthetic Genes in Higher Plants. Crit. Rev. Plant Sci. 2003, 22, 417–452. [Google Scholar] [CrossRef]
- Fankhauser, C.; Chory, J. Light Control of Plant Development. Annu. Rev. Cell Dev. Biol. 1997, 13, 203–229. [Google Scholar] [CrossRef]
- Baldoni, E.; Genga, A.; Cominelli, E. Plant MYB Transcription Factors: Their Role in Drought Response Mechanisms. Int. J. Mol. Sci. 2015, 16, 15811–15851. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Hou, Z.; He, Q.; Zhang, X.; Yan, K.; Han, R.; Liang, Z. Genome-Wide Characterization and Expression Analysis of BZIP Gene Family Under Abiotic Stress in Glycyrrhiza uralensis. Front. Genet. 2021, 12, 754237. [Google Scholar] [CrossRef] [PubMed]
- Baillo, E.H.; Hanif, M.S.; Guo, Y.; Zhang, Z.; Xu, P.; Algam, S.A. Genome-Wide Identification of WRKY Transcription Factor Family Members in Sorghum (Sorghum bicolor (L.) Moench). PLoS ONE 2020, 15, e0236651. [Google Scholar] [CrossRef] [PubMed]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene Ontology: Tool for the Unification of Biology. The Gene Ontology Consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Kolodziej, M.C.; Singla, J.; Sanchez-Martin, J.; Zbinden, H.; Simkova, H.; Karafiatova, M.; Dolezel, J.; Gronnier, J.; Poretti, M.; Glauser, G.; et al. A Membrane-Bound Ankyrin Repeat Protein Confers Race-Specific Leaf Rust Disease Resistance in Wheat. Nat. Commun. 2021, 12, 956. [Google Scholar] [CrossRef]
- Yu, S.; Liao, F.; Wang, F.; Wen, W.; Li, J.; Mei, H.; Luo, L. Identification of Rice Transcription Factors Associated with Drought Tolerance Using the Ecotilling Method. PLoS ONE 2012, 7, e30765. [Google Scholar] [CrossRef]
- Daetwyler, H.D.; Pong-Wong, R.; Villanueva, B.; Woolliams, J.A. The Impact of Genetic Architecture on Genome-Wide Evaluation Methods. Genetics 2010, 185, 1021–1031. [Google Scholar] [CrossRef]
- Tsukada, Y.; Fang, J.; Erdjument-Bromage, H.; Warren, M.E.; Borchers, C.H.; Tempst, P.; Zhang, Y. Histone Demethylation by a Family of JmjC Domain-Containing Proteins. Nature 2006, 439, 811–816. [Google Scholar] [CrossRef]
- Yaschenko, A.E.; Fenech, M.; Mazzoni-Putman, S.; Alonso, J.M.; Stepanova, A.N. Deciphering the Molecular Basis of Tissue-Specific Gene Expression in Plants: Can Synthetic Biology Help? Curr. Opin. Plant Biol. 2022, 68, 102241. [Google Scholar] [CrossRef]
- Duan, H.; Zhu, Y.; Li, J.; Ding, W.; Wang, H.; Jiang, L.; Zhou, Y. Effects of Drought Stress on Growth and Development of Wheat Seedlings. Int. J. Agric. Biol. 2017, 19, 1119–1124. [Google Scholar] [CrossRef]
- Qin, F.; Sakuma, Y.; Tran, L.-S.P.; Maruyama, K.; Kidokoro, S.; Fujita, Y.; Fujita, M.; Umezawa, T.; Sawano, Y.; Miyazono, K.; et al. Arabidopsis DREB2A-Interacting Proteins Function as RING E3 Ligases and Negatively Regulate Plant Drought Stress-Responsive Gene Expression. Plant Cell 2008, 20, 1693–1707. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Yang, C.; Chen, H.; Luo, L.; Leng, Q.; Li, S.; Han, Z.; Li, X.; Song, C.; Zhang, X.; et al. Exploring Transcription Factors Reveals Crucial Members and Regulatory Networks Involved in Different Abiotic Stresses in Brassica napus L. BMC Plant Biol. 2018, 18, 202. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Yang, Q.; Zhang, X.; Zhang, X.; Yu, T.; Wu, Y.; Fang, Y.; Xue, D. Genome-Wide Identification of the HMA Gene Family and Expression Analysis under Cd Stress in Barley. Plants 2021, 10, 1849. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Cao, H.; Zhang, X.; Gui, L.; Chen, Q.; Qian, G.; Xiao, J.; Li, Z. Genome-Wide Identification and Expression Analysis of Tomato ADK Gene Family during Development and Stress. Int. J. Mol. Sci. 2021, 22, 7708. [Google Scholar] [CrossRef] [PubMed]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree: Computing Large Minimum Evolution Trees with Profiles Instead of a Distance Matrix. Mol. Biol. Evol. 2009, 26, 1641–1650. [Google Scholar] [CrossRef]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2-Approximately Maximum-Likelihood Trees for Large Alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A Toolkit for Detection and Evolutionary Analysis of Gene Synteny and Collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Sharma, A.; Sharma, H.; Rajput, R.; Pandey, A.; Upadhyay, S.K. Molecular Characterization Revealed the Role of Thaumatin-Like Proteins of Bread Wheat in Stress Response. Front. Plant Sci. 2022, 12, 807448. [Google Scholar] [CrossRef]
- Gaut, B.S.; Morton, B.R.; McCaig, B.C.; Clegg, M.T. Substitution Rate Comparisons between Grasses and Palms: Synonymous Rate Differences at the Nuclear Gene Adh Parallel Rate Differences at the Plastid Gene RbcL. Proc. Natl. Acad. Sci. USA 1996, 93, 10274–10279. [Google Scholar] [CrossRef]
- Letunic, I.; Khedkar, S.; Bork, P. SMART: Recent Updates, New Developments and Status in 2020. Nucleic Acids Res. 2021, 49, D458–D460. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Wang, M.; Wu, J.; Guo, W.; Chen, Y.; Li, G.; Wang, Y.; Shi, W.; Xia, G.; Fu, D.; et al. WheatOmics: A Platform Combining Multiple Omics Data to Accelerate Functional Genomics Studies in Wheat. Mol. Plant 2021, 14, 1965–1968. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, Y.; Fan, C.; Wei, Y.; Meng, J.; Li, Z.; Zhong, C. Genome-Wide Analysis of MYB Transcription Factors and Their Responses to Salt Stress in Casuarina equisetifolia. BMC Plant Biol. 2021, 21, 328. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, Z.; Shen, S.; Wang, Y.; Tao, W.; Zhao, Z.; Hu, X.; Yu, P. Genome-Wide Identification and Expression Analysis of the Zinc Finger Protein Gene Subfamilies under Drought Stress in Triticum aestivum. Plants 2022, 11, 2511. https://doi.org/10.3390/plants11192511
Wu Z, Shen S, Wang Y, Tao W, Zhao Z, Hu X, Yu P. Genome-Wide Identification and Expression Analysis of the Zinc Finger Protein Gene Subfamilies under Drought Stress in Triticum aestivum. Plants. 2022; 11(19):2511. https://doi.org/10.3390/plants11192511
Chicago/Turabian StyleWu, Zhaoming, Shenghai Shen, Yueduo Wang, Weiqi Tao, Ziqi Zhao, Xiangli Hu, and Pei Yu. 2022. "Genome-Wide Identification and Expression Analysis of the Zinc Finger Protein Gene Subfamilies under Drought Stress in Triticum aestivum" Plants 11, no. 19: 2511. https://doi.org/10.3390/plants11192511
APA StyleWu, Z., Shen, S., Wang, Y., Tao, W., Zhao, Z., Hu, X., & Yu, P. (2022). Genome-Wide Identification and Expression Analysis of the Zinc Finger Protein Gene Subfamilies under Drought Stress in Triticum aestivum. Plants, 11(19), 2511. https://doi.org/10.3390/plants11192511






