Transcripts Expressed during Germination Sensu Stricto Are Associated with Vigor in Soybean Seeds
Abstract
:1. Introduction
2. Results
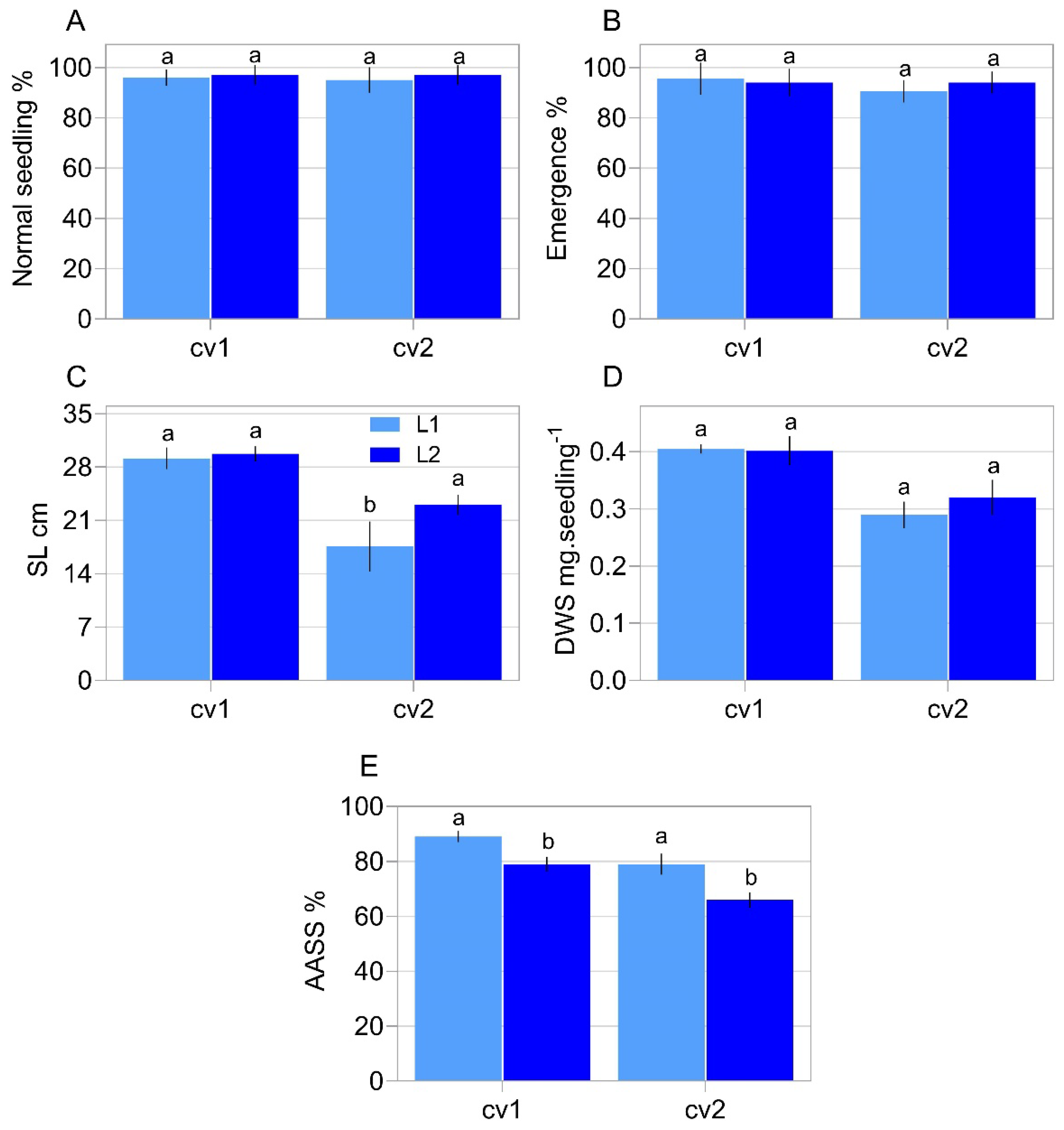
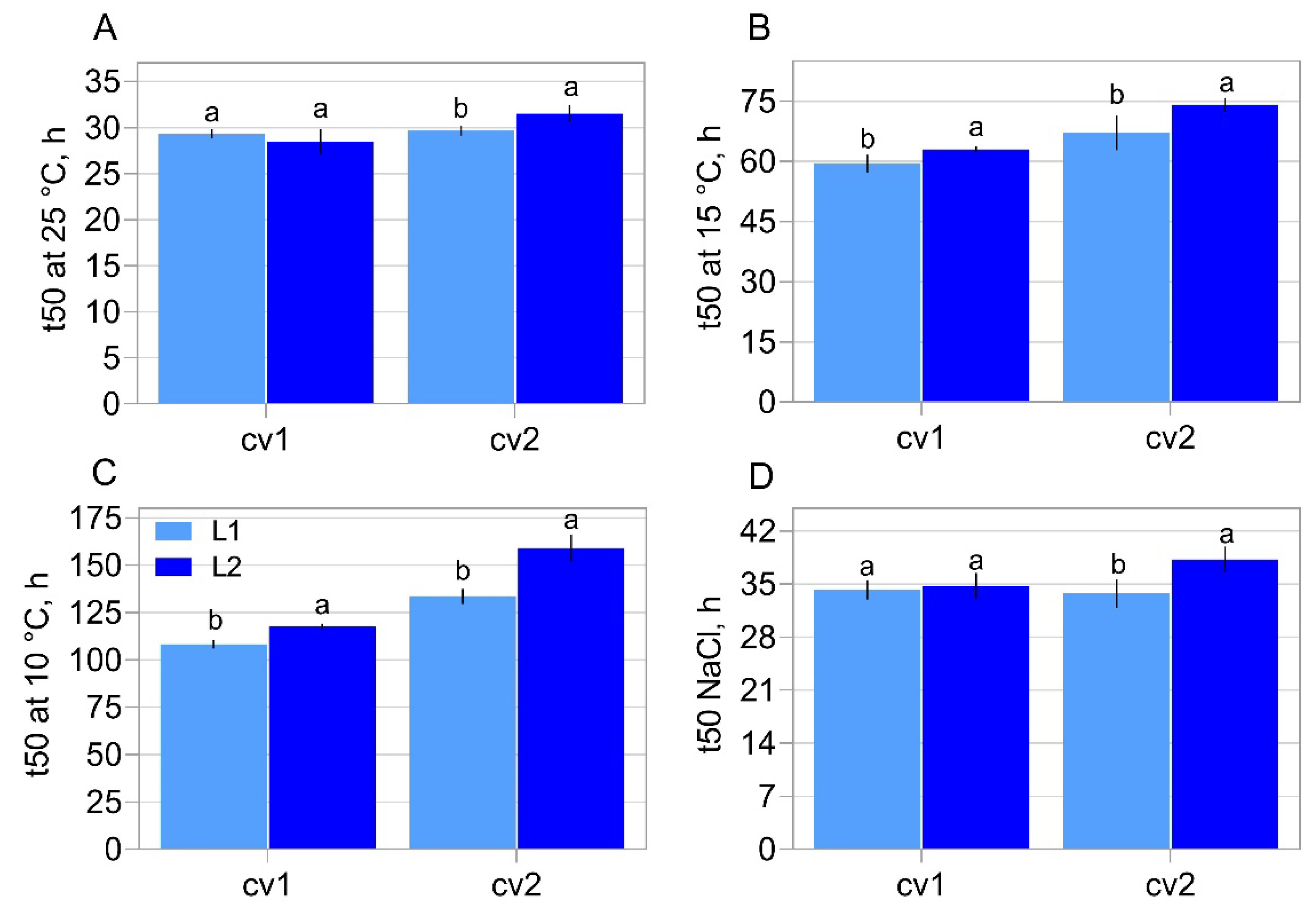
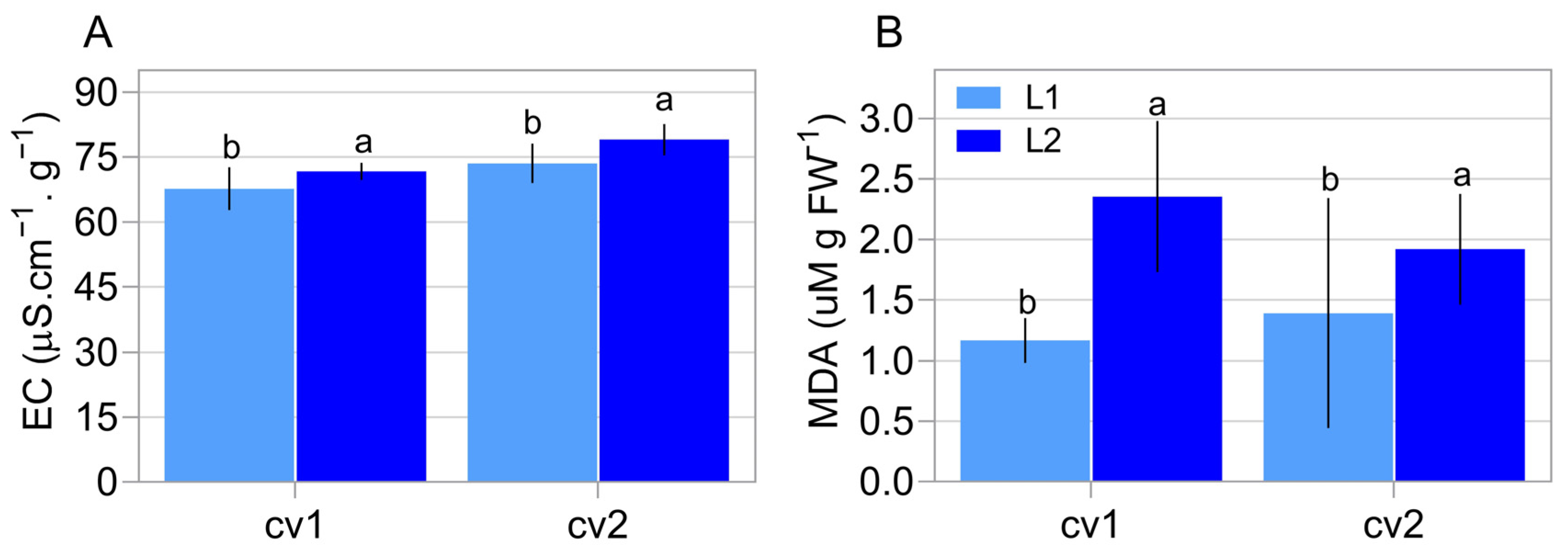
2.1. Characterization of Seed Properties
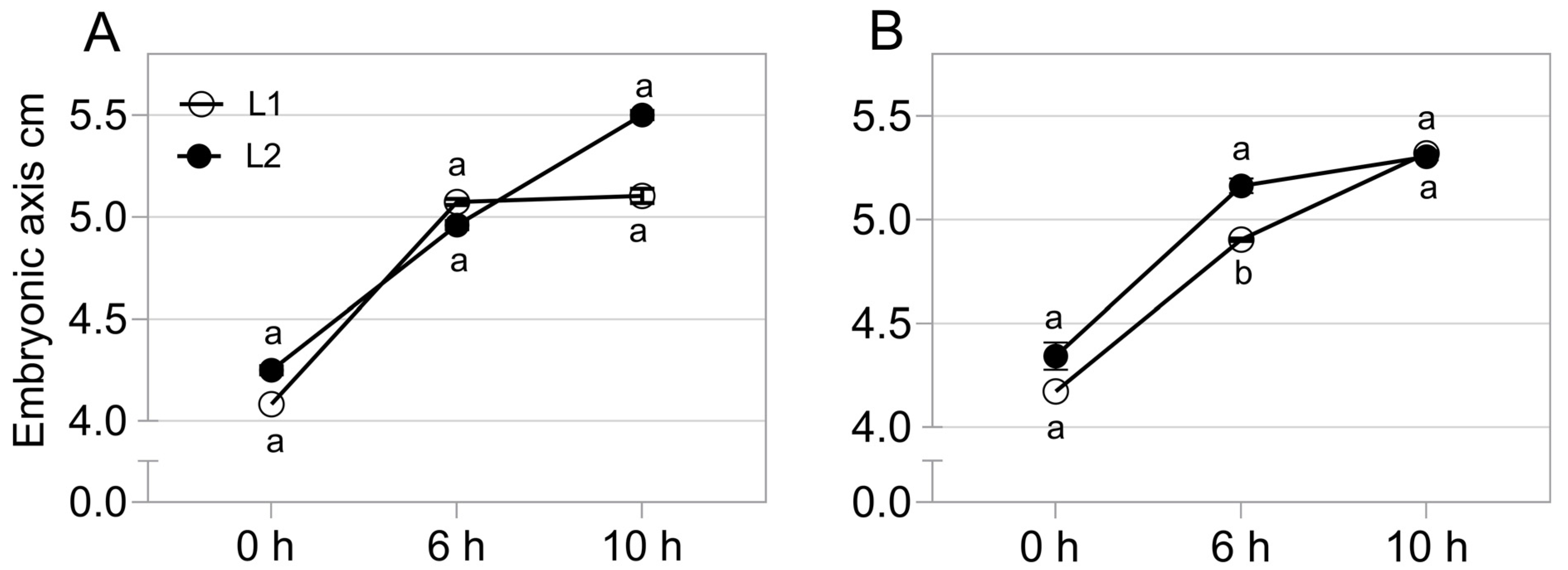
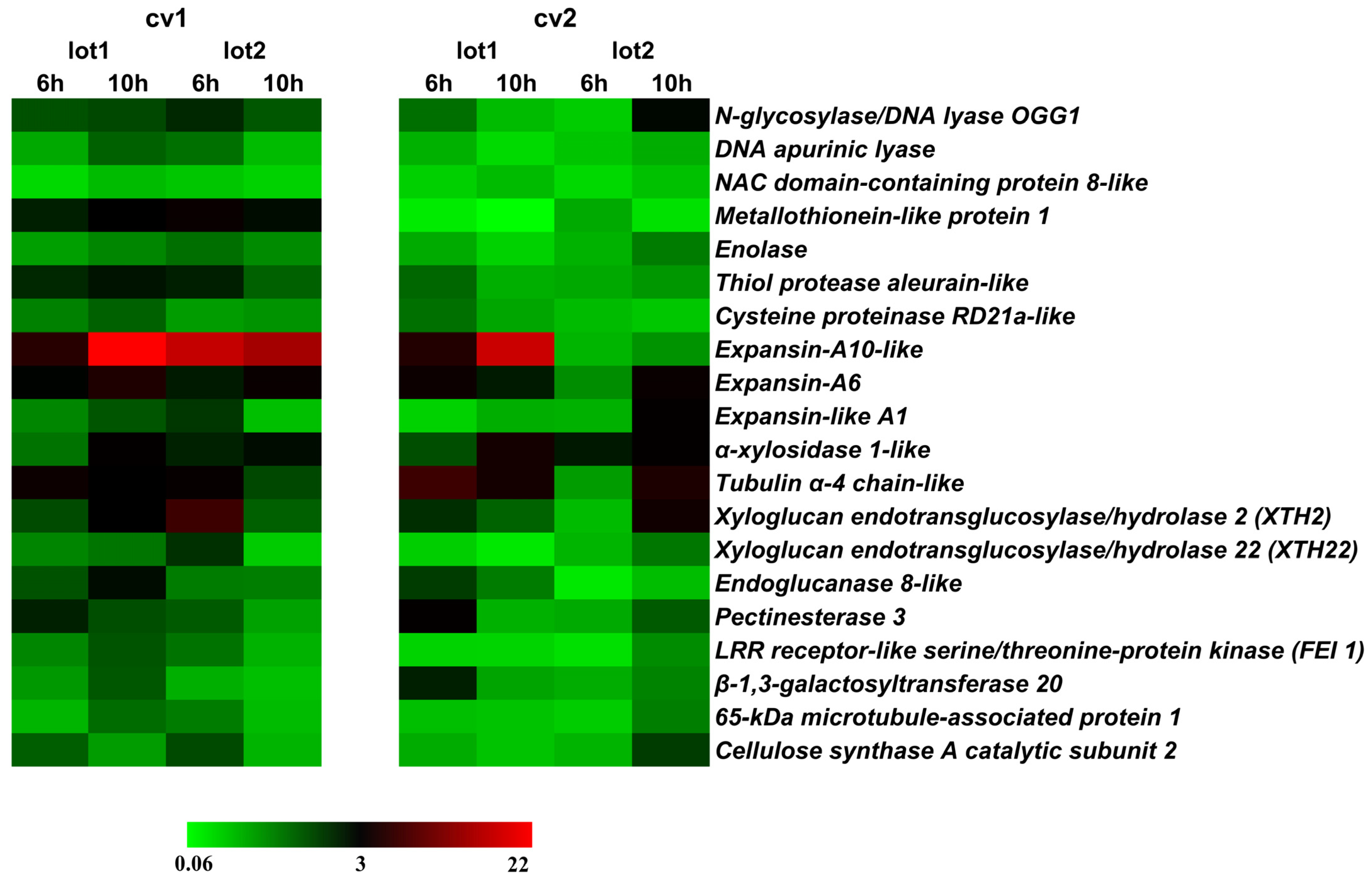
2.2. Gene Expression Related to Growth in the Embryonic Axis during Germination
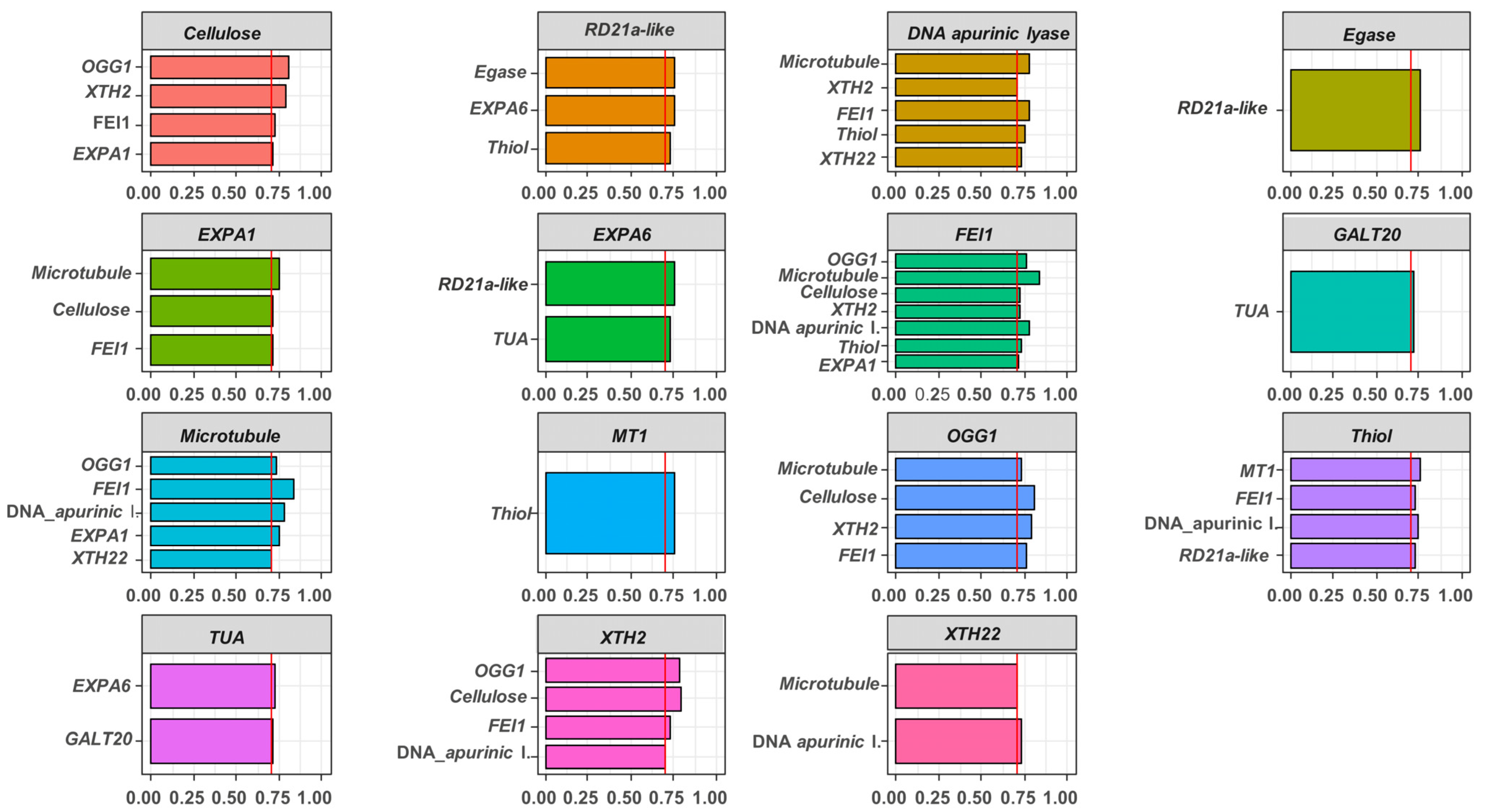
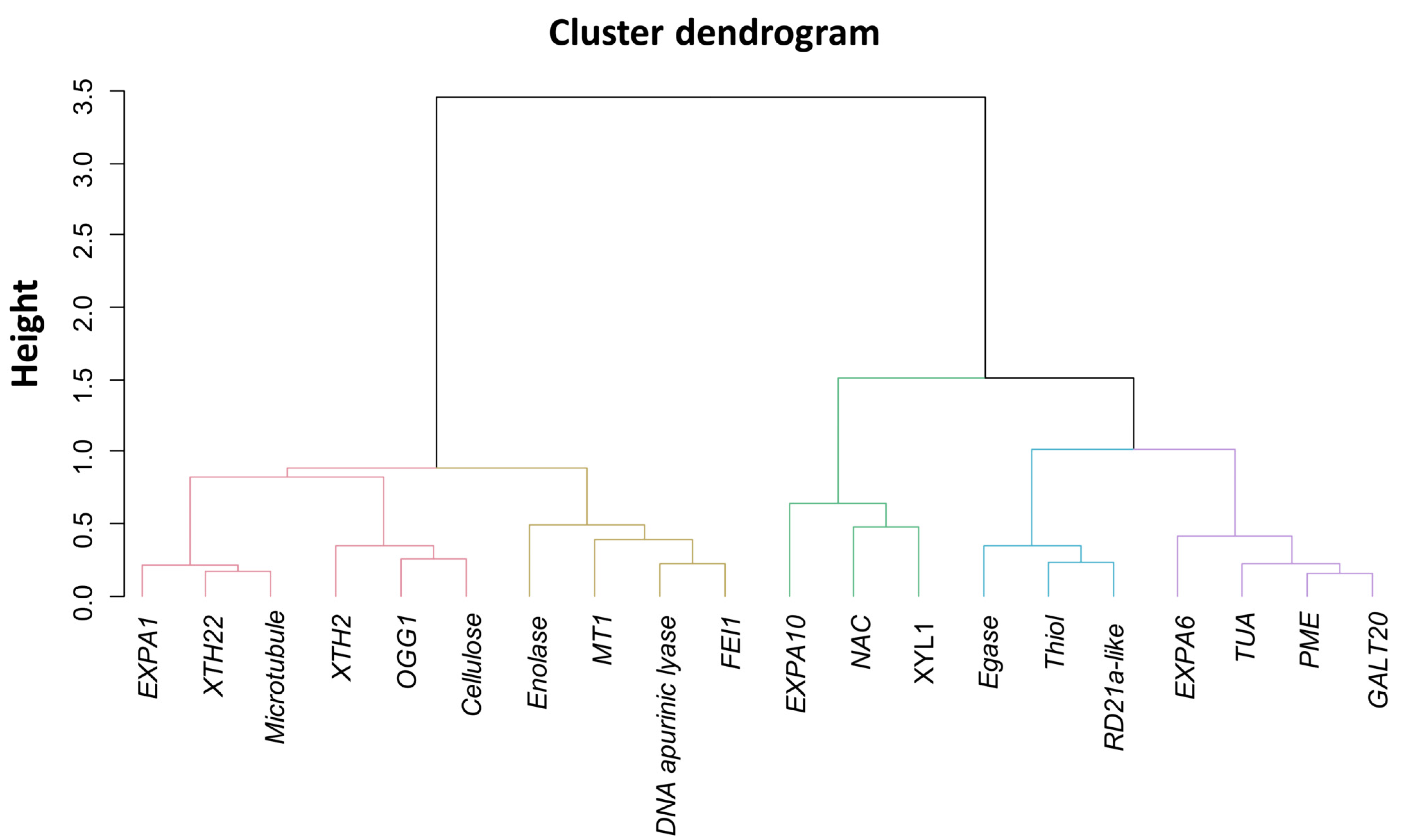
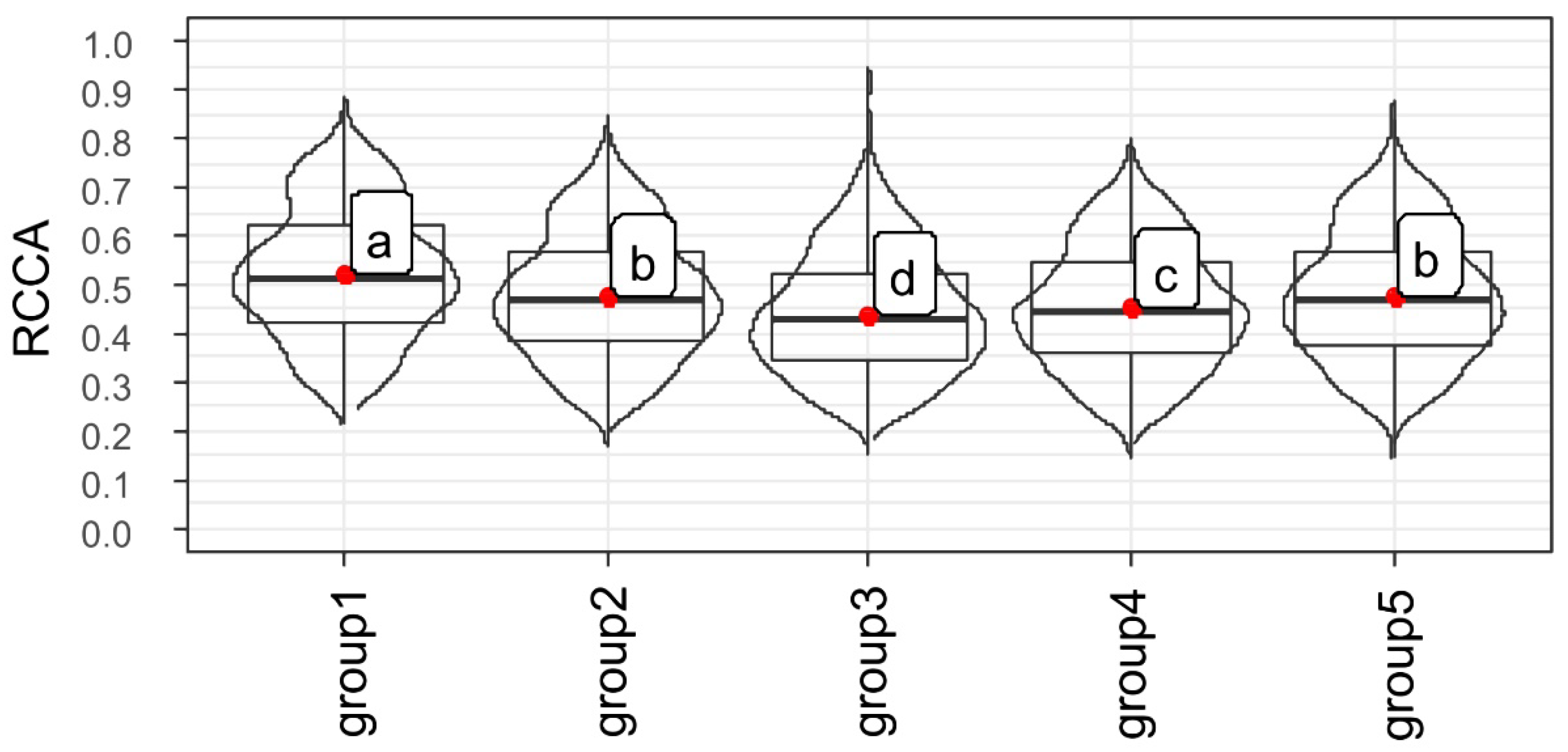
2.3. Association between Gene Expressions during Germination Sensu Stricto with Seed Vigor
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Seed Samples
5.2. Physiological Assays
5.3. Gene Expression during Germination Sensu Stricto
5.4. Statistical Design
Supplementary Materials
Author Contributions
Funding
Institutional review board statement
Informed consent statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Soystats. 25 July 2012. Available online: http://soystats.com (accessed on 20 September 2021).
- Haggag, W.M.; Abouziena, H.F.; Abd-El-Kreem, F.; El-Habbasha, S. Agriculture biotechnology for management of multiple biotic and abiotic environmental stress in crops. J. Chem. Pharm. Res. 2015, 7, 882–889. [Google Scholar]
- Fonseca de Oliveira, G.R.; Mastrangelo, C.B.; Hirai, W.Y.; Batista, T.B.; Sudki, J.M.; Petronilio, A.C.P.; Crusciol, C.A.C.; Amaral da Silva, E.A. An Approach Using Emerging Optical Technologies and Artificial Intelligence Brings New Markers to Evaluate Peanut Seed Quality. Front. Plant Sci. 2022, 13, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Ebone, L.A.; Caverzan, A.; Tagliari, A.; Chiomento, J.L.T.; Silveira, D.C.; Chavarri, G. Soybean seed vigor: Uniformity and growth as key factors to improve yield. Agronomy. 2020, 10, 545. [Google Scholar] [CrossRef]
- Bewley, J.D.; Bradford, K.J.; Hilhorst, H.W.M.; Nonogaki, H. Seeds: Physiology of Development, Germination and Dormancy, 3rd ed.; Springer International Publishing: New York, NY, USA, 2013; 408p. [Google Scholar]
- Ribeiro-Oliveira, J.P.; Bosseli, M.A.; Silva, E.A.A. Acceleration in germination Sensu stricto plays a central role on seedling vigor in post-germination. Plants 2021, 10, 2151. [Google Scholar] [CrossRef] [PubMed]
- Perry, D.A. Report of the vigour test committee 1974–1977. Seed Sci. Technol. 1978, 159–181. [Google Scholar]
- Perry, D.A. The concept of seed vigour and its relevance to seed production techniques. In Seed Production, 1st ed.; Hebblethwaite, P.D., Ed.; Butterworths: London, UK, 1980; pp. 585–591. [Google Scholar]
- Finch-savage, W.E.; Bassel, G.W. Seed vigour and crop establishment: Extending performance beyond adaptation. J. Exp. Bot. 2016, 67, 567–591. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barboza da Silva, C.; Oliveira, N.M.; de Carvalho, M.E.A.; de Medeiros, A.D.; de Lima Nogueira, M.; dos Reis, A.R. Autofluorescence-spectral imaging as an innovative method for rapid, non-destructive and reliable assessing of soybean seed quality. Sci. Rep. 2021, 11, 1–12. [Google Scholar] [CrossRef]
- Basso, D.P.; Hoshino-Bezerra, A.A.; Sartori, M.M.P.; Buitink, J.; Leprince, O.; Silva, E.A.A. Late seed maturation improves the preservation of seedling emergence during storage in soybean. J. Seed Sci. 2018, 40, 185–192. [Google Scholar] [CrossRef]
- Lima, J.J.P.; Buitink, J.; Lalanne, D.; Rossi, R.F.; Pelletier, S.; Silva, E.A.A.; Leprince, O. Molecular characterization of the acquisition of longevity during seed maturation in soybean. PLoS ONE 2017, 12, e0180282. [Google Scholar] [CrossRef]
- Bellieny-Rabelo, D.; Oliveira, E.A.G.; Ribeiro, E.S.; Costa, E.P.; Oliveira, A.E.A.; Venancio, T.M. Transcriptome analysis uncovers key regulatory and metabolic aspects of soybean embryonic axes during germination. Sci. Rep. 2016, 6, 36009. [Google Scholar] [CrossRef] [Green Version]
- Chen, H.; Chu, P.; Zhou, Y.; Li, Y.; Liu, J.; Tsang, E.W.T.; Jiang, L.; Wu, K.; Huang, S. Overexpression of AtOGG1, a DNA glycosylase/AP lyase, enhances seed longevity and abiotic stress tolerance in Arabidopsis. J. Exp. Bot. 2012, 63, 4107–4121. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cosgrove, D.J. Growth of the plant cell wall. Nat. Rev. Mol. Cell Biol. 2005, 6, 850–861. [Google Scholar] [CrossRef] [PubMed]
- Grennan, A.K. Metallothioneins, a Diverse Protein Family. Plant Physiol. 2011, 155, 1750–1751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marowa, P.; Ding, A.; Kong, Y. Expansins: Roles in plant growth and potential applications in crop improvement. Plant Cell Rep. 2016, 35, 949–965. [Google Scholar] [CrossRef] [Green Version]
- Sliwinska, E.; Bassel, G.W.; Bewley, J.D. Germination of Arabidopsis thaliana seeds is not completed as a result of elongation of the radicle but of the adjacent transition zone and lower hypocotyl. J. Exp. Bot. 2009, 60, 3587–3594. [Google Scholar] [CrossRef] [Green Version]
- Souza, N.M.; Topham, A.T.; Bassel, G.W. Quantitative analysis of the 3D cell shape changes driving soybean germination. J. Exp. Bot. 2017, 68, 1532–1537. [Google Scholar] [CrossRef] [Green Version]
- Chavent, M.; Kuentz-Simonet, V.; Liquet, B.; Saracco, J. ClustOfVar: An R Package for the Clustering of Variables. J. Sta. Sof. 2012, 50, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Smolikova, G.; Strygina, K.; Krylova, E.; Leonova, T.; Frolov, A.; Khlestkina, E.; Medvedev, S. Transition from seeds to seedlings: Hormonal and epigenetic aspects. Plants 2021, 10, 1884. [Google Scholar] [CrossRef]
- Bassel, G.W.; Lan, H.; Glaab, E.; Gibbs, D.J.; Gerjets, T.; Krasnogor, N.; Bonner, A.J.; Holdsworth, M.J.; Provart, N.J. Genome-wide network model capturing seed germination reveals coordinated regulation of plant cellular phase transitions. Proc. Natl. Acad. Sci. USA 2011, 108, 9709–9714. [Google Scholar] [CrossRef] [Green Version]
- Waterworth, W.M.; Bray, C.M.; West, C.E. The importance of safeguarding genome integrity in germination and seed longevity. J. Exp. Bot. 2015, 66, 3549–3558. [Google Scholar] [CrossRef] [Green Version]
- Matthews, S.; Khajeh-Hosseini, M. Length of the lag period of germination and metabolic repair explain vigour differences in seed lots of maize (Zea mays). Seed Sci. Technol. 2007, 35, 200–212. [Google Scholar] [CrossRef]
- Li, W.; Niu, Y.; Zheng, Y.; Wang, Z. Advances in the understanding of reactive oxygen species-dependent regulation on seed dormancy, germination, and deterioration in crops. Front. Plant Sci. 2022, 13, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Yan, A.; Wu, M.; Yan, L.; Hu, R.; Ali, I.; Gan, Y. AtEXP2 is involved in seed germination and abiotic stress response in Arabidopsis. PLoS ONE 2014, 9, e85208. [Google Scholar] [CrossRef] [PubMed]
- Shen, Z.; Chen, M. Deciphering novel transcriptional regulators of soybean hypocotyl elongation based on gene co-expression network analysis. Front. Plant Sci. 2022, 13, 837130. [Google Scholar] [CrossRef]
- Vivas, P.G.; Resende, L.S.; Braga, R.A.; Guimarães, R.M.; Azevedo, R.; Silva, E.A.A.; Toorop, P.E. Biospeckle activity in coffee seeds is associated non-destructively with seedling quality. Ann. Appl. Biol. 2017, 170, 141–149. [Google Scholar] [CrossRef]
- Brasil. Regras Para Análise de Sementes; MAPA/ACS: Brasília, Brazil, 2009; 399p. [Google Scholar]
- Joosen, R.V.L.; Kodde, J.; Willems, L.A.J.; Ligterink, W.; van der Plas, L.H.W.; Hilhorst, H.W.M. Germinator: A software package for high-throughput scoring and curve fitting of Arabidopsis seed germination. Plant J. 2010, 62, 148–159. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods. 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Nakagawa, J. Testes de vigor baseados no desempenho de plântulas. In Vigor de Sementes: Conceitos e Testes, 1st ed.; Krzyzanowski, F.C., Vieira, R.D., França Neto, J.B., Eds.; ABRATES: Londrina, Brazil, 1999; pp. 1–24. [Google Scholar]
- Nakagawa, J. Testes de vigor baseados na avaliação das plântulas. In Testes de Vigor em Sementes, 1st ed.; Vieira, R.D., Carvalho, N.M., Eds.; FUNEP: Jaboticabal, Brazil, 1994; pp. 49–85. [Google Scholar]
- Marcos-Filho, J. Teste de envelhecimento acelerado. In Vigor de Sementes: Conceitos e Testes, 1st ed.; Krzyzanowski, F.C., Vieira, R.D., França Neto, J.B., Eds.; ABRATES: Londrina, Brazil, 1999; pp. 3.1–3.24. [Google Scholar]
- Vieira, R.D.; Krzyzanowski, F.C. Teste de condutividade elétrica. In Vigor de Sementes: Conceitos e Testes, 1st ed.; Krzyzanowski, F.C., Vieira, R.D., França Neto, J.B., Eds.; ABRATES: Londrina, Brazil, 1999; pp. 1–26. [Google Scholar]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Teixeira, R.N.; Ligterink, W.; França-Neto, J.; Hilhorst, H.W.M.; Silva, E.A.A. Gene expression profiling of the green seed problem in soybean. BMC Plant Bio. 2016, 16, 37. [Google Scholar] [CrossRef] [Green Version]
- Pfaffl, M.W.; Horgan, G.W.; Dempfle, L. Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res. 2002, 30, e36. [Google Scholar] [CrossRef]
- Leurgans, S.E.; Moyeed, R.A.; Silverman, B.W. Canonical correlation analysis when the data are curves. J. Roy. Sta. Soc. Series B (Methodol.) 1993, 55, 725–740. [Google Scholar] [CrossRef]
- Vinod, H.D. Canonical ridge and econometrics of joint production. J. Econ. 1976, 4, 147–166. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ducatti, K.R.; Batista, T.B.; Hirai, W.Y.; Luccas, D.A.; Moreno, L.d.A.; Guimarães, C.C.; Bassel, G.W.; da Silva, E.A.A. Transcripts Expressed during Germination Sensu Stricto Are Associated with Vigor in Soybean Seeds. Plants 2022, 11, 1310. https://doi.org/10.3390/plants11101310
Ducatti KR, Batista TB, Hirai WY, Luccas DA, Moreno LdA, Guimarães CC, Bassel GW, da Silva EAA. Transcripts Expressed during Germination Sensu Stricto Are Associated with Vigor in Soybean Seeds. Plants. 2022; 11(10):1310. https://doi.org/10.3390/plants11101310
Chicago/Turabian StyleDucatti, Karina Renostro, Thiago Barbosa Batista, Welinton Yoshio Hirai, Daiani Ajala Luccas, Leticia de Aguila Moreno, Cristiane Carvalho Guimarães, George W. Bassel, and Edvaldo Aparecido Amaral da Silva. 2022. "Transcripts Expressed during Germination Sensu Stricto Are Associated with Vigor in Soybean Seeds" Plants 11, no. 10: 1310. https://doi.org/10.3390/plants11101310
APA StyleDucatti, K. R., Batista, T. B., Hirai, W. Y., Luccas, D. A., Moreno, L. d. A., Guimarães, C. C., Bassel, G. W., & da Silva, E. A. A. (2022). Transcripts Expressed during Germination Sensu Stricto Are Associated with Vigor in Soybean Seeds. Plants, 11(10), 1310. https://doi.org/10.3390/plants11101310






