Abstract
Spontaneous tumors can develop in different organs of various plant species without any pathogen infection and, as a rule, appear in plants with a certain genotype: Mutants, interspecific hybrids, etc. In particular, among the inbred lines of radish (Raphanus sativus L.), lines that form spontaneous tumors on the taproot during the flowering period were obtained many years ago. In this work, we analyzed the differential gene expression in the spontaneous tumors of radish versus the lateral roots using the RNA-seq method. Data were obtained indicating the increased expression of genes associated with cell division and growth (especially genes that regulate G2-M transition and cytokinesis) in the spontaneous tumor. Among genes downregulated in the tumor tissue, genes participating in the response to stress and wounding, mainly involved in the biosynthesis of jasmonic acid and glucosinolates, were enriched. Our data will help elucidate the mechanisms of spontaneous tumor development in higher plants.
1. Introduction
The tumor (syn.: Neoplasm) is a pathological structure emerging as a result of uncontrolled proliferation of a group of cells leaving the systemic control of growth rate, cell differentiation, and proliferation. Therefore, the elucidation of the mechanisms of tumor formation may help identify the key regulators of systemic mechanisms controlling cell proliferation and differentiation. Tumor-like structures are found in almost all multicellular organisms, including higher plants. Pathogen-induced tumors, which make up the majority of neoplasms in higher plants, develop under the influence of infectious agents (bacteria, viruses, fungi, nematodes, insects, etc.), which create a niche for their own habitation in the host plant’s organism mostly by shifting the phytohormonal balance and sometimes activating the meristematic competence of plant cells or modulation of the plant cell cycle. At the same time, much rarer spontaneous tumors of higher plants are formed in plants with certain genotypes (mutants, interspecific hybrids, inbred lines) in the absence of any pathogen, which make them closer to animal tumors [1].
The object of our research is spontaneous tumorigenesis in inbred lines of radish (Raphanus sativus var. radicula Pers.). The genetic collection of radish was created in St. Petersburg State University by selfing individual plants of four cultivars [2], and now it contains 32 highly inbred lines [3]. Eleven radish lines of different origins are characterized by the spontaneous formation of tumors on the roots of plants at the beginning of the flowering stage [3] (Figure 1). Spontaneous tumors in radish originate from the pericycle and cambium cells [4,5], which brings them closer to the lateral and adventitious roots. Moreover, young growing tumors contain meristematic foci in the periphery, which resemble root apical meristems due to the presence of indole-3-acetic acid (IAA) response maxima and RsWOX5 expression [5]. The genetic analysis revealed that the spontaneous tumor formation in radish is a polygenic trait, however, it is also inherited as a monogenic recessive trait in several hybrid combinations [6]. As with most examples of spontaneous tumors in plants, the exact cause of spontaneous tumor formation in radish inbred lines is unknown, but the most likely reason is associated with an increase of free cytokinins (CKs) content, which was found in the roots of tumor-producing radish lines [6].
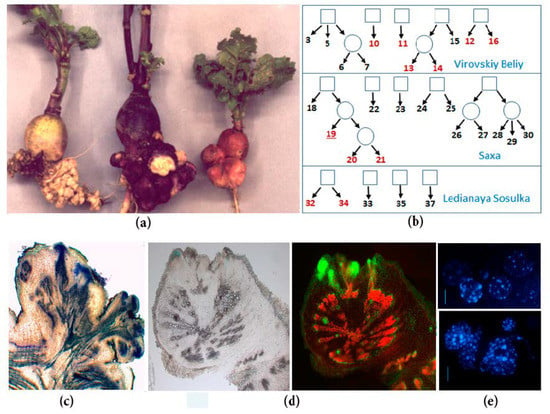
Figure 1.
Spontaneous tumors of radish inbred lines: (a) Spontaneous tumors on the taproots of different inbred lines of SPbU genetic collection of Raphanus sativus (from left to right: Lines 34, 16, and 19); (b) the origin of inbred radish lines of SPbU genetic collection. Lines with spontaneous tumor formation are marked in red; (c) anatomy of mature spontaneous tumors in radish: Mass of undifferentiated cells in the periphery of tumor, vascularization of the proximal part of tumor, and the connection of tumor and plant vascular systems; (d) analysis of cell proliferation intensity in the radish taproot with young tumors: Cells with active DNA synthesis were incorporated EdU (5-ethynyl-2’-deoxyuridine) and fluorescently labeled with Alexa Fluor-488 (right) [5]; (e) cytological analysis of roots and tumors in radish inbred line 19 (DAPI staining): Tumor cells (below) have an increased number of chromocenters, which indicate an increased level of ploidy in comparison with taproot cells (above) [7].
In the present study, we used RNA-seq to analyze the changes in gene expression at the early stages of development of spontaneous tumors in radish inbred line 19, while the lateral roots of the same plants were used as the control. The data we obtained revealed the differential expression of more than 1600 genes. We performed the analysis of GO, KEGG, and GSEA categories and also individual transcripts and revealed the upregulation of genes of pathways associated with cell division and expansion, while among the downregulated, the genes involved in stress response were overrepresented. Our findings were confirmed with qPCR for selected transcripts. Since the examples of spontaneous tumors in plants are very rare, and the mechanisms of their development are poorly understood, our study contributes to the understanding of the genetic control of spontaneous tumors development in radish inbred lines in particular and in higher plants in general.
2. Results
2.1. RNA-seq of Radish Spontaneous Tumors
Spontaneous tumors of radish, such as lateral roots, are of pericyclic origin and, possibly, represent some modification of the lateral root development program [5]. To compare transcriptomes of spontaneous tumors and lateral roots of radish, we isolated the RNA from tumors and lateral roots of radish plants of tumor-producing line 19 and subjected it to sequencing. Total RNA from three replicates of root and tumor samples was sequenced with an Illumina HiSeq2500 sequencer resulting in 190.3 million paired-end reads. After all adapter trimming and contamination removal steps, 73.8 million paired-end reads were used for differential expression estimation and all downstream analyses.
We have found that 425 genes were significantly upregulated in young tumors compared to lateral roots, while 1203 genes were significantly downregulated (adjusted p-value < 0.05). Data on differential expression of genes were further analyzed using the gene set enrichment analysis (GSEA) method with the fgsea package. Three hundred and forty-four groups of genes (out of 9831 groups taken for the analysis) were significantly enriched among the genes with increased expression in radish tumor samples, and 132 groups of genes were significantly enriched among the genes with reduced expression in the tumor compared to the lateral root (adjusted p-value < 0.05) (Table S2). The gene ontology enrichment analysis performed with the clusterProfiler package on significantly differentially expressed genes has shown that 76 biological process terms were enriched among upregulated genes and 99 were enriched among downregulated genes with adjusted p-value < 0.05 (Figure 2, Table S2), whereas the KEGG enrichment analysis has shown that 8 and 15 pathways were enriched among upregulated and downregulated genes, respectively (adjusted p-value < 0.05) (Table S2).
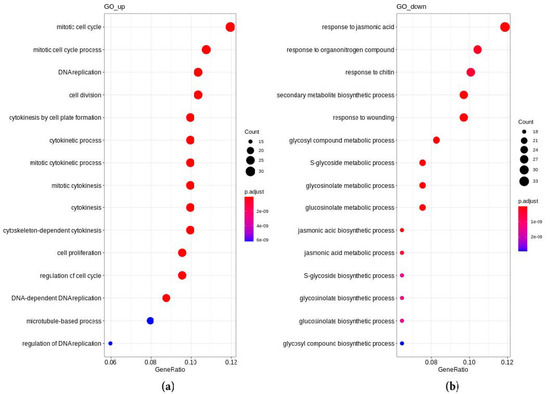
Figure 2.
Overrepresented “biological process” GO pathways in genes upregulated (a) and downregulated (b) in the spontaneous tumors of radish in comparison with lateral roots. The count is the number of DEGs included in the respective pathway.
2.2. Enriched Pathways in Genes Differentially Expressed between Spontaneous Tumors and Lateral Roots
We analyzed GO, GSEA, and KEGG pathways in lateral roots and spontaneous tumors of radish line 19. The data are presented in Table S2 and in Figure 2 and Figure 3.
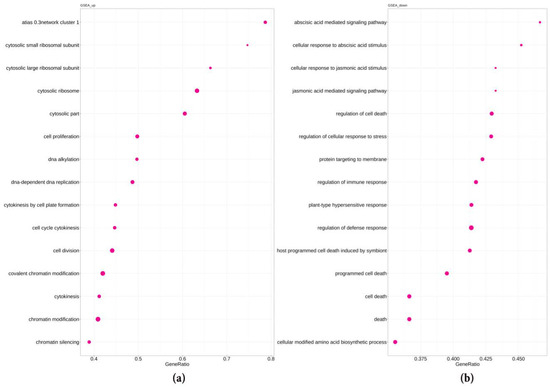
Figure 3.
Overrepresented “biological process” GSEA pathways in genes upregulated (a) and downregulated (b) in the spontaneous tumors of radish in comparison with lateral roots. The count is the number of DEGs included in the respective pathway, the adjusted p-value is 3.4 × 10−9.
According to the GO pathways analysis, among the upregulated in young spontaneous tumors (Figure 2a), the most enriched were pathways associated with the regulation of hyperplasia: Cell cycle (GO:1903047, GO:0000278, GO:0008283, GO:0051301, GO:0051301, GO:0051726, GO:0010389, GO:1902749, GO:0000086, GO:1901987, GO:1901990, GO:0044770, GO:0044772, GO:0007346, GO:0010564, GO:0045786), DNA replication (GO:0006260, GO:0006261, GO:0006275, GO:0006270, GO:0042023, GO:0044786), cytokinesis (GO:0000911, GO:0032506, GO:1902410, GO:0000281, GO:0000910, GO:0061640), spindle assembly (GO:0051225, GO:0007051), cytoskeleton rearrangement (GO:0007017, GO:0000226, GO:0007010), cell expansion (GO:0009825), and cell wall modifications (GO:0009828, GO:0042547, GO:0009831, GO:0009505), which reflect the activation of cell division and growth in the developing tumor in comparison with the lateral root. The upregulation was noted for pathways involved in chromatin modification (GO:0016570, GO:0034968, GO:0016569, GO:0016571, GO:0016572, GO:0051567, GO:0061647, GO:0006305, GO:0006306, GO:0044728, GO:0006304, GO:0000785) and gene silencing (GO:0016458), response to hormones (CK—GO:0009736, GO:0071368, and gibberellic acid (GA)—GO:0009739), numerous pathways associated with amino acid and protein modifications (GO:0018022, GO:0006479, GO:0008213, GO:0018205, GO:0018193), DNA metabolism (GO:0051052), organelle development (GO:0070925, GO:0031225, GO:0022626, GO:0044445), and morphogenetic processes (GO:0010374, GO:0048646, GO:0048831).
At the same time, among downregulated (Figure 2b) genes, there were numerous stress-related pathways such as response to wounding (GO:0009611), response to abiotic stress (GO:0080135, GO:0006979, GO:0010193), and immune response (GO:0045088, GO:0050776, GO:0002682, GO:0050832, GO:0009625, GO:0031348, GO:0010200), in particular, of hypersensitive response (HR) GO:0010363, GO:0009626) and systemic acquired resistance (SAR) (GO:0009862) reactions, and cell death (GO:0043067, GO:0010941, GO:0012501, GO:0034050, GO:0008219). Among downregulated genes, there were also pathways of biosynthesis, metabolism, and signaling of different phytohormones, which can also participate in plant stress response: jasmonic acid (JA) (GO:0009753, GO:0009695, GO:0009694, GO:0009867, GO:0071395), salycilic acid (SA) (GO:0009751, GO:0009863, GO:0071446), ethylene (GO:0009692, GO:0009693, GO:0009723), IAA (GO:0009684, GO:0009683, GO:0042435, GO:0009851, GO:0009850, GO:0009733, GO:0042436), and even karrikins (GO:0080167). The activation of some metabolic pathways, such as biosynthesis and metabolism of ROS compounds (GO:0010310, GO:2000377, GO:0016702, GO:0016701), phenylpropanoids (GO:0009698, GO:0009699), flavonoids (GO:0009812, GO:0009813, GO:0009963, GO:0009962), glucosinolates (GO:0019757, GO:0019760, GO:0019758, GO:0019761), and polyamines (GO:0006595, GO:0006598), may also be associated with the response to stress. The downregulation was also found for pathways associated with amino acid metabolism and transport (GO:0019344, GO:0006534, GO:0009070, GO:0009069, GO:0003333, GO:0006865), and also the transport of organic acids (GO:0015849, GO:0046942) and anions (GO:0015711, GO:0015698).
The data obtained in the analysis of the GSEA and KEGG pathways (Table S2), in general confirm the data obtained for the GO pathways. For instance, among upregulated GSEA pathways in spontaneous tumors (Figure 3, Table S2), those associated with cell division and chromatin modification were presented. Moreover, among the enriched, there were GSEA pathways associated with the ribosome biogenesis and also the development of meristems and floral organs. At the same time, among downregulated GSEA pathways in the spontaneous tumors, the pathways associated with the response to JA, ABA, as well as stress and immune responses were prevailed.
Among the most enriched KEGG pathways in the tumor tissue, there were also those associated with cell division (e.g., DNA replication and mismatch repair), and among the downregulated, the most enriched were the pathways of biosynthesis of glucosinolates and phenylpropanoides. Thus, according to the data on enriched pathways determined by different classifications (GO, GSEA, and KEGG pathways analyses), most of the upregulated pathways in spontaneous tumors were related with cell division and chromatin modification, and most of the downregulated pathways were associated with stress response.
2.3. Individual Transcripts Differentially Expressed between Spontaneous Tumors and Lateral Roots
In our experiment, numerous genes that are differentially expressed were identified between the spontaneous tumor and the lateral root: 425 genes were significantly upregulated and 1203 genes were downregulated in the spontaneous tumors of radish (Table S2).
As noted above, the large number of genes upregulated in spontaneous tumors control cell division and elongation, while genes involved in the stress response were widely represented among the downregulated ones. The exact functions of these genes in radish are unknown, however, the functions of their closest homologues in Arabidopsis were previously studied. Due to the high conservatism of the pathways of cell cycle control [8], as well as JA biosynthesis and stress response [9] in higher plants, we can assume that the radish DEGs identified by us probably perform functions similar to those of their homologues in Arabidopsis.
2.3.1. Cell Division and Cell Expansion Genes: Mostly Upregulated
As it was shown in anatomical studies, the development of tumors of various types in plants is associated with the intensification of cell division and cell elongation [1,10]. According to this, a lot of genes associated with mitotic cell cycle and most of the genes associated with cell growth were upregulated in spontaneous tumors (Figure 4) reflecting the increased level of cell proliferation during the formation of neoplasm.
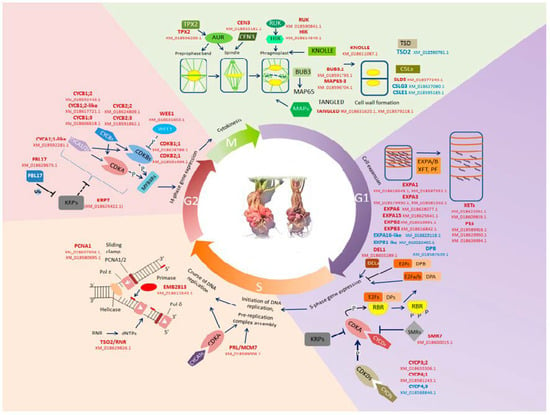
Figure 4.
Scheme of genetic control of the plant cell cycle; genes whose homologues were differentially expressed in spontaneous tumors of radish are marked in color (red—upregulated genes, blue—downregulated genes). See the text for an explanation.
As in the analysis of pathways, among the individual upregulated transcripts, there were radish homologs of Arabidopsis genes that are involved in the control of different stages of the cell cycle and cytokinesis: Genes encoding cyclins of different classes, cyclin-dependent kinase B (CDKB), E2F and DP transcription factors, WEE1 kinase, cell cycle inhibitors of KRP and SMR families, proteins involved in the DNA replication and in the cytoskeleton dynamics during mitosis, and also genes, encoding cell wall loosening enzymes, which are essential for cell expansion (Figure 4). An increase in the expression levels of these genes in the spontaneous tumors may be the cause for the intensification of cell divisions in the tumor tissue.
At the G1 phase of plant cell cycle, the growth of plant cell takes place requiring cell wall destabilization, which depends on the activity of cell wall modifying enzymes such as expansins (EXP), pectinesterases (PE), and xyloglucan endotransglucosylases/hydrolases (XET) [11]. Radish homologs of Arabidopsis EXPA1, EXPA3, EXPA6, EXPA15, EXPB2, EXPB3, along with three PE genes and two XET genes were upregulated in the spontaneous tumors, and only two EXP genes were downregulated. In addition, TUMOROUS SHOOT DEVELOPMENT 2/QUASIMODO 2 (TSD2/QUA2) encoding pectin methyltranspherase, which is essential for cell adhesion was downregulated in the tumor tissue [12].
The G1-S transition of plant cell cycle is under the control of A-class cyclin-dependent kinases (CDKA) and D-class cyclins (CYCDs) [13,14]: CDKA-CYCDs phosphorylate the retinoblastoma-related (RBR) protein and release transcription factors (TFs) of E2F/DP family from an interaction with RBR [13,14]. The E2F and DP proteins form heterodimers, while DP-E2F-like (DEL) proteins do not dimerize [15]. The E2F-DP and DEL TFs regulate the expression of S phase genes [16,17]. Among DEGs in spontaneous tumors, there were radish homologs of Arabidopsis E2FE/DEL1 and DPB genes: homolog of DEL1 was upregulated in the tumors, while homolog of DPB was strongly downregulated.
The main event of S phase of the plant cell cycle, the course of which is under the control of CDKA-CYCA3 complexes [18], is DNA replication. In the spontaneous tumors of radish, several genes encoding important regulators of DNA replication were upregulated. Among them were radish homologs of PROLIFERA (PRL), which encode a key component of the pre-replication complex [19], PROLIFERATING CELLULAR NUCLEAR ANTIGEN 1 (PCNA1) whose product forms the DNA clamp [20], TSO2 whose product catalyzes the synthesis of deoxyribonucleosides [21], EMBRYO DEFECTIVE 2813 (EMB2813) encoding a large subunit of DNA primase [22], and RPA32B (AT3G02920) which encodes one of three subunits of replication protein A (RPA)—an ssDNA-binding protein [23].
The G2-M transition of plant cell cycle is under the control of CDKAs and CDKBs, which interact with B-class and certain A-class cyclins [24], and the main targets of CDKs at G2-M are MYB3R TFs, which regulate the expression of M phase genes [25]. In the spontaneous tumors, the upregulation of genes homologous to Arabidopsis CDKB1;1, CDKB2;1, CYCA1;1-like, CYCB1;2, CYCB1;2-like, CYCB1;3, and CYCB2;2 was noticed.
The activity of CDK-cyclin complexes at all phases of the plant cell cycle depends on other proteins: CDKDs, activators of CDK, which interact with CYCPs acting in the nutritional control of cell cycle [26,27], and inhibitors of CDK—SIAMESE-RELATED (SMR) [28] and Kip-related proteins (KRPs) [29]. In turn, the stability of KRPs depends on the F-box protein FBL17 [30]. At G2-M, the CDK activity is also under the control of the WEE1 kinase, which phosphorylates and inhibits CDKs [31]. Among genes which were upregulated in the spontaneous tumors of radish, there were homologs of Arabidopsis CYCP3;2, CYCP4;1, SMR7, KRP7, FBL17, and WEE1, while among downregulated genes, there was a homolog of CYCP4;3.
At the M phase of cell cycle, various regulators of spindle formation and cytokinesis play a key role. Radish homologs of several genes from these groups were upregulated in the spontaneous tumors, including genes encoding microtubule-associated proteins TANGLED and MAP65-3/PLEIADE, which play a role at early and late mitosis [32,33], BUDDING UNINHIBITED BY BENZYMIDAZOL3.2 (BUB3.2), which encodes the mitotic checkpoint protein regulating the localization of MAP65-3 in the phragmoplast [34], TARGETING PROTEIN FOR XKLP2 (TPX2) whose product is an upstream regulator of Aurora A kinase and a participant of mitotic spindle assembly [35], CENTROMERIC HISTONE H3 (CEN3) encoding an assembly site for the kinetochore complex [36], KNOLLE encoding cytokinesis-specific syntaxin [37], RUNKEL (RUK) encoding microtubule-associated kinase [38] together with the gene of RUK target HINKEL (HIK), a kinesin required for phragmoplast expansion [39], and also SLD5 encoding cellulose synthase, which functions in cell plate formation [40].
It was previously reported that the homologs of cell division and cell expansion genes, which were upregulated in the radish spontaneous tumors can participate in the control of meristem activity and organ size, and also in the development of other types of plant tumors. For example, CDKB2;1 whose radish homolog is strongly upregulated in the spontaneous tumors is involved in cell cycle progression and determines the meristem size [41]. The protein kinase WEE1 which represses the cell cycle progress by inactivating CDKs dose-dependently decreases the meristem size and regenerative capacity in Arabidopsis [42], but the expression of Arabidopsis WEE1 is associated with actively proliferating cells [31]. Therefore, its upregulation in the tumor tissue may be a part of the negative feedback regulation of cells proliferation. It was also reported that the increase of the expression levels of cell cycle and cell expansion genes, such as DEL1, WEE1, KRP7, and certain EXPAs and EXPBs takes place during the development of galls induced by nematodes [43,44,45,46]. In our previous work, the same radish homolog of EXPA1 was revealed as one of the most upregulated genes in the transcriptome of Agrobacterium-induced crown gall tumor [47]. The list of downregulated genes in the spontaneous tumor includes the radish homolog of TSD2/QUA2, while the tsd2/qua2 mutant belongs to a specific small group of monogenic spontaneous tumor-producing mutants of Arabidopsis [1,12]. Therefore, downregulation of TSD2 homolog may be one of the causes of spontaneous tumor formation in radish.
2.3.2. Stress Response Genes: Mostly Downregulated
Among the downregulated in the spontaneous tumors, the stress-related genes, namely genes involved in the biotic stress and wounding response, are widely represented (Figure 5).

Figure 5.
Scheme of several pathways of the plant stress response; genes whose homologues were differentially expressed in spontaneous tumors of radish are marked in color (red—upregulated genes, blue—downregulated genes). See the text for an explanation.
A significant part of these genes is involved in the biosynthesis and signal transduction of JA, an oxylipin phytohormone involved in the response to biotic and abiotic stress, wounding, pathogenesis, and herbivorous insects [9]. The JA biosynthesis is initiated in chloroplasts by the release of α-linolenic acid (α-LeA) from galactolipids of chloroplast membranes by phospholipase A1 (PLA1)/ DEFECTIVE IN ANTHER DEHISCENCE 1 (DAD1) [48]. The second step of JA biosynthesis is catalyzed by lipoxigenases (LOX)—enzymes which oxidize the α-linolenic acid of chloroplast membranes to the (13S)-hydroperoxyoctadecatrienoic acid (13-HPOT). Among LOX family enzymes, only the allene oxide synthase (AOS) branch, which includes LOX2, LOX3, LOX4, and LOX6, is involved in the JA biosynthesis [49]. In the chloroplasts, AOS enzymes work in tight communication with allene oxide cyclases (AOCs), which cyclize the unstable product of α-linolenic acid oxidation to obtain the 12-oxo-phytodienoic acid (OPDA). The third step of JA biosynthesis is the conversion of OPDA in the peroxisome to 3-oxo-2-(2′-[Z]-pentenyl)cyclopentane-1-octanoic acid (OPC-8), which subsequently undergoes three rounds of beta-oxidation to yield JA [50].
The ubiquitin-dependent degradation of JA-zim-domain (JAZ) proteins plays a central role in the response to JA. These proteins are transcriptional repressors which bind and inhibit the TF MYC2 to repress the JA response, and as a negative feedback in the JA response, JAZ transcript levels rise in response to a JA stimulus [51].
Among genes which were downregulated in radish spontaneous tumors, there were homologs of genes which regulate key steps of JA biosynthesis: DAD1-like Lipase 3 (DALL3), LOX3, LOX4, LOX6, and AOC3, as well as homologs of OPCL1 gene encoding enzyme, which activate JA biosynthetic precursors in the peroxisomes [50] and CYP94B3 which encodes a key enzyme in the oxidative catabolism of JA [52]. Among downregulated genes, there were also upstream regulators of JA biosynthesis such as the gene encoding octadecanoid-responsive AP2/ERF-domain TF ORA47 [53]. Among downregulated in the spontaneous tumors, there were also homologs of genes encoding TFs involved in the response to JA MYC2 and AIF1, as well as JA-responsive genes of transcriptional repressors JAZ6, JAZ8, and JAZ9 [51,54].
Moreover, among genes which were downregulated in the spontaneous tumor, there were many stress-responsive genes acting probably independently of JA. These genes encode quite different proteins, and some of them are involved in the metabolism, transport or signaling of other stress-related hormones such as ABA, SA, ethylene, as well as genes involved in the response to drought, salt stress, and also local and systemic immune response.
The targets of the stress response in a plant cell can be genes encoding proteins with protective functions and enzymes responsible for the synthesis of antimicrobial and protective substances, as well as detoxification of toxic metabolites [55]. Accordingly, among genes which were downregulated in the spontaneous tumors, there were radish homologs of several genes encoding stress-related enzymes such as metacaspase AMC6 involved in the cell death and hypersensitivity response (HR) to pathogens [56], and GLUTATHIONE-S-TRANSFERASE GSTF11, which play a role in herbicide detoxification and responses to biotic and abiotic stress [57].
Examples of plant metabolites with an antimicrobial or barrier activity that are synthesized in response to stress are lignin, which provides physical and chemical protection for plants against pathogen invasion, and a number of substances that act as natural bacterio-, fungi-, and/or insecticides, such as glucosinolates.
Lignin is a heterogeneous polymer of monolignols, which is polymerized at the surface of the cell walls. It is extremely important for terrestrial plants: lignin provides structural support for the upward growth and also forms a physical barrier to block pathogen invasion, and prevents the ingress or diffusion of toxins from pathogens [58]. Monolignols, structural units of lignin polymers, are synthesized from phenylalanine via the phenylpropanoid pathway. Thus, enzymes of the phenylpropanoid biosynthetic pathway are critical for lignin biosynthesis [59]. In the spontaneous tumors in radish, we revealed the downregulation of radish homologs of genes encoding enzymes, which act at all stages of the phenylpropanoid pathway: genes encoding phenylalanine ammonia-lyases ATPAL1 and ATPAL2, CCR2 encoding cinnamoyl CoA reductase, gene encoding cinnamyl alcohol dehydrogenase ATCAD6, and also DIR5, which encodes the dirigent-like family protein involved in the synthesis of lignans from two molecules of coniferyl alcohol. In addition, among the downregulated, there was a radish homolog of NAC SECONDARY WALL THICKENING PROMOTING FACTOR 1 (NST1) encoding TF, which regulates secondary cell wall thickening via lignification [60].
Among plant antimicrobial substances, glucosinolates are specific for plants of the Brassicaceae family [61]. Glucosinolates have direct antimicrobial properties, in addition, upon damage to plants, e.g., by chewing insects, glucosinolates are enzymatically converted into a range of compounds with antimicrobe and insecticide activities, such as isothiocyanates [62]. In the tumor tissue, we revealed the downregulation of radish homologs of genes for glucosinolate biosynthesis enzymes: Cytochrome P450s CYP83A1 and SPS1/BUS1, methylthioalkylmalate synthases IMS3/MAM1 and IMS2/MAM3, isopropylmalate isomerase LEUD1, methionine-oxo-acid transaminase BCAT4, C-S lyase SUR1/ALF1, and 3-isopropylmalate dehydrogenase ATIMD1 [63]. Another downregulated gene, the radish homolog of MYB28 encoding TF involved in the positive regulation of aliphatic glucosinolate production [64], was presented. It is interesting that in Arabidopsis, genes SUR1/ALF1 (SUPERROOT/ABERRANT LATERAL ROOT FORMATION 1) and SPS1/BUS1 (SUPERSHOOT/BUSHY 1) encoding enzymes of glucosinolate biosynthesis, are also involved in IAA production and play a role in the development of root and shoot systems [65,66]. At the same time, the situation with a negative regulation of stress-related genes in the spontaneous tumor is not absolute: among upregulated genes, there were also several radish homologs of genes encoding enzymes of JA and SA biosynthesis, such as AZELAIC ACID INDUCED 1 (AZI1), which is involved in the priming of systemic immunity [67], and one of lipoxygenase genes, LOX2.
2.3.3. Genes Related with Cytokinins, Auxins, and Gibberellins
CK, IAA, and GA are plant hormones, which play major roles in the control of cell proliferation and expansion, as well as in the meristem development and organogenesis. It is well known that GA and CK exhibit antagonistic effects on various developmental and molecular processes during plant growth and can inhibit signaling of each other [68,69]. For instance, in the SAM and cambium, GA stimulates differentiation of specialized cell types, while CK stimulates stem cells proliferation [70,71]. In lateral root formation, however, both CK and GA inhibit this process influencing the polar transport of IAA [72,73]. IAA is a well-known positive regulator of lateral root formation, which is necessary for the expression of key “root genes” [74], and this process is under the antagonistic control of IAA and CK [75]. Among DEGs which were identified in the spontaneous tumors in radish, there were numerous genes involved in the biosynthesis of IAA, CK, and GA and the response to these hormones.
It is interesting that among upregulated genes in the tumor tissue, there were genes involved in the positive regulation of GA and negative regulation of CK content and signaling, while among downregulated genes, the situation was opposite. For instance, among downregulated genes, there were radish homologs of IPT5 and IPT7 genes encoding adenylate isopentenyltransferases, key enzymes of CK biosynthesis [76], and also genes encoding CK-activating enzymes of LONELY GUY family [77]—LOG1, LOG3-like, LOG4, and LOG8. Among the downregulated, there was also a radish homolog of ARR1 gene encoding key CK-responsive TF [78].
At the same time, homologs of genes encoding CK dehydrogenases, which are involved in the degradation of active CK [79]—CKX3 and CKX3-like, and also genes encoding repressors of CK signaling [80]—ARR4 and ARR5, were upregulated in the spontaneous tumors. The only downregulated gene encoding CK-inactivating enzyme was the radish homolog of CKX1.
The opposite situation was observed for GA: the radish homologs of KAO2 encoding ent-kaurenoic acid oxidase, enzyme of GA biosynthesis [81], and also gene GA-20 oxidase-3-like (probably involved in the production of active GA [82]), were strongly upregulated in the spontaneous tumors. At the same time, among downregulated genes, there was the GA2OX6 encoding protein of GA 2-oxidase family, which is a key enzyme of GA inactivation [83].
The IAA is the main positive regulator of lateral root development, and the genes involved in the biosynthesis, conjugation, transport, and signaling of IAA were downregulated in the tumor tissue compared to the lateral root. In the list of downregulated genes, there were regulators of polar IAA transport: radish homologs of genes encoding IAA efflux protein PIN7, which establishes an apical-basal axis in the embryo and is involved in the pattern specification during root development [84], as well as AUX1 and LAX3 IAA influx transporters [85]. Among genes involved in the IAA signaling and downregulated in tumors, there were radish homologs of AUXIN RESPONSE FACTOR 9 (ARF9), encoding IAA-regulated TF, and also IAA2 and IAA18 genes encoding transcriptional repressors of Aux/IAA family involved in root development [86,87]. Moreover, among downregulated genes in the spontaneous tumors, there were radish homologs of genes, connecting the IAA biosynthesis with the stress response: YUCCA8 encoding enzyme of IAA biosynthesis which acts in the JA-dependent regulation of IAA homeostasis [88], acyl acid amido synthetase gene GH3.5, whose product conjugates IAA with amino acids but also can conjugate SA to modulate both auxin and pathogen response pathways [89], and IAR1 encoding IAA-Alanin hydrolase, an enzyme which releases IAA from conjugates with amino acids [90].
2.3.4. Genes Involved in Photomorphogenesis and Flowering
Among DEGs in the spontaneous tumor, there were also several key regulators of circadian clock and photomorphogenesis, and their main targets—regulators of the development of inflorescence meristem and flower organs. Members of this system function in the leaves and SAM, and the identification of their homologs among DEGs between the lateral root and the tumor in the root, was rather unexpected.
First, among downregulated genes, there were radish homologs of CICRCADIAN CLOCK ASSOCIATED (CCA1) and LATE ELONGATED HYPOCOTYL (LHY): in Arabidopsis, the transcriptional repressor CCA1 forms a heterodimer with the homeodomain-like superfamily TF LHY and binds to the promoter of TIMING OF CAB 1 (TOC1) gene forming a regulatory feedback loop in the circadian oscillator [91]. The target of this regulatory loop is the CONSTANS (CO) gene encoding TF with a zinc finger domain [92]. The CO-like (COL) TF family includes about 20 proteins involved in the flowering time control in Arabidopsis and radish [93], and RsCOL5, a radish homolog of Arabidopsis CO-like 5 was downregulated in the spontaneous tumor. The main target of TF CO in the leaf is the FLOWERING LOCUS T (FT) gene, which encodes a short mobile peptide known as florigene [94]. In Arabidopsis and other plant species, there are many FT-like peptides each of them playing a specific role in the day length-dependent control of flowering (there are florigenes and antiflorigenes among them), and also in some other processes such as the development of adventitious buds, potato tubers, and bulbs [95]. Among downregulated genes in the spontaneous tumor, there was a radish homolog of BROTHER OF FT AND TFL1 (BFT) gene encoding FT-like protein, which functions as a repressor of flowering [96]. In the shoot apical meristem, FT-like proteins interact with FLOWERING LOCUS D (FD) family TFs, and the targets of this complex are inflorescence meristem identity genes such as the SUPRESSOR OF OVEREXPRESSION OF CONSTANS 1 (SOC1) and APETALA1 (AP1). In turn, the TFs SOC1 together with AGAMOUS-LIKE 24 (AGL24) positively regulate the expression of floral meristem identity gene LEAFY [97]. The radish homologs of key inflorescence meristem identity gene SOC1 and one of the indirect targets of SOC1 TF in the flower, PISTILLATA (PI) [97], were downregulated in the spontaneous tumors. Moreover, among genes downregulated in spontaneous tumors, there were homologs of RADIALIS-like3 and DIVARICATA genes, encoding TFs which regulate flower symmetry [98]. The expression of these genes in the radish root and their downregulation in the spontaneous tumor allow us to suppose the broader function of the flowering control system, e.g., in the regulation of root development.
2.4. Verification of Differential Expression Analysis Data with qPCR
Next, we performed the qPCR expression analysis for some genes that showed an increase or decrease in their expression levels in the spontaneous tumors of radish according to the transcriptome analysis. In total, we analyzed 12 upregulated and 12 downregulated genes by qPCR. To order to conduct qPCR, we selected genes involved in different pathways and associated with varied functions.
Among upregulated genes, we analyzed the expression of radish homologs of genes involved in the control of cell cycle (CYCA1;1 (XM_018592281.1), CYCB1;2 (XM_018617721.1), DEL1 (XM_018603289.1)) and cell growth (EXPA3 (XM_018581553.1)), biosynthesis of GA (GA20OX3 (XM_018633395.1)), negative regulation of cell response to cytokinin (ARR4 (XM_018624193.1) and ARR5 (XM_018581651.1)), genes encoding peptide phytohormone CLE46 (XM_018604748.1), TFs of different families (LBD25 (XM_018597567.1), LBD38 (XM_018581284.1), WRKY9 (XM_018604382.1)), and also the homolog of unique bifunctional gene ENO2/MBP1 (XM_018580867.1) encoding glycolytic enzyme enolase ENO2 and TF MBP1, which alternatively translated from the same transcript [99]—this gene was in the first place among those upregulated in spontaneous tumors.
Among genes which were downregulated in the tumor tissue, we took into analysis radish homologs of genes involved in the cell cycle regulation (DPB (XM_018587639.1)), genes encoding enzymes of biosynthesis of JA (AOC3 (XM_018598854.1)), IAA (YUC8 (XM_018594055.1)), CK (IPT5 (XM_018587590.1) and IPT7 (XM_018633594.1)), and glucosinolates (IMS3 (XM_018608445.1)), TF-encoding genes (TCP2 (XM_018608972.1), RL3 (XM_018614337.1), NAC090 (XM_018588875.1), SOC1 (XM_018623442.1), BEL-like 4 (XM_018603518.1)), and also the gene encoding FT-like protein BFT (XM_018581284.1).
In general, the obtained results of qPCR confirmed the transcriptome data: genes that have been identified as upregulated and downregulated in the RNA-seq experiment, demonstrated a significant increase or decrease of expression levels in the qPCR analysis, respectively (Figure 6).

Figure 6.
Gene expression analysis by qPCR: (a) Genes which were upregulated in the RNA-seq experiment; (b) genes which were downregulated in the RNA-seq experiment. Error bars indicate the standard deviation of three technical repeats (p-value < 0.01—**, p-value < 0.001—***).
3. Discussion
Tumors of higher plants are insufficiently studied, but represent a very interesting phenomenon, since they arise due to the release of a population of cells out of a very reliably arranged systemic control of the proliferation of plant cells. Spontaneous tumors in higher plants are especially rare, and the exact mechanisms of development of most of them remain unclear. In recent years, technologies of next-generation sequencing (NGS) have proven to be instrumental for studying various biological phenomena in different plants [100,101]. RNA sequencing (RNA-seq) is used to assess the functional state of biological tissue based on the variety and abundance of RNA molecules in the sample, so RNA-seq makes it possible to identify candidates for the role of regulators of plant tumor formation. To date, RNA-seq approaches were used for studying the gene expression during the development of only one type of plant tumor, the crown gall caused by A. tumefaciens [47,102]. In our work, we used RNA-seq to analyze the changes in gene expression in the spontaneous tumor of the radish inbred line compared with the lateral root of the same line of radish.
We obtained data demonstrating the significant difference between spontaneous tumors and lateral roots in gene expression profiles: both by pathways and by individual transcripts. The data that seemed most interesting to us were also verified by qPCR. In particular, in the spontaneous tumors, we observed a significant activation of genes associated with the control of cell division: among the pathways, the most enriched were those associated with the control of the cell cycle, DNA replication, cytokinesis, and cell growth. Among numerous upregulated genes involved in cell division, the genes controlling the processes of the G2-M transition and cytokinesis were most widely represented. Moreover, among them there were radish homologs of genes encoding the central participants in these processes in Arabidopsis, such as genes of CDKBs and its interacting partners, B-class cyclins, which provide the G2-M transition [103], as well as such important regulators of cytokinesis as centromere-associated histone H3 variant CEN3, which represents the assembly site for the kinetochore complex of active centromeres [36] or syntaxin KNOLLE, which is essential for phragmoplast formation [37]. The upregulation of genes involved in the control of the cell cycle and cytokinesis reflects the high frequency of cell divisions in spontaneous tumors (Figure 1). In addition, among the upregulated “cell division genes”, there were also inhibitors of CDKs, an increase in the expression of which is usually associated with an increase in the frequency of endomitoses, e.g., during the formation of one of the types of plant tumors—root galls induced by nematodes [43,44]. Indeed, in the spontaneous radish tumors, a high level of polyploid cells was noted compared to normal root tissues (Figure 1). Finally, in our early studies using qPCR, we revealed an increase in the expression levels of genes associated with cell division, including cell cycle genes and some of meristem regulators, in the spontaneous tumors of radish [5,104]. Thus, data on the upregulation of genes involved in the control of cell division in spontaneous tumors of radish are not unexpected for us.
At the same time, the pathways associated with the response to biotic and abiotic stresses were mostly downregulated in the spontaneous tumors. Among genes involved in these pathways, the most widely represented were genes controlling the biosynthesis of JA and the response to them, as well as genes that regulate the synthesis of specific secondary metabolites that are synthesized when plants are wounded or eaten by insects, for example, glucosinolates and lignin [105,106]. Thus, among downregulated genes, there were radish homologs of the Arabidopsis genes that regulate all stages of the JA biosynthesis from the beginning until the end [9], as well as genes that regulate the biosynthesis of glucosinolates [61] and common precursors of glucosinolates and IAA [65,66], and biosynthesis of lignin [58].
These data were also not very unexpected, since the intensification of cell division can lead to tissue juvenilization. The work of age-dependent resistance mechanisms leads to a gain or reinforcement of disease resistance in mature plant organs, whereas juvenile organs with a high level of cell divisions are more susceptible to pathogens [107]. In the adult plant, the activation of age-dependent resistance takes place under the control of genes involved in the regulation of flowering, such as LEAFY [107]. Therefore, the downregulation of genes involved in the flowering control which was revealed in the tumor tissue, may also contribute to the suppression of the age-dependent resistance in the tumor.
The results on changes in the expression of genes that regulate the level of CK and GA and the response to them were quite unexpected for us: based on the data obtained, the balance of these hormones in tumors should sharply shift towards GA. CK and GA are known to antagonistically interact in the control of meristem development, where CK promotes cell division, and GA is responsible for cell growth and differentiation in the periphery [68,69]. Therefore, one should rather expect the activation of CK biosynthesis and suppression of GA biosynthesis in the tumor. Moreover, our previous data indicate the sharp increase of free CK content in the spontaneous tumor of radish [6], and CK were shown to induce tumor formation in radish in vitro [4,108]. It is possible that the downregulation of “CK genes” and an upregulation of “GA genes” revealed by RNA-seq was the result of the work of hypothetical negative feedbacks that reduce the hyperactivation of CK biosynthesis in the radish tumor.
Finally, in the spontaneous tumors, the downregulation of genes that are involved in the control of photomorphogenesis and flowering was revealed. These data were rather unexpected and we can explain the expression of flowering-associated genes in the radish root only by suggesting the broader functions of these genes, e.g., in the radish storage root development. On the other hand, it was reported that individual circadian clock regulators can also participate in providing the plant stress response and affecting SA and JA signaling [109,110]. Therefore, these photomorphogenesis- and flowering-associated genes may be downregulated in the tumor together with other stress-related genes.
Thus, for the first time, data were obtained, which reflect changes in the expression of various gene families in spontaneous tumors in radish lines. A further study of the functions of revealed up- and downregulated genes will help elucidate the mechanisms of plant tumors development.
4. Materials and Methods
4.1. Plant Material
Line 19 from the genetic collection of R. sativus was used in this study. This line is approximately the 40th inbred generation and originated from Saxa cultivar (European group of radish varieties). Line 19 is characterized by a spontaneous tumor formation trait: at the flowering stage, nearly 100% of the plants of line 19 form tumors on the root and lower part of the stem [3].
For RNA sequencing of spontaneous tumors and lateral roots, radish plants of tumor-producing line 19 were grown in field conditions until the flowering stage (about 70 days). Plant material (lateral roots and spontaneous tumors) was harvested on the 15th day from the beginning of flowering, when the tumors became large and clearly visible. To harvest tumor and lateral root samples, plants were removed from the soil and rinsed with a large amount of water. Tumors and root samples were harvested by dissection with a scalpel and immediately preserved in the RNA-later solution (Sigma-Aldrich, Darmstadt, Germany) for further RNA extraction. For both lateral root and tumor samples, three biological replicates, each including material from three individual plants, were used.
4.2. RNA Isolation, Library Preparation, and Sequencing
Total RNA from radish plants and seedlings was isolated according to the modified phenol-chloroform method [111]. The cDNA was synthesized with the Mint-2 kit (Evrogen, Moscow, Russia). The NEBNext® Ultra™ DNA Library Prep Kit for Illumina (New England Biolabs, Hitchin, UK) was used to prepare sequencing libraries. Double barcoding was performed using the NEBNext® Ultra™ DNA Index Prep Kit for Illumina and the NEBNext® Multiplex Oligos® Illumina® (Dual Index Primers Set 1). Libraries were sequenced on the Illumina HiSeq2500 sequencer.
4.3. Bioinformatic Processing of Sequencing Results
Fastq reads were filtered with bbduk utility from the bbtools suite (v. 38.44) (URL: https://jgi.doe.gov/data-and-tools/bbtools/, accessed on 5 March 2020). To decontaminate reads, we used sequences of mitochondrial, plastid, and ribosomal DNA of radish. Quality filtered and decontaminated reads were additionally trimmed with cutadapt (v. 2.10) and Trimmomatic (v. 0.39) with “ILLUMINACLIP:{ADAPTERS}:2:30:10 LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36” options [112]. The quality control of reads was performed with fastqc (v. 0.11.8) (URL: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/, accessed on 20 March 2020). Filtered reads were quantified with kallisto (v. 0.46.1) [113] using RNA sequences from the representative genome assembly of R. sativus (NCBI assembly ID GCF_000801105.1). The differential expression between tumor and non-tumor samples was analyzed with the DESeq2 package (v. 1.26.0) [114] for R (v. 3.6.2). Genes with s-value < 0.05 and |logFoldChange| > 1 were considered differentially expressed. For functional annotation and subsequent enrichment analysis of transcripts, genes were compared with Arabidopsis thaliana cDNA sequences from the Araport v. 11 [115] base using blast+ (v. 2.6.0) [116]. The R packages ClusterProfiler v. 3.14.0 [117] and DOSE 3.12.0 [118] were applied to perform the GO, KEGG, and gene set enrichment analyses. Gene lists from plantGSEA [119] were used as a reference. FDR < 0.05 was selected as the cut-off value for the functional and pathway enrichment analysis of differentially expressed genes. The code for all the analysis steps of read data is available at https://github.com/castrofiber/radish2_RNAseq (accessed on 10 March 2021). SRA data can be downloaded from http://www.ncbi.nlm.nih.gov/bioproject/686790 (accessed on 1 March 2021).
4.4. The qPCR
The data on the differential expression obtained by RNA-seq were verified by the qPCR of selected genes. For qPCR, total RNA was extracted by the Purezol reagent (Bio-Rad, Hercules, CA, USA), purified with chloroform, and precipitated with isopropanol. The RNA pellet was washed three times with 80% ethanol, dried under air flow in a laminar box, and dissolved in sterile deionized water. The DNase treatment was done using the Rapid Out DNA Removal Kit (Thermo Fisher Scientific, Waltham, MA, USA). The RNA concentration was measured by the NanoDrop 2000 UV spectrophotometer (Thermo Fisher Scientific) at 260 nm. For reverse transcription, 500 ng of RNA were used in all the samples. The RNA reverse transcription was performed using the Revert Aid Reverse Transcriptase kit (Thermo Fisher Scientific). To check the DNase treatment efficacy, the qRT-PCR analysis of control samples without the reverse transcriptase was performed. The qRT-PCR experiments were done on a CFX-96 real-time PCR detection system with the C1000 thermal cycler (Bio-Rad), and Eva Green intercalating dye (Syntol, Moscow, Russia) was used for detection. Primers for qRT-PCR (Table S1) were designed to amplify 140–200 bp fragments and were synthesized by Evrogen. The specificity of PCR amplification was confirmed based on the melting curve (55–95 °C). All the reactions were performed in technical triplicate and averaged. Cycle threshold values were obtained with the accompanying CFX manager software, and data were analyzed by the 2−ΔΔCt method [120]. The relative expression was normalized against constitutively expressed R. sativus ubiquitin (RsUBQ11) and glyceraldehyde-3-phosphate dehydrogenase (RsGAPDH) genes [121]. Experiments were repeated three times with independent biological samples, whose results were then averaged.
Supplementary Materials
The following materials are available online at https://www.mdpi.com/article/10.3390/plants10050919/s1. Table S1: List of primers used for qPCR, Table S2: Results of differential expression analysis of tumor samples in comparison with lateral roots.
Author Contributions
Conceptualization, L.L. and I.D.; plant material, A.T. and K.K.; RNA isolation, A.T. and O.P.; bioinformatic processing of sequencing results, A.T., A.P., and O.P.; qPCR, I.D.; writing—original draft preparation, I.D.; writing—review and editing, L.L., V.T., and I.D.; visualization, K.K.; supervision, L.L. All authors have read and agreed to the published version of the manuscript. Please turn to the CRediT taxonomy for the term explanation. Authorship must be limited to those who have contributed substantially to the work reported.
Funding
The article was made with the support of the Ministry of Science and Higher Education of the Russian Federation in accordance with agreement number 075-15-2020-922, date 16 November 2020, on providing a grant in the form of subsidies from the Federal budget of Russian Federation. The grant was provided for state support for the creation and development of a World-Class Scientific Center “Agrotechnologies for the Future”.
Data Availability Statement
The scripts, programs, and pipeline used for the RNA-seq analysis are available at https://github.com/castrofiber/radish2_RNAseq (accessed on 22 April 2021). Raw reads are available at https://www.ncbi.nlm.nih.gov/bioproject/686790 (accessed on 22 April 2021).
Acknowledgments
The authors are grateful to Irina Buzovkina (Saint-Peresburg State University), Maria Lebedeva (Saint-Peresburg State University), Mikail Burlakovskiy (Saint-Peresburg State University), and Alexander Betekhtin (University of Silesia) for the provided photos illustrating the morphology, anatomy, and cytology of spontaneous radish tumors.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
ABA—abscisic acid, CK—cytokinin, DEG—differentially expressed gene, GA—gibberellic acid, IAA—indole-3-acetic acid, JA—jasmonic acid, SA—salycilic acid, TF—transcription factor.
References
- Dodueva, I.E.; Lebedeva, M.A.; Kuznetsova, K.A.; Gancheva, M.S.; Paponova, S.S.; Lutova, L.L. Plant Tumors: A Hundred Years of Study. Planta 2020, 251, 82. [Google Scholar] [CrossRef]
- Narbut, S.I. Genetic tumor in a radish obtained by inbreeding. Vestn. Leningr. Univ. Biol. 1967, 3, 144–149. [Google Scholar] [PubMed]
- Buzovkina, I.S.; Lutova, L.A. The Genetic Collection of Radish Inbred Lines: History and Prospects. Russ. J. Genet. 2007, 43, 1181–1192. [Google Scholar] [CrossRef]
- Ilina, E.; Dodueva, I.; Ivanova, N.; Lutova, L. The Effect of Cytokinins on in Vitro Cultured Inbred Lines of Raphanus sativus Var. Radicula Pers. with Genetically Determined Tumorigenesis. Russ. J. Plant Physiol. 2006, 53, 514–522. [Google Scholar] [CrossRef]
- Lebedeva Osipova, M.A.; Tvorogova, V.E.; Vinogradova, A.P.; Gancheva, M.S.; Azarakhsh, M.; Ilina, E.L.; Demchenko, K.N.; Dodueva, I.E.; Lutova, L.A. Initiation of Spontaneous Tumors in Radish (Raphanus sativus): Cellular, Molecular and Physiological Events. J. Plant Physiol. 2015, 173, 97–104. [Google Scholar] [CrossRef] [PubMed]
- Matveeva, T.V.; Frolova, N.V.; Smets, R.; Dodueva, I.E.; Buzovkina, I.S.; Van Onckelen, H.; Lutova, L.A. Hormonal Control of Tumor Formation in Radish. J. Plant Growth Regul. 2004, 23, 37–43. [Google Scholar] [CrossRef]
- Betekhtin, A.; Siwińska, D.; Hasterok, R.; Dodueva, I.; Ilina, E.; Lutova, L. Cytological and Histological Analysis of Radish Root Tumors. In Proceedings of the VII Meeting of Russian Society of Plant Physiologists, International Conference “Plant Physiology—Fundamentals of Ecology and Biotechnology Innovation”, Nizhny Novgorod, Russia, 4–10 July 2011. [Google Scholar]
- Banerjee, G.; Singh, D.; Sinha, A.K. Plant Cell Cycle Regulators: Mitogen-Activated Protein Kinase, a New Regulating Switch? Plant Sci. 2020, 301, 110660. [Google Scholar] [CrossRef]
- Wasternack, C.; Strnad, M. Jasmonates Are Signals in the Biosynthesis of Secondary Metabolites—Pathways, Transcription Factors and Applied Aspects—A Brief Review. New Biotechnol. 2019, 48, 1–11. [Google Scholar] [CrossRef]
- Ahuja, M.R. Genetic Tumors in Nicotiana and Other Plants. Q. Rev. Biol. 1998, 73, 439–462. [Google Scholar] [CrossRef]
- Cosgrove, D.J. Catalysts of Plant Cell Wall Loosening. F1000Research 2016, 5. [Google Scholar] [CrossRef]
- Krupková, E.; Immerzeel, P.; Pauly, M.; Schmülling, T. The TUMOROUS SHOOT DEVELOPMENT2 Gene of Arabidopsis Encoding a Putative Methyltransferase Is Required for Cell Adhesion and Co-Ordinated Plant Development. Plant J. Cell Mol. Biol. 2007, 50, 735–750. [Google Scholar] [CrossRef] [PubMed]
- Sorrell, D.A.; Menges, M.; Healy, J.M.S.; Deveaux, Y.; Amano, C.; Su, Y.; Nakagami, H.; Shinmyo, A.; Doonan, J.H.; Sekine, M.; et al. Cell Cycle Regulation of Cyclin-Dependent Kinases in Tobacco Cultivar Bright Yellow-2 Cells. Plant Physiol. 2001, 126, 1214–1223. [Google Scholar] [CrossRef]
- Dewitte, W.; Riou-Khamlichi, C.; Scofield, S.; Healy, J.M.S.; Jacqmard, A.; Kilby, N.J.; Murray, J.A.H. Altered Cell Cycle Distribution, Hyperplasia, and Inhibited Differentiation in Arabidopsis Caused by the D-Type Cyclin CYCD3. Plant Cell 2003, 15, 79–92. [Google Scholar] [CrossRef]
- De Jager, S.M.; Maughan, S.; Dewitte, W.; Scofield, S.; Murray, J.A.H. The Developmental Context of Cell-Cycle Control in Plants. Semin. Cell Dev. Biol. 2005, 16, 385–396. [Google Scholar] [CrossRef]
- Kosugi, S.; Ohashi, Y. Interaction of the Arabidopsis E2F and DP Proteins Confers Their Concomitant Nuclear Translocation and Transactivation. Plant Physiol. 2002, 128, 833–843. [Google Scholar] [CrossRef]
- Kosugi, S.; Ohashi, Y. E2Ls, E2F-like Repressors of Arabidopsis That Bind to E2F Sites in a Monomeric Form. J. Biol. Chem. 2002, 277, 16553–16558. [Google Scholar] [CrossRef]
- Takahashi, I.; Kojima, S.; Sakaguchi, N.; Umeda-Hara, C.; Umeda, M. Two Arabidopsis Cyclin A3s Possess G1 Cyclin-like Features. Plant Cell Rep. 2010, 29, 307–315. [Google Scholar] [CrossRef]
- Springer, P.S.; Holding, D.R.; Groover, A.; Yordan, C.; Martienssen, R.A. The Essential Mcm7 Protein PROLIFERA Is Localized to the Nucleus of Dividing Cells during the G(1) Phase and Is Required Maternally for Early Arabidopsis Development. Dev. Camb. Engl. 2000, 127, 1815–1822. [Google Scholar]
- Strzalka, W.; Aggarwal, C. Arabidopsis Thaliana: Proliferating Cell Nuclear Antigen 1 and 2 Possibly Form Homo- and Hetero-Trimeric Complexes in the Plant Cell. Plant Signal. Behav. 2013, 8, e24837. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wang, C.; Liu, Z. Arabidopsis Ribonucleotide Reductases Are Critical for Cell Cycle Progression, DNA Damage Repair, and Plant Development. Plant Cell 2006, 18, 350–365. [Google Scholar] [CrossRef]
- Wu, L.; Liu, Y.; Kong, D. Mechanism of Chromosomal DNA Replication Initiation and Replication Fork Stabilization in Eukaryotes. Sci. China Life Sci. 2014, 57, 482–487. [Google Scholar] [CrossRef] [PubMed]
- Osman, K.; Sanchez-Moran, E.; Mann, S.C.; Jones, G.H.; Franklin, F.C.H. Replication Protein A (AtRPA1a) Is Required for Class I Crossover Formation but Is Dispensable for Meiotic DNA Break Repair. EMBO J. 2009, 28, 394–404. [Google Scholar] [CrossRef]
- De Veylder, L.; Joubès, J.; Inzé, D. Plant Cell Cycle Transitions. Curr. Opin. Plant Biol. 2003, 6, 536–543. [Google Scholar] [CrossRef]
- Ito, M. Conservation and Diversification of Three-Repeat Myb Transcription Factors in Plants. J. Plant Res. 2005, 118, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, M.; Kato, H.; Yoshida, S.; Yamamura, S.; Uchimiya, H.; Umeda, M. Control of in Vitro Organogenesis by Cyclin-Dependent Kinase Activities in Plants. Proc. Natl. Acad. Sci. USA 2003, 100, 8019–8023. [Google Scholar] [CrossRef]
- Torres Acosta, J.A.; de Almeida Engler, J.; Raes, J.; Magyar, Z.; De Groodt, R.; Inzé, D.; De Veylder, L. Molecular Characterization of Arabidopsis PHO80-like Proteins, a Novel Class of CDKA;1-Interacting Cyclins. Cell. Mol. Life Sci. CMLS 2004, 61, 1485–1497. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Ndathe, R.W.; Kumar, N.; Zeringue, E.A.; Kato, N.; Larkin, J.C. The CDK Inhibitor SIAMESE Targets Both CDKA;1 and CDKB1 Complexes to Establish Endoreplication in Trichomes. Plant Physiol. 2020, 184, 165–175. [Google Scholar] [CrossRef] [PubMed]
- Wen, B.; Nieuwland, J.; Murray, J.A.H. The Arabidopsis CDK Inhibitor ICK3/KRP5 Is Rate Limiting for Primary Root Growth and Promotes Growth through Cell Elongation and Endoreduplication. J. Exp. Bot. 2013, 64, 1135–1144. [Google Scholar] [CrossRef] [PubMed]
- Gentric, N.; Masoud, K.; Journot, R.P.; Cognat, V.; Chabouté, M.-E.; Noir, S.; Genschik, P. The F-Box-Like Protein FBL17 Is a Regulator of DNA-Damage Response and Colocalizes with RETINOBLASTOMA RELATED1 at DNA Lesion Sites. Plant Physiol. 2020, 183, 1295–1305. [Google Scholar] [CrossRef]
- Sorrell, D.A.; Marchbank, A.; McMahon, K.; Dickinson, J.R.; Rogers, H.J.; Francis, D. A WEE1 Homologue from Arabidopsis Thaliana. Planta 2002, 215, 518–522. [Google Scholar] [CrossRef]
- Walker, K.L.; Müller, S.; Moss, D.; Ehrhardt, D.W.; Smith, L.G. Arabidopsis TANGLED Identifies the Division Plane throughout Mitosis and Cytokinesis. Curr. Biol. CB 2007, 17, 1827–1836. [Google Scholar] [CrossRef] [PubMed]
- Steiner, A.; Rybak, K.; Altmann, M.; McFarlane, H.E.; Klaeger, S.; Nguyen, N.; Facher, E.; Ivakov, A.; Wanner, G.; Kuster, B.; et al. Cell Cycle-Regulated PLEIADE/AtMAP65-3 Links Membrane and Microtubule Dynamics during Plant Cytokinesis. Plant J. Cell Mol. Biol. 2016, 88, 531–541. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Deng, X.; Sun, B.; Lee Van, S.; Kang, Z.; Lin, H.; Lee, Y.-R.J.; Liu, B. Role of the BUB3 Protein in Phragmoplast Microtubule Reorganization during Cytokinesis. Nat. Plants 2018, 4, 485–494. [Google Scholar] [CrossRef] [PubMed]
- Vos, J.W.; Pieuchot, L.; Evrard, J.-L.; Janski, N.; Bergdoll, M.; de Ronde, D.; Perez, L.H.; Sardon, T.; Vernos, I.; Schmit, A.-C. The Plant TPX2 Protein Regulates Prospindle Assembly before Nuclear Envelope Breakdown. Plant Cell 2008, 20, 2783–2797. [Google Scholar] [CrossRef] [PubMed]
- Lermontova, I.; Rutten, T.; Schubert, I. Deposition, Turnover, and Release of CENH3 at Arabidopsis Centromeres. Chromosoma 2011, 120, 633–640. [Google Scholar] [CrossRef] [PubMed]
- Touihri, S.; Knöll, C.; Stierhof, Y.-D.; Müller, I.; Mayer, U.; Jürgens, G. Functional Anatomy of the Arabidopsis Cytokinesis-Specific Syntaxin KNOLLE. Plant J. Cell Mol. Biol. 2011, 68, 755–764. [Google Scholar] [CrossRef] [PubMed]
- Krupnova, T.; Sasabe, M.; Ghebreghiorghis, L.; Gruber, C.W.; Hamada, T.; Dehmel, V.; Strompen, G.; Stierhof, Y.-D.; Lukowitz, W.; Kemmerling, B.; et al. Microtubule-Associated Kinase-like Protein RUNKEL Needed [Corrected] for Cell Plate Expansion in Arabidopsis Cytokinesis. Curr. Biol. CB 2009, 19, 518–523. [Google Scholar] [CrossRef] [PubMed]
- Sasabe, M.; Ishibashi, N.; Haruta, T.; Minami, A.; Kurihara, D.; Higashiyama, T.; Nishihama, R.; Ito, M.; Machida, Y. The Carboxyl-Terminal Tail of the Stalk of Arabidopsis NACK1/HINKEL Kinesin Is Required for Its Localization to the Cell Plate Formation Site. J. Plant Res. 2015, 128, 327–336. [Google Scholar] [CrossRef] [PubMed]
- Gu, F.; Bringmann, M.; Combs, J.R.; Yang, J.; Bergmann, D.C.; Nielsen, E. Arabidopsis CSLD5 Functions in Cell Plate Formation in a Cell Cycle-Dependent Manner. Plant Cell 2016, 28, 1722–1737. [Google Scholar] [CrossRef] [PubMed]
- Andersen, S.U.; Buechel, S.; Zhao, Z.; Ljung, K.; Novák, O.; Busch, W.; Schuster, C.; Lohmann, J.U. Requirement of B2-Type Cyclin-Dependent Kinases for Meristem Integrity in Arabidopsis Thaliana. Plant Cell 2008, 20, 88–100. [Google Scholar] [CrossRef] [PubMed]
- Spadafora, N.; Perrotta, L.; Nieuwland, J.; Albani, D.; Bitonti, M.B.; Herbert, R.J.; Doonan, J.H.; Marchbank, A.M.; Siciliano, I.; Lentz Grønlund, A.; et al. Gene Dosage Effect of WEE1 on Growth and Morphogenesis from Arabidopsis Hypocotyl Explants. Ann. Bot. 2012, 110, 1631–1639. [Google Scholar] [CrossRef] [PubMed]
- De Almeida Engler, J.; Gheysen, G. Nematode-Induced Endoreduplication in Plant Host Cells: Why and How? Mol. Plant Microbe Interact. 2012, 26, 17–24. [Google Scholar] [CrossRef]
- Cabral, D.; Banora, M.Y.; Antonino, J.D.; Rodiuc, N.; Vieira, P.; Coelho, R.R.; Chevalier, C.; Eekhout, T.; Engler, G.; Veylder, L.D.; et al. The Plant WEE1 Kinase Is Involved in Checkpoint Control Activation in Nematode-Induced Galls. New Phytol. 2020, 225, 430–447. [Google Scholar] [CrossRef] [PubMed]
- Coelho, R.R.; Vieira, P.; Antonino de Souza Júnior, J.D.; Martin-Jimenez, C.; De Veylder, L.; Cazareth, J.; Engler, G.; Grossi-de-Sa, M.F.; de Almeida Engler, J. Exploiting Cell Cycle Inhibitor Genes of the KRP Family to Control Root-Knot Nematode Induced Feeding Sites in Plants. Plant Cell Environ. 2017, 40, 1174–1188. [Google Scholar] [CrossRef] [PubMed]
- Wieczorek, K.; Golecki, B.; Gerdes, L.; Heinen, P.; Szakasits, D.; Durachko, D.M.; Cosgrove, D.J.; Kreil, D.P.; Puzio, P.S.; Bohlmann, H.; et al. Expansins Are Involved in the Formation of Nematode-Induced Syncytia in Roots of Arabidopsis Thaliana. Plant J. Cell Mol. Biol. 2006, 48, 98–112. [Google Scholar] [CrossRef]
- Tkachenko, A.A.; Gancheva, M.S.; Tvorogova, V.E.; Danilov, L.G.; Predeus, A.V.; Dodueva, I.E.; Lutova, L.L. Transcriptome Analysis of Crown Gall in Radish (Raphanus sativus L.) Inbred Lines. Ann. Appl. Biol. 2021, 178, 527–548. [Google Scholar] [CrossRef]
- Ishiguro, S.; Kawai-Oda, A.; Ueda, J.; Nishida, I.; Okada, K. The DEFECTIVE IN ANTHER DEHISCIENCE Gene Encodes a Novel Phospholipase A1 Catalyzing the Initial Step of Jasmonic Acid Biosynthesis, Which Synchronizes Pollen Maturation, Anther Dehiscence, and Flower Opening in Arabidopsis. Plant Cell 2001, 13, 2191–2209. [Google Scholar] [CrossRef]
- Christensen, S.A.; Huffaker, A.; Kaplan, F.; Sims, J.; Ziemann, S.; Doehlemann, G.; Ji, L.; Schmitz, R.J.; Kolomiets, M.V.; Alborn, H.T.; et al. Maize Death Acids, 9-Lipoxygenase–Derived Cyclopente(a)Nones, Display Activity as Cytotoxic Phytoalexins and Transcriptional Mediators. Proc. Natl. Acad. Sci. USA 2015, 112, 11407–11412. [Google Scholar] [CrossRef] [PubMed]
- Koo, A.J.K.; Chung, H.S.; Kobayashi, Y.; Howe, G.A. Identification of a Peroxisomal Acyl-Activating Enzyme Involved in the Biosynthesis of Jasmonic Acid in Arabidopsis. J. Biol. Chem. 2006, 281, 33511–33520. [Google Scholar] [CrossRef]
- Chini, A.; Fonseca, S.; Fernández, G.; Adie, B.; Chico, J.M.; Lorenzo, O.; García-Casado, G.; López-Vidriero, I.; Lozano, F.M.; Ponce, M.R.; et al. The JAZ Family of Repressors Is the Missing Link in Jasmonate Signalling. Nature 2007, 448, 666–671. [Google Scholar] [CrossRef]
- Kitaoka, N.; Matsubara, T.; Sato, M.; Takahashi, K.; Wakuta, S.; Kawaide, H.; Matsui, H.; Nabeta, K.; Matsuura, H. Arabidopsis CYP94B3 Encodes Jasmonyl-L-Isoleucine 12-Hydroxylase, a Key Enzyme in the Oxidative Catabolism of Jasmonate. Plant Cell Physiol. 2011, 52, 1757–1765. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.-Y.; Hsieh, E.-J.; Cheng, M.-C.; Chen, C.-Y.; Hwang, S.-Y.; Lin, T.-P. ORA47 (Octadecanoid-Responsive AP2/ERF-Domain Transcription Factor 47) Regulates Jasmonic Acid and Abscisic Acid Biosynthesis and Signaling through Binding to a Novel Cis-Element. New Phytol. 2016, 211, 599–613. [Google Scholar] [CrossRef]
- Pozo, M.J.; Van Der Ent, S.; Van Loon, L.C.; Pieterse, C.M.J. Transcription Factor MYC2 Is Involved in Priming for Enhanced Defense during Rhizobacteria-Induced Systemic Resistance in Arabidopsis Thaliana. New Phytol. 2008, 180, 511–523. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, N.; Rivero, R.M.; Shulaev, V.; Blumwald, E.; Mittler, R. Abiotic and Biotic Stress Combinations. New Phytol. 2014, 203, 32–43. [Google Scholar] [CrossRef]
- Watanabe, N.; Lam, E. Two Arabidopsis Metacaspases AtMCP1b and AtMCP2b Are Arginine/Lysine-Specific Cysteine Proteases and Activate Apoptosis-like Cell Death in Yeast. J. Biol. Chem. 2005, 280, 14691–14699. [Google Scholar] [CrossRef]
- Wagner, U.; Edwards, R.; Dixon, D.P.; Mauch, F. Probing the Diversity of the Arabidopsis Glutathione S-Transferase Gene Family. Plant Mol. Biol. 2002, 49, 515–532. [Google Scholar] [CrossRef]
- Sattler, S.E.; Funnell-Harris, D.L. Modifying Lignin to Improve Bioenergy Feedstocks: Strengthening the Barrier against Pathogens? Front. Plant Sci. 2013, 4, 70. [Google Scholar] [CrossRef] [PubMed]
- Xie, M.; Zhang, J.; Tschaplinski, T.J.; Tuskan, G.A.; Chen, J.-G.; Muchero, W. Regulation of Lignin Biosynthesis and Its Role in Growth-Defense Tradeoffs. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef]
- Mitsuda, N.; Ohme-Takagi, M. NAC Transcription Factors NST1 and NST3 Regulate Pod Shattering in a Partially Redundant Manner by Promoting Secondary Wall Formation after the Establishment of Tissue Identity. Plant J. Cell Mol. Biol. 2008, 56, 768–778. [Google Scholar] [CrossRef]
- Blažević, I.; Montaut, S.; Burčul, F.; Olsen, C.E.; Burow, M.; Rollin, P.; Agerbirk, N. Glucosinolate Structural Diversity, Identification, Chemical Synthesis and Metabolism in Plants. Phytochemistry 2020, 169, 112100. [Google Scholar] [CrossRef]
- Winde, I.; Wittstock, U. Insect Herbivore Counteradaptations to the Plant Glucosinolate–Myrosinase System. Phytochemistry 2011, 72, 1566–1575. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Mawhinney, T.P.; Preuss, M.L.; Schroeder, A.C.; Chen, B.; Abraham, L.; Jez, J.M.; Chen, S. A Redox-Active Isopropylmalate Dehydrogenase Functions in the Biosynthesis of Glucosinolates and Leucine in Arabidopsis. Plant J. Cell Mol. Biol. 2009, 60, 679–690. [Google Scholar] [CrossRef]
- Gigolashvili, T.; Yatusevich, R.; Berger, B.; Müller, C.; Flügge, U.-I. The R2R3-MYB Transcription Factor HAG1/MYB28 Is a Regulator of Methionine-Derived Glucosinolate Biosynthesis in Arabidopsis Thaliana. Plant J. Cell Mol. Biol. 2007, 51, 247–261. [Google Scholar] [CrossRef] [PubMed]
- Reintanz, B.; Lehnen, M.; Reichelt, M.; Gershenzon, J.; Kowalczyk, M.; Sandberg, G.; Godde, M.; Uhl, R.; Palme, K. Bus, a Bushy Arabidopsis CYP79F1 Knockout Mutant with Abolished Synthesis of Short-Chain Aliphatic Glucosinolates. Plant Cell 2001, 13, 351–367. [Google Scholar] [CrossRef]
- Mikkelsen, M.D.; Naur, P.; Halkier, B.A. Arabidopsis Mutants in the C-S Lyase of Glucosinolate Biosynthesis Establish a Critical Role for Indole-3-Acetaldoxime in Auxin Homeostasis. Plant J. Cell Mol. Biol. 2004, 37, 770–777. [Google Scholar] [CrossRef]
- Wang, X.-Y.; Li, D.-Z.; Li, Q.; Ma, Y.-Q.; Yao, J.-W.; Huang, X.; Xu, Z.-Q. Metabolomic Analysis Reveals the Relationship between AZI1 and Sugar Signaling in Systemic Acquired Resistance of Arabidopsis. Plant Physiol. Biochem. PPB 2016, 107, 273–287. [Google Scholar] [CrossRef] [PubMed]
- Greenboim-Wainberg, Y.; Maymon, I.; Borochov, R.; Alvarez, J.; Olszewski, N.; Ori, N.; Eshed, Y.; Weiss, D. Cross Talk between Gibberellin and Cytokinin: The Arabidopsis GA Response Inhibitor SPINDLY Plays a Positive Role in Cytokinin Signaling. Plant Cell 2005, 17, 92–102. [Google Scholar] [CrossRef]
- Fleishon, S.; Shani, E.; Ori, N.; Weiss, D. Negative Reciprocal Interactions between Gibberellin and Cytokinin in Tomato. New Phytol. 2011, 190, 609–617. [Google Scholar] [CrossRef] [PubMed]
- Jasinski, S.; Piazza, P.; Craft, J.; Hay, A.; Woolley, L.; Rieu, I.; Phillips, A.; Hedden, P.; Tsiantis, M. KNOX Action in Arabidopsis Is Mediated by Coordinate Regulation of Cytokinin and Gibberellin Activities. Curr. Biol. CB 2005, 15, 1560–1565. [Google Scholar] [CrossRef]
- Oles, V.; Panchenko, A.; Smertenko, A. Modeling Hormonal Control of Cambium Proliferation. PLoS ONE 2017, 12, e0171927. [Google Scholar] [CrossRef]
- Bielach, A.; Podlesáková, K.; Marhavy, P.; Duclercq, J.; Cuesta, C.; Müller, B.; Grunewald, W.; Tarkowski, P.; Benková, E. Spatiotemporal Regulation of Lateral Root Organogenesis in Arabidopsis by Cytokinin. Plant Cell 2012, 24, 3967–3981. [Google Scholar] [CrossRef] [PubMed]
- Gou, J.; Strauss, S.H.; Tsai, C.J.; Fang, K.; Chen, Y.; Jiang, X.; Busov, V.B. Gibberellins Regulate Lateral Root Formation in Populus through Interactions with Auxin and Other Hormones. Plant Cell 2010, 22, 623–639. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Scheres, B. Lateral Root Formation and the Multiple Roles of Auxin. J. Exp. Bot. 2018, 69, 155–167. [Google Scholar] [CrossRef]
- Jing, H.; Strader, L.C. Interplay of Auxin and Cytokinin in Lateral Root Development. Int. J. Mol. Sci. 2019, 20, 486. [Google Scholar] [CrossRef]
- Miyawaki, K.; Tarkowski, P.; Matsumoto-Kitano, M.; Kato, T.; Sato, S.; Tarkowska, D.; Tabata, S.; Sandberg, G.; Kakimoto, T. Roles of Arabidopsis ATP/ADP Isopentenyltransferases and TRNA Isopentenyltransferases in Cytokinin Biosynthesis. Proc. Natl. Acad. Sci. USA 2006, 103, 16598–16603. [Google Scholar] [CrossRef]
- Kuroha, T.; Tokunaga, H.; Kojima, M.; Ueda, N.; Ishida, T.; Nagawa, S.; Fukuda, H.; Sugimoto, K.; Sakakibara, H. Functional Analyses of LONELY GUY Cytokinin-Activating Enzymes Reveal the Importance of the Direct Activation Pathway in Arabidopsis. Plant Cell 2009, 21, 3152–3169. [Google Scholar] [CrossRef]
- Hill, K.; Mathews, D.E.; Kim, H.J.; Street, I.H.; Wildes, S.L.; Chiang, Y.-H.; Mason, M.G.; Alonso, J.M.; Ecker, J.R.; Kieber, J.J.; et al. Functional Characterization of Type-B Response Regulators in the Arabidopsis Cytokinin Response. Plant Physiol. 2013, 162, 212–224. [Google Scholar] [CrossRef]
- Avalbaev, A.M.; Somov, K.A.; Yuldashev, R.A.; Shakirova, F.M. Cytokinin Oxidase Is Key Enzyme of Cytokinin Degradation. Biochem. Mosc. 2012, 77, 1354–1361. [Google Scholar] [CrossRef]
- To, J.P.C.; Haberer, G.; Ferreira, F.J.; Deruère, J.; Mason, M.G.; Schaller, G.E.; Alonso, J.M.; Ecker, J.R.; Kieber, J.J. Type-A Arabidopsis Response Regulators Are Partially Redundant Negative Regulators of Cytokinin Signaling. Plant Cell 2004, 16, 658–671. [Google Scholar] [CrossRef]
- Regnault, T.; Davière, J.-M.; Heintz, D.; Lange, T.; Achard, P. The Gibberellin Biosynthetic Genes AtKAO1 and AtKAO2 Have Overlapping Roles throughout Arabidopsis Development. Plant J. Cell Mol. Biol. 2014, 80, 462–474. [Google Scholar] [CrossRef] [PubMed]
- Plackett, A.R.G.; Powers, S.J.; Fernandez-Garcia, N.; Urbanova, T.; Takebayashi, Y.; Seo, M.; Jikumaru, Y.; Benlloch, R.; Nilsson, O.; Ruiz-Rivero, O.; et al. Analysis of the Developmental Roles of the Arabidopsis Gibberellin 20-Oxidases Demonstrates That GA20ox1, -2, and -3 Are the Dominant Paralogs. Plant Cell 2012, 24, 941–960. [Google Scholar] [CrossRef] [PubMed]
- Rieu, I.; Eriksson, S.; Powers, S.J.; Gong, F.; Griffiths, J.; Woolley, L.; Benlloch, R.; Nilsson, O.; Thomas, S.G.; Hedden, P.; et al. Genetic Analysis Reveals That C19-GA 2-Oxidation Is a Major Gibberellin Inactivation Pathway in Arabidopsis. Plant Cell 2008, 20, 2420–2436. [Google Scholar] [CrossRef]
- Jang, G.; Yoon, Y.; Choi, Y.D. Jasmonic Acid Modulates Xylem Development by Controlling Expression of PIN-FORMED 7. Plant Signal. Behav. 2019, 14. [Google Scholar] [CrossRef] [PubMed]
- Péret, B.; Swarup, K.; Ferguson, A.; Seth, M.; Yang, Y.; Dhondt, S.; James, N.; Casimiro, I.; Perry, P.; Syed, A.; et al. AUX/LAX Genes Encode a Family of Auxin Influx Transporters That Perform Distinct Functions during Arabidopsis Development. Plant Cell 2012, 24, 2874–2885. [Google Scholar] [CrossRef]
- Taniguchi, M.; Nakamura, M.; Tasaka, M.; Morita, M.T. Identification of Gravitropic Response Indicator Genes in Arabidopsis Inflorescence Stems. Plant Signal. Behav. 2014, 9, e29570. [Google Scholar] [CrossRef] [PubMed]
- Uehara, T.; Okushima, Y.; Mimura, T.; Tasaka, M.; Fukaki, H. Domain II Mutations in CRANE/IAA18 Suppress Lateral Root Formation and Affect Shoot Development in Arabidopsis Thaliana. Plant Cell Physiol. 2008, 49, 1025–1038. [Google Scholar] [CrossRef]
- Hentrich, M.; Böttcher, C.; Düchting, P.; Cheng, Y.; Zhao, Y.; Berkowitz, O.; Masle, J.; Medina, J.; Pollmann, S. The Jasmonic Acid Signaling Pathway Is Linked to Auxin Homeostasis through the Modulation of YUCCA8 and YUCCA9 Gene Expression. Plant J. Cell Mol. Biol. 2013, 74, 626–637. [Google Scholar] [CrossRef] [PubMed]
- Westfall, C.S.; Sherp, A.M.; Zubieta, C.; Alvarez, S.; Schraft, E.; Marcellin, R.; Ramirez, L.; Jez, J.M. Arabidopsis Thaliana GH3.5 Acyl Acid Amido Synthetase Mediates Metabolic Crosstalk in Auxin and Salicylic Acid Homeostasis. Proc. Natl. Acad. Sci. USA 2016, 113, 13917–13922. [Google Scholar] [CrossRef]
- Widemann, E.; Miesch, L.; Lugan, R.; Holder, E.; Heinrich, C.; Aubert, Y.; Miesch, M.; Pinot, F.; Heitz, T. The Amidohydrolases IAR3 and ILL6 Contribute to Jasmonoyl-Isoleucine Hormone Turnover and Generate 12-Hydroxyjasmonic Acid upon Wounding in Arabidopsis Leaves. J. Biol. Chem. 2013, 288, 31701–31714. [Google Scholar] [CrossRef]
- Lu, S.X.; Knowles, S.M.; Andronis, C.; Ong, M.S.; Tobin, E.M. CIRCADIAN CLOCK ASSOCIATED1 and LATE ELONGATED HYPOCOTYL Function Synergistically in the Circadian Clock of Arabidopsis. Plant Physiol. 2009, 150, 834–843. [Google Scholar] [CrossRef]
- Nakamichi, N.; Kita, M.; Niinuma, K.; Ito, S.; Yamashino, T.; Mizoguchi, T.; Mizuno, T. Arabidopsis Clock-Associated Pseudo-Response Regulators PRR9, PRR7 and PRR5 Coordinately and Positively Regulate Flowering Time through the Canonical CONSTANS-Dependent Photoperiodic Pathway. Plant Cell Physiol. 2007, 48, 822–832. [Google Scholar] [CrossRef]
- Hu, T.; Wei, Q.; Wang, W.; Hu, H.; Mao, W.; Zhu, Q.; Bao, C. Genome-Wide Identification and Characterization of CONSTANS-like Gene Family in Radish (Raphanus Sativus). PLoS ONE 2018, 13, e0204137. [Google Scholar] [CrossRef]
- Luo, X.; Gao, Z.; Wang, Y.; Chen, Z.; Zhang, W.; Huang, J.; Yu, H.; He, Y. The NUCLEAR FACTOR-CONSTANS Complex Antagonizes Polycomb Repression to de-Repress FLOWERING LOCUS T Expression in Response to Inductive Long Days in Arabidopsis. Plant J. Cell Mol. Biol. 2018, 95, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Lebedeva, M.A.; Dodueva, I.E.; Gancheva, M.S.; Tvorogova, V.E.; Kuznetsova, K.A.; Lutova, L.A. The Evolutionary Aspects of Flowering Control: Florigens and Anti-Florigens. Russ. J. Genet. 2020, 56, 1323–1344. [Google Scholar] [CrossRef]
- Yoo, S.J.; Chung, K.S.; Jung, S.H.; Yoo, S.Y.; Lee, J.S.; Ahn, J.H. BROTHER OF FT AND TFL1 (BFT) Has TFL1-like Activity and Functions Redundantly with TFL1 in Inflorescence Meristem Development in Arabidopsis. Plant J. Cell Mol. Biol. 2010, 63, 241–253. [Google Scholar] [CrossRef]
- Lee, J.; Lee, I. Regulation and Function of SOC1, a Flowering Pathway Integrator. J. Exp. Bot. 2010, 61, 2247–2254. [Google Scholar] [CrossRef]
- Corley, S.B.; Carpenter, R.; Copsey, L.; Coen, E. Floral Asymmetry Involves an Interplay between TCP and MYB Transcription Factors in Antirrhinum. Proc. Natl. Acad. Sci. USA 2005, 102, 5068–5073. [Google Scholar] [CrossRef] [PubMed]
- Eremina, M.; Rozhon, W.; Yang, S.; Poppenberger, B. ENO2 Activity Is Required for the Development and Reproductive Success of Plants, and Is Feedback-Repressed by AtMBP-1. Plant J. 2015, 81, 895–906. [Google Scholar] [CrossRef] [PubMed]
- Hearn, D.J.; O’Brien, P.; Poulsen, T.M. Comparative Transcriptomics Reveals Shared Gene Expression Changes during Independent Evolutionary Origins of Stem and Hypocotyl/Root Tubers in Brassica (Brassicaceae). PLoS ONE 2018, 13, e0197166. [Google Scholar] [CrossRef]
- McNeil, M.D.; Bhuiyan, S.A.; Berkman, P.J.; Croft, B.J.; Aitken, K.S. Analysis of the Resistance Mechanisms in Sugarcane during Sporisorium Scitamineum Infection Using RNA-Seq and Microscopy. PLoS ONE 2018, 13, e0197840. [Google Scholar] [CrossRef]
- Willig, C.J.; Duan, K.; Zhang, Z.J. Transcriptome Profiling of Plant Genes in Response to Agrobacterium Tumefaciens-Mediated Transformation. Curr. Top. Microbiol. Immunol. 2018, 418, 319–348. [Google Scholar] [CrossRef] [PubMed]
- Weimer, A.K.; Biedermann, S.; Harashima, H.; Roodbarkelari, F.; Takahashi, N.; Foreman, J.; Guan, Y.; Pochon, G.; Heese, M.; Van Damme, D.; et al. The Plant-Specific CDKB1-CYCB1 Complex Mediates Homologous Recombination Repair in Arabidopsis. EMBO J. 2016, 35, 2068–2086. [Google Scholar] [CrossRef] [PubMed]
- Lutova, L.A.; Dodueva, I.E. Role of Meristem-Specific Genes of Plants in Formation of Genetic Tumors. Russ. J. Dev. Biol. 2007, 38, 350–362. [Google Scholar] [CrossRef]
- Johansson, O.N.; Fantozzi, E.; Fahlberg, P.; Nilsson, A.K.; Buhot, N.; Tör, M.; Andersson, M.X. Role of the Penetration-Resistance Genes PEN1, PEN2 and PEN3 in the Hypersensitive Response and Race-Specific Resistance in Arabidopsis Thaliana. Plant J. 2014, 79, 466–476. [Google Scholar] [CrossRef]
- Kumar, D. Salicylic Acid Signaling in Disease Resistance. Plant Sci. 2014, 228, 127–134. [Google Scholar] [CrossRef]
- Hu, L.; Yang, L. Time to Fight: Molecular Mechanisms of Age-Related Resistance. Phytopathology 2019, 109, 1500–1508. [Google Scholar] [CrossRef]
- Lutova, L.; Dodueva, I. Genetic Control of Regeneration Processes of Radish Plants In Vitro: From Phenotype to Genotype. Biol. Commun. 2019, 64, 124–132. [Google Scholar] [CrossRef]
- Zhang, C.; Gao, M.; Seitz, N.C.; Angel, W.; Hallworth, A.; Wiratan, L.; Darwish, O.; Alkharouf, N.; Dawit, T.; Lin, D.; et al. LUX ARRHYTHMO Mediates Crosstalk between the Circadian Clock and Defense in Arabidopsis. Nat. Commun. 2019, 10, 2543. [Google Scholar] [CrossRef]
- Yamaura, S.; Yamauchi, Y.; Makihara, M.; Yamashino, T.; Ishikawa, A. CCA1 and LHY Contribute to Nonhost Resistance to Pyricularia Oryzae (Syn. Magnaporthe Oryzae) in Arabidopsis Thaliana. Biosci. Biotechnol. Biochem. 2020, 84, 76–84. [Google Scholar] [CrossRef]
- Siebert, P.D.; Chenchik, A. Modified Acid Guanidinium Thiocyanate-Phenol-Chloroform RNA Extraction Method Which Greatly Reduces DNA Contamination. Nucleic Acids Res. 1993, 21, 2019–2020. [Google Scholar] [CrossRef][Green Version]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Bray, N.L.; Pimentel, H.; Melsted, P.; Pachter, L. Near-Optimal Probabilistic RNA-Seq Quantification. Nat. Biotechnol. 2016, 34, 525–527. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Krishnakumar, V.; Hanlon, M.R.; Contrino, S.; Ferlanti, E.S.; Karamycheva, S.; Kim, M.; Rosen, B.D.; Cheng, C.-Y.; Moreira, W.; Mock, S.A.; et al. Araport: The Arabidopsis Information Portal. Nucleic Acids Res. 2015, 43, D1003–D1009. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic Local Alignment Search Tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.-G.; Han, Y.; He, Q.-Y. ClusterProfiler: An R Package for Comparing Biological Themes among Gene Clusters. Omics J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Wang, L.-G.; Yan, G.-R.; He, Q.-Y. DOSE: An R/Bioconductor Package for Disease Ontology Semantic and Enrichment Analysis. Bioinform. Oxf. Engl. 2015, 31, 608–609. [Google Scholar] [CrossRef]
- Yi, X.; Du, Z.; Su, Z. PlantGSEA: A Gene Set Enrichment Analysis Toolkit for Plant Community. Nucleic Acids Res. 2013, 41, W98–W103. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods San Diego Calif. 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Dodueva, I.E.; Kiryushkin, A.S.; Yurlova, E.V.; Osipova, M.A.; Buzovkina, I.S.; Lutova, L.A. Effect of Cytokinins on Expression of Radish CLE Genes. Russ. J. Plant Physiol. 2013, 60, 388–395. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).