Expression Analyses in the Rachis Hint towards Major Cell Wall Modifications in Grape Clusters Showing Berry Shrivel Symptoms
Abstract
1. Introduction
2. Results
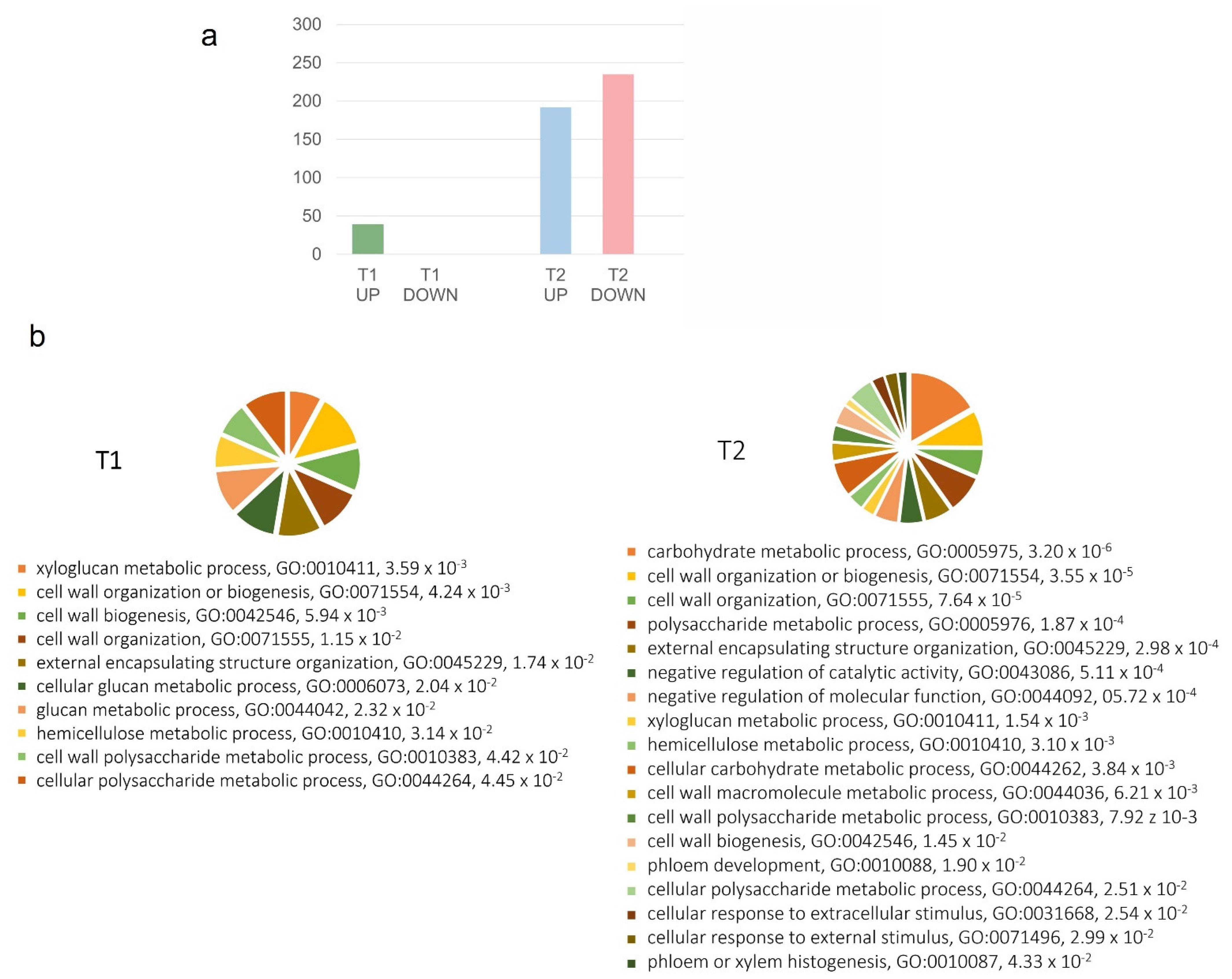
2.1. Gene Expression Modulation in the Rachis of BS-Affected Grape Clusters
2.2. Berry Cell Wall Modifications Induced by the Disorder
3. Discussion
4. Materials and Methods
4.1. Plant Material Sampling
4.2. RNA Extraction and cDNA Preparation for qPCR
4.3. Microarray Analyses of Rachis Tissue and qPCR
4.4. RNASeq Analyses of Grape Berries
4.5. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- van Leeuwen, C.; Destrac-Irvine, A. Modified Grape Composition under Climate Change Conditions Requires Adaptations in the Vineyard. Oeno One 2017, 51, 147–154. [Google Scholar] [CrossRef]
- Ollat, N.; van Leeuwen, C.; de Cortazar-Atauri, I.G.; Touzard, J.-M. The Challenging Issue of Climate Change for Sustainable Grape and Wine Production. Oeno One 2017, 51, 59–60. [Google Scholar] [CrossRef]
- van Leeuwen, C.; Destrac-Irvine, A.; Dubernet, M.; Duchêne, E.; Gowdy, M.; Marguerit, E.; Pieri, P.; Parker, A.; De Rességuier, L.; Ollat, N. An Update on the Impact of Climate Change in Viticulture and Potential Adaptations. Agronomy 2019, 9, 514. [Google Scholar] [CrossRef]
- Gomès, É.; Maillot, P.; Duchêne, É. Molecular Tools for Adapting Viticulture to Climate Change. Front. Plant Sci. 2021, 12, 633846. [Google Scholar] [CrossRef]
- Coombe, B.G. Research on development and ripening of the grape berry. Am. J. Enol. Vitic. 1992, 43, 101–110. [Google Scholar]
- Serrano, A.; Espinoza, C.; Armijo, G.; Inostroza-Blancheteau, C.; Poblete, E.; Meyer-Regueiro, C.; Arce, A.; Parada, F.; Santibáñez, C.; Arce-Johnson, P. Omics Approaches for Understanding Grapevine Berry Development: Regulatory Networks Associated with Endogenous Processes and Environmental Responses. Front. Plant Sci. 2017, 8, 1486. [Google Scholar] [CrossRef]
- Fasoli, M.; Richter, C.L.; Zenoni, S.; Bertini, E.; Vitulo, N.; Santo, S.D.; Dokoozlian, N.; Pezzotti, M.; Tornielli, G.B. Timing and Order of the Molecular Events Marking the Onset of Berry Ripening in Grapevine. Plant Physiol. 2018, 178, 1187–1206. [Google Scholar] [CrossRef]
- Tornielli, G.B.; Sandri, M.; Fasoli, M.; Amato, A.; Pezzotti, M.; Zuccolotto, P.; Zenoni, S. A Molecular Phenology Scale of Fruit Development. bioRxiv 2022. [Google Scholar] [CrossRef]
- Rienth, M.; Torregrosa, L.; Sarah, G.; Ardisson, M.; Brillouet, J.-M.; Romieu, C. Temperature Desynchronizes Sugar and Organic Acid Metabolism in Ripening Grapevine Fruits and Remodels Their Transcriptome. BMC Plant Biol. 2016, 16, 164. [Google Scholar] [CrossRef]
- Shahood, R.; Torregrosa, L.; Savoi, S.; Romieu, C. First Quantitative Assessment of Growth, Sugar Accumulation and Malate Breakdown in a Single Ripening Berry. Oeno One 2020, 54, 1077–1092. [Google Scholar] [CrossRef]
- Burbidge, C.A.; Ford, C.M.; Melino, V.J.; Wong, D.C.J.; Jia, Y.; Jenkins, C.L.D.; Soole, K.L.; Castellarin, S.D.; Darriet, P.; Rienth, M.; et al. Biosynthesis and Cellular Functions of Tartaric Acid in Grapevines. Front. Plant Sci. 2021, 12, 643024. [Google Scholar] [CrossRef]
- Suter, B.; Destrac Irvine, A.; Gowdy, M.; Dai, Z.; van Leeuwen, C. Adapting Wine Grape Ripening to Global Change Requires a Multi-Trait Approach. Front. Plant Sci. 2021, 12, 624867. [Google Scholar] [CrossRef]
- Rienth, M.; Vigneron, N.; Darriet, P.; Sweetman, C.; Burbidge, C.; Bonghi, C.; Walker, R.P.; Famiani, F.; Castellarin, S.D. Grape Berry Secondary Metabolites and Their Modulation by Abiotic Factors in a Climate Change Scenario—A Review. Front. Plant Sci. 2021, 12, 643258. [Google Scholar] [CrossRef]
- Alem, H.; Rigou, P.; Schneider, R.; Ojeda, H.; Torregrosa, L. Impact of Agronomic Practices on Grape Aroma Composition: A Review. J. Sci. Food Agric. 2019, 99, 975–985. [Google Scholar] [CrossRef]
- Savoi, S.; Wong, D.C.J.; Arapitsas, P.; Miculan, M.; Bucchetti, B.; Peterlunger, E.; Fait, A.; Mattivi, F.; Castellarin, S.D. Transcriptome and Metabolite Profiling Reveals That Prolonged Drought Modulates the Phenylpropanoid and Terpenoid Pathway in White Grapes (Vitis vinifera L.). BMC Plant Biol. 2016, 16, 67. [Google Scholar] [CrossRef]
- Pinasseau, L.; Vallverdú-Queralt, A.; Verbaere, A.; Roques, M.; Meudec, E.; Le Cunff, L.; Péros, J.-P.; Ageorges, A.; Sommerer, N.; Boulet, J.-C.; et al. Cultivar Diversity of Grape Skin Polyphenol Composition and Changes in Response to Drought Investigated by LC-MS Based Metabolomics. Front. Plant Sci. 2017, 8, 1826. [Google Scholar] [CrossRef]
- Wu, J.; Drappier, J.; Hilbert, G.; Guillaumie, S.; Dai, Z.; Geny, L.; Delrot, S.; Darriet, P.; Thibon, C.; Pieri, P. The Effects of a Moderate Grape Temperature Increase on Berry Secondary Metabolites: This Article Is Published in Cooperation with the 21th GIESCO International Meeting, June 23-28 2019, Thessaloniki, Greece. Guests Editors: Stefanos Koundouras and Laurent Torregrosa. Oeno One 2019, 53, 321–333. [Google Scholar] [CrossRef]
- Lecourieux, D.; Kappel, C.; Claverol, S.; Pieri, P.; Feil, R.; Lunn, J.E.; Bonneu, M.; Wang, L.; Gomès, E.; Delrot, S.; et al. Proteomic and Metabolomic Profiling Underlines the Stage- and Time-Dependent Effects of High Temperature on Grape Berry Metabolism. J. Integr. Plant Biol. 2020, 62, 1132–1158. [Google Scholar] [CrossRef]
- Rienth, M.; Vigneron, N.; Walker, R.P.; Castellarin, S.D.; Sweetman, C.; Burbidge, C.A.; Bonghi, C.; Famiani, F.; Darriet, P. Modifications of Grapevine Berry Composition Induced by Main Viral and Fungal Pathogens in a Climate Change Scenario. Front. Plant Sci. 2021, 12, 717223. [Google Scholar] [CrossRef]
- Dal Santo, S.; Tucker, M.R.; Tan, H.-T.; Burbidge, C.A.; Fasoli, M.; Böttcher, C.; Boss, P.K.; Pezzotti, M.; Davies, C. Auxin Treatment of Grapevine (Vitis Vinifera L.) Berries Delays Ripening Onset by Inhibiting Cell Expansion. Plant Mol. Biol. 2020, 103, 91–111. [Google Scholar] [CrossRef]
- Davies, C.; Böttcher, C.; Nicholson, E.L.; Burbidge, C.A.; Boss, P.K. Timing of Auxin Treatment Affects Grape Berry Growth, Ripening Timing and the Synchronicity of Sugar Accumulation. Aust. J. Grape Wine Res. 2022, 28, 232–241. [Google Scholar] [CrossRef]
- Zhang, X.Y.; Wang, X.L.; Wang, X.F.; Xia, G.H.; Pan, Q.H.; Fan, R.C.; Wu, F.Q.; Yu, X.C.; Zhang, D.P. A Shift of Phloem Unloading from Symplasmic to Apoplasmic Pathway Is Involved in Developmental Onset of Ripening in Grape Berry. Plant Physiol. 2006, 142, 220–232. [Google Scholar] [CrossRef] [PubMed]
- Glissant, D.; Dédaldéchamp, F.; Delrot, S. Transcriptomic Analysis of Grape Berry Softening during Ripening. Oeno One 2008, 42, 1–13. [Google Scholar] [CrossRef]
- Gambetta, G.A.; Matthews, M.A.; Shaghasi, T.H.; McElrone, A.J.; Castellarin, S.D. Sugar and Abscisic Acid Signaling Orthologs Are Activated at the Onset of Ripening in Grape. Planta 2010, 232, 219–234. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Montes, E.; Zhang, Y.; Chang, B.-M.; Shcherbatyuk, N.; Keller, M. Soft, Sweet, and Colorful: Stratified Sampling Reveals Sequence of Events at the Onset of Grape Ripening. Am. J. Enol. Vitic. 2021, 72, 137–151. [Google Scholar] [CrossRef]
- Castellarin, S.D.; Gambetta, G.A.; Wada, H.; Krasnow, M.N.; Cramer, G.R.; Peterlunger, E.; Shackel, K.A.; Matthews, M.A. Characterization of Major Ripening Events during Softening in Grape: Turgor, Sugar Accumulation, Abscisic Acid Metabolism, Colour Development, and Their Relationship with Growth. J. Exp. Bot. 2016, 67, 709–722. [Google Scholar] [CrossRef]
- Chervin, C.; Tira-umphon, A.; Terrier, N.; Zouine, M.; Severac, D.; Roustan, J.-P. Stimulation of the Grape Berry Expansion by Ethylene and Effects on Related Gene Transcripts, over the Ripening Phase. Physiol. Plant. 2008, 134, 534–546. [Google Scholar] [CrossRef]
- Savoi, S.; Torregrosa, L.; Romieu, C. Transcripts Switched off at the Stop of Phloem Unloading Highlight the Energy Efficiency of Sugar Import in the Ripening V. Vinifera Fruit. Hortic. Res. 2021, 8, 1–15. [Google Scholar] [CrossRef]
- Höfte, H.; Voxeur, A. Plant Cell Walls. Curr. Biol. 2017, 27, R865–R870. [Google Scholar] [CrossRef]
- Somerville, C.; Bauer, S.; Brininstool, G.; Facette, M.; Hamann, T.; Milne, J.; Osborne, E.; Paredez, A.; Persson, S.; Raab, T.; et al. Toward a Systems Approach to Understanding Plant Cell Walls. Science 2004, 306, 2206–2211. [Google Scholar] [CrossRef]
- Arnous, A.; Meyer, A.S. Quantitative Prediction of Cell Wall Polysaccharide Composition in Grape (Vitis Vinifera L.) and Apple (Malus Domestica) Skins from Acid Hydrolysis Monosaccharide Profiles. J. Agric. Food Chem. 2009, 57, 3611–3619. [Google Scholar] [CrossRef]
- Nunan, K.J.; Sims, I.M.; Bacic, A.; Robinson, S.P.; Fincher, G.B. Changes in Cell Wall Composition during Ripening of Grape Berries. Plant Physiol. 1998, 118, 783–792. [Google Scholar] [CrossRef]
- Fasoli, M.; Dell’Anna, R.; Dal Santo, S.; Balestrini, R.; Sanson, A.; Pezzotti, M.; Monti, F.; Zenoni, S. Pectins, Hemicelluloses and Celluloses Show Specific Dynamics in the Internal and External Surfaces of Grape Berry Skin during Ripening. Plant Cell Physiol. 2016, 57, 1332–1349. [Google Scholar] [CrossRef]
- Tello, J.; Forneck, A. A Double-Sigmoid Model for Grapevine Bunch Compactness Development. Oeno One 2018, 52, 307–316. [Google Scholar] [CrossRef]
- Grimplet, J.; Ibáñez, S.; Baroja, E.; Tello, J.; Ibáñez, J. Phenotypic, Hormonal, and Genomic Variation among Vitis Vinifera Clones with Different Cluster Compactness and Reproductive Performance. Front. Plant Sci. 2019, 9, 1917. [Google Scholar] [CrossRef]
- Tello, J.; Torres-Pérez, R.; Flutre, T.; Grimplet, J.; Ibáñez, J. VviUCC1 Nucleotide Diversity, Linkage Disequilibrium and Association with Rachis Architecture Traits in Grapevine. Genes 2020, 11, 598. [Google Scholar] [CrossRef]
- Keller, M.; Smith, J.P.; Bondada, B.R. Ripening Grape Berries Remain Hydraulically Connected to the Shoot. J. Exp. Bot. 2006, 57, 2577–2587. [Google Scholar] [CrossRef]
- Choat, B.; Gambetta, G.A.; Shackel, K.A.; Matthews, M.A. Vascular Function in Grape Berries across Development and Its Relevance to Apparent Hydraulic Isolation. Plant Physiol. 2009, 151, 1677–1687. [Google Scholar] [CrossRef]
- Zhang, Y.; Keller, M. Discharge of Surplus Phloem Water May Be Required for Normal Grape Ripening. J. Exp. Bot. 2017, 68, 585–595. [Google Scholar] [CrossRef][Green Version]
- Cuéllar, T.; Azeem, F.; Andrianteranagna, M.; Pascaud, F.; Verdeil, J.-L.; Sentenac, H.; Zimmermann, S.; Gaillard, I. Potassium Transport in Developing Fleshy Fruits: The Grapevine Inward K+ Channel VvK1.2 Is Activated by CIPK–CBL Complexes and Induced in Ripening Berry Flesh Cells. Plant J. 2013, 73, 1006–1018. [Google Scholar] [CrossRef]
- Keller, M.; Shrestha, P.M. Solute Accumulation Differs in the Vacuoles and Apoplast of Ripening Grape Berries. Planta 2014, 239, 633–642. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Dong, C.; Wei, J.; Guo, L.; Meng, Y.; Wu, B.; Chen, J. A Transcriptional Study of the Effects of Nitric Oxide on Rachis Browning in Table Grapes Cv. Thompson Seedless. Postharvest Biol. Technol. 2021, 175, 111471. [Google Scholar] [CrossRef]
- Hall, G.E.; Bondada, B.R.; Keller, M. Loss of Rachis Cell Viability Is Associated with Ripening Disorders in Grapes. J. Exp. Bot. 2011, 62, 1145–1153. [Google Scholar] [CrossRef] [PubMed]
- Zufferey, V.; Spring, J.-L.; Voinesco, F.; Viret, O.; Gindro, K. Physiological and Histological Approaches to Study Berry Shrivel in Grapes. Oeno One 2015, 49, 113–125. [Google Scholar] [CrossRef]
- Crespo-Martínez, S.; Sobczak, M.; Różańska, E.; Forneck, A.; Griesser, M. The Role of the Secondary Phloem during the Development of the Grapevine Berry Shrivel Ripening Disorder. Micron 2019, 116, 36–45. [Google Scholar] [CrossRef]
- Hoff, R.T.; Bondada, B.R.; Keller, M. Onset and Progression of the Berry Shrivel Ripening Disorder in Grapes. Aust. J. Grape Wine Res. 2021, 27, 280–289. [Google Scholar] [CrossRef]
- Griesser, M.; Eder, R.; Besser, S.; Forneck, A. Berry Shrivel of Grapes in Austria–Aspects of the Physiological Disorder with Cultivar Zweigelt (Vitis Vinifera L.). Sci. Hortic. 2012, 145, 87–93. [Google Scholar] [CrossRef]
- Griesser, M.; Martinez, S.C.; Eitle, M.W.; Warth, B.; Andre, C.M.; Schuhmacher, R.; Forneck, A. The Ripening Disorder Berry Shrivel Affects Anthocyanin Biosynthesis and Sugar Metabolism in Zweigelt Grape Berries. Planta 2018, 247, 471–481. [Google Scholar] [CrossRef]
- Savoi, S.; Herrera, J.C.; Forneck, A.; Griesser, M. Transcriptomics of the Grape Berry Shrivel Ripening Disorder. Plant Mol. Biol. 2019, 100, 285–301. [Google Scholar] [CrossRef]
- Knoll, M.; Achleitner, D.; Redl, H. Sugar Accumulation in ’Zweigelt ’ Grapes as Affected by “Traubenwelke”. Vitis 2010, 49, 101–106. [Google Scholar]
- Krasnow, M.; Matthews, M.; Smith, R.; Benz, J.; Weber, E.; Shackel, K. Distinctive Symptoms Differentiate Four Common Types of Berry Shrivel Disorder in Grape. Calif. Agric. 2010, 64, 155–159. [Google Scholar] [CrossRef]
- Bondada, B.R.; Keller, M. Not All Shrivels Are Created Equal—Morpho-Anatomical and Compositional Characteristics Differ among Different Shrivel Types That Develop during Ripening of Grape (Vitis Vinifera L.) Berries. Am. J. Plant Sci. 2012, 3, 879. [Google Scholar] [CrossRef]
- Krasnow, M.; Matthews, M.; Shackel, K. Analysis of Cell Viability during Ripening in Healthy and SAD-Affected Grape Berries. Am. J. Enol. Vitic. 2012, 63, 460A. [Google Scholar]
- Griesser, M.; Savoi, S.; Supapvanich, S.; Dobrev, P.; Vankova, R.; Forneck, A. Phytohormone Profiles Are Strongly Altered during Induction and Symptom Development of the Physiological Ripening Disorder Berry Shrivel in Grapevine. Plant Mol. Biol. 2020, 103, 141–157. [Google Scholar] [CrossRef]
- Cosgrove, D.J. Growth of the Plant Cell Wall. Nat. Rev. Mol. Cell Biol. 2005, 6, 850–861. [Google Scholar] [CrossRef]
- Daras, G.; Templalexis, D.; Avgeri, F.; Tsitsekian, D.; Karamanou, K.; Rigas, S. Updating Insights into the Catalytic Domain Properties of Plant Cellulose Synthase (CesA) and Cellulose Synthase-like (Csl) Proteins. Molecules 2021, 26, 4335. [Google Scholar] [CrossRef]
- Cosgrove, D.J. Plant Cell Wall Extensibility: Connecting Plant Cell Growth with Cell Wall Structure, Mechanics, and the Action of Wall-Modifying Enzymes. J. Exp. Bot. 2016, 67, 463–476. [Google Scholar] [CrossRef]
- Sharples, S.C.; Nguyen-Phan, T.C.; Fry, S.C. Xyloglucan Endotransglucosylase/Hydrolases (XTHs) Are Inactivated by Binding to Glass and Cellulosic Surfaces, and Released in Active Form by a Heat-Stable Polymer from Cauliflower Florets. J. Plant Physiol. 2017, 218, 135–143. [Google Scholar] [CrossRef]
- Supapvanich, S.; Tucker, G. Cell Wall Hydrolysis in Netted Melon Fruit (Cucumis melo Var. reticulatus L. Naud) during Storage . Chiang Mai J. Sci. 2013, 40, 447–458. [Google Scholar]
- Forlani, S.; Masiero, S.; Mizzotti, C. Fruit Ripening: The Role of Hormones, Cell Wall Modifications, and Their Relationship with Pathogens. J. Exp. Bot. 2019, 70, 2993–3006. [Google Scholar] [CrossRef]
- Uluisik, S.; Seymour, G.B. Pectate Lyases: Their Role in Plants and Importance in Fruit Ripening. Food Chem. 2020, 309, 125559. [Google Scholar] [CrossRef]
- Marowa, P.; Ding, A.; Kong, Y. Expansins: Roles in Plant Growth and Potential Applications in Crop Improvement. Plant Cell Rep. 2016, 35, 949–965. [Google Scholar] [CrossRef]
- Nunan, K.J.; Davies, C.; Robinson, S.P.; Fincher, G.B. Expression Patterns of Cell Wall-Modifying Enzymes during Grape Berry Development. Planta 2001, 214, 257–264. [Google Scholar] [CrossRef]
- Bashline, L.; Lei, L.; Li, S.; Gu, Y. Cell Wall, Cytoskeleton, and Cell Expansion in Higher Plants. Mol. Plant 2014, 7, 586–600. [Google Scholar] [CrossRef]
- Dal Santo, S.; Vannozzi, A.; Tornielli, G.B.; Fasoli, M.; Venturini, L.; Pezzotti, M.; Zenoni, S. Genome-Wide Analysis of the Expansin Gene Superfamily Reveals Grapevine-Specific Structural and Functional Characteristics. PLoS ONE 2013, 8, e62206. [Google Scholar] [CrossRef]
- Fischer, R.L.; Bennett, A.B. Role of Cell Wall Hydrolases in Fruit Ripening. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1991, 42, 675–703. [Google Scholar] [CrossRef]
- Wormit, A.; Usadel, B. The Multifaceted Role of Pectin Methylesterase Inhibitors (PMEIs). Int. J. Mol. Sci. 2018, 19, 2878. [Google Scholar] [CrossRef]
- Goujon, T.; Minic, Z.; El Amrani, A.; Lerouxel, O.; Aletti, E.; Lapierre, C.; Joseleau, J.-P.; Jouanin, L. AtBXL1, a Novel Higher Plant (Arabidopsis Thaliana) Putative Beta-Xylosidase Gene, Is Involved in Secondary Cell Wall Metabolism and Plant Development. Plant J. 2003, 33, 677–690. [Google Scholar] [CrossRef]
- Lee, E.-J.; Matsumura, Y.; Soga, K.; Hoson, T.; Koizumi, N. Glycosyl Hydrolases of Cell Wall are Induced by Sugar Starvation in Arabidopsis. Plant Cell Physiol. 2007, 48, 405–413. [Google Scholar] [CrossRef]
- Arias, M.C.; Pelletier, S.; Hilliou, F.; Wattebled, F.; Renou, J.-P.; D’Hulst, C. From Dusk till Dawn: The Arabidopsis Thaliana Sugar Starving Responsive Network. Front. Plant Sci. 2014, 5, 482. [Google Scholar] [CrossRef]
- Bondada, B.; Keller, M. Structural and Fruit Compositional Anomalies Related to Various Shrivel Types Developing during Ripening of Grape Berries. Acta Hortic. 2017, 1157, 49–54. [Google Scholar] [CrossRef]
- Sengupta, S.; Mukherjee, S.; Basak, P.; Majumder, A.L. Significance of Galactinol and Raffinose Family Oligosaccharide Synthesis in Plants. Front. Plant Sci. 2015, 6, 656. [Google Scholar] [CrossRef] [PubMed]
- Lü, J.; Sui, X.; Ma, S.; Li, X.; Liu, H.; Zhang, Z. Suppression of Cucumber Stachyose Synthase Gene (CsSTS) Inhibits Phloem Loading and Reduces Low Temperature Stress Tolerance. Plant Mol. Biol. 2017, 95, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Herrera-Rodríguez, M.B.; Pérez-Vicente, R.; Maldonado, J.-M. Expression of Asparagine Synthetase Genes in Sunflower (Helianthus Annuus) under Various Environmental Stresses. Plant Physiol. Biochem. 2007, 45, 33–38. [Google Scholar] [CrossRef]
- Gaufichon, L.; Rothstein, S.J.; Suzuki, A. Asparagine Metabolic Pathways in Arabidopsis. Plant Cell Physiol. 2016, 57, 675–689. [Google Scholar] [CrossRef]
- Castellarin, S.D.; Gambetta, G.A.; Wada, H.; Shackel, K.A.; Matthews, M.A. Fruit Ripening in Vitis Vinifera: Spatiotemporal Relationships among Turgor, Sugar Accumulation, and Anthocyanin Biosynthesis. J. Exp. Bot. 2011, 62, 4345–4354. [Google Scholar] [CrossRef]
- Schlosser, J.; Olsson, N.; Weis, M.; Reid, K.; Peng, F.; Lund, S.; Bowen, P. Cellular Expansion and Gene Expression in the Developing Grape (Vitis Vinifera L.). Protoplasma 2008, 232, 255. [Google Scholar] [CrossRef]
- Khan, N.; Fatima, F.; Haider, M.S.; Shazadee, H.; Liu, Z.; Zheng, T.; Fang, J. Genome-Wide Identification and Expression Profiling of the Polygalacturonase (PG) and Pectin Methylesterase (PME) Genes in Grapevine (Vitis Vinifera L.). Int. J. Mol. Sci. 2019, 20, 3180. [Google Scholar] [CrossRef]
- Shin, Y.; Chane, A.; Jung, M.; Lee, Y. Recent Advances in Understanding the Roles of Pectin as an Active Participant in Plant Signaling Networks. Plants 2021, 10, 1712. [Google Scholar] [CrossRef]
- Gou, J.-Y.; Miller, L.M.; Hou, G.; Yu, X.-H.; Chen, X.-Y.; Liu, C.-J. Acetylesterase-Mediated Deacetylation of Pectin Impairs Cell Elongation, Pollen Germination, and Plant Reproduction. Plant Cell 2012, 24, 50–65. [Google Scholar] [CrossRef]
- Krasnow, M.; Matthews, M.; Shackel, K. Evidence for Substantial Maintenance of Membrane Integrity and Cell Viability in Normally Developing Grape (Vitis Vinifera L.) Berries throughout Development. J. Exp. Bot. 2008, 59, 849–859. [Google Scholar] [CrossRef] [PubMed]
- Krasnow, M.; Weis, N.; Smith, R.J.; Benz, M.J.; Matthews, M.; Shackel, K. Inception, Progression, and Compositional Consequences of a Berry Shrivel Disorder. Am. J. Enol. Vitic. 2009, 60, 24–34. [Google Scholar]
- Tilbrook, J.; Tyerman, S.D. Cell Death in Grape Berries: Varietal Differences Linked to Xylem Pressure and Berry Weight Loss. Funct. Plant Biol. 2008, 35, 173–184. [Google Scholar] [CrossRef]
- Fuentes, S.; Sullivan, W.; Tilbrook, J.; Tyerman, S. A Novel Analysis of Grapevine Berry Tissue Demonstrates a Variety-Dependent Correlation between Tissue Vitality and Berry Shrivel. Aust. J. Grape Wine Res. 2010, 16, 327–336. [Google Scholar] [CrossRef]
- Griesser, M.; Crespo Martinez, S.; Weidinger, M.L.; Kandler, W.; Forneck, A. Challenging the Potassium Deficiency Hypothesis for Induction of the Ripening Disorder Berry Shrivel in Grapevine. Sci. Hortic. 2017, 216, 141–147. [Google Scholar] [CrossRef]
- Reid, K.E.; Olsson, N.; Schlosser, J.; Peng, F.; Lund, S.T. An Optimized Grapevine RNA Isolation Procedure and Statistical Determination of Reference Genes for Real-Time RT-PCR during Berry Development. BMC Plant Biol. 2006, 6, 27. [Google Scholar] [CrossRef]
- Griesser, M.; Stralis-Pavese, N.; Lawo, N.C.; Kreil, D.; Forneck, A. Developing a Genome-Scale Assay to Probe the Expression Response of Phylloxera (Daktulosphaira Vitifoliae Fitch)-Induced Root Galls of Vitis Ssp. Acta Hortic. 2014, 1045, 21–27. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma Powers Differential Expression Analyses for RNA-Sequencing and Microarray Studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Smyth, G.K. Linear Models and Empirical Bayes Methods for Assessing Differential Expression in Microarray Experiments. Stat. Appl. Genet. Mol. Biol. 2004, 3, 1–26. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Smyth, G.K.; Michaud, J.; Scott, H.S. Use of Within-Array Replicate Spots for Assessing Differential Expression in Microarray Experiments. Bioinforma. Oxf. Engl. 2005, 21, 2067–2075. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, D.J.; Smyth, G.K. Testing Significance Relative to a Fold-Change Threshold Is a TREAT. Bioinformatics 2009, 25, 765–771. [Google Scholar] [CrossRef] [PubMed]
- Efron, B.; Tibshirani, R. On Testing the Significance of Sets of Genes. Ann. Appl. Stat. 2007, 1, 107–129. [Google Scholar] [CrossRef]
- Hellemans, J.; Mortier, G.; De Paepe, A.; Speleman, F.; Vandesompele, J. QBase Relative Quantification Framework and Software for Management and Automated Analysis of Real-Time Quantitative PCR Data. Genome Biol. 2007, 8, R19. [Google Scholar] [CrossRef]
- Raudvere, U.; Kolberg, L.; Kuzmin, I.; Arak, T.; Adler, P.; Peterson, H.; Vilo, J. G: Profiler: A Web Server for Functional Enrichment Analysis and Conversions of Gene Lists (2019 Update). Nucleic Acids Res. 2019, 47, W191–W198. [Google Scholar] [CrossRef]


| Functional Group | 12Xv1 ID | logFC T1 | q-Value | Rank | Log FC T2 | q-Value | Rank | Annotation |
|---|---|---|---|---|---|---|---|---|
| Cell wall biosynthesis | VIT_02s0025g01750 | −0.39 | n.s. | −1.35 | 6.6 × 10−3 | 384 | Cellulose synthase CSLG3 | |
| VIT_04s0008g02830 | 0.21 | n.s. | 1.83 | 2.4 × 10−8 | 237 | Galactokinase like protein | ||
| VIT_07s0005g04110 | 0.54 | n.s. | −1.30 | 3.7 × 10−2 | 422 | Cellulose synthase CESA4 | ||
| VIT_14s0083g01100 | 0.58 | n.s. | 1.49 | 2.6 × 10−6 | 290 | Alpha-1,4-glucan-protein synthase 1 | ||
| VIT_16s0039g02020 | 1.41 | 2.1 × 10−3 | 33 | −0.69 | n.s. | Cellulose synthase CSLD3 | ||
| VIT_18s0122g00120 | 0.05 | n.s. | −1.81 | 5.7 × 10−13 | 158 | Cellulose synthase CESA2 | ||
| Cell wall degradation | VIT_04s0044g01000 | 0.21 | n.s. | −1.34 | 3.2 × 10−2 | 418 | Pectinesterase family | |
| VIT_05s0077g01280 | 0.49 | n.s. | 4.60 | 1.4 × 10−42 | 3 | Glycosyl hydrolase family 3-β-xylosidase BXL1 | ||
| VIT_08s0007g04820 | 0.84 | n.s. | 1.51 | 1.7 × 10−4 | 340 | Pectate lyase | ||
| VIT_08s0007g07690 | 0.40 | n.s. | 1.57 | 1.4 × 10−6 | 283 | Polygalacturonase inhibiting protein PGIP1 | ||
| VIT_08s0007g07880 | 0.31 | n.s. | 1.30 | 1.4 × 10−2 | 395 | Polygalacturonase GH28 | ||
| VIT_09s0002g00320 | 0.81 | n.s. | 1.20 | 2.2 × 10−2 | 408 | Pectinesterase PME3 | ||
| VIT_11s0016g03020 | −0.20 | n.s. | −1.42 | 3.6 × 10−6 | 294 | Pectinesterase family | ||
| VIT_16s0050g01110 | 0.36 | n.s. | 1.90 | 1.2 × 10−12 | 165 | Polygalacturonase GH28 | ||
| VIT_16s0098g01900 | −0.12 | n.s. | −1.35 | 8.4 × 10−8 | 247 | Pectinesterase family | ||
| Cell wall modification | VIT_00s0323g00050 | 0.03 | n.s. | 1.53 | 5.4 × 10−10 | 205 | Invertase/pectin methylesterase inhibitor | |
| VIT_00s0386g00050 | 0.44 | n.s. | −2.26 | 3.3 × 10−17 | 105 | XET/XTH | ||
| VIT_01s0026g00200 | 0.22 | n.s. | −2.25 | 1.2 × 10−22 | 68 | Xyloglucan endotransglucosylase/hydrolase 28 | ||
| VIT_03s0088g00650 | 0.48 | n.s. | 2.44 | 1.8 × 10−25 | 51 | Xyloglucan/xyloglucosyl transferase | ||
| VIT_05s0062g00480 | 0.28 | n.s. | −1.66 | 3.7 × 10−5 | 324 | Xyloglucan endo-transglycosylase | ||
| VIT_06s0004g04860 | 2.01 | 3.6 × 10−18 | 10 | 1.71 | 7.9 × 10−13 | 160 | Expansin A (VvEXPA06) | |
| VIT_06s0061g00550 | 1.47 | 5.3 × 10−8 | 20 | 2.88 | 3.5 × 10−33 | 22 | [SWITCH] XTH 32 | |
| VIT_11s0052g01180 | 1.60 | 3.5 × 10−8 | 19 | −0.53 | n.s. | XET/XTH | ||
| VIT_11s0052g01250 | 0.04 | n.s. | −1.87 | 5.3 × 10−17 | 108 | XET/XTH | ||
| VIT_11s0052g01270 | 1.42 | 2.6 × 10−3 | 34 | −0.86 | n.s. | XET 6 | ||
| VIT_11s0052g01280 | 2.03 | 1.4 × 10−12 | 12 | −0.92 | n.s. | XET/XTH | ||
| VIT_12s0134g00160 | 0.14 | n.s. | −2.59 | 2.2 × 10−26 | 48 | XET/XTH | ||
| Hydrolytic enzyme | VIT_00s0455g00040 | 0.23 | n.s. | −1.39 | 1.7 × 10−2 | 402 | Glycosyl transferase family 8 protein | |
| VIT_03s0017g02240 | 0.28 | n.s. | 1.56 | 2.7 × 10−7 | 264 | Endo-1,3-β-glucosidase precursor | ||
| VIT_03s0180g00280 | -0.07 | n.s. | 2.50 | 2.1 × 10−20 | 79 | Indole-3-acetate β-glucosyltransferase | ||
| VIT_05s0062g00310 | 1.11 | n.s. | −1.66 | 9.3 × 10−9 | 231 | UDP-glucoronosyl/UDP-glucosyl transferase UGT75C1 | ||
| VIT_06s0004g01430 | 0.04 | n.s. | 2.39 | 1.0 × 10−20 | 76 | ABA glucosidase | ||
| VIT_06s0004g07230 | 0.30 | n.s. | 3.20 | 2.8 × 10−33 | 21 | Indole-3-acetate β-glucosyltransferase | ||
| VIT_06s0061g00120 | −0.39 | n.s. | 2.68 | 2.5 × 10−28 | 38 | β-1,3-glucanase | ||
| VIT_08s0007g06040 | −0.06 | n.s. | 3.30 | 1.2 × 10−38 | 7 | [SWITCH] β-1,3-glucanase | ||
| VIT_08s0040g01470 | −0.18 | n.s. | 1.66 | 6.7 × 10−9 | 229 | Cis-zeatin O-β-D-glucosyltransferase | ||
| VIT_11s0016g02200 | 0.74 | n.s. | 3.98 | 4.8 × 10−36 | 13 | β-galactosidase | ||
| VIT_12s0028g00050 | 0.49 | n.s. | 1.29 | 1.5 × 10−3 | 362 | β-1,3 glucanase | ||
| VIT_18s0001g06090 | 0.88 | n.s. | −1.40 | 4.7 × 10−5 | 326 | Cis-zeatin O-β-D-glucosyltransferase | ||
| VIT_19s0014g03240 | −0.05 | n.s. | −1.49 | 1.3 × 10−5 | 312 | β-mannosidase 4 |
| Analyses Performed | Year of Sampling | Grape Cluster Tissue | Description | DAA | EL-Scale | Sampling Date (DD/MM/YYYY) |
|---|---|---|---|---|---|---|
| Microarray | 2011 | rachis | Pre-symptomatic T1 | 49 and 55 | EL-34 and EL-35 | 29 July 2011 & 4 August 2011 |
| Microarray | 2011 | rachis | Symptomatic T2 | 69 and 75 | EL-36 and EL-37 | 17 August 2011 & 24 August 2011 |
| qPCR | 2013 | rachis | Pre-symptomatic | 30 | EL-32 | 11 July 2013 |
| qPCR | 2013 | rachis | Pre-symptomatic | 44 | EL-33 | 25 July 2013 |
| qPCR | 2013 | rachis | Pre-symptomatic | 51 | EL-34 | 1 August 2013 |
| qPCR | 2013 | rachis | Veraison, pre-symptomatic | 58 | EL-35 | 8 August 2013 |
| qPCR | 2013 | rachis | Symptomatic | 65 | EL-36/1 | 15 August 2013 |
| qPCR | 2013 | rachis | Symptomatic | 72 | EL-36/2 | 22 August 2013 |
| RNASeq | 2013 | berries | Pre-symptomatic | 30 | EL-32 | 11 July 2013 |
| RNASeq | 2013 | berries | Pre-symptomatic | 44 | EL-33 | 25 July 2013 |
| RNASeq | 2013 | berries | Pre-symptomatic | 51 | EL-34 | 1 August 2013 |
| RNASeq | 2013 | berries | Veraison, pre-symptomatic | 58 | EL-35 | 8 August 2013 |
| RNASeq | 2013 | berries | Symptomatic | 65 | EL-36/1 | 15 August 2013 |
| RNASeq | 2013 | berries | Symptomatic | 72 | EL-36/2 | 22 August 2013 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Savoi, S.; Supapvanich, S.; Hildebrand, H.; Stralis-Pavese, N.; Forneck, A.; Kreil, D.P.; Griesser, M. Expression Analyses in the Rachis Hint towards Major Cell Wall Modifications in Grape Clusters Showing Berry Shrivel Symptoms. Plants 2022, 11, 2159. https://doi.org/10.3390/plants11162159
Savoi S, Supapvanich S, Hildebrand H, Stralis-Pavese N, Forneck A, Kreil DP, Griesser M. Expression Analyses in the Rachis Hint towards Major Cell Wall Modifications in Grape Clusters Showing Berry Shrivel Symptoms. Plants. 2022; 11(16):2159. https://doi.org/10.3390/plants11162159
Chicago/Turabian StyleSavoi, Stefania, Suriyan Supapvanich, Heinrich Hildebrand, Nancy Stralis-Pavese, Astrid Forneck, David P. Kreil, and Michaela Griesser. 2022. "Expression Analyses in the Rachis Hint towards Major Cell Wall Modifications in Grape Clusters Showing Berry Shrivel Symptoms" Plants 11, no. 16: 2159. https://doi.org/10.3390/plants11162159
APA StyleSavoi, S., Supapvanich, S., Hildebrand, H., Stralis-Pavese, N., Forneck, A., Kreil, D. P., & Griesser, M. (2022). Expression Analyses in the Rachis Hint towards Major Cell Wall Modifications in Grape Clusters Showing Berry Shrivel Symptoms. Plants, 11(16), 2159. https://doi.org/10.3390/plants11162159







