Herbicide Resistance: Another Hot Agronomic Trait for Plant Genome Editing
Abstract
1. Introduction
2. Chemical/Herbicides Method of Weed Control
3. Mechanism and Approaches for Development of Herbicide Resistant Plants/Crops
3.1. Mutagenesis/Mutation Breeding
3.2. Transgenic Approach Including the Over-expression
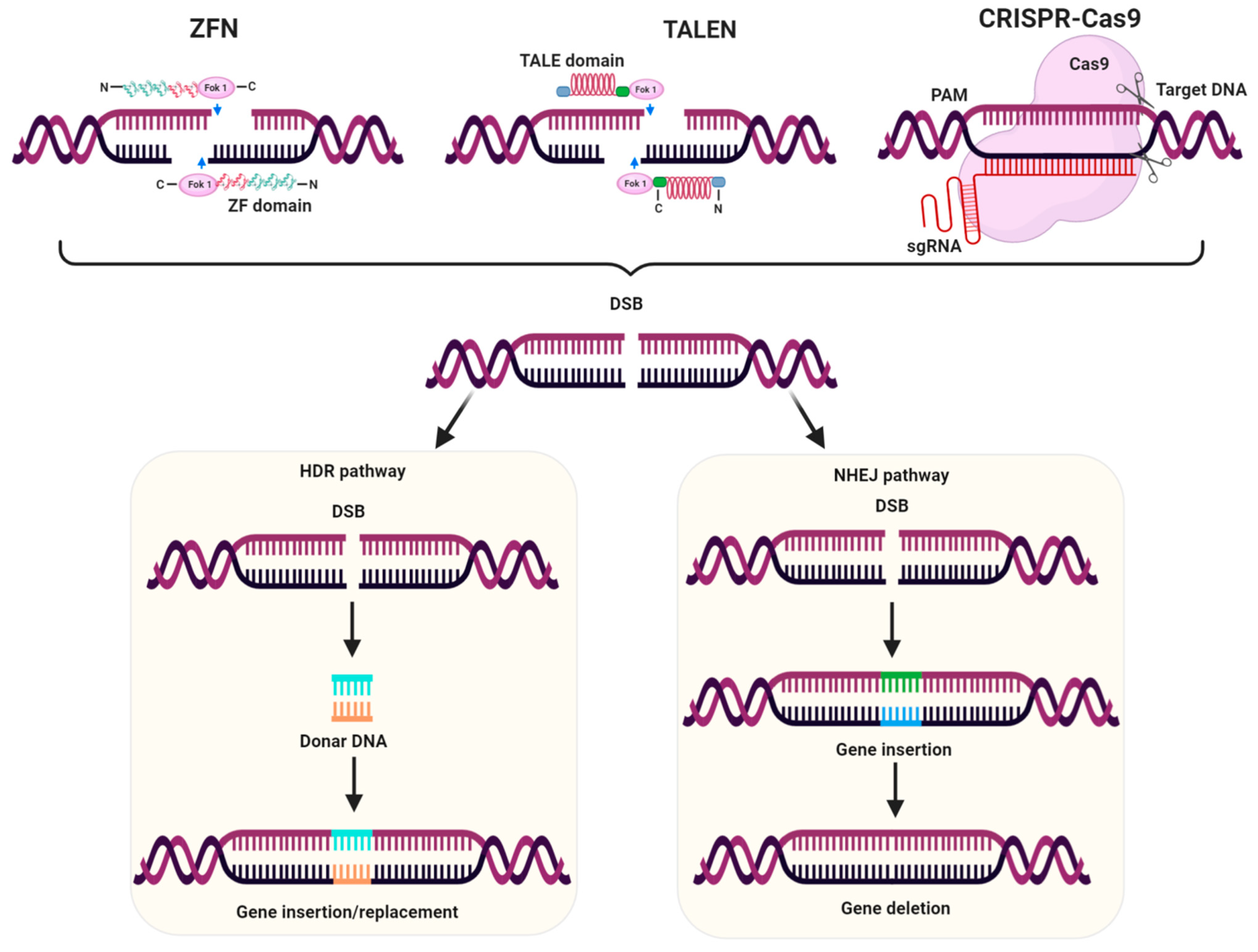
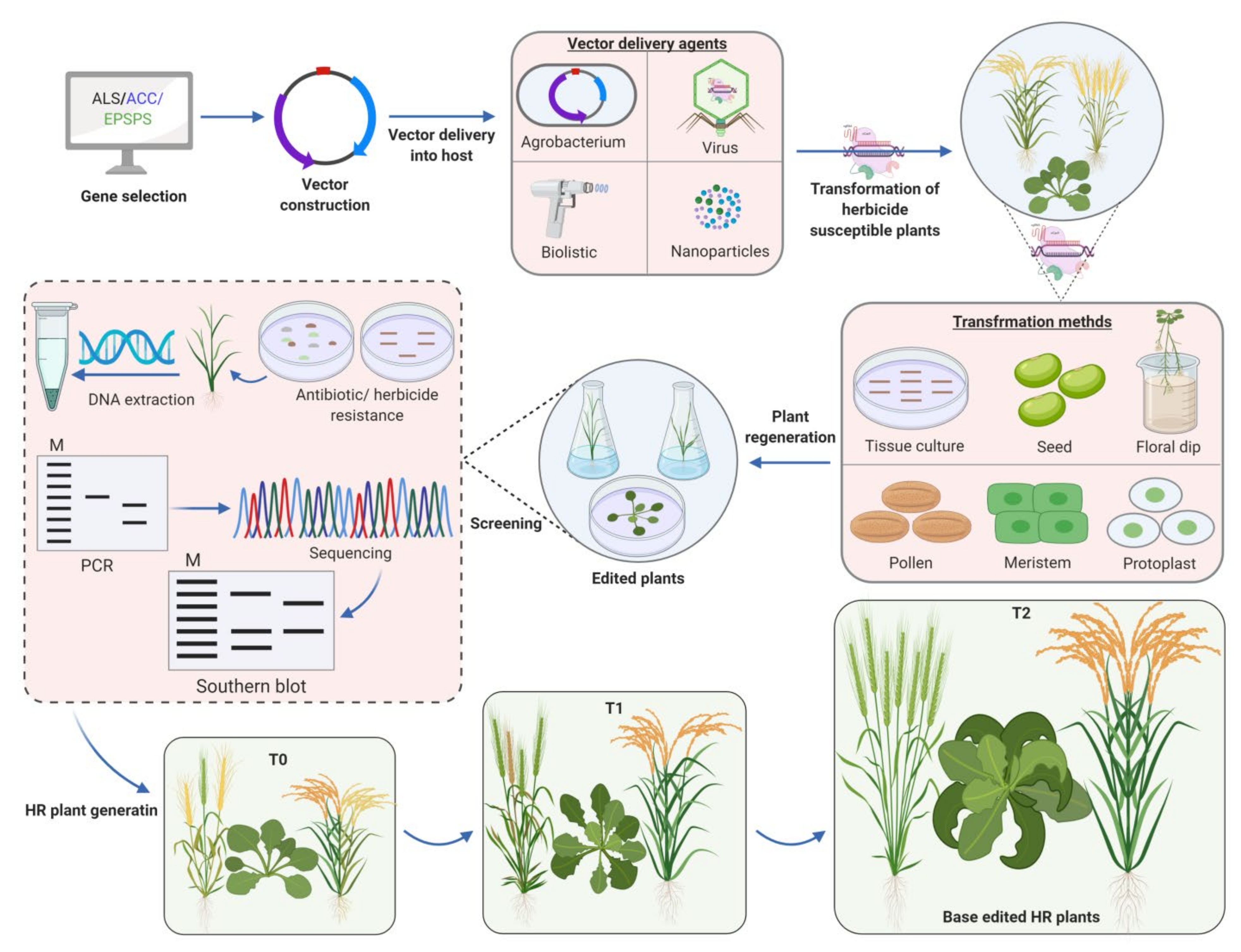
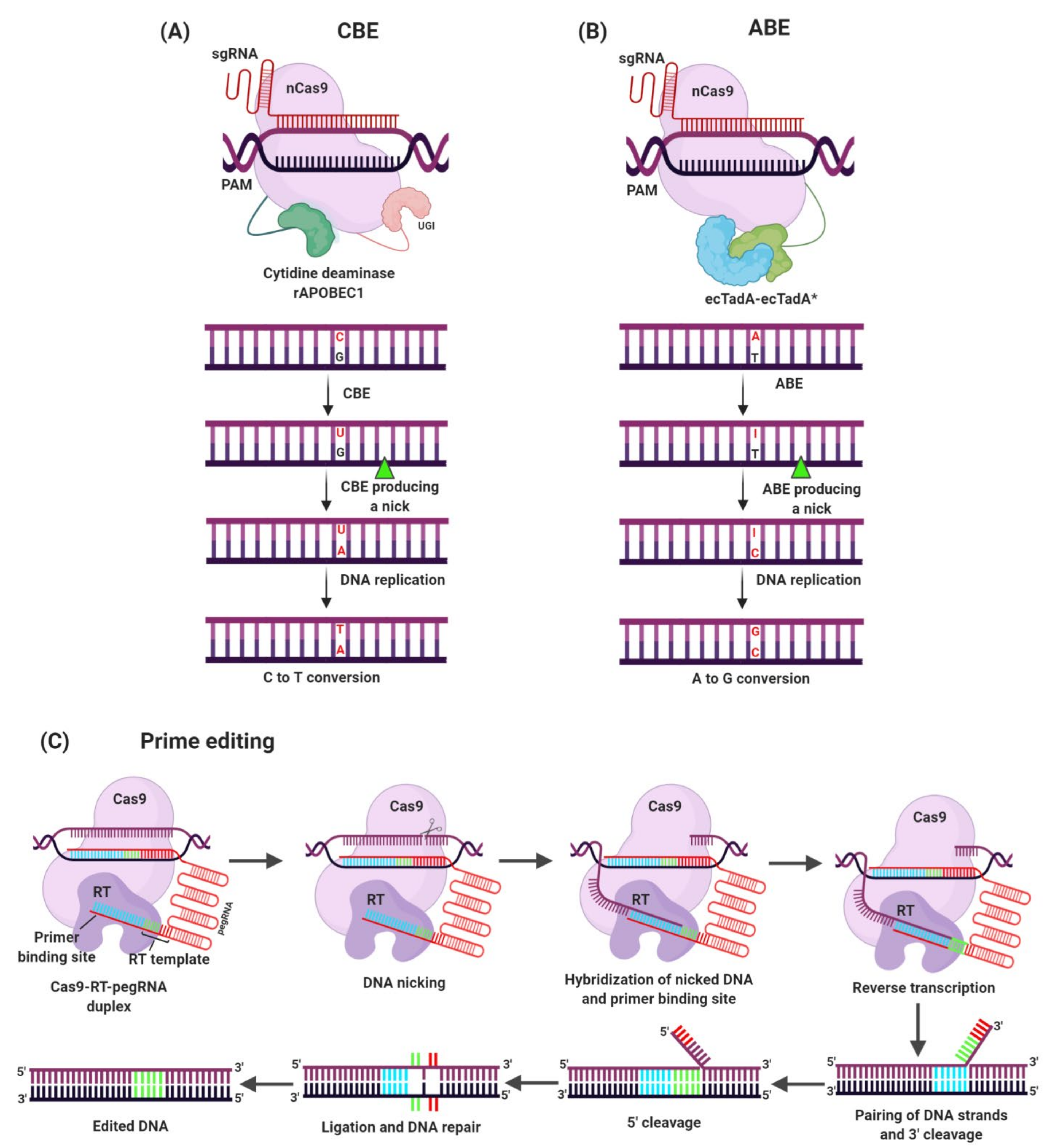
3.3. Genome Editing for Development of Herbicide Resistant Crops
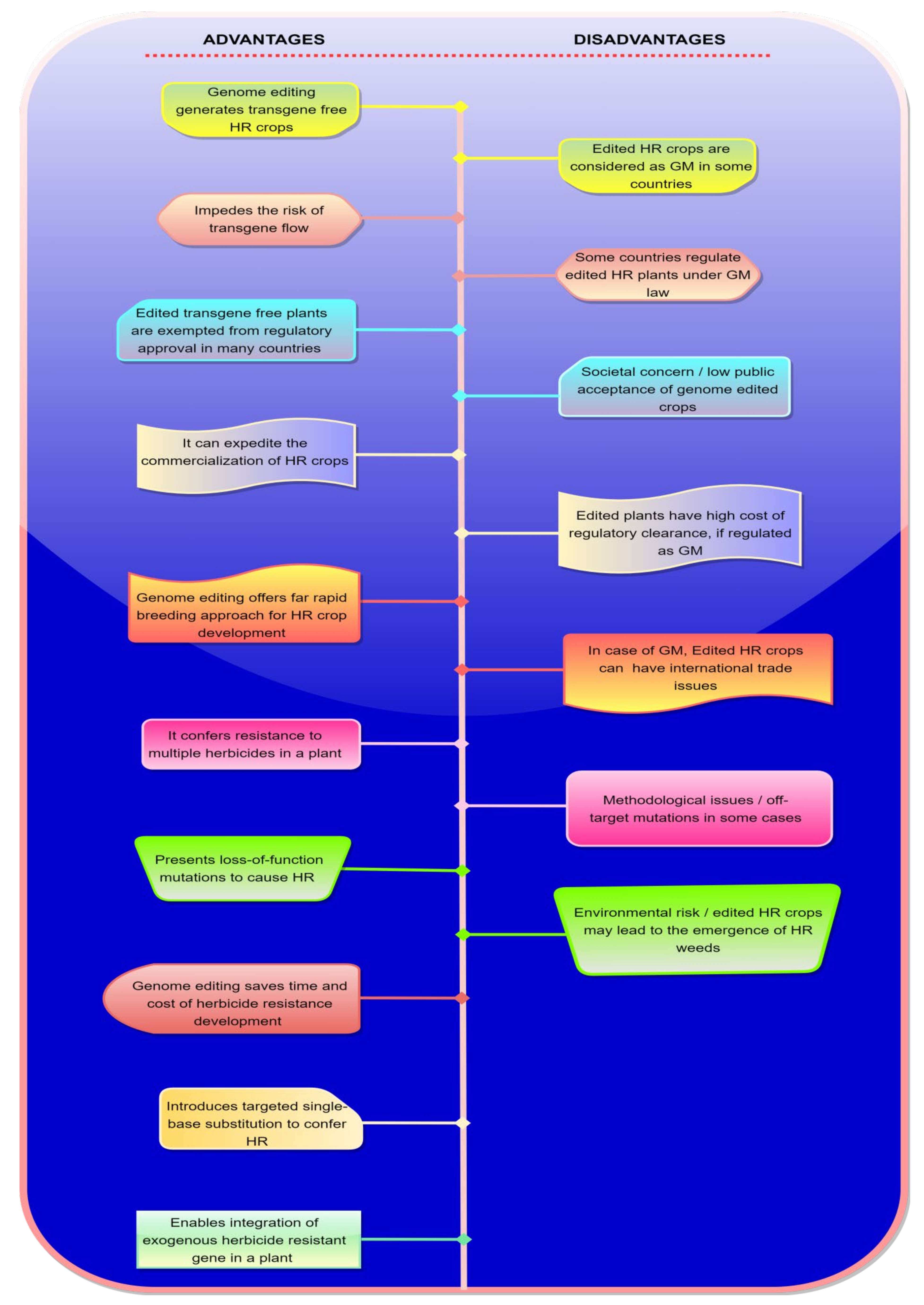
4. Advantages and Disadvantages of Genome Editing in Herbicide Resistance
5. Impact of Genome-Edited Herbicide Resistant Crops in Agriculture
6. Concluding Remarks and Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ganie, Z.A.; Obrigawitch, T.; Kang, I.H.; Copeland, D.; Gutteridge, S.; May, J.; Schrage, B.; Albright, R.; Chandi, A. An outlook of FMC’s current and future herbicide-resistance management strategies. Pest Manag. Sci. 2020, 77, 1559–1563. [Google Scholar] [CrossRef] [PubMed]
- Hickey, L.T.; Hafeez, A.N.; Robinson, H.; Jackson, S.A.; Leal-Bertioli, S.C.; Tester, M.; Gao, C.; Godwin, I.D.; Hayes, B.J.; Wulff, B.B. Breeding crops to feed 10 billion. Nat. Biotechnol. 2019, 37, 744–754. [Google Scholar] [CrossRef]
- Food and Agriculture Organization of the United Nations (FAO). Available online: www.fao.org (accessed on 9 August 2020).
- Food and Agriculture Organization of the United Nations (FAO). Recommendations for Improved Weed Management. Available online: www.fao.org/3/a0884e/a0884e.pdf (accessed on 11 August 2020).
- Adetunji, C.O.; Oloke, J.K.; Bello, O.M.; Pradeep, M.; Jolly, R.S. Innovation. Isolation, structural elucidation and bioherbicidal activity of an eco-friendly bioactive 2-(hydroxymethyl) phenol, from Pseudomonas aeruginosa (C1501) and its ecotoxicological evaluation on soil. Environ. Technol. Innov. 2019, 13, 304–317. [Google Scholar] [CrossRef]
- Matloob, A.; Safdar, M.E.; Abbas, T.; Aslam, F.; Khaliq, A.; Tanveer, A.; Rehman, A.; Chadhar, A.R. Challenges and prospects for weed management in Pakistan: A review. Crop Prot. 2019, 134, 104724. [Google Scholar] [CrossRef]
- Chauhan, B.S. Grand challenges in weed management. Front. Agron. 2020, 1, 3. [Google Scholar] [CrossRef]
- Brun, T.; Rabuske, J.E.; Confortin, T.C.; Luft, L.; Todero, I.; Fischer, M.; Zabot, G.L.; Mazutti, M.A. Weed control by metabolites produced from Diaporthe schini. Environ. Technol. 2020, 1–26. [Google Scholar] [CrossRef]
- Alvarado-Serrano, D.F.; Van Etten, M.L.; Chang, S.-M.; Baucom, R.S. The relative contribution of natural landscapes and human-mediated factors on the connectivity of a noxious invasive weed. Heredity 2019, 122, 29–40. [Google Scholar] [CrossRef]
- Bahadur, S.; Verma, S.; Prasad, S.; Madane, A.; Maurya, S.; Gaurav, V.V.; Sihag, S.J. Eco-friendly weed management for sustainable crop production-A review. J. Crop Weed 2015, 11, 181–189. [Google Scholar]
- Abouziena, H.; Haggag, W. Weed control in clean agriculture: A review1. Planta Daninha 2016, 34, 377–392. [Google Scholar] [CrossRef]
- Jabran, K.; Mahajan, G.; Sardana, V.; Chauhan, B.S. Allelopathy for weed control in agricultural systems. Crop Prot. 2015, 72, 57–65. [Google Scholar] [CrossRef]
- Zimdahl, R.L. Fundamentals of Weed Science, 3rd ed.; Academic Press: San Diego, CA, USA, 2013; pp. 259–309. [Google Scholar]
- Del Rey, J.C.; Lizana, A.R.; Agüera-Vega, J.; Ruiz, M.P. Field sprayer for inter and intra-row weed control: Performance and labor savings. Span. J. Agric. Res. 2013, 11, 642–651. [Google Scholar]
- Bond, W.; Grundy, A. Non-chemical weed management in organic farming systems. Weed Res. 2001, 41, 383–405. [Google Scholar] [CrossRef]
- Guo, Y.; Cheng, L.; Long, W.; Gao, J.; Zhang, J.; Chen, S.; Pu, H.; Hu, M. Synergistic mutations of two rapeseed AHAS genes confer high resistance to sulfonylurea herbicides for weed control. Theor. Appl. Genet. 2020, 133, 2811–2824. [Google Scholar] [CrossRef]
- Andrew, I.; Storkey, J.; Sparkes, D. A review of the potential for competitive cereal cultivars as a tool in integrated weed management. Weed Res. 2015, 55, 239–248. [Google Scholar] [CrossRef]
- Duke, S.O. Perspectives on transgenic, herbicide-resistant crops in the United States almost 20 years after introduction. Pest Manag. Sci. 2015, 71, 652–657. [Google Scholar] [CrossRef]
- Menendez, J.; Rojano-Delgado, M.; De Prado, R. Differences in herbicide uptake, translocation, and distribution as sources of herbicide resistance in weeds. In Retention, Uptake, and Translocation of Agrochemicals in Plants; Myung, K., Sctchivi, N.M., Kingston, C.K., Eds.; ACS Publications: Washington, DC, USA, 2014; Volume 1171, pp. 141–157. [Google Scholar]
- Eichert, T.; Burkhardt, J. Quantification of stomatal uptake of ionic solutes using a new model system. J. Exp. Bot. 2001, 52, 771–781. [Google Scholar] [CrossRef] [PubMed]
- Gaines, T.A.; Duke, S.O.; Morran, S.; Rigon, C.A.; Tranel, P.J.; Küpper, A.; Dayan, F.E. Mechanisms of evolved herbicide resistance. J. Biol. Chem. 2020, 295, 10307–10330. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.S.; Tranel, P.J.; Stewart, C.N., Jr. Non-target-site herbicide resistance: A family business. Trends Plant Sci. 2007, 12, 6–13. [Google Scholar] [CrossRef]
- Van Eerd, L.L.; Hoagland, R.E.; Zablotowicz, R.M.; Hall, J.C. Pesticide metabolism in plants and microorganisms. Weed Sci. 2003, 51, 472–495. [Google Scholar] [CrossRef]
- Carvalho, S.J.P.d.; Nicolai, M.; Ferreira, R.R.; Figueira, A.V.d.O.; Christoffoleti, P.J. Herbicide selectivity by differential metabolism: Considerations for reducing crop damages. Sci. Agric. 2009, 66, 136–142. [Google Scholar] [CrossRef]
- Edwards, R.; Brazier-Hicks, M.; Dixon, D.P.; Cummins, I. Chemical manipulation of antioxidant defences in plants. Adv. Bot. Res. 2005, 42, 1–32. [Google Scholar]
- Zhang, R.; Liu, J.; Chai, Z.; Chen, S.; Bai, Y.; Zong, Y.; Chen, K.; Li, J.; Jiang, L.; Gao, C. Generation of herbicide tolerance traits and a new selectable marker in wheat using base editing. Nat. Plants 2019, 5, 480–485. [Google Scholar] [CrossRef]
- Weed Science Society of America (WSSA). Available online: https://wssa.net/wssa/weed/resistance/herbicide-resistance-and-herbicide-tolerance-definitions/ (accessed on 19 February 2021).
- Lombardo, L.; Coppola, G.; Zelasco, S. New technologies for insect-resistant and herbicide-tolerant plants. Trends Biotechnol. 2016, 34, 49–57. [Google Scholar] [CrossRef]
- Ni, Z.; Han, Q.; He, Y.Q.; Huang, S. Application of genome-editing technology in crop improvement. Cereal Chem. 2018, 95, 35–48. [Google Scholar] [CrossRef]
- Govindaraj, M.; Vetriventhan, M.; Srinivasan, M. Importance of genetic diversity assessment in crop plants and its recent advances: An overview of its analytical perspectives. Genet. Res. Int. 2015, 2015, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Hussain, A.; Manghwar, H.; Xie, K.; Xie, S.; Zhao, S.; Larkin, R.M.; Qing, P.; Jin, S.; Ding, F. Genome editing with the CRISPR-Cas system: An art, ethics and global regulatory perspective. Plant Biotechnol. J. 2020, 18, 1651–1669. [Google Scholar] [CrossRef] [PubMed]
- Hartung, F.; Schiemann, J. Precise plant breeding using new genome editing techniques: Opportunities, safety and regulation in the EU. Plant J. 2014, 78, 742–752. [Google Scholar] [CrossRef]
- Mishra, R.; Zhao, K. Genome editing technologies and their applications in crop improvement. Plant Biotechnol. Rep. 2018, 12, 57–68. [Google Scholar] [CrossRef]
- Shah, T.; Andleeb, T.; Lateef, S.; Noor, M.A. Genome editing in plants: Advancing crop transformation and overview of tools. Plant Physiol. Biochem. 2018, 131, 12–21. [Google Scholar] [CrossRef] [PubMed]
- Mao, Y.; Botella, J.R.; Liu, Y.; Zhu, J.-K. Gene editing in plants: Progress and challenges. Natl. Sci. Rev. 2019, 6, 421–437. [Google Scholar] [CrossRef]
- Shukla, V.K.; Doyon, Y.; Miller, J.C.; DeKelver, R.C.; Moehle, E.A.; Worden, S.E.; Mitchell, J.C.; Arnold, N.L.; Gopalan, S.; Meng, X. Precise genome modification in the crop species Zea mays using zinc-finger nucleases. Nature 2009, 459, 437–441. [Google Scholar] [CrossRef]
- Li, T.; Liu, B.; Chen, C.Y.; Yang, B.J. TALEN-mediated homologous recombination produces site-directed DNA base change and herbicide-resistant rice. J. Genet. Genomics 2016, 43, 297–305. [Google Scholar] [CrossRef]
- Li, M.; Li, X.; Zhou, Z.; Wu, P.; Fang, M.; Pan, X.; Lin, Q.; Luo, W.; Wu, G.; Li, H. Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas9 system. Front. Plant Sci. 2016, 7, 377. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Xu, Y.; Wan, J. Crop genome editing: A way to breeding by design. Crop J. 2020, 8, 379–383. [Google Scholar] [CrossRef]
- Butt, H.; Rao, G.S.; Sedeek, K.; Aman, R.; Kamel, R.; Mahfouz, M.J. Engineering herbicide resistance via prime editing in rice. Plant Biotechnol. J. 2020, 18, 2370–2372. [Google Scholar] [CrossRef]
- Naso, G.; Petrova, A. CRISPR/Cas9 gene editing for genodermatoses: Progress and perspectives. Emerging Top. Life Sci. 2019, 3, 313–326. [Google Scholar]
- Ahmadi, A.R.; Shahbazi, S.; Diyanat, M. Efficacy of five herbicides for weed control in rain-fed lentil (Lens culinaris Medik.). Weed Technol. 2016, 30, 448–455. [Google Scholar] [CrossRef]
- Dayan, F.E. Current status and future prospects in herbicide discovery. Plants 2019, 8, 341. [Google Scholar] [CrossRef] [PubMed]
- Herbicide Resistance Action Committee (HRAC). New Mode of Action of Herbicides. Available online: https://hracglobal.com/tools/hrac-moa-2020-revision-description-and-master-herbicide-list (accessed on 18 February 2021).
- Shamkuwar, S.; Baral, S.; Budhe, V.; Gupta, P.; Swarnkar, R. A critical study on weed control techniques. Int. J. Adv. Agric. Sci. Technol. 2019, 6, 1–12. [Google Scholar]
- Vats, S. Herbicides: History, classification and genetic manipulation of plants for herbicide resistance. In Sustainable Agriculture Reviews; Lichtfouse, E., Ed.; Springer: Cham, Switzerland, 2015; Volume 15, pp. 153–192. [Google Scholar]
- Staub, J.M.; Brand, L.; Tran, M.; Kong, Y.; Rogers, S.G. Bacterial glyphosate resistance conferred by overexpression of an E. coli membrane efflux transporter. J. Ind. Microbiol. Biotechnol. 2012, 39, 641–647. [Google Scholar] [CrossRef] [PubMed]
- Schönbrunn, E.; Eschenburg, S.; Shuttleworth, W.A.; Schloss, J.V.; Amrhein, N.; Evans, J.N.; Kabsch, W. Interaction of the herbicide glyphosate with its target enzyme 5-enolpyruvylshikimate 3-phosphate synthase in atomic detail. Proc. Natl. Acad. Sci. USA 2001, 98, 1376–1380. [Google Scholar] [CrossRef]
- Van Almsick, A. New HPPD-inhibitors–a proven mode of action as a new hope to solve current weed problems. Outlooks Pest Manag. 2009, 20, 27–30. [Google Scholar] [CrossRef]
- Godar, A.S.; Varanasi, V.K.; Nakka, S.; Prasad, P.V.; Thompson, C.R.; Mithila, J. Physiological and molecular mechanisms of differential sensitivity of Palmer amaranth (Amaranthus palmeri) to mesotrione at varying growth temperatures. PLoS ONE 2015, 10, e0126731. [Google Scholar] [CrossRef]
- Duke, S.O. Biotechnology: Herbicide-resistant crops. In Encyclopedia of Agriculture and Food Systems; Academic Press: San Diego, CA, USA, 2014; pp. 94–116. [Google Scholar]
- Takano, H.K.; Beffa, R.; Preston, C.; Westra, P.; Dayan, F.E. Reactive oxygen species trigger the fast action of glufosinate. Plants 2019, 249, 1837–1849. [Google Scholar] [CrossRef] [PubMed]
- Dilipkumar, M.; Chuah, T.S.; Goh, S.S.; Sahid, I. Weed management issues, challenges, and opportunities in Malaysia. Crop Prot. 2020, 134, 104347. [Google Scholar] [CrossRef]
- Marble, S.C.; Koeser, A.K.; Hasing, G. A review of weed control practices in landscape planting beds: Part II—Chemical weed control methods. Hortscience 2015, 50, 857–862. [Google Scholar] [CrossRef]
- Heap, I. The International Herbicide-Resistant Weed Database. Available online: www.weedscience.org (accessed on 17 February 2021).
- Yan, Y.; Liu, Q.; Zang, X.; Yuan, S.; Bat-Erdene, U.; Nguyen, C.; Gan, J.; Zhou, J.; Jacobsen, S.E.; Tang, Y. Resistance-gene-directed discovery of a natural-product herbicide with a new mode of action. Nature 2018, 559, 415–418. [Google Scholar] [CrossRef]
- Duke, S.O. The history and current status of glyphosate. Pest Mang. Sci. 2018, 74, 1027–1034. [Google Scholar] [CrossRef] [PubMed]
- Dayan, F.E.; Rimando, A.M.; Pan, Z.; Baerson, S.R.; Gimsing, A.L.; Duke, S.O. Sorgoleone. Phytochemistry 2010, 71, 1032–1039. [Google Scholar] [CrossRef]
- Tibugari, H.; Manyeruke, N.; Mafere, G.; Chakavarika, M.; Nyamuzuwe, L.; Marumahoko, P.; Mandumbu, R. Allelopathic effect of stressing sorghum on weed growth. Cogent. Biol. 2019, 5, 1684865. [Google Scholar] [CrossRef]
- Einhellig, F.A.; Rasmussen, J.A.; Hejl, A.M.; Souza, I.F. Effects of root exudate sorgoleone on photosynthesis. J. Chem. Ecol. 1993, 19, 369–375. [Google Scholar] [CrossRef]
- Meazza, G.; Scheffler, B.E.; Tellez, M.R.; Rimando, A.M.; Romagni, J.G.; Duke, S.O.; Nanayakkara, D.; Khan, I.A.; Abourashed, E.A.; Dayan, F.E. The inhibitory activity of natural products on plant p-hydroxyphenylpyruvate dioxygenase. Phytochemistry 2002, 60, 281–288. [Google Scholar] [CrossRef]
- Hejl, A.M.; Koster, K.L. The allelochemical sorgoleone inhibits root H+-ATPase and water uptake. J. Chem. Ecol. 2004, 30, 2181–2191. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Liu, W.; Jin, T.; Peng, X.; Zhang, L.; Wang, J. Bipyrazone: A new HppD-inhibiting herbicide in wheat. Sci. Rep. 2020, 10, 1–10. [Google Scholar] [CrossRef]
- Food and Agricuture Organization of the United States (FAO). What Is the Integrated Weed Management. Available online: http://www.fao.org/agriculture/crops/thematic-sitemap/theme/spi/scpi-home/managing-ecosystems/integrated-weed-management/iwm-what/en/#a (accessed on 15 February 2021).
- Harker, K.N.; O’Donovan, J.T. Recent weed control, weed management, and integrated weed management. Weed Technol. 2013, 27, 1–11. [Google Scholar] [CrossRef]
- Green, J.M. The benefits of herbicide-resistant crops. Pest Manag. Sci. 2012, 68, 1323–1331. [Google Scholar] [CrossRef]
- Awan, M.; Abass, M.; Muzaffar, A.; Ali, A.; Tabassum, B.; Rao, A.; Ahmad Nasir, I.; Husnain, T. Transformation of insect and herbicide resistance genes in cotton (Gossypium hirsutum L.). J. Agric. Sci. Technol. 2015, 17, 287–298. [Google Scholar]
- Green, J.M.; Owen, M.D. Herbicide-resistant crops: Utilities and limitations for herbicide-resistant weed management. J. Agric. Food Chem. 2011, 59, 5819–5829. [Google Scholar] [CrossRef] [PubMed]
- Duke, S.O.; Cerdeira, A.L. Transgenic crops for herbicide resistance. In Transgenic Crop Plants; Kole, C., Michler, C.H., Abbott, A.G., Hall, T.C., Eds.; Springer: Berlin, Germany, 2010; pp. 133–166. [Google Scholar]
- Schütte, G.; Eckerstorfer, M.; Rastelli, V.; Reichenbecher, W.; Restrepo-Vassalli, S.; Ruohonen-Lehto, M.; Saucy, A.-G.W.; Mertens, M. Herbicide resistance and biodiversity: Agronomic and environmental aspects of genetically modified herbicide-resistant plants. Environ. Sci. Eur. 2017, 29, 3. [Google Scholar] [CrossRef] [PubMed]
- Roychowdhury, R.; Tah, J. Mutagenesis—A potential approach for crop improvement. In Crop Improvement; Hakeem, K., Ahmad, P., Ozturk, M., Eds.; Springer: Boston, MA, USA, 2013; pp. 149–187. [Google Scholar]
- Zoller, M.J. New molecular biology methods for protein engineering. Curr. Opin. Biotechnol. 1991, 2, 526–531. [Google Scholar] [CrossRef]
- Ho, S.N.; Hunt, H.D.; Horton, R.M.; Pullen, J.K.; Pease, L.R. Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene 1989, 77, 51–59. [Google Scholar] [CrossRef]
- Oladosu, Y.; Rafii, M.Y.; Abdullah, N.; Hussin, G.; Ramli, A.; Rahim, H.A.; Miah, G.; Usman, M. Principle and application of plant mutagenesis in crop improvement: A review. Biotechnol. Biotechnol. Equip. 2016, 30, 1–16. [Google Scholar] [CrossRef]
- Forster, B.; Shu, Q. Plant mutation breeding and biotechnology. In Plant Mutagenesis in Crop Improvement: Basic Terms and Applications; Shu, Q.Y., Forster, B.P., Nakagawa, H., Eds.; CABI: Wallingford, UK, 2012; pp. 9–20. [Google Scholar]
- Shu, Q.; Forster, B.; Nakagawa, H. Principles and applications of plant mutation breeding. In Plant Mutagenesis in Crop Improvement: Basic Terms and Applications; Shu, Q.Y., Forster, B.P., Nakagawa, H., Eds.; CABI: Wallingford, UK, 2012; pp. 301–325. [Google Scholar]
- Jung, C.; Capistrano-Gossmann, G.; Braatz, J.; Sashidhar, N.; Melzer, S. Recent developments in genome editing and applications in plant breeding. Plant Breed. 2018, 137, 1–9. [Google Scholar] [CrossRef]
- Sebastian, S.; Fader, G.; Ulrich, J.; Forney, D.; Chaleff, R. Semidominant soybean mutation for resistance to sulfonylurea herbicides. Crop Sci. 1989, 29, 1403–1408. [Google Scholar] [CrossRef]
- Sala, C.A.; Bulos, M.; Altieri, E.; Ramos, M.L. Genetics and breeding of herbicide tolerance in sunflower. Helia 2012, 35, 57–70. [Google Scholar]
- Newhouse, K.E.; Smith, W.A.; Starrett, M.A.; Schaefer, T.J.; Singh, B.K.J.P.P. Tolerance to imidazolinone herbicides in wheat. Plant Physiol. 1992, 100, 882–886. [Google Scholar] [CrossRef]
- Tan, S.; Bowe, S.J. Developing herbicide-tolerant crops from mutations. In Induced Plant Mutations in the Genomics Era; Shu, Q.Y., Ed.; Joint FAO/IAEA Program: Rome, Italy, 2009; p. 315. [Google Scholar]
- Lenser, T.; Theißen, G.J. Molecular mechanisms involved in convergent crop domestication. Trends Plant Sci. 2013, 18, 704–714. [Google Scholar] [CrossRef]
- Georges, F.; Ray, H. Genome editing of crops: A renewed opportunity for food security. GM Crops Food 2017, 8, 1–12. [Google Scholar] [CrossRef] [PubMed]
- International Service for the Acquisition of Agri-Biotech Applications (ISAAA). Pocket K No. 17. Genetic Engineering and GM Crops. Available online: https://www.isaaa.org/resources/publications/pocketk/document/Doc-Pocket%20K17.pdf (accessed on 8 October 2020).
- Beckie, H.J.; Hall, L.M. Genetically-modified herbicide-resistant (GMHR) crops a two-edged sword? An Americas perspective on development and effect on weed management. Crop Prot. 2014, 66, 40–45. [Google Scholar] [CrossRef]
- Busi, R.; Powles, S.B. Transgenic glyphosate-resistant canola (Brassica napus) can persist outside agricultural fields in Australia. Agric. Ecosyst. Environ. 2016, 220, 28–34. [Google Scholar] [CrossRef]
- International Service for the Acquisition of Agri-Biotech Applications (ISAAA). Pocket K No. 16. Biotech Crop Highlights in 2018. Available online: https://www.isaaa.org/resources/publications/pocketk/document/Doc-Pocket%20K16-2019.pdf (accessed on 8 October 2020).
- Fartyal, D.; Agarwal, A.; James, D.; Borphukan, B.; Ram, B.; Sheri, V.; Yadav, R.; Manna, M.; Varakumar, P.; Achary, V.M.M. Co-expression of P173S mutant rice EPSPS and igrA genes results in higher glyphosate tolerance in transgenic rice. Front. Plant Sci. 2018, 9, 144. [Google Scholar] [CrossRef]
- Dun, B.; Wang, X.; Lu, W.; Chen, M.; Zhang, W.; Ping, S.; Wang, Z.; Zhang, B.; Lin, M. Development of highly glyphosate-tolerant tobacco by coexpression of glyphosate acetyltransferase gat and EPSPS G2-aroA genes. Crop J. 2014, 2, 164–169. [Google Scholar] [CrossRef]
- Guo, B.; Guo, Y.; Hong, H.; Jin, L.; Zhang, L.; Chang, R.-Z.; Lu, W.; Lin, M.; Qiu, L.J. Co-expression of G2-EPSPS and glyphosate acetyltransferase GAT genes conferring high tolerance to glyphosate in soybean. Front. Plant Sci. 2015, 6, 847. [Google Scholar] [CrossRef] [PubMed]
- Xiao, P.Y.; Yi, L.; Cao, Y.P. Overexpression of G10-EPSPS in soybean provides high glyphosate tolerance. J. Integr. Agric. 2019, 18, 1851–1858. [Google Scholar] [CrossRef]
- Achary, V.M.M.; Sheri, V.; Manna, M.; Panditi, V.; Borphukan, B.; Ram, B.; Agarwal, A.; Fartyal, D.; Teotia, D.; Masakapalli, S.K. Overexpression of improved EPSPS gene results in field level glyphosate tolerance and higher grain yield in rice. Plant Biotechnol. J. 2020, 18, 2504–2519. [Google Scholar] [CrossRef]
- Sanvido, O.; Romeis, J.; Gathmann, A.; Gielkens, M.; Raybould, A.; Bigler, F. Evaluating environmental risks of genetically modified crops: Ecological harm criteria for regulatory decision-making. Environ. Sci. Policy 2012, 15, 82–91. [Google Scholar] [CrossRef]
- Kim, J.; Kim, J.S. Bypassing GMO regulations with CRISPR gene editing. Nat. Biotechnol. 2016, 34, 1014–1015. [Google Scholar] [CrossRef]
- Li, Y.; Zhu, J.; Wu, H.; Liu, C.; Huang, C.; Lan, J.; Zhao, Y.; Xie, C. Precise base editing of non-allelic acetolactate synthase genes confers sulfonylurea herbicide resistance in maize. Crop J. 2020, 8, 449–456. [Google Scholar] [CrossRef]
- Wang, Q.; Alariqi, M.; Wang, F.; Li, B.; Ding, X.; Rui, H.; Li, Y.; Xu, Z.; Qin, L.; Sun, L. The application of a heat-inducible CRISPR/Cas12b (C2c1) genome editing system in tetraploid cotton (G. hirsutum) plants. Plant Biotechnol. J. 2020, 18, 2436–2443. [Google Scholar] [CrossRef]
- Li, B.; Rui, H.; Li, Y.; Wang, Q.; Alariqi, M.; Qin, L.; Sun, L.; Ding, X.; Wang, F.; Zou, J. Robust CRISPR/Cpf1 (Cas12a)-mediated genome editing in allotetraploid cotton (Gossypium hirsutum). Plant Biotechnol. J. 2019, 17, 1862. [Google Scholar] [CrossRef]
- Manghwar, H.; Lindsey, K.; Zhang, X.; Jin, S. CRISPR/Cas system: Recent advances and future prospects for genome editing. Trends Plant Sci. 2019, 24, 1102–1125. [Google Scholar] [CrossRef]
- Ricroch, A. Global developments of genome editing in agriculture. Transgenic Res. 2019, 28, 45–52. [Google Scholar] [CrossRef]
- Wang, P.; Zhang, J.; Sun, L.; Ma, Y.; Xu, J.; Liang, S.; Deng, J.; Tan, J.; Zhang, Q.; Tu, L. High efficient multisites genome editing in allotetraploid cotton (Gossypium hirsutum) using CRISPR/Cas9 system. Plant Biotechnol. J. 2018, 16, 137–150. [Google Scholar] [CrossRef] [PubMed]
- Gaj, T.; Gersbach, C.A.; Barbas, C.F., III. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 2013, 31, 397–405. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Kaur, R.; Singh, A. Recent advances in CRISPR/Cas mediated genome editing for crop improvement. Plant Biotechnol. Rep. 2017, 11, 193–207. [Google Scholar] [CrossRef]
- Puchta, H. Applying CRISPR/Cas for genome engineering in plants: The best is yet to come. Curr. Opin. Plant Biol. 2017, 36, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Lowder, L.G.; Zhang, T.; Malzahn, A.A.; Zheng, X.; Voytas, D.F.; Zhong, Z.; Chen, Y.; Ren, Q.; Li, Q. A CRISPR–Cpf1 system for efficient genome editing and transcriptional repression in plants. Nat. Plants 2017, 3, 1–5. [Google Scholar]
- Doudna, J.A.; Charpentier, E. The new frontier of genome engineering with CRISPR-Cas9. Science 2014, 346, 1258096-1–1258096-9. [Google Scholar] [CrossRef] [PubMed]
- Manghwar, H.; Li, B.; Ding, X.; Hussain, A.; Lindsey, K.; Zhang, X.; Jin, S. CRISPR/Cas Systems in Genome Editing: Methodologies and tools for sgRNA design, off-target evaluation, and strategies to mitigate off-target effects. Adv. Sci. 2020, 7, 1902312. [Google Scholar] [CrossRef]
- Chneiweiss, H.; Hirsch, F.; Montoliu, L.; Müller, A.M.; Fenet, S.; Abecassis, M.; Merchant, J.; Baertschi, B.; Botbol-Baum, M.; Houghton, J.A. Fostering responsible research with genome editing technologies: A European perspective. Transgenic Res. 2017, 26, 709–713. [Google Scholar] [CrossRef]
- Li, B.; Liang, S.; Alariqi, M.; Wang, F.; Wang, G.; Wang, Q.; Xu, Z.; Yu, L.; Zafar, M.N.; Sun, L. The application of temperature sensitivity CRISPR/LbCpf1 (LbCas12a) mediated genome editing in allotetraploid cotton (G. hirsutum) and creation of nontransgenic, gossypol-free cotton. Plant Biotechnol. J. 2020, 19, 221–223. [Google Scholar] [CrossRef]
- Kumar, V.; Jain, M.J. The CRISPR–Cas system for plant genome editing: Advances and opportunities. J. Exp. Bot. 2015, 66, 47–57. [Google Scholar] [CrossRef] [PubMed]
- Svitashev, S.; Young, J.K.; Schwartz, C.; Gao, H.; Falco, S.C.; Cigan, A.M. Targeted mutagenesis, precise gene editing, and site-specific gene insertion in maize using Cas9 and guide RNA. Plant Physiol. 2015, 169, 931–945. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Zhang, X.; Wu, C.; He, Y.; Ma, Y.; Hou, H.; Guo, X.; Du, W.; Zhao, Y.; Xia, L. Engineering herbicide-resistant rice plants through CRISPR/Cas9-mediated homologous recombination of acetolactate synthase. Mol. Plant 2016, 9, 628–631. [Google Scholar] [CrossRef]
- Wang, F.; Xu, Y.; Li, W.; Chen, Z.; Wang, J.; Fan, F.; Tao, Y.; Jiang, Y.; Zhu, Q.-H.; Yang, J. Creating a novel herbicide-tolerance OsALS allele using CRISPR/Cas9-mediated gene editing. Crop J. 2020. [Google Scholar] [CrossRef]
- Shimatani, Z.; Fujikura, U.; Ishii, H.; Matsui, Y.; Suzuki, M.; Ueke, Y.; Taoka, K.-I.; Terada, R.; Nishida, K.; Kondo, A. Inheritance of co-edited genes by CRISPR-based targeted nucleotide substitutions in rice. Plant Physiol. Biochem. 2018, 131, 78–83. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Kuang, Y.; Yan, F.; Li, S.; Ren, B.; Gosavi, G.; Spetz, C.; Li, X.; Wang, X.; Zhou, X. Developing a novel artificial rice germplasm for dinitroaniline herbicide resistance by base editing of OsTubA2. Plant Biotechnol. J. 2020, 19, 5–7. [Google Scholar] [CrossRef]
- Liu, X.; Qin, R.; Li, J.; Liao, S.; Shan, T.; Xu, R.; Wu, D.; Wei, P. A CRISPR-Cas9-mediated domain-specific base-editing screen enables functional assessment of ACCase variants in rice. Plant Biotechnol. J. 2020, 18, 1845–1847. [Google Scholar] [CrossRef]
- Kuang, Y.; Li, S.; Ren, B.; Yan, F.; Spetz, C.; Li, X.; Zhou, X.; Zhou, H. Base-editing-mediated artificial evolution of OsALS1 in planta to develop novel herbicide-tolerant rice germplasms. Mol. Plant 2020, 13, 565–572. [Google Scholar] [CrossRef]
- Li, C.; Zhang, R.; Meng, X.; Chen, S.; Zong, Y.; Lu, C.; Qiu, J.-L.; Chen, Y.H.; Li, J.; Gao, C. Targeted, random mutagenesis of plant genes with dual cytosine and adenine base editors. Nat. Biotechnol. 2020, 38, 875–882. [Google Scholar] [CrossRef]
- Li, J.; Meng, X.; Zong, Y.; Chen, K.; Zhang, H.; Liu, J.; Li, J.; Gao, C. Gene replacements and insertions in rice by intron targeting using CRISPR–Cas9. Nat. Plants 2016, 2, 1–6. [Google Scholar] [CrossRef]
- Li, C.; Zong, Y.; Wang, Y.; Jin, S.; Zhang, D.; Song, Q.; Zhang, R.; Gao, C. Expanded base editing in rice and wheat using a Cas9-adenosine deaminase fusion. Genome Biol. 2018, 19, 59. [Google Scholar] [CrossRef]
- Hua, K.; Jiang, Y.; Tao, X.; Zhu, J.K. Precision genome engineering in rice using prime editing system. Plant Biotechnol. J. 2020, 18, 2167–2169. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Li, J.; Liu, X.; Shan, T.; Qin, R.; Wei, P. Development of Plant Prime-Editing Systems for Precise Genome Editing. Plant Commun. 2020, 1, 100043. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.; Jiang, L.; Cui, X.; Zhang, J.; Guo, S.; Li, M.; Zhang, H.; Ren, Y.; Gong, G.; Zong, M. Engineering herbicide-resistant watermelon variety through CRISPR/Cas9-mediated base-editing. Plant Cell Rep. 2018, 37, 1353–1356. [Google Scholar] [CrossRef] [PubMed]
- Zong, Y.; Song, Q.; Li, C.; Jin, S.; Zhang, D.; Wang, Y.; Qiu, J.-L.; Gao, C. Efficient C-to-T base editing in plants using a fusion of nCas9 and human APOBEC3A. Nat. Biotechnol. 2018, 36, 950–953. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Wang, Z.; Ni, H.; Xu, Y.; Chen, Q.; Jiang, L.J.S.C.L.S. CRISPR/Cas9-mediated base-editing system efficiently generates gain-of-function mutations in Arabidopsis. Sci. China Life Sci. 2017, 60, 520–523. [Google Scholar] [CrossRef]
- Wu, J.; Chen, C.; Xian, G.; Liu, D.; Lin, L.; Yin, S.; Sun, Q.; Fang, Y.; Zhang, H.; Wang, Y. Engineering herbicide-resistant oilseed rape by CRISPR/Cas9-mediated cytosine base-editing. Plant Biotechnol. J. 2020, 18, 1857–1859. [Google Scholar] [CrossRef] [PubMed]
- Veillet, F.; Perrot, L.; Chauvin, L.; Kermarrec, M.-P.; Guyon-Debast, A.; Chauvin, J.-E.; Nogué, F.; Mazier, M. Transgene-free genome editing in tomato and potato plants using agrobacterium-mediated delivery of a CRISPR/Cas9 cytidine base editor. Int. J. Mol. Sci. 2019, 20, 402. [Google Scholar] [CrossRef]
- Li, Z.; Liu, Z.-B.; Xing, A.; Moon, B.P.; Koellhoffer, J.P.; Huang, L.; Ward, R.T.; Clifton, E.; Falco, S.C.; Cigan, A.M. Cas9-guide RNA directed genome editing in soybean. Plant Physiol. 2015, 169, 960–970. [Google Scholar] [CrossRef]
- Sauer, N.J.; Narváez-Vásquez, J.; Mozoruk, J.; Miller, R.B.; Warburg, Z.J.; Woodward, M.J.; Mihiret, Y.A.; Lincoln, T.A.; Segami, R.E.; Sanders, S.L. Oligonucleotide-mediated genome editing provides precision and function to engineered nucleases and antibiotics in plants. Plant Physiol. 2016, 170, 1917–1928. [Google Scholar] [CrossRef]
- Ortega, J.L.; Rajapakse, W.; Bagga, S.; Apodaca, K.; Lucero, Y.; Sengupta-Gopalan, C. An intragenic approach to confer glyphosate resistance in chile (Capsicum annuum) by introducing an in vitro mutagenized chile EPSPS gene encoding for a glyphosate resistant EPSPS protein. PLoS ONE 2018, 13, e0194666. [Google Scholar] [CrossRef]
- Hummel, A.W.; Chauhan, R.D.; Cermak, T.; Mutka, A.M.; Vijayaraghavan, A.; Boyher, A.; Starker, C.G.; Bart, R.; Voytas, D.F.; Taylor, N.J. Allele exchange at the EPSPS locus confers glyphosate tolerance in cassava. Plant Biotechnol. J. 2018, 16, 1275–1282. [Google Scholar] [CrossRef]
- Butler, N.M.; Baltes, N.J.; Voytas, D.F.; Douches, D.S. Geminivirus-mediated genome editing in potato (Solanum tuberosum L.) using sequence-specific nucleases. Front. Plant Sci. 2016, 7, 1045. [Google Scholar] [CrossRef]
- Baerson, S.R.; Rodriguez, D.J.; Tran, M.; Feng, Y.; Biest, N.A.; Dill, G.M. Glyphosate-resistant goosegrass. Identification of a mutation in the target enzyme 5-enolpyruvylshikimate-3-phosphate synthase. Plant Physiol. 2002, 129, 1265–1275. [Google Scholar] [CrossRef]
- Yu, Q.; Jalaludin, A.; Han, H.; Chen, M.; Sammons, R.D.; Powles, S.B. Evolution of a double amino acid substitution in the 5-enolpyruvylshikimate-3-phosphate synthase in Eleusine indica conferring high-level glyphosate resistance. Plant Physiol. 2015, 167, 1440–1447. [Google Scholar] [CrossRef] [PubMed]
- Ali, Z.; Shami, A.; Sedeek, K.; Kamel, R.; Alhabsi, A.; Tehseen, M.; Hassan, N.; Butt, H.; Kababji, A.; Hamdan, S.M. Fusion of the Cas9 endonuclease and the VirD2 relaxase facilitates homology-directed repair for precise genome engineering in rice. Commun. Biol. 2020, 3, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Butt, H.; Eid, A.; Ali, Z.; Atia, M.A.; Mokhtar, M.M.; Hassan, N.; Lee, C.M.; Bao, G.; Mahfouz, M.M. Efficient CRISPR/Cas9-mediated genome editing using a chimeric single-guide RNA molecule. Front. Plant Sci. 2017, 8, 1441. [Google Scholar] [CrossRef] [PubMed]
- Shan, Q.; Wang, Y.; Li, J.; Zhang, Y.; Chen, K.; Liang, Z.; Zhang, K.; Liu, J.; Xi, J.J.; Qiu, J.L. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 2013, 31, 686–688. [Google Scholar] [CrossRef]
- Bharat, S.S.; Li, S.; Li, J.; Yan, L.; Xia, L. Base editing in plants: Current status and challenges. Crop J. 2020, 8, 384–395. [Google Scholar] [CrossRef]
- Qin, L.; Li, J.; Wang, Q.; Xu, Z.; Sun, L.; Alariqi, M.; Manghwar, H.; Wang, G.; Li, B.; Ding, X. High-efficient and precise base editing of C• G to T• A in the allotetraploid cotton (Gossypium hirsutum) genome using a modified CRISPR/Cas9 system. Plant Biotechnol. J. 2020, 18, 45–56. [Google Scholar] [CrossRef] [PubMed]
- Packer, M.S.; Liu, D.R. Methods for the directed evolution of proteins. Nat. Rev. Genet. 2015, 16, 379–394. [Google Scholar] [CrossRef] [PubMed]
- Komor, A.C.; Kim, Y.B.; Packer, M.S.; Zuris, J.A.; Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 2016, 533, 420–424. [Google Scholar] [CrossRef]
- Nishida, K.; Arazoe, T.; Yachie, N.; Banno, S.; Kakimoto, M.; Tabata, M.; Mochizuki, M.; Miyabe, A.; Araki, M.; Hara, K.Y. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 2016, 353, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Yu, Q.; Powles, S.B. Resistance to AHAS inhibitor herbicides: Current understanding. Pest Manag. Sci. 2014, 70, 1340–1350. [Google Scholar] [CrossRef] [PubMed]
- Baud, S.; Bellec, Y.; Miquel, M.; Bellini, C.; Caboche, M.; Lepiniec, L.; Faure, J.D.; Rochat, C. 11and pasticcino3 mutants affected in embryo development are impaired in acetyl-CoA carboxylase. EMBO Rep. 2004, 5, 515–520. [Google Scholar] [CrossRef] [PubMed]
- Jang, S.; Marjanovic, J.; Gornicki, P. Resistance to herbicides caused by single amino acid mutations in acetyl-C o A carboxylase in resistant populations of grassy weeds. New Phytol. 2013, 197, 1110–1116. [Google Scholar] [CrossRef]
- Shimatani, Z.; Kashojiya, S.; Takayama, M.; Terada, R.; Arazoe, T.; Ishii, H.; Teramura, H.; Yamamoto, T.; Komatsu, H.; Miura, K. Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat. Biotechnol. 2017, 35, 441–443. [Google Scholar] [CrossRef]
- Ma, Y.; Zhang, J.; Yin, W.; Zhang, Z.; Song, Y.; Chang, X. Targeted AID-mediated mutagenesis (TAM) enables efficient genomic diversification in mammalian cells. Nat. Methods 2016, 13, 1029–1035. [Google Scholar] [CrossRef]
- Anzalone, A.V.; Randolph, P.B.; Davis, J.R.; Sousa, A.A.; Koblan, L.W.; Levy, J.M.; Chen, P.J.; Wilson, C.; Newby, G.A.; Raguram, A. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 2019, 576, 149–157. [Google Scholar] [CrossRef]
- Lin, Q.; Zong, Y.; Xue, C.; Wang, S.; Jin, S.; Zhu, Z.; Wang, Y.; Anzalone, A.V.; Raguram, A.; Doman, J.L. Prime genome editing in rice and wheat. Nat. Biotechnol. 2020, 38, 582–585. [Google Scholar] [CrossRef]
- Han, Y.J.; Kim, J.I. Application of CRISPR/Cas9-mediated gene editing for the development of herbicide-resistant plants. Plant Biotechnol. Rep. 2019, 13, 447–457. [Google Scholar] [CrossRef]
- Zhang, Y.; Malzahn, A.A.; Sretenovic, S.; Qi, Y. The emerging and uncultivated potential of CRISPR technology in plant science. Nat. Plants 2019, 5, 778–794. [Google Scholar] [CrossRef]
- Huang, S.; Weigel, D.; Beachy, R.N.; Li, J. A proposed regulatory framework for genome-edited crops. Nat. Genet. 2016, 48, 109–111. [Google Scholar] [CrossRef] [PubMed]
- Lassoued, R.; Macall, D.M.; Hesseln, H.; Phillips, P.W.; Smyth, S.J. Benefits of genome-edited crops: Expert opinion. Transgenic Res. 2019, 28, 247–256. [Google Scholar] [CrossRef] [PubMed]
- Ishii, T.; Araki, M. A future scenario of the global regulatory landscape regarding genome-edited crops. GM Crops Food 2017, 8, 44–56. [Google Scholar] [CrossRef]
- Waltz, E. Gene-edited CRISPR mushroom escapes US regulation. Nat. News 2016, 532, 293. [Google Scholar] [CrossRef] [PubMed]
- Grohmann, L.; Keilwagen, J.; Duensing, N.; Dagand, E.; Hartung, F.; Wilhelm, R.; Bendiek, J.; Sprink, T. Detection and identification of genome editing in plants: Challenges and opportunities. Front. Plant Sci. 2019, 10, 236. [Google Scholar] [CrossRef] [PubMed]
- Jansing, J.; Schiermeyer, A.; Schillberg, S.; Fischer, R.; Bortesi, L. Genome editing in agriculture: Technical and practical considerations. Int. J. Mol. Sci. 2019, 20, 2888. [Google Scholar] [CrossRef]
- Pineda, M.; Lear, A.; Collins, J.P.; Kiani, S. Safe CRISPR: Challenges and possible solutions. Trends Biotechnol. 2019, 37, 389–401. [Google Scholar] [CrossRef]
- El-Mounadi, K.; Morales-Floriano, M.L.; Garcia-Ruiz, H. Principles, Applications, and Biosafety of Plant Genome Editing Using CRISPR-Cas9. Front. Plant Sci. 2020, 11, 56. [Google Scholar] [CrossRef]
- Garcia Ruiz, M.T.; Knapp, A.N.; Garcia-Ruiz, H. Profile of genetically modified plants authorized in Mexico. GM Crops Food 2018, 9, 152–168. [Google Scholar] [CrossRef] [PubMed]
- Eckerstorfer, M.F.; Engelhard, M.; Heissenberger, A.; Simon, S.; Teichmann, H. Plants developed by new genetic modification techniques—comparison of existing regulatory frameworks in the EU and non-EU countries. Front. Bioeng. Biotechnol. 2019, 7, 26. [Google Scholar] [CrossRef] [PubMed]
- Burgos, N.R.; Singh, V.; Tseng, T.M.; Black, H.; Young, N.D.; Huang, Z.; Hyma, K.E.; Gealy, D.R.; Caicedo, A.L. The impact of herbicide-resistant rice technology on phenotypic diversity and population structure of United States weedy rice. Plant Physiol. 2014, 166, 1208–1220. [Google Scholar] [CrossRef]
- Ishii, T.; Araki, M. Consumer acceptance of food crops developed by genome editing. GM Crops Food 2016, 35, 1507–1518. [Google Scholar] [CrossRef]
- Voytas, D.F.; Gao, C. Precision genome engineering and agriculture: Opportunities and regulatory challenges. PLoS Biol. 2014, 12, e1001877. [Google Scholar] [CrossRef] [PubMed]
- Wallace, J.G.; Rodgers-Melnick, E.; Buckler, E.S. On the road to breeding 4.0: Unraveling the good, the bad, and the boring of crop quantitative genomics. Annu. Rev. Genet. 2018, 52, 421–444. [Google Scholar] [PubMed]
- Nakka, S.; Jugulam, M.; Peterson, D.; Asif, M. Herbicide resistance: Development of wheat production systems and current status of resistant weeds in wheat cropping systems. Crop J. 2019, 7, 750–760. [Google Scholar] [CrossRef]
- Green, J.M. The rise and future of glyphosate and glyphosate-resistant crops. Pest Manag. Sci. 2018, 74, 1035–1039. [Google Scholar] [CrossRef] [PubMed]
- Metje-Sprink, J.; Sprink, T.; Hartung, F. Genome-edited plants in the field. Curr. Opin. Biotechnol. 2020, 61, 1–6. [Google Scholar] [CrossRef]
- CIBUS. Our Crops. Available online: https://www.cibus.com/crops.php (accessed on 17 September 2020).
- Gullickson, G. New Herbicide Site of Action, Gene-Edited Short-Stature Corn, Xtendflex Coming from Bayer. Available online: https://www.agriculture.com/news/crops/new-herbicide-site-of-action-gene-edited-short-stature-corn-xtendflex-coming-from-bayer (accessed on 21 October 2020).
- Zhang, Y.; Pribil, M.; Palmgren, M.; Gao, C. A CRISPR way for accelerating improvement of food crops. Nat. Food 2020, 1, 200–205. [Google Scholar] [CrossRef]
- Sun, J.L.; Nakagawa, H.; Karita, S.; Ohmiya, K.; Hattori, T. Rice embryo globulins: Amino-terminal amino acid sequences, cDNA cloning and expression. Plant Cell Physiol. 1996, 37, 612–620. [Google Scholar] [CrossRef]
- Mariani, C.; De Beuckeleer, M.; Truettner, J.; Leemans, J.; Goldberg, R.B. Induction of male sterility in plants by a chimaeric ribonuclease gene. Nature 1990, 347, 737–741. [Google Scholar] [CrossRef]
- Hu, J.; Wang, K.; Huang, W.; Liu, G.; Gao, Y.; Wang, J.; Huang, Q.; Ji, Y.; Qin, X.; Wan, L. The rice pentatricopeptide repeat protein RF5 restores fertility in Hong-Lian cytoplasmic male-sterile lines via a complex with the glycine-rich protein GRP162. Plant Cell 2012, 24, 109–122. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Zhu, M.; Wang, L.; Wu, J.; Wang, Q.; Wang, R.; Zhao, Y. Programmed self elimination of the CRISPR/Cas9 construct greatly accelerates the isolation of edited and transgene-free rice plants. Mol. Plant 2018, 11, 1210–1213. [Google Scholar] [CrossRef] [PubMed]
| Plant | Genome Editing System | Delivery Method | Target Gene | Target Herbicide | Reference |
|---|---|---|---|---|---|
| Rice | CRISPR-Cas9 | Particle bombardment | ALS | Bispyribac- sodium | [111] |
| CRISPR-Cas9 | Agrobacterium-mediated transformation | ALS | Imazethapyr | [112] | |
| TALEN | Ballistic delivery | OsALS | Bispyribac- sodium | [37] | |
| Target-AID | Agrobacterium-mediated transformation | ALS | Imazamox | [113] | |
| ABE | Agrobacterium-mediated transformation | OsTubA2 | Dinitroaniline | [114] | |
| CBE | Agrobacterium-mediated transformation | OsACCase | Gallant | [115] | |
| BEMGE | Agrobacterium-mediated and Particle bombardment | OsALS1 | Bispyribac-sodium | [116] | |
| STEMEs | Agrobacterium-mediated transformation | ACCase | Haloxyfop | [117] | |
| CRISPR–Cas9 | Particle bombardment | EPSPS | Glyphosate | [118] | |
| ABE | Agrobacterium-mediated transformation | ACCase | Haloxyfop | [119] | |
| Prime editing | Agrobacterium-mediated transformation | OsALS | Bispyribac sodium | [40] | |
| Prime editing | Agrobacterium-mediated transformation | ALS | Imidazolinone | [120] | |
| Prime editing | Agrobacterium-mediated transformation | OsACCase1 | Haloxyfop | [121] | |
| Watermelon | CBE | Agrobacterium-mediated transformation | ClALS | Tribenuron | [122] |
| Wheat | n/dCas9-PBE | Particle bombardment | TaALS, ACCase | Sulfonylurea, Imidazolinone and Aryloxyphenoxy propionate | [26] |
| n/dCas9-PBE | Particle bombardment | TaALS | Nicosulfuron | [123] | |
| Maize | CBE | Agrobacterium-mediated transformation | ZmALS | Sulfonylurea | [95] |
| Cas9-gRNA | Particle bombardment | ALS | Chlorsulfuron | [110] | |
| Arabidopsis | CBE | Agrobacterium-mediated transformation | ALS | Tribenuron-methyl | [124] |
| Oilseed rape | CBE | Agrobacterium-mediated transformation | BnALS1 | Tribenuron-methyl | [125] |
| Tomato and Potato | CBE | Agrobacterium-mediated transformation | ALS | Chlorsulfuron | [126] |
| Soybean | CRISPR–Cas9 | Particle bombardment | ALS1 | Chlorsulfuron | [127] |
| Flax | ssODN and CRISPR/Cas9 | Protoplast transfection | EPSPS | Glyphosate | [128] |
| Chile peeper | Intragenic method | Agrobacterium-mediated transformation | EPSPS | Glyphosate | [129] |
| Cassava | Cas9-gRNA | Agrobacterium-mediated transformation | EPSPS | Glyphosate | [130] |
| Potato | GVR | Agrobacterium-mediated transformation | ALS1 | Imidazolinone | [131] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hussain, A.; Ding, X.; Alariqi, M.; Manghwar, H.; Hui, F.; Li, Y.; Cheng, J.; Wu, C.; Cao, J.; Jin, S. Herbicide Resistance: Another Hot Agronomic Trait for Plant Genome Editing. Plants 2021, 10, 621. https://doi.org/10.3390/plants10040621
Hussain A, Ding X, Alariqi M, Manghwar H, Hui F, Li Y, Cheng J, Wu C, Cao J, Jin S. Herbicide Resistance: Another Hot Agronomic Trait for Plant Genome Editing. Plants. 2021; 10(4):621. https://doi.org/10.3390/plants10040621
Chicago/Turabian StyleHussain, Amjad, Xiao Ding, Muna Alariqi, Hakim Manghwar, Fengjiao Hui, Yapei Li, Junqi Cheng, Chenglin Wu, Jinlin Cao, and Shuangxia Jin. 2021. "Herbicide Resistance: Another Hot Agronomic Trait for Plant Genome Editing" Plants 10, no. 4: 621. https://doi.org/10.3390/plants10040621
APA StyleHussain, A., Ding, X., Alariqi, M., Manghwar, H., Hui, F., Li, Y., Cheng, J., Wu, C., Cao, J., & Jin, S. (2021). Herbicide Resistance: Another Hot Agronomic Trait for Plant Genome Editing. Plants, 10(4), 621. https://doi.org/10.3390/plants10040621










