Marker-Assisted Introgression and Stacking of Major QTLs Controlling Grain Number (Gn1a) and Number of Primary Branching (WFP) to NERICA Cultivars
Abstract
1. Introduction
2. Results
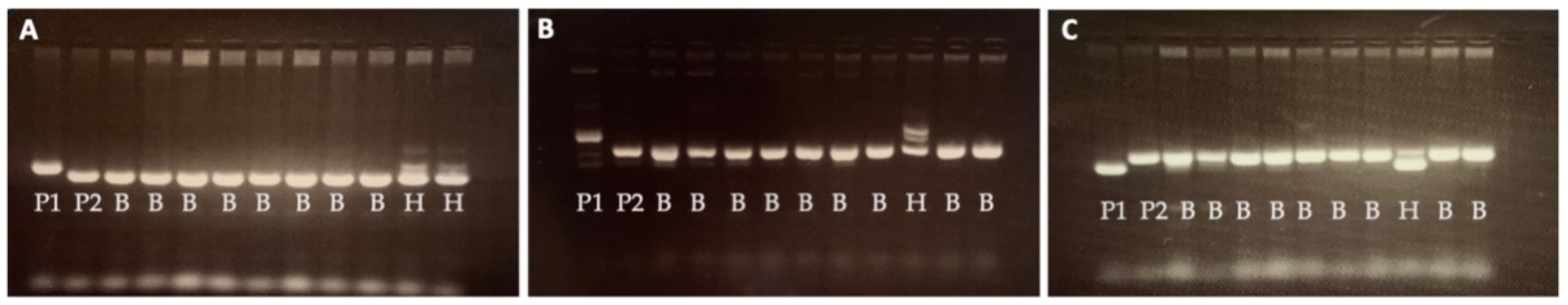
2.1. Marker-Assisted Backcrossing for Development of WISH Lines
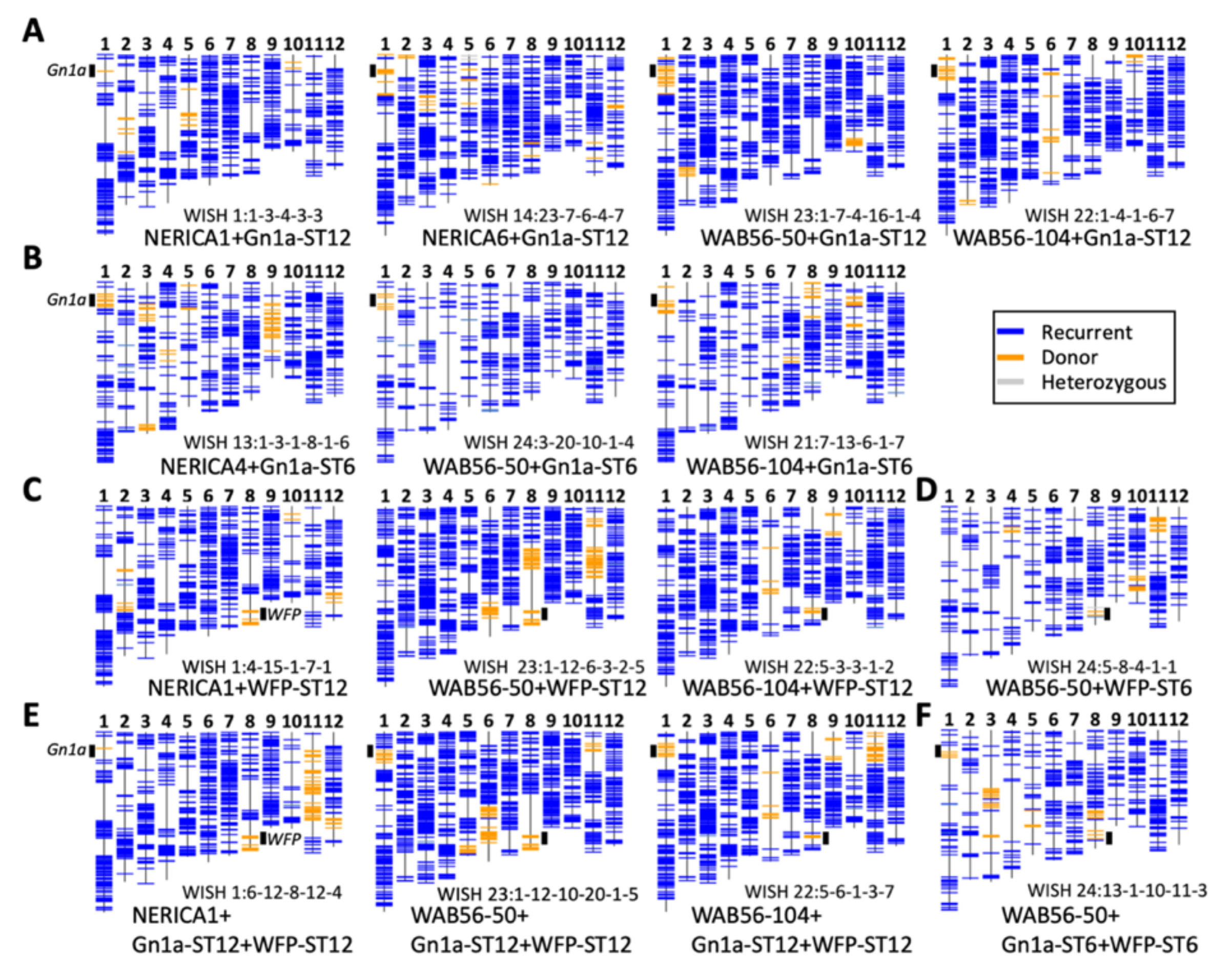
2.2. Recurrent Parent Genome Recovery Analysis on Selected Improved Lines
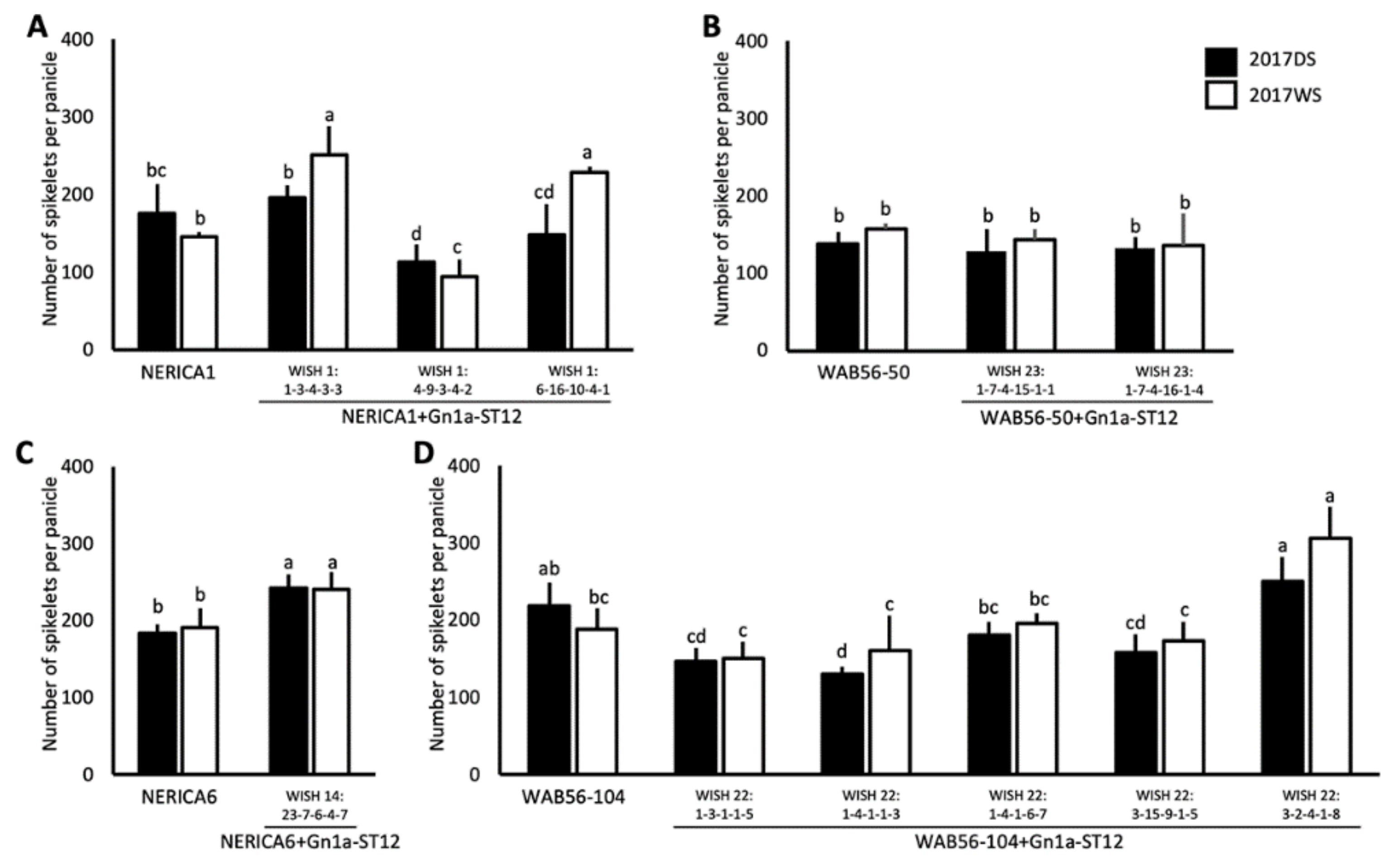
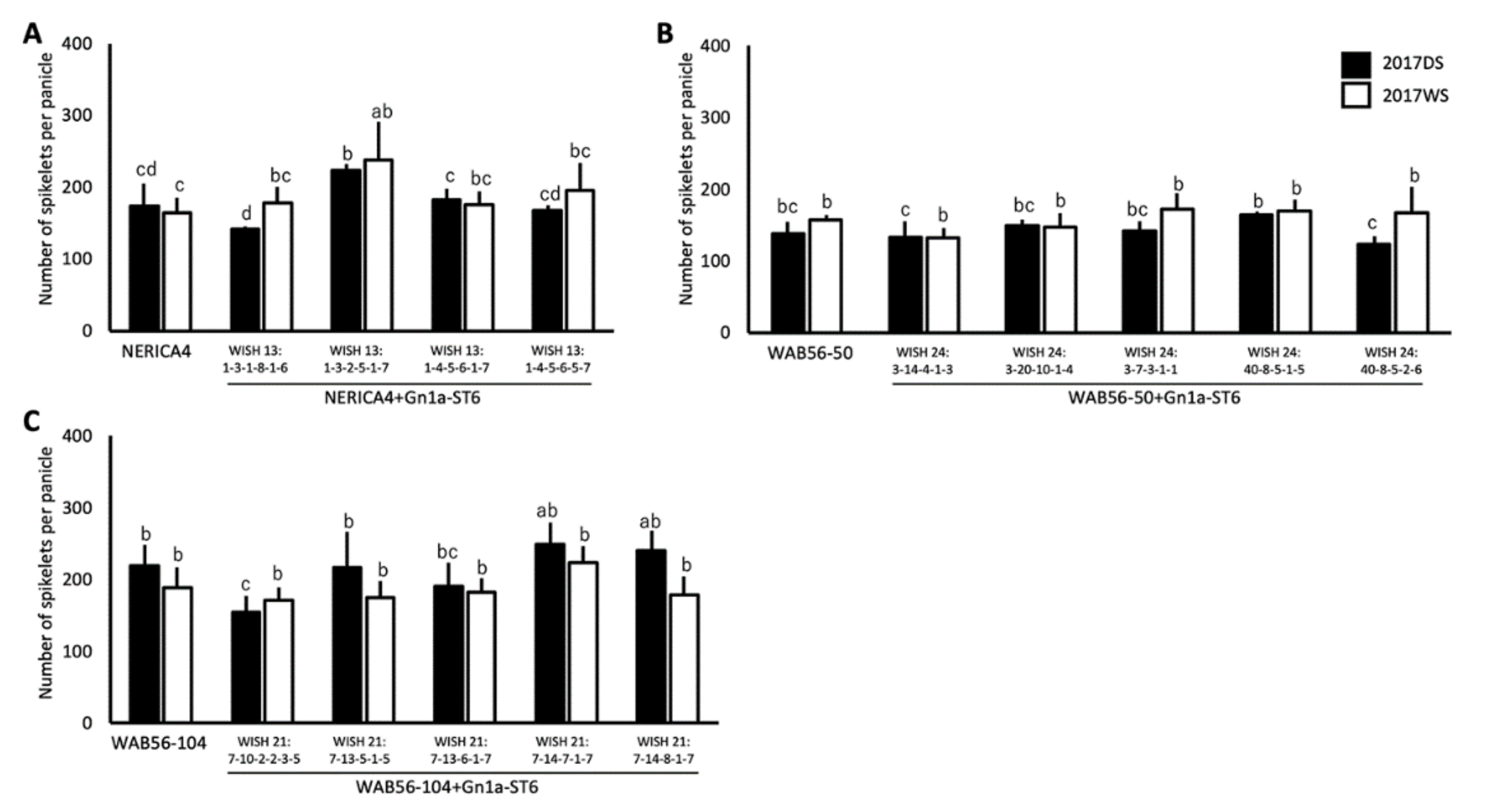
2.3. Effects of Gn1a Allele on Grain Number
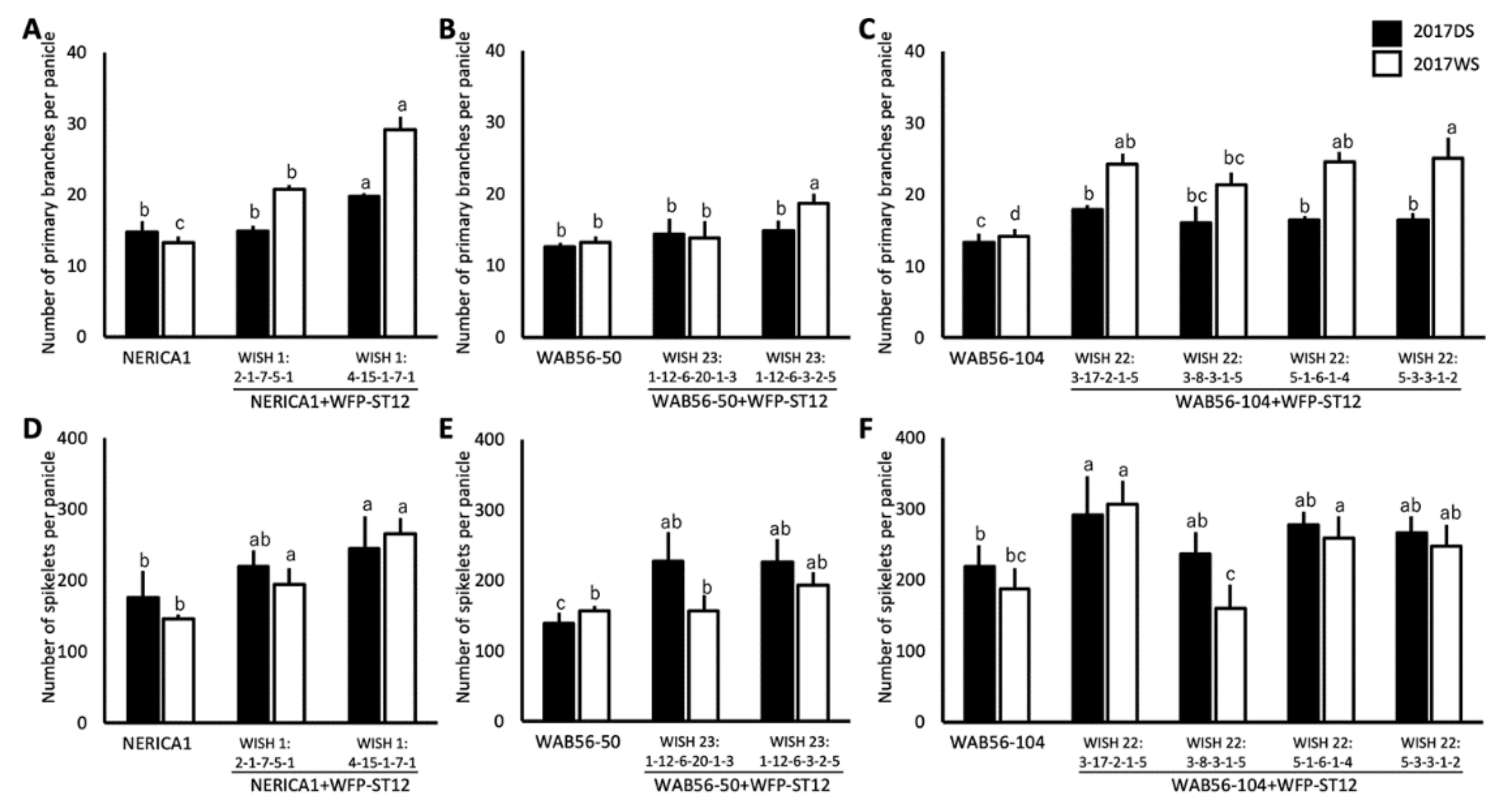
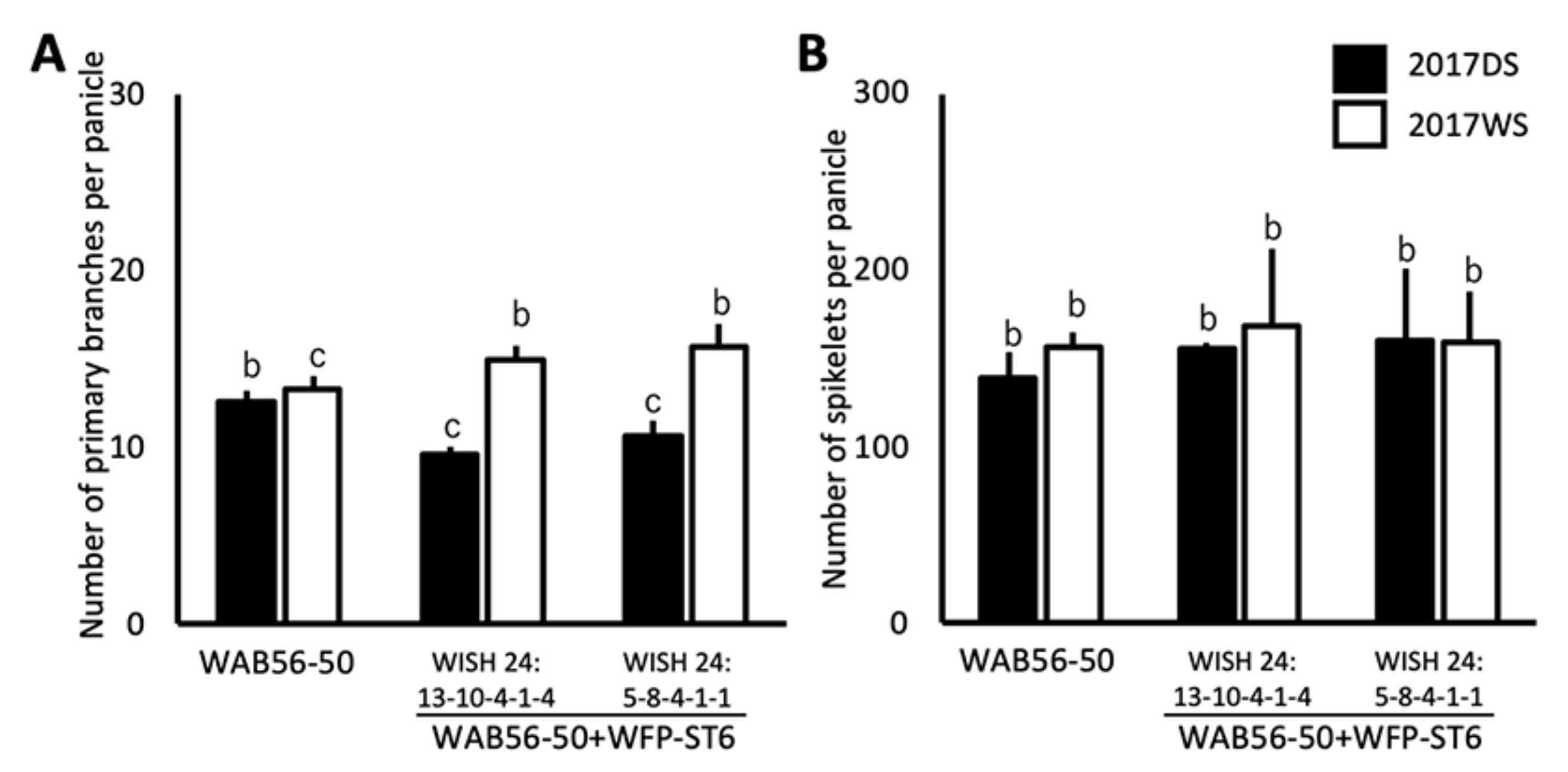
2.4. Effects of WFP Allele on Primary Branching Number and Grain Number
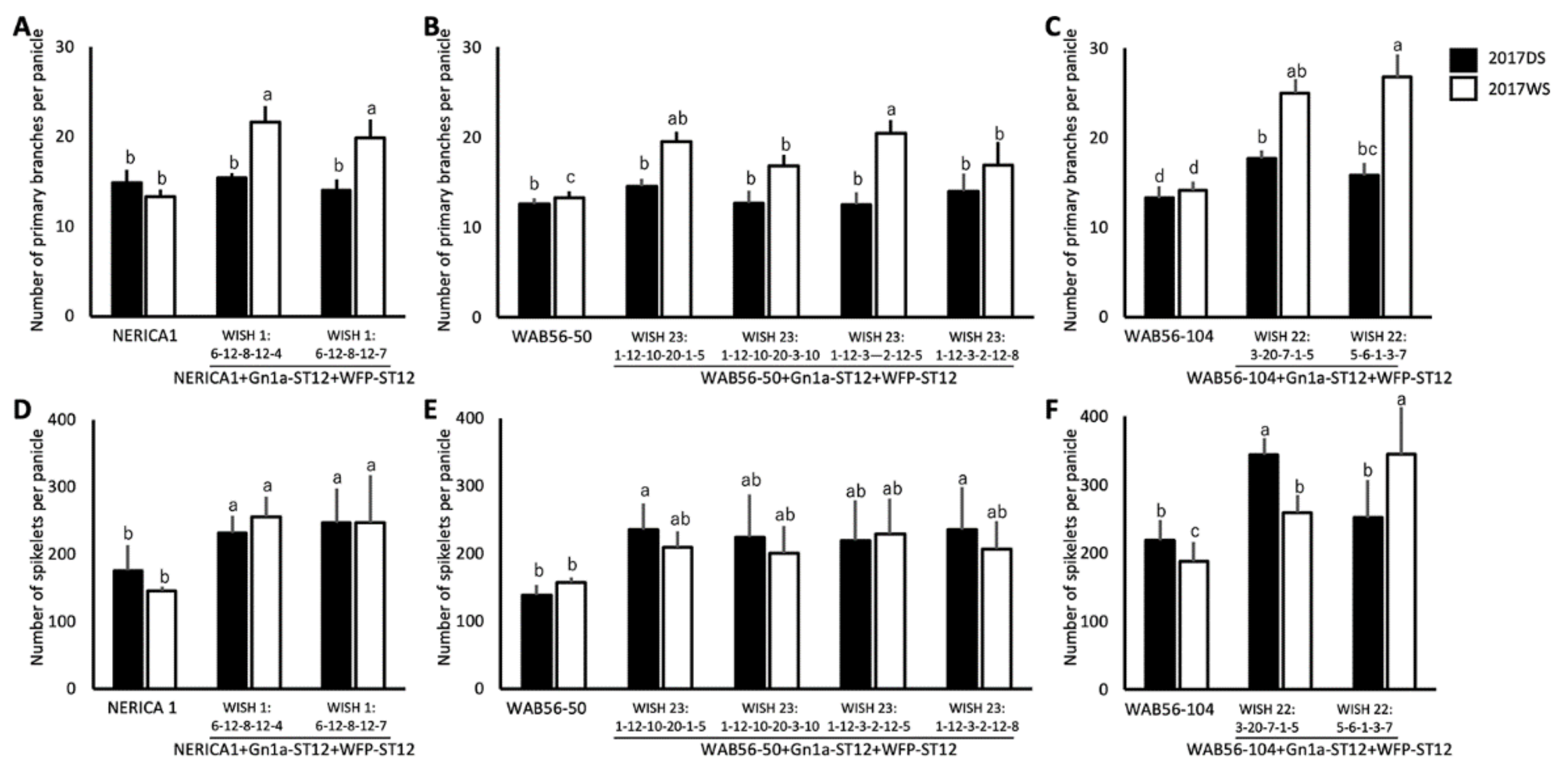
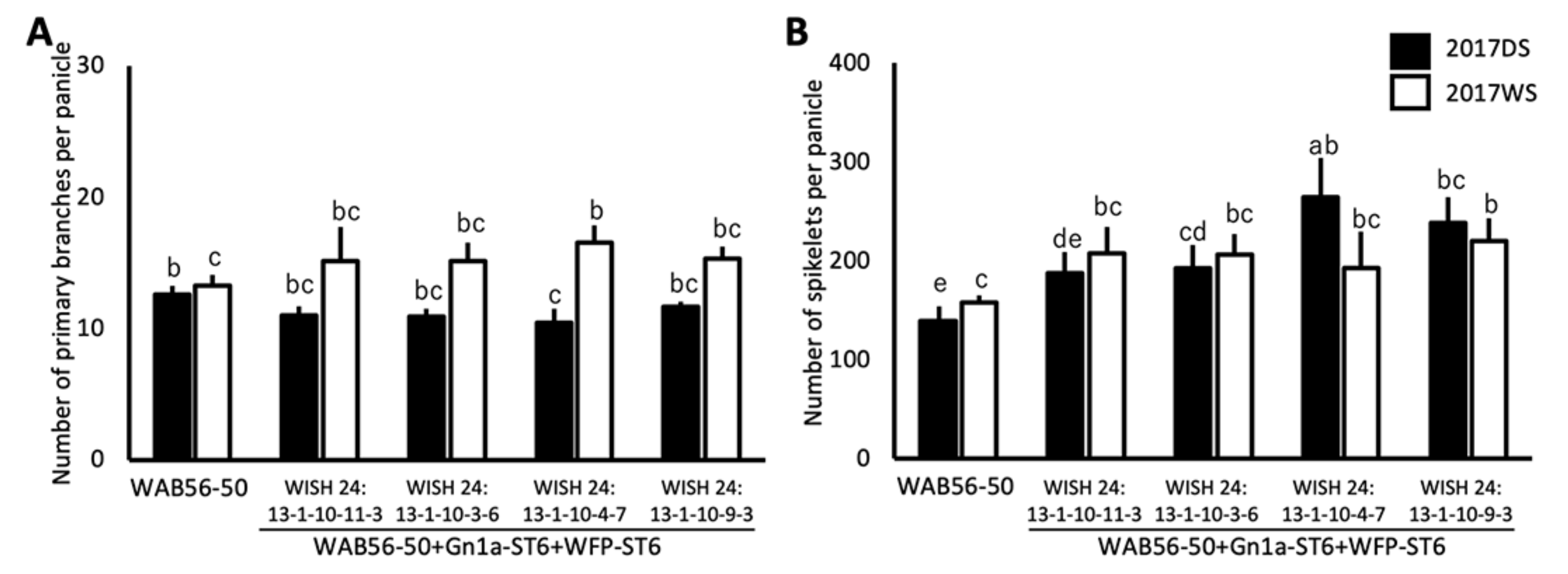
2.5. Effects of Stacking of Gn1a + WFP on Spikelet Number and Primary Branching
2.6. Agronomic Performance of WISH Introgression Lines
3. Discussion
4. Materials and Methods
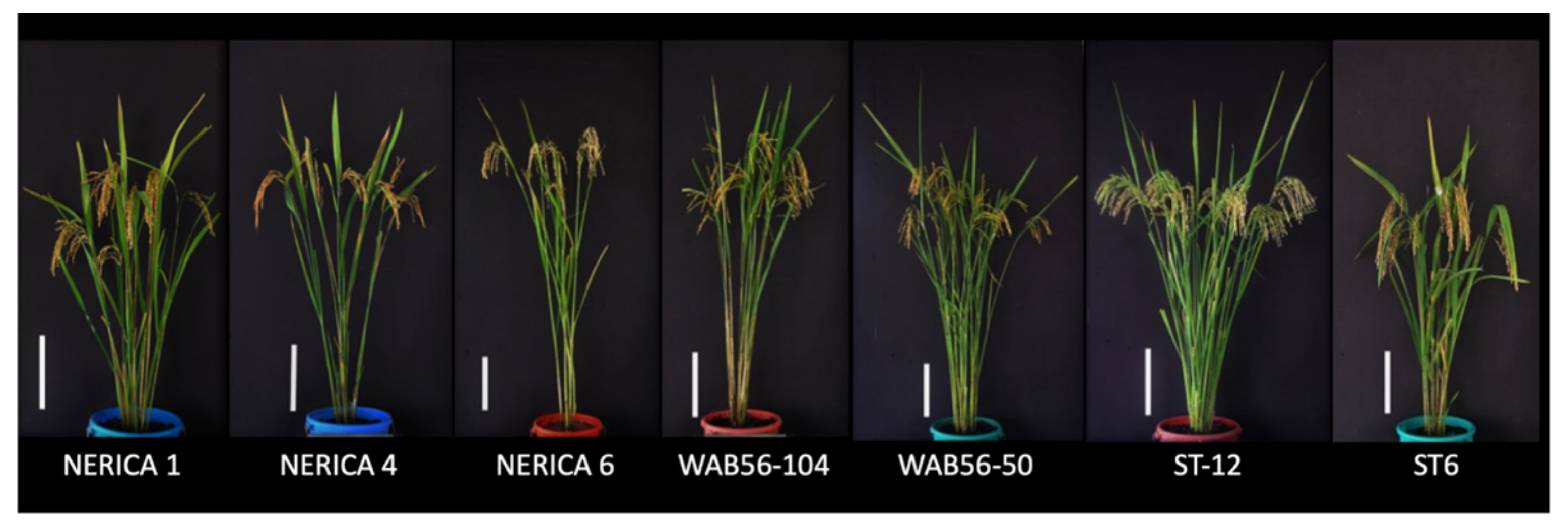
4.1. Plant Materials
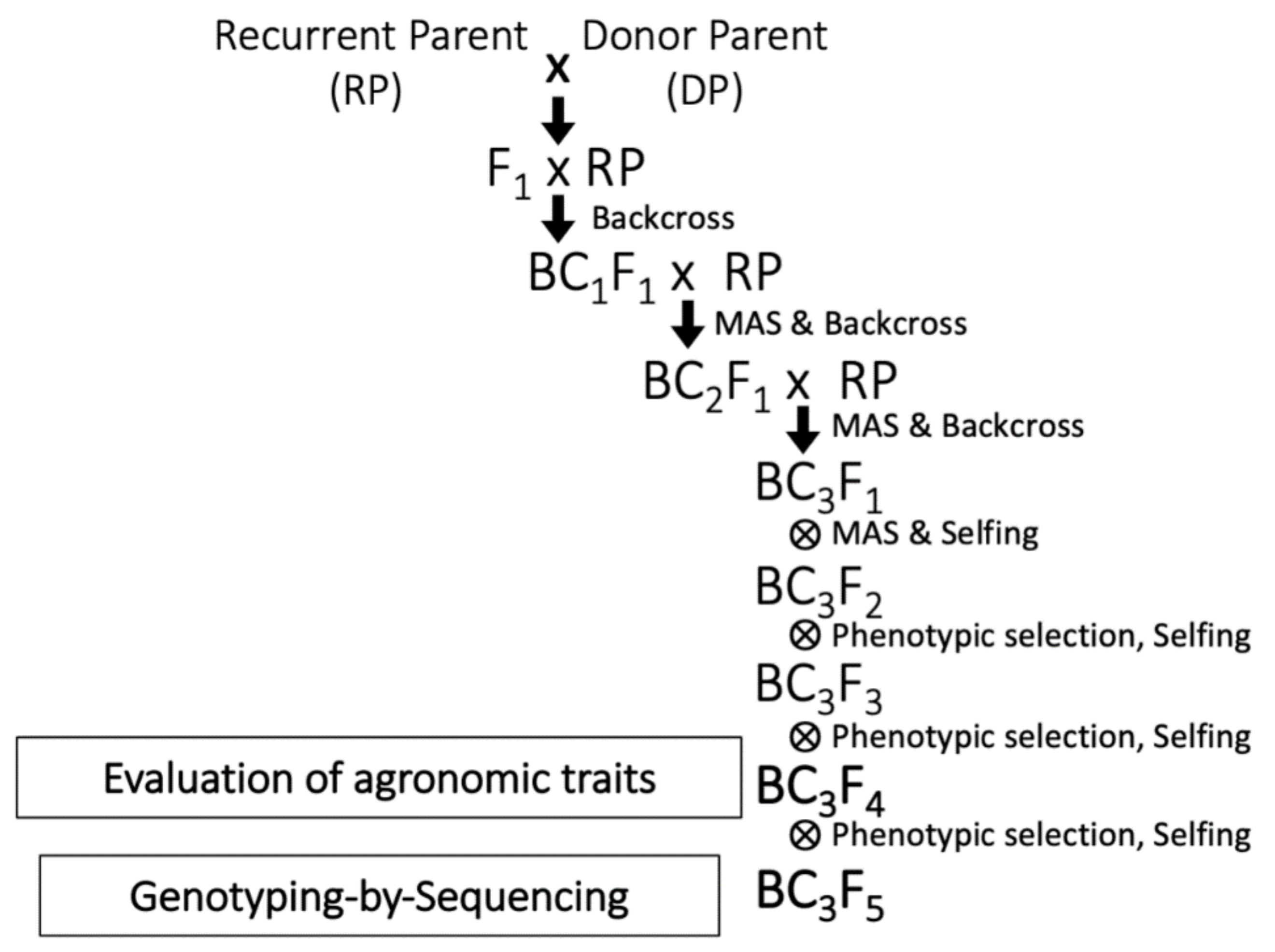
4.2. Development of WISH Lines by MAS
4.3. Foreground Selection for Gn1a and WFP Alleles
4.4. Genotyping-by-Sequencing (GBS)
4.5. Recurrent Parent Genome Recovery (RPGR) Analysis
4.6. Field Trials and Agronomic Evaluation
4.7. Data Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Maclean, J.L.; Dawe, D.; Hardy, B.; Hettel, G.P. Rice Almanac: Source Book for One of the Most Important Economic Activities on Earth, 3rd ed.; IRRI: Manila, Philippines, 2013; pp. 216–217. ISBN 0851996361. [Google Scholar]
- Zeigler, R.S.; Barclay, A. The relevance of rice. Rice 2008, 1, 3–10. [Google Scholar] [CrossRef]
- Xing, Y.; Zhang, Q. Genetic and molecular bases of rice yield. Annu. Rev. Plant. Biol. 2010, 61, 421–442. [Google Scholar] [CrossRef] [PubMed]
- Miura, K.; Ashikari, M.; Matsuoka, M. The role of QTLs in the breeding of high-yielding rice. Trends Plant. Sci. 2011, 16, 319–326. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, T.; Wu, J.; Kanamori, H.; Katayose, Y.; Fujisawa, M.; Namiki, N.; Mizuno, H.; Yamamoto, K.; Antonio, B.A.; Baba, T.; et al. The map-based sequence of the rice genome. Nature 2005, 436, 793–800. [Google Scholar] [CrossRef]
- Song, S.; Tian, D.; Zhang, Z.; Hu, S.; Yu, J. Rice genomics: Over the past two decades and into the future. Genom. Proteom. Bioinform. 2018, 16, 397–404. [Google Scholar] [CrossRef]
- Yano, M.; Katayose, Y.; Ashikari, M.; Yamanouchi, U.; Monna, L.; Fuse, T.; Baba, T.; Yamamoto, K.; Umehara, Y.; Nagamura, Y.; et al. Hd1 a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the arabidopsis flowering time gene CONSTANS. Plant Cell 2000, 12, 2473–2483. [Google Scholar] [CrossRef]
- Xue, W.; Xing, Y.; Weng, X.; Zhao, Y.; Tang, W.; Wang, L.; Zhou, H.; Yu, S.; Xu, C.; Li, X.; et al. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat. Genet. 2008, 40, 761–767. [Google Scholar] [CrossRef]
- Doi, K.; Izawa, T.; Fuse, T.; Yamanouchi, U.; Kubo, T.; Shimatani, Z.; Yano, M.; Yoshimura, A. Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev. 2004, 18, 926–936. [Google Scholar] [CrossRef]
- Fan, C.; Xing, Y.; Mao, H.; Lu, T.; Han, B.; Xu, C.; Li, X.; Zhang, Q. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor. Appl. Genet. 2006, 112, 1164–1171. [Google Scholar] [CrossRef]
- Song, X.J.; Huang, W.; Shi, M.; Zhu, M.Z.; Lin, H.X. A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat. Genet. 2007, 39, 623–630. [Google Scholar] [CrossRef]
- Weng, J.; Gu, S.; Wan, X.; Gao, H.; Guo, T.; Su, N.; Lei, C.; Zhang, X.; Cheng, Z.; Guo, X.; et al. Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res. 2008, 18, 1199–1209. [Google Scholar] [CrossRef]
- Ashikari, M.; Sakakibara, H.; Lin, S.; Yamamoto, T.; Takashi, T.; Nishimura, A.; Angeles, E.R.; Qian, Q.; Kitano, H.; Matsuoka, M. Cytokinin oxidase regulates rice grain production. Science 2005, 309, 741–745. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Qian, Q.; Liu, Z.; Sun, H.; He, S.; Luo, D.; Xia, G.; Chu, C.; Li, J.; Fu, X. Natural variation at the DEP1 locus enhances grain yield in rice. Nat. Genet. 2009, 41, 494–497. [Google Scholar] [CrossRef] [PubMed]
- Miura, K.; Ikeda, M.; Matsubara, A.; Song, X.J.; Ito, M.; Asano, K.; Matsuoka, M.; Kitano, H.; Ashikari, M. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 2010, 42, 545–549. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.-R.; Ramos, J.M.; Hizon, R.J.M.; Ashikari, M.; Virk, P.S.; Torres, E.A.; Nissila, E.; Jena, K.K. Introgression of a functional epigenetic OsSPL14/WFP allele into elite indica rice genomes greatly improved panicle traits and grain yield. Sci. Rep. 2018, 8, 3833. [Google Scholar] [CrossRef] [PubMed]
- Yamada, S.; Kurokawa, Y.; Nagai, K.; Angeles-Shim, R.B.; Yasui, H.; Furuya, N.; Yoshimura, A.; Doi, K.; Ashikari, M.; Sunohara, H. Evaluation of backcrossed pyramiding lines of the yield-related gene and the bacterial leaf Blight resistant genes. J. Intl. Cooper Agric. Dev. 2020, 18, 18–28. [Google Scholar]
- Feng, X.; Wang, C.; Nan, J.; Zhang, X.; Wang, R.; Jiang, G.; Yuan, Q.; Lin, S. Updating the elite rice variety Kongyu 131 by improving the Gn1a locus. Rice 2017, 10, 35. [Google Scholar] [CrossRef] [PubMed]
- Hasan, M.M.; Rafii, M.Y.; Ismail, M.R.; Mahmood, M.; Rahim, H.A.; Alam, M.A.; Ashkani, S.; Malek, M.A.; Latif, M.A. Marker-assisted backcrossing: A useful method for rice improvement. Biotechnol. Biotechnol. Equip. 2015, 29, 237–254. [Google Scholar] [CrossRef] [PubMed]
- Jena, K.K.; Mackill, D.J. Molecular markers and their use in marker-assisted selection in rice. Crop. Sci. 2008, 48, 1266–1276. [Google Scholar] [CrossRef]
- Chen, S.; Lin, X.H.; Xu, C.G.; Zhang, Q. Improvement of bacterial blight resistance of ‘Minghui 63’, an elite restorer line of hybrid rice, by molecular marker-assisted selection. Crop. Sci. 2000, 40, 239–244. [Google Scholar] [CrossRef]
- Angeles-Shim, R.B.; Reyes, V.P.; del Valle, M.M.; Lapis, R.S.; Shim, J.; Sunohara, H.; Jena, K.K.; Ashikari, M.; Doi, K. Marker-assisted introgression of quantitative resistance gene Pi21 confers broad spectrum resistance to rice blast. Rice Sci. 2020, 27, 113–123. [Google Scholar] [CrossRef]
- Septiningsih, E.M.; Pamplona, A.M.; Sanchez, D.L.; Neeraja, C.N.; Vergara, G.V.; Heuer, S.; Ismail, A.M.; Mackill, D.J. Development of submergence-tolerant rice cultivars: The Sub1 locus and beyond. Ann. Bot. 2009, 103, 151–160. [Google Scholar] [CrossRef] [PubMed]
- McCouch, S.R.; Zhao, K.; Wright, M.; Tung, C.-W.; Ebana, K.; Thomson, M.; Reynolds, A.; Wang, D.; DeClerck, G.; Ali, M.L.; et al. Development of genome-wide SNP assays for rice. Breed. Sci. 2010, 60, 524–535. [Google Scholar] [CrossRef]
- Thomson, M.J.; Zhao, K.; Wright, M.; McNally, K.L.; Rey, J.; Tung, C.-W.; Reynolds, A.; Scheffler, B.; Eizenga, G.; McClung, A.; et al. High-throughput single nucleotide polymorphism genotyping for breeding applications in rice using the BeadXpress platform. Mol. Breed. 2012, 29, 875–886. [Google Scholar] [CrossRef]
- Elshire, R.J.; Glaubitz, J.C.; Sun, Q.; Poland, J.A.; Kawamoto, K.; Buckler, E.S.; Mitchell, S.E. A robust, simple Genotyping-by-Sequencing (GBS) approach for high diversity species. PLoS ONE 2011, 6, e19379. [Google Scholar] [CrossRef]
- Arbelaez, J.D.; Moreno, L.T.; Singh, N.; Tung, C.-W.; Maron, L.G.; Ospina, Y.; Martinez, C.P.; Grenier, C.; Lorieux, M.; McCouch, S. Development and GBS-genotyping of introgression lines (ILs) using two wild species of rice, O. meridionalis and O. rufipogon, in a common recurrent parent, O. sativa cv. curinga. Mol. Breed. 2015, 35, 81. [Google Scholar] [CrossRef]
- Begum, H.; Spindel, J.E.; Lalusin, A.; Borromeo, T.; Gregorio, G.; Hernandez, J.; Virk, P.; Collard, B.; McCouch, S.R. Genome-wide association mapping for yield and other agronomic traits in an elite breeding population of tropical rice (Oryza sativa). PLoS ONE 2015, 10, e0119873. [Google Scholar] [CrossRef]
- Liao, C.Y.; Wu, P.; Hu, B.; Yi, K.K. Effects of genetic background and environment on QTLs and epistasis for rice (Oryza sativa L.) panicle number. Theor. Appl. Genet. 2001, 103, 104–111. [Google Scholar] [CrossRef]
- Pabuayon, I.C.M.; Trinidad, J.L.; Angeles-Shim, R.B.; Kohli, A. Systems biology of crop improvement: Drought tolerance as a model to integrate molecular biology, physiology, and breeding. In Advancement in Crop Improvement Techniques; Woodhead Publishing: Southston, UK, 2020; pp. 209–231. [Google Scholar]
- Ikeda, M.; Hirose, Y.; Takashi, T.; Shibata, Y.; Yamamura, T.; Komura, T.; Doi, K.; Ashikari, M.; Matsuoka, M.; Kitano, H. Analysis of rice panicle traits and detection of QTLs using an image analyzing method. Breed. Sci. 2010, 60, 55–64. [Google Scholar] [CrossRef]
- McCouch, S.R. Development and mapping of 2240 new ssr markers for rice (Oryza sativa L.). DNA Res. 2002, 9, 199–207. [Google Scholar] [CrossRef]
- Poland, J.A.; Brown, P.J.; Sorrells, M.E.; Jannink, J.L. Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PLoS ONE 2012, 7, e32253. [Google Scholar] [CrossRef]
- Jones, M.P.; Dingkuhn, M.; Aluko, G.K.; Semon, M. Interspecific Oryza sativa L. X O. glaberrima Steud. progenies in upland rice improvement. Euphytica 1997, 94, 237–246. [Google Scholar] [CrossRef]
- Sakamoto, T.; Matsuoka, M. Identifying and exploiting grain yield genes in rice. Curr. Opin. Plant. Biol. 2008, 11, 209–214. [Google Scholar] [CrossRef]
- Jiao, Y.; Wang, Y.; Xue, D.; Wang, J.; Yan, M.; Liu, G.; Dong, G.; Zeng, D.; Lu, Z.; Zhu, X.; et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 2010, 42, 541–544. [Google Scholar] [CrossRef]
- Lu, Z.; Yu, H.; Xiong, G.; Wang, J.; Jiao, Y.; Liu, G.; Jing, Y.; Meng, X.; Hu, X.; Qian, Q.; et al. Genome-wide binding analysis of the transcription activator ideal plant architecture1 reveals a complex network regulating rice plant architecture. Plant. Cell 2013, 25, 3743–3759. [Google Scholar] [CrossRef]
- Xu, H.; Zhao, M.; Zhang, Q.; Xu, Z.; Xu, Q. The DENSE AND ERECT PANICLE 1 (DEP1) gene offering the potential in the breeding of high-yielding rice. Breed. Sci. 2016, 66, 659–667. [Google Scholar] [CrossRef]
- Chu, J.; Zhao, Y.; Beier, S.; Schulthess, A.W.; Stein, N.; Philipp, N.; Röder, M.S.; Reif, J.C. Suitability of single-nucleotide polymorphism arrays versus genotyping-by-sequencing for genebank genomics in wheat. Front. Plant. Sci. 2020, 11, 42. [Google Scholar] [CrossRef]
- Kurokawa, Y.; Noda, T.; Yamagata, Y.; Angeles-Shim, R.; Sunohara, H.; Uehara, K.; Furuta, T.; Nagai, K.; Jena, K.K.; Yasui, H.; et al. Construction of a versatile SNP array for pyramiding useful genes of rice. Plant. Sci. 2016, 242, 131–139. [Google Scholar] [CrossRef]
- Chukwu, S.C.; Rafii, M.Y.; Ramlee, S.I.; Ismail, S.I.; Oladosu, Y.; Muhammad, I.; Musa, I.; Ahmed, M.; Jatto, M.I.; Yusuf, B.R. Recovery of recurrent parent genome in a marker-assisted backcrossing against rice blast and blight infections using functional markers and SSRs. Plants 2020, 9, 1411. [Google Scholar] [CrossRef]
- Khush, G.S. Breaking the yield frontier of rice. GeoJournal 1995, 35, 329–332. [Google Scholar] [CrossRef]
- Kim, S.R.; Ramos, J.; Ashikari, M.; Virk, P.S.; Torres, E.A.; Nissila, E.; Hechanova, S.L.; Mauleon, R.; Jena, K.K. Development and validation of allele-specific snp/indel markers for eight yield-enhancing genes using whole-genome sequencing strategy to increase yield potential of rice, Oryza Sativa L. Rice 2016, 9, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Miura, K.; Agetsuma, M.; Kitano, H.; Yoshimura, A.; Matsuoka, M.; Jacobsen, S.E.; Ashikari, M. A metastable DWARF1 epigenetic mutant affecting plant stature in rice. Proc. Natl. Acad. Sci. USA 2009, 106, 11218–11223. [Google Scholar] [CrossRef] [PubMed]
- Bradbury, P.J.; Zhang, Z.; Kroon, D.E.; Casstevens, T.M.; Ramdoss, Y.; Buckler, E.S. TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics 2007, 23, 2633–2635. [Google Scholar] [CrossRef] [PubMed]
- Kawahara, Y.; de la Bastide, M.; Hamilton, J.P.; Kanamori, H.; McCombie, W.R.; Ouyang, S.; Schwartz, D.C.; Tanaka, T.; Wu, J.; Zhou, S.; et al. Improvement of the Oryza sativa nipponbare reference genome using next generation sequence and optical map data. Rice 2013, 6, 4. [Google Scholar] [CrossRef]
- Li, H.; Durbin, R. Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Danecek, P.; Auton, A.; Abecasis, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef]
- Van Berloo, R. GGT 2.0: Versatile software for visualization and analysis of genetic data. J. Hered. 2008, 99, 232–236. [Google Scholar] [CrossRef]
- Microsoft Corporation. Microsoft Excel. 2018. Available online: https://office.microsoft.com/excel (accessed on 21 January 2021).
- IRRI. Statistical Tool for Agricultural Research Software (STAR); International Rice Research Institute: Los Banos, Philippins, 2013. [Google Scholar]

| Genetic Background | Total Number of SNPs | Recurrent Parent Genome Recovery Range of BC3F4 Population (%) |
|---|---|---|
| NERICA1 × ST12 | 1418 | 85.4–98 |
| NERICA4 × ST6 | 1175 | 84.5–87.7 |
| NERICA6 × ST12 | 1658 | 94.8 |
| WAB56-104 × ST6 | 1614 | 77.7–96.9 |
| WAB56-104 × ST12 | 1064 | 80.6–93.1 |
| WAB56-50 × ST12 | 1873 | 88.3–96.7 |
| WAB56-50 × ST6 | 828 | 81.2–95.3 |
| Marker | Forward Primer Sequence | Reverse Primer Sequence | Target Gene |
|---|---|---|---|
| RM3360 | ACTTACACAAGGCCGGGAAAGG | TGGTAGTGGTAACTCTACTCCGATGG | Gn1a |
| RM3452 | TGGACTTGGTCTCTCCAAACTCC | CAGTATGTGTTGGTGGGTCAAGC | WFP |
| RM5493 | GCGGTAACAAACCAACCAACC | AAAGCAGGACACAGTCACACAGG | WFP |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reyes, V.P.; Angeles-Shim, R.B.; Mendioro, M.S.; Manuel, M.C.C.; Lapis, R.S.; Shim, J.; Sunohara, H.; Nishiuchi, S.; Kikuta, M.; Makihara, D.; et al. Marker-Assisted Introgression and Stacking of Major QTLs Controlling Grain Number (Gn1a) and Number of Primary Branching (WFP) to NERICA Cultivars. Plants 2021, 10, 844. https://doi.org/10.3390/plants10050844
Reyes VP, Angeles-Shim RB, Mendioro MS, Manuel MCC, Lapis RS, Shim J, Sunohara H, Nishiuchi S, Kikuta M, Makihara D, et al. Marker-Assisted Introgression and Stacking of Major QTLs Controlling Grain Number (Gn1a) and Number of Primary Branching (WFP) to NERICA Cultivars. Plants. 2021; 10(5):844. https://doi.org/10.3390/plants10050844
Chicago/Turabian StyleReyes, Vincent P., Rosalyn B. Angeles-Shim, Merlyn S. Mendioro, Ma. Carmina C. Manuel, Ruby S. Lapis, Junghyun Shim, Hidehiko Sunohara, Shunsaku Nishiuchi, Mayumi Kikuta, Daigo Makihara, and et al. 2021. "Marker-Assisted Introgression and Stacking of Major QTLs Controlling Grain Number (Gn1a) and Number of Primary Branching (WFP) to NERICA Cultivars" Plants 10, no. 5: 844. https://doi.org/10.3390/plants10050844
APA StyleReyes, V. P., Angeles-Shim, R. B., Mendioro, M. S., Manuel, M. C. C., Lapis, R. S., Shim, J., Sunohara, H., Nishiuchi, S., Kikuta, M., Makihara, D., Jena, K. K., Ashikari, M., & Doi, K. (2021). Marker-Assisted Introgression and Stacking of Major QTLs Controlling Grain Number (Gn1a) and Number of Primary Branching (WFP) to NERICA Cultivars. Plants, 10(5), 844. https://doi.org/10.3390/plants10050844







